Young+Seok+Ju
-
 Unraveling Mitochondrial DNA Mutations in Human Cells
Throughout our lifetime, cells accumulate DNA mutations, which contribute to genetic diversity, or “mosaicism”, among cells. These genomic mutations are pivotal for the aging process and the onset of various diseases, including cancer. Mitochondria, essential cellular organelles involved in energy metabolism and apoptosis, possess their own DNA, which are susceptible to mutations. However, studies on mtDNA mutations and mosaicism have been limited due to a variety of technical challenges.
Genomic scientists from KAIST have revealed the genetic mosaicism characterized by variations in mitochondrial DNA (mtDNA) across normal human cells. This study provides fundamental insights into understanding human aging and disease onset mechanisms.
The study, “Mitochondrial DNA mosaicism in normal human somatic cells,” was published in Nature Genetics on July 22. It was led by graduate student Jisong An under the supervision of Professor Young Seok Ju from the Graduate School of Medical Science and Engineering.
Researchers from Seoul National University College of Medicine, Yonsei University College of Medicine, Korea University College of Medicine, Washington University School of Medicine National Cancer Center, Seoul National University Hospital, Gangnam Severance Hospital and KAIST faculty startup company Inocras Inc. also participated in this study.
< Figure 1. a. Flow of experiment. b. Schematic diagram illustrating the origin and dynamics of mtDNA alterations across a lifetime. >
The study involved a bioinformatic analysis of whole-genome sequences from 2,096 single cells obtained from normal human colorectal epithelial tissue, fibroblasts, and blood collected from 31 individuals. The study highlights an average of three significant mtDNA differences between cells, with approximately 6% of these variations confirmed to be inherited as heteroplasmy from the mother.
Moreover, mutations significantly increased during tumorigenesis, with some mutations contributing to instability in mitochondrial RNA. Based on these findings, the study illustrates a computational model that comprehensively elucidates the evolution of mitochondria from embryonic development to aging and tumorigenesis.
This study systematically reveals the mechanisms behind mitochondrial DNA mosaicism in normal human cells, establishing a crucial foundation for understanding the impact of mtDNA on aging and disease onset.
Professor Ju remarked, “By systematically utilizing whole-genome big data, we can illuminate previously unknown phenomena in life sciences.” He emphasized the significance of the study, adding, “For the first time, we have established a method to systematically understand mitochondrial DNA changes occurring during human embryonic development, aging, and cancer development.”
This work was supported by the National Research Foundation of Korea and the Suh Kyungbae Foundation.
2024.07.24 View 5430
Unraveling Mitochondrial DNA Mutations in Human Cells
Throughout our lifetime, cells accumulate DNA mutations, which contribute to genetic diversity, or “mosaicism”, among cells. These genomic mutations are pivotal for the aging process and the onset of various diseases, including cancer. Mitochondria, essential cellular organelles involved in energy metabolism and apoptosis, possess their own DNA, which are susceptible to mutations. However, studies on mtDNA mutations and mosaicism have been limited due to a variety of technical challenges.
Genomic scientists from KAIST have revealed the genetic mosaicism characterized by variations in mitochondrial DNA (mtDNA) across normal human cells. This study provides fundamental insights into understanding human aging and disease onset mechanisms.
The study, “Mitochondrial DNA mosaicism in normal human somatic cells,” was published in Nature Genetics on July 22. It was led by graduate student Jisong An under the supervision of Professor Young Seok Ju from the Graduate School of Medical Science and Engineering.
Researchers from Seoul National University College of Medicine, Yonsei University College of Medicine, Korea University College of Medicine, Washington University School of Medicine National Cancer Center, Seoul National University Hospital, Gangnam Severance Hospital and KAIST faculty startup company Inocras Inc. also participated in this study.
< Figure 1. a. Flow of experiment. b. Schematic diagram illustrating the origin and dynamics of mtDNA alterations across a lifetime. >
The study involved a bioinformatic analysis of whole-genome sequences from 2,096 single cells obtained from normal human colorectal epithelial tissue, fibroblasts, and blood collected from 31 individuals. The study highlights an average of three significant mtDNA differences between cells, with approximately 6% of these variations confirmed to be inherited as heteroplasmy from the mother.
Moreover, mutations significantly increased during tumorigenesis, with some mutations contributing to instability in mitochondrial RNA. Based on these findings, the study illustrates a computational model that comprehensively elucidates the evolution of mitochondria from embryonic development to aging and tumorigenesis.
This study systematically reveals the mechanisms behind mitochondrial DNA mosaicism in normal human cells, establishing a crucial foundation for understanding the impact of mtDNA on aging and disease onset.
Professor Ju remarked, “By systematically utilizing whole-genome big data, we can illuminate previously unknown phenomena in life sciences.” He emphasized the significance of the study, adding, “For the first time, we have established a method to systematically understand mitochondrial DNA changes occurring during human embryonic development, aging, and cancer development.”
This work was supported by the National Research Foundation of Korea and the Suh Kyungbae Foundation.
2024.07.24 View 5430 -
 Genome Sequencing Unveils Mutational Impacts of Radiation on Mammalian Cells
Recent release of the waste water from Japan's Fukushima nuclear disaster stirred apprehension regarding the health implications of radiation exposure. Classified as a Group 1 carcinogen, ionizing radiation has long been associated with various cancers and genetic disorders, as evidenced by survivors and descendants of atomic bombings and the Chernobyl disaster. Despite much smaller amount, we remain consistently exposed to low levels of radiation in everyday life and medical procedures.
Radiation, whether in the form of high-energy particles or electromagnetic waves, is conventionally known to break our cellular DNA, leading to cancer and genetic disorders. Yet, our understanding of the quantitative and qualitative mutational impacts of ionizing radiation has been incomplete.
On the 14th, Professor Young Seok Ju and his research team from KAIST, in collaboration with Dr. Tae Gen Son from the Dongnam Institute of Radiological and Medical Science, and Professors Kyung Su Kim and Ji Hyun Chang from Seoul National University, unveiled a breakthrough. Their study, led by joint first authors Drs. Jeonghwan Youk, Hyun Woo Kwon, Joonoh Lim, Eunji Kim and Tae-Woo Kim, titled "Quantitative and qualitative mutational impact of ionizing radiation on normal cells," was published in Cell Genomics.
Employing meticulous techniques, the research team comprehensively analyzed the whole-genome sequences of cells pre- and post-radiation exposure, pinpointing radiation-induced DNA mutations. Experiments involving cells from different organs of humans and mice exposed to varying radiation doses revealed mutation patterns correlating with exposure levels. (Figure 1)
Notably, exposure to 1 Gray (Gy) of radiation resulted in on average 14 mutations in every post-exposure cell. (Figure 2) Unlike other carcinogens, radiation-induced mutations primarily comprised short base deletions and a set of structural variations including inversions, translocations, and various complex genomic rearrangements. (Figure 3) Interestingly, experiments subjecting cells to low radiation dose rate over 100 days demonstrated that mutation quantities, under equivalent total radiation doses, mirrored those of high-dose exposure.
"Through this study, we have clearly elucidated the effects of radiation on cells at the molecular level," said Prof. Ju at KAIST. "Now we understand better how radiation changes the DNA of our cells," he added.
Dr. Son from the Dongnam Institute of Radiological and Medical Science stated, "Based on this study, we will continue to research the effects of very low and very high doses of radiation on the human body," and further remarked, "We will advance the development of safe and effective radiation therapy techniques."
Professors Kim and Chang from Seoul National University College of Medicine expressed their views, saying, "Through this study, we believe we now have a tool to accurately understand the impact of radiation on human DNA," and added, "We hope that many subsequent studies will emerge using the research methodologies employed in this study."
This research represents a significant leap forward in radiation studies, made possible through collaborative efforts and interdisciplinary approaches. This pioneering research engaged scholars from diverse backgrounds, spanning from the Genetic Engineering Research Institute at Seoul National University, the Cambridge Stem Cell Institute in the UK, the Institute for Molecular Biotechnology in Austria (IMBA), and the Genome Insight Inc. (a KAIST spin-off start-up). This study was supported by various institutions including the National Research Foundation of Korea, Dongnam Institute of Radiological and Medical Science (supported by Ministry of Science and ICT, the government of South Korea), the Suh Kyungbae Foundation, the Human Frontier Science Program (HFSP), and the Korea University Anam Hospital Korea Foundation for the Advancement of Science and Creativity, the Ministry of Science and ICT, and the National R&D Program.
2024.02.15 View 8855
Genome Sequencing Unveils Mutational Impacts of Radiation on Mammalian Cells
Recent release of the waste water from Japan's Fukushima nuclear disaster stirred apprehension regarding the health implications of radiation exposure. Classified as a Group 1 carcinogen, ionizing radiation has long been associated with various cancers and genetic disorders, as evidenced by survivors and descendants of atomic bombings and the Chernobyl disaster. Despite much smaller amount, we remain consistently exposed to low levels of radiation in everyday life and medical procedures.
Radiation, whether in the form of high-energy particles or electromagnetic waves, is conventionally known to break our cellular DNA, leading to cancer and genetic disorders. Yet, our understanding of the quantitative and qualitative mutational impacts of ionizing radiation has been incomplete.
On the 14th, Professor Young Seok Ju and his research team from KAIST, in collaboration with Dr. Tae Gen Son from the Dongnam Institute of Radiological and Medical Science, and Professors Kyung Su Kim and Ji Hyun Chang from Seoul National University, unveiled a breakthrough. Their study, led by joint first authors Drs. Jeonghwan Youk, Hyun Woo Kwon, Joonoh Lim, Eunji Kim and Tae-Woo Kim, titled "Quantitative and qualitative mutational impact of ionizing radiation on normal cells," was published in Cell Genomics.
Employing meticulous techniques, the research team comprehensively analyzed the whole-genome sequences of cells pre- and post-radiation exposure, pinpointing radiation-induced DNA mutations. Experiments involving cells from different organs of humans and mice exposed to varying radiation doses revealed mutation patterns correlating with exposure levels. (Figure 1)
Notably, exposure to 1 Gray (Gy) of radiation resulted in on average 14 mutations in every post-exposure cell. (Figure 2) Unlike other carcinogens, radiation-induced mutations primarily comprised short base deletions and a set of structural variations including inversions, translocations, and various complex genomic rearrangements. (Figure 3) Interestingly, experiments subjecting cells to low radiation dose rate over 100 days demonstrated that mutation quantities, under equivalent total radiation doses, mirrored those of high-dose exposure.
"Through this study, we have clearly elucidated the effects of radiation on cells at the molecular level," said Prof. Ju at KAIST. "Now we understand better how radiation changes the DNA of our cells," he added.
Dr. Son from the Dongnam Institute of Radiological and Medical Science stated, "Based on this study, we will continue to research the effects of very low and very high doses of radiation on the human body," and further remarked, "We will advance the development of safe and effective radiation therapy techniques."
Professors Kim and Chang from Seoul National University College of Medicine expressed their views, saying, "Through this study, we believe we now have a tool to accurately understand the impact of radiation on human DNA," and added, "We hope that many subsequent studies will emerge using the research methodologies employed in this study."
This research represents a significant leap forward in radiation studies, made possible through collaborative efforts and interdisciplinary approaches. This pioneering research engaged scholars from diverse backgrounds, spanning from the Genetic Engineering Research Institute at Seoul National University, the Cambridge Stem Cell Institute in the UK, the Institute for Molecular Biotechnology in Austria (IMBA), and the Genome Insight Inc. (a KAIST spin-off start-up). This study was supported by various institutions including the National Research Foundation of Korea, Dongnam Institute of Radiological and Medical Science (supported by Ministry of Science and ICT, the government of South Korea), the Suh Kyungbae Foundation, the Human Frontier Science Program (HFSP), and the Korea University Anam Hospital Korea Foundation for the Advancement of Science and Creativity, the Ministry of Science and ICT, and the National R&D Program.
2024.02.15 View 8855 -
 'Jumping Genes' Found to Alter Human Colon Genomes, Offering Insights into Aging and Tumorigenesis
The Korea Advanced Institute of Science and Technology (KAIST) and their collaborators have conducted a groundbreaking study targeting 'jumping genes' in the entire genomes of the human large intestine. Published in Nature on May 18 2023, the research unveils the surprising activity of 'Long interspersed nuclear element-1 (L1),' a type of jumping gene previously thought to be mostly dormant in human genomes. The study shows that L1 genes can become activated and disrupt genomic functions throughout an individual's lifetime, particularly in the colorectal epithelium.
(Paper Title: Widespread somatic L1 retrotransposition in normal colorectal epithelium, https://www.nature.com/articles/s41586-023-06046-z)
With approximately 500,000 L1 jumping genes, accounting for 17% of the human genome, they have long been recognized for their contribution to the evolution of the human species by introducing 'disruptive innovation' to genome sequences. Until now, it was believed that most L1 elements had lost their ability to jump in normal tissues of modern humans. However, this study reveals that some L1 jumping genes can be widely activated in normal cells, leading to the accumulation of genomic mutations over an individual's lifetime. The rate of L1 jumping and resulting genomic changes vary among different cell types, with a notable concentration observed in aged colon epithelial cells. The study illustrates that every colonic epithelial cell experiences an L1 jumping event by the age of 40 on average.
The research, led by co-first authors Chang Hyun Nam (a graduate student at KAIST) and Dr. Jeonghwan Youk (former graduate student at KAIST and assistant clinical professor at Seoul National University Hospital), involved the analysis of whole-genome sequences from 899 single cells obtained from skin (fibroblasts), blood, and colon epithelial tissues collected from 28 individuals. The study uncovers the activation of L1 jumping genes in normal cells, resulting in the gradual accumulation of genomic mutations over time. Additionally, the team explored epigenomic (DNA methylation) sequences to understand the mechanism behind L1 jumping gene activation. They found that cells with activated L1 jumping genes exhibit epigenetic instability, suggesting the critical role of epigenetic changes in regulating L1 jumping gene activity. Most of these epigenomic instabilities were found to arise during the early stages of embryogenesis. The study provides valuable insights into the aging process and the development of diseases in human colorectal tissues.
"This study illustrates that genomic damage in normal cells is acquired not only through exposure to carcinogens but also through the activity of endogenous components whose impact was previously unclear. Genomes of apparently healthy aged cells, particularly in the colorectal epithelium, become mosaic due to the activity of L1 jumping genes," said Prof. Young Seok Ju at KAIST.
"We emphasize the essential and ongoing collaboration among researchers in clinical medicine and basic medical sciences," said Prof. Min Jung Kim of the Department of Surgery at Seoul National University Hospital. "This case highlights the critical role of systematically collected human tissues from clinical settings in unraveling the complex process of disease development in humans."
"I am delighted that the research team's advancements in single-cell genome technology have come to fruition. We will persistently strive to lead in single-cell genome technology," said Prof. Hyun Woo Kwon of the Department of Nuclear Medicine at Korea University School of Medicine.
The research team received support from the Research Leader Program and the Young Researcher Program of the National Research Foundation of Korea, a grant from the MD-PhD/Medical Scientist Training Program through the Korea Health Industry Development Institute, and the Suh Kyungbae Foundation.
< Figure 1. Experimental design of the study >
< Figure 2. Schematic diagram illustrating factors influencing the soL1R landscape. >
Genetic composition of rc-L1s is inherited from the parents. The methylation landscape of rc-L1 promoters is predominantly determined by global DNA demethylation, followed by remethylation processes in the developmental stages. Then, when an rc-L1 is promoter demethylated in a specific cell lineage, the source expresses L1 transcripts thus making possible the induction of soL1Rs.
2023.05.22 View 9389
'Jumping Genes' Found to Alter Human Colon Genomes, Offering Insights into Aging and Tumorigenesis
The Korea Advanced Institute of Science and Technology (KAIST) and their collaborators have conducted a groundbreaking study targeting 'jumping genes' in the entire genomes of the human large intestine. Published in Nature on May 18 2023, the research unveils the surprising activity of 'Long interspersed nuclear element-1 (L1),' a type of jumping gene previously thought to be mostly dormant in human genomes. The study shows that L1 genes can become activated and disrupt genomic functions throughout an individual's lifetime, particularly in the colorectal epithelium.
(Paper Title: Widespread somatic L1 retrotransposition in normal colorectal epithelium, https://www.nature.com/articles/s41586-023-06046-z)
With approximately 500,000 L1 jumping genes, accounting for 17% of the human genome, they have long been recognized for their contribution to the evolution of the human species by introducing 'disruptive innovation' to genome sequences. Until now, it was believed that most L1 elements had lost their ability to jump in normal tissues of modern humans. However, this study reveals that some L1 jumping genes can be widely activated in normal cells, leading to the accumulation of genomic mutations over an individual's lifetime. The rate of L1 jumping and resulting genomic changes vary among different cell types, with a notable concentration observed in aged colon epithelial cells. The study illustrates that every colonic epithelial cell experiences an L1 jumping event by the age of 40 on average.
The research, led by co-first authors Chang Hyun Nam (a graduate student at KAIST) and Dr. Jeonghwan Youk (former graduate student at KAIST and assistant clinical professor at Seoul National University Hospital), involved the analysis of whole-genome sequences from 899 single cells obtained from skin (fibroblasts), blood, and colon epithelial tissues collected from 28 individuals. The study uncovers the activation of L1 jumping genes in normal cells, resulting in the gradual accumulation of genomic mutations over time. Additionally, the team explored epigenomic (DNA methylation) sequences to understand the mechanism behind L1 jumping gene activation. They found that cells with activated L1 jumping genes exhibit epigenetic instability, suggesting the critical role of epigenetic changes in regulating L1 jumping gene activity. Most of these epigenomic instabilities were found to arise during the early stages of embryogenesis. The study provides valuable insights into the aging process and the development of diseases in human colorectal tissues.
"This study illustrates that genomic damage in normal cells is acquired not only through exposure to carcinogens but also through the activity of endogenous components whose impact was previously unclear. Genomes of apparently healthy aged cells, particularly in the colorectal epithelium, become mosaic due to the activity of L1 jumping genes," said Prof. Young Seok Ju at KAIST.
"We emphasize the essential and ongoing collaboration among researchers in clinical medicine and basic medical sciences," said Prof. Min Jung Kim of the Department of Surgery at Seoul National University Hospital. "This case highlights the critical role of systematically collected human tissues from clinical settings in unraveling the complex process of disease development in humans."
"I am delighted that the research team's advancements in single-cell genome technology have come to fruition. We will persistently strive to lead in single-cell genome technology," said Prof. Hyun Woo Kwon of the Department of Nuclear Medicine at Korea University School of Medicine.
The research team received support from the Research Leader Program and the Young Researcher Program of the National Research Foundation of Korea, a grant from the MD-PhD/Medical Scientist Training Program through the Korea Health Industry Development Institute, and the Suh Kyungbae Foundation.
< Figure 1. Experimental design of the study >
< Figure 2. Schematic diagram illustrating factors influencing the soL1R landscape. >
Genetic composition of rc-L1s is inherited from the parents. The methylation landscape of rc-L1 promoters is predominantly determined by global DNA demethylation, followed by remethylation processes in the developmental stages. Then, when an rc-L1 is promoter demethylated in a specific cell lineage, the source expresses L1 transcripts thus making possible the induction of soL1Rs.
2023.05.22 View 9389 -
 Genomic Data Reveals New Insights into Human Embryonic Development
KAIST researchers have used whole-genome sequencing to track the development from a single fertilized-egg to a human body
Genomic scientists at KAIST have revealed new insights into the process of human embryonic development using large-scale, whole-genome sequencing of cells and tissues from adult humans. The study, published in Nature on Aug.25, is the first to analyse somatic mutations in normal tissue across multiple organs within and between humans.
An adult human body comprises trillions of cells of more than 200 types. How a human develops from a single fertilized egg to a fully grown adult is a fundamental question in biomedical science. Due to the ethical challenges of performing studies on human embryos, however, the details of this process remain largely unknown.
To overcome these issues, the research team took a different approach. They analysed genetic mutations in cells taken from adult human post-mortem tissue. Specifically, they identified mutations that occur spontaneously in early developmental cell divisions. These mutations, also called genomic scars, act like unique genetic fingerprints that can be used to trace the embryonic development process.
The study, which looked at 334 single-cell colonies and 379 tissue samples from seven recently deceased human body donors, is the largest single-cell, whole-genome analysis carried out to date. The researchers examined the genomic scars of each individual in order to reconstruct their early embryonic cellular dynamics.
The result revealed several key characteristics of the human embryonic development process. Firstly, mutation rates are higher in the first cell division, but then decrease to approximately one mutation per cell during later cell division. Secondly, early cells contributed unequally to the development of the embryo in all informative donors, for example, at the two-cell stage, one of the cells always left more progeny cells than the other. The ratio of this was different from person to person, implying that the process varies between individuals and is not fully deterministic.
The researchers were also able to deduce the timing of when cells begin to differentiate into individual organ-specific cells. They found that within three days of fertilization, embryonic cells began to be distributed asymmetrically into tissues for the left and right sides of the body, followed by differentiation into three germ layers, and then differentiation into specific tissues and organs.
“It is an impressive scientific achievement that, within 20 years of the completion of human genome project, genomic technology has advanced to the extent that we are now able to accurately identify mutations in a single-cell genome,” said Professor Young Seok Ju from the Graduate School of Medical Science and Engineering at KAIST. “This technology will enable us to track human embryogenesis at even higher resolutions in the future.”
The techniques used in this study could be used to improve our understanding of rare diseases caused by abnormalities in embryonic development, and to design new precision diagnostics and treatments for patients.
The research was completed in collaboration with Kyungpook National University Hospital, the Korea Institute of Science and Technology Information, Catholic University of Korea School of Medicine, Genome Insights Inc, and Immune Square Inc. This work was supported by the Suh Kyungbae Foundation, the Ministry of Health and Welfare of Korea, the National Research Foundastion of Korea.
-PublicationSeongyeol Park, Nanda Mali, Ryul Kim et al. ‘Clonal dynamics in early human embryogenesis inferred from somatic mutation’ Nature Online ahead of print, Aug. 25, 2021 (https://doi.org/10.1038/s41586-021-03786-8)
-ProfileProfessor Young Seok JuLab of Cancer Genomics (https://www.julab.kaist.ac.kr/)Graduate School of Medical Science and EngineeringKAIST
2021.08.31 View 10174
Genomic Data Reveals New Insights into Human Embryonic Development
KAIST researchers have used whole-genome sequencing to track the development from a single fertilized-egg to a human body
Genomic scientists at KAIST have revealed new insights into the process of human embryonic development using large-scale, whole-genome sequencing of cells and tissues from adult humans. The study, published in Nature on Aug.25, is the first to analyse somatic mutations in normal tissue across multiple organs within and between humans.
An adult human body comprises trillions of cells of more than 200 types. How a human develops from a single fertilized egg to a fully grown adult is a fundamental question in biomedical science. Due to the ethical challenges of performing studies on human embryos, however, the details of this process remain largely unknown.
To overcome these issues, the research team took a different approach. They analysed genetic mutations in cells taken from adult human post-mortem tissue. Specifically, they identified mutations that occur spontaneously in early developmental cell divisions. These mutations, also called genomic scars, act like unique genetic fingerprints that can be used to trace the embryonic development process.
The study, which looked at 334 single-cell colonies and 379 tissue samples from seven recently deceased human body donors, is the largest single-cell, whole-genome analysis carried out to date. The researchers examined the genomic scars of each individual in order to reconstruct their early embryonic cellular dynamics.
The result revealed several key characteristics of the human embryonic development process. Firstly, mutation rates are higher in the first cell division, but then decrease to approximately one mutation per cell during later cell division. Secondly, early cells contributed unequally to the development of the embryo in all informative donors, for example, at the two-cell stage, one of the cells always left more progeny cells than the other. The ratio of this was different from person to person, implying that the process varies between individuals and is not fully deterministic.
The researchers were also able to deduce the timing of when cells begin to differentiate into individual organ-specific cells. They found that within three days of fertilization, embryonic cells began to be distributed asymmetrically into tissues for the left and right sides of the body, followed by differentiation into three germ layers, and then differentiation into specific tissues and organs.
“It is an impressive scientific achievement that, within 20 years of the completion of human genome project, genomic technology has advanced to the extent that we are now able to accurately identify mutations in a single-cell genome,” said Professor Young Seok Ju from the Graduate School of Medical Science and Engineering at KAIST. “This technology will enable us to track human embryogenesis at even higher resolutions in the future.”
The techniques used in this study could be used to improve our understanding of rare diseases caused by abnormalities in embryonic development, and to design new precision diagnostics and treatments for patients.
The research was completed in collaboration with Kyungpook National University Hospital, the Korea Institute of Science and Technology Information, Catholic University of Korea School of Medicine, Genome Insights Inc, and Immune Square Inc. This work was supported by the Suh Kyungbae Foundation, the Ministry of Health and Welfare of Korea, the National Research Foundastion of Korea.
-PublicationSeongyeol Park, Nanda Mali, Ryul Kim et al. ‘Clonal dynamics in early human embryogenesis inferred from somatic mutation’ Nature Online ahead of print, Aug. 25, 2021 (https://doi.org/10.1038/s41586-021-03786-8)
-ProfileProfessor Young Seok JuLab of Cancer Genomics (https://www.julab.kaist.ac.kr/)Graduate School of Medical Science and EngineeringKAIST
2021.08.31 View 10174 -
 'Mini-Lungs' Reveal Early Stages of SARS-CoV-2 Infection
Researchers in Korea and the UK have successfully grown miniature models of critical lung structures called alveoli, and used them to study how the coronavirus that causes COVID-19 infects the lungs.
To date, there have been more than 40 million cases of COVID-19 and almost 1.13 million deaths worldwide. The main target tissues of SARS-CoV-2, the virus that causes COVID-19, especially in patients that develop pneumonia, appear to be alveoli – tiny air sacs in the lungs that take up the oxygen we breathe and exchange it with carbon dioxide to exhale.
To better understand how SARS-CoV-2 infects the lungs and causes disease, a team of Professor Young Seok Ju from the Graduate School of Medical Science and Engineering at KAIST in collaboration with the Wellcome-MRC Cambridge Stem Cell Institute at the University of Cambridge turned to organoids – ‘mini-organs’ grown in three dimensions to mimic the behaviour of tissue and organs.
The team used tissue donated to tissue banks at the Royal Papworth Hospital NHS Foundation Trust and Addenbrooke’s Hospital, Cambridge University NHS Foundations Trust, UK, and Seoul National University Hospital to extract a type of lung cell known as human lung alveolar type 2 cells. By reprogramming these cells back to their earlier ‘stem cell’ stage, they were able to grow self-organizing alveolar-like 3D structures that mimic the behaviour of key lung tissue.
“The research community now has a powerful new platform to study precisely how the virus infects the lungs, as well as explore possible treatments,” said Professor Ju, co-senior author of the research.
Dr. Joo-Hyeon Lee, another co-senior author at the Wellcome-MRC Cambridge Stem Cell Institute, said: “We still know surprisingly little about how SARS-CoV-2 infects the lungs and causes disease. Our approach has allowed us to grow 3D models of key lung tissue – in a sense, ‘mini-lungs’ – in the lab and study what happens when they become infected.”
The team infected the organoids with a strain of SARS-CoV-2 taken from a patient in Korea who was diagnosed with COVID-19 on January 26 after traveling to Wuhan, China. Using a combination of fluorescence imaging and single cell genetic analysis, they were able to study how the cells responded to the virus.
When the 3D models were exposed to SARS-CoV-2, the virus began to replicate rapidly, reaching full cellular infection just six hours after infection. Replication enables the virus to spread throughout the body, infecting other cells and tissue.
Around the same time, the cells began to produce interferons – proteins that act as warning signals to neighbouring cells, telling them to activate their antiviral defences. After 48 hours, the interferons triggered the innate immune response – its first line of defence – and the cells started fighting back against infection.
Sixty hours after infection, a subset of alveolar cells began to disintegrate, leading to cell death and damage to the lung tissue.
Although the researchers observed changes to the lung cells within three days of infection, clinical symptoms of COVID-19 rarely occur so quickly and can sometimes take more than ten days after exposure to appear. The team say there are several possible reasons for this. It may take several days from the virus first infiltrating the upper respiratory tract to it reaching the alveoli. It may also require a substantial proportion of alveolar cells to be infected or for further interactions with immune cells resulting in inflammation before a patient displays symptoms.
“Based on our model we can tackle many unanswered key questions, such as understanding genetic susceptibility to SARS-CoV-2, assessing relative infectivity of viral mutants, and revealing the damage processes of the virus in human alveolar cells,” said Professor Ju. “Most importantly, it provides the opportunity to develop and screen potential therapeutic agents against SARS-CoV-2 infection.”
“We hope to use our technique to grow these 3D models from cells of patients who are particularly vulnerable to infection, such as the elderly or people with diseased lungs, and find out what happens to their tissue,” added Dr. Lee.
The research was a collaboration involving scientists from KAIST, the University of Cambridge, Korea National Institute of Health, Institute for Basic Science (IBS), Seoul National University Hospital and Genome Insight in Korea.
- ProfileProfessor Young Seok JuLaboratory of Cancer Genomics https://julab.kaist.ac.kr the Graduate School of Medical Science and EngineeringKAIST
2020.10.26 View 12875
'Mini-Lungs' Reveal Early Stages of SARS-CoV-2 Infection
Researchers in Korea and the UK have successfully grown miniature models of critical lung structures called alveoli, and used them to study how the coronavirus that causes COVID-19 infects the lungs.
To date, there have been more than 40 million cases of COVID-19 and almost 1.13 million deaths worldwide. The main target tissues of SARS-CoV-2, the virus that causes COVID-19, especially in patients that develop pneumonia, appear to be alveoli – tiny air sacs in the lungs that take up the oxygen we breathe and exchange it with carbon dioxide to exhale.
To better understand how SARS-CoV-2 infects the lungs and causes disease, a team of Professor Young Seok Ju from the Graduate School of Medical Science and Engineering at KAIST in collaboration with the Wellcome-MRC Cambridge Stem Cell Institute at the University of Cambridge turned to organoids – ‘mini-organs’ grown in three dimensions to mimic the behaviour of tissue and organs.
The team used tissue donated to tissue banks at the Royal Papworth Hospital NHS Foundation Trust and Addenbrooke’s Hospital, Cambridge University NHS Foundations Trust, UK, and Seoul National University Hospital to extract a type of lung cell known as human lung alveolar type 2 cells. By reprogramming these cells back to their earlier ‘stem cell’ stage, they were able to grow self-organizing alveolar-like 3D structures that mimic the behaviour of key lung tissue.
“The research community now has a powerful new platform to study precisely how the virus infects the lungs, as well as explore possible treatments,” said Professor Ju, co-senior author of the research.
Dr. Joo-Hyeon Lee, another co-senior author at the Wellcome-MRC Cambridge Stem Cell Institute, said: “We still know surprisingly little about how SARS-CoV-2 infects the lungs and causes disease. Our approach has allowed us to grow 3D models of key lung tissue – in a sense, ‘mini-lungs’ – in the lab and study what happens when they become infected.”
The team infected the organoids with a strain of SARS-CoV-2 taken from a patient in Korea who was diagnosed with COVID-19 on January 26 after traveling to Wuhan, China. Using a combination of fluorescence imaging and single cell genetic analysis, they were able to study how the cells responded to the virus.
When the 3D models were exposed to SARS-CoV-2, the virus began to replicate rapidly, reaching full cellular infection just six hours after infection. Replication enables the virus to spread throughout the body, infecting other cells and tissue.
Around the same time, the cells began to produce interferons – proteins that act as warning signals to neighbouring cells, telling them to activate their antiviral defences. After 48 hours, the interferons triggered the innate immune response – its first line of defence – and the cells started fighting back against infection.
Sixty hours after infection, a subset of alveolar cells began to disintegrate, leading to cell death and damage to the lung tissue.
Although the researchers observed changes to the lung cells within three days of infection, clinical symptoms of COVID-19 rarely occur so quickly and can sometimes take more than ten days after exposure to appear. The team say there are several possible reasons for this. It may take several days from the virus first infiltrating the upper respiratory tract to it reaching the alveoli. It may also require a substantial proportion of alveolar cells to be infected or for further interactions with immune cells resulting in inflammation before a patient displays symptoms.
“Based on our model we can tackle many unanswered key questions, such as understanding genetic susceptibility to SARS-CoV-2, assessing relative infectivity of viral mutants, and revealing the damage processes of the virus in human alveolar cells,” said Professor Ju. “Most importantly, it provides the opportunity to develop and screen potential therapeutic agents against SARS-CoV-2 infection.”
“We hope to use our technique to grow these 3D models from cells of patients who are particularly vulnerable to infection, such as the elderly or people with diseased lungs, and find out what happens to their tissue,” added Dr. Lee.
The research was a collaboration involving scientists from KAIST, the University of Cambridge, Korea National Institute of Health, Institute for Basic Science (IBS), Seoul National University Hospital and Genome Insight in Korea.
- ProfileProfessor Young Seok JuLaboratory of Cancer Genomics https://julab.kaist.ac.kr the Graduate School of Medical Science and EngineeringKAIST
2020.10.26 View 12875 -
 Professor Ju, to Receive Grants from HFSP
(Professor Young Seok Ju)
Professor Young Seok Ju from the Graduate School of Medical Science and Engineering was selected as a young investigator to receive research funds from the Human Frontiers Science Program.
The Human Frontiers Science Program (HFSP) was founded in 1989 with members of the G7 and European Union to stimulate innovative research in the field of life sciences.
Professor Ju placed third out of the eight teams that were selected from 158 applicants representing 60 countries. He is now the fourth Korean to receive a research grant as a young investigator. Professor Jae Kyoung Kim from the Department of Mathematical Sciences also received this prize last year, hence KAIST has produced grant recipients for two consecutive years.
Professor Ju is a medical doctor specializing in cancer genomics and computer biology. He has been studying somatic mutations and their functional consequences in human cancer in a bioinformatics way. He has published papers in international journals including Nature, Science, Genome Research, and Journal of Clinical Oncology.
With a title ‘Tracing AID/APOBEC- and MSI-mediated hyper-mutagenesis in the clonal evolution of gastric cancer,’ Professor Ju will receive 1.05 million dollars for three years along with Professor Bon-Kyoung Koo from the Institute of Molecular Biotechnology at Austrian Academy of Sciences, and Sinppert Hugo from University Medical Center Utrecht.
Professor Ju said, “As a young investigator, it is my great honor to receive this research fund from this organization. Through this internationally collaborative research, I will carry out groundbreaking research to understand the pathophysiology of cancers at a molecular level.”
2018.04.24 View 9949
Professor Ju, to Receive Grants from HFSP
(Professor Young Seok Ju)
Professor Young Seok Ju from the Graduate School of Medical Science and Engineering was selected as a young investigator to receive research funds from the Human Frontiers Science Program.
The Human Frontiers Science Program (HFSP) was founded in 1989 with members of the G7 and European Union to stimulate innovative research in the field of life sciences.
Professor Ju placed third out of the eight teams that were selected from 158 applicants representing 60 countries. He is now the fourth Korean to receive a research grant as a young investigator. Professor Jae Kyoung Kim from the Department of Mathematical Sciences also received this prize last year, hence KAIST has produced grant recipients for two consecutive years.
Professor Ju is a medical doctor specializing in cancer genomics and computer biology. He has been studying somatic mutations and their functional consequences in human cancer in a bioinformatics way. He has published papers in international journals including Nature, Science, Genome Research, and Journal of Clinical Oncology.
With a title ‘Tracing AID/APOBEC- and MSI-mediated hyper-mutagenesis in the clonal evolution of gastric cancer,’ Professor Ju will receive 1.05 million dollars for three years along with Professor Bon-Kyoung Koo from the Institute of Molecular Biotechnology at Austrian Academy of Sciences, and Sinppert Hugo from University Medical Center Utrecht.
Professor Ju said, “As a young investigator, it is my great honor to receive this research fund from this organization. Through this internationally collaborative research, I will carry out groundbreaking research to understand the pathophysiology of cancers at a molecular level.”
2018.04.24 View 9949 -
 First Mutations in Human Life Discovered
The earliest mutations of human life have been observed by research team led by the Wellcome Trust Sanger Institute and their collaborators. Analyzing genomes from adult cells, the scientists could look back in time to reveal how each embryo developed.
Research team of the Sanger Institute including Professor Young Seok Ju of the Graduate School of Medical Science and Engineering at KAIST published an article of “Somatic Mutations Reveal Asymmetric Cellular Dynamics in the Early Human Embryo” in Nature on March 22.
The study shows that from the two-cell stage of the human embryo, one of these cells becomes more dominant than the other and leads to a higher proportion of the adult body.
A longstanding question for researchers has been what happens in the very early human development as this has proved impossible to study directly. Now, researchers have analyzed the whole genome sequences of blood samples (collected from 279 individuals with breast cancer) and discovered 163 mutations that occurred very early in the embryonic development of those people.
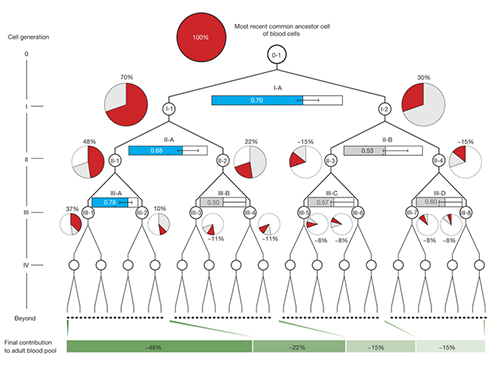
Once identified, the researchers used mutations from the first, second and third divisions of the fertilized egg to calculate which proportion of adult cells resulted from each of the first two cells in the embryo. They found that these first two cells contribute differently to the whole body. One cell gives rise to about 70 percent of the adult body tissues, whereas the other cell has a more minor contribution, leading to about 30percent of the tissues. This skewed contribution continues for some cells in the second and third generation too.
Originally pinpointed in normal blood cells from cancer patients, the researchers then looked for these mutations in cancer samples that had been surgically removed from the patients during treatment. Unlike normal tissues composed of multiple somatic cell clones, a cancer develops from one mutant cell. Therefore, each proposed embryonic mutation should either be present in all of the cancer cells in a tumor, or none of them. This proved to be the case, and by using these cancer samples, the researchers were able to validate that the mutations had originated during early development.
Dr. Young Seok Ju, first author from the Wellcome Trust Sanger Institute and KAIST, said: "This is the first time that anyone has seen where mutations arise in the very early human development. It is like finding a needle in a haystack. There are just a handful of these mutations, compared with millions of inherited genetic variations, and finding them allowed us to track what happened during embryogenesis."
Dr. Inigo Martincorena, from the Sanger Institute, said: "Having identified the mutations, we were able to use statistical analysis to better understand cell dynamics during embryo development. We determined the relative contribution of the first embryonic cells to the adult blood cell pool and found one dominant cell - that led to 70 percent of the blood cells - and one minor cell. We also sequenced normal lymph and breast cells, and the results suggested that the dominant cell also contributes to these other tissues at a similar level. This opens an unprecedented window into the earliest stages of human development."
During this study, the researchers were also able to measure the rate of mutation in early human development for the first time, up to three generations of cell division. Previous researchers had estimated one mutation per cell division, but this study measured three mutations for each cell doubling, in every daughter cell.
Mutations during the development of the embryo occur by two processes - known as mutational signatures 1 and 5. These mutations are fairly randomly distributed through the genome, and the vast majority of them will not affect the developing embryo. However, a mutation that occurs in an important gene can lead to disease such as developmental disorders.
Professor Sir Mike Stratton, lead author on the paper and Director of the Sanger Institute, said: "This is a significant step forward in widening the range of biological insights that can be extracted using genome sequences and mutations. Essentially, the mutations are archaeological traces of embryonic development left in our adult tissues, so if we can find and interpret them, we can understand human embryology better. This is just one early insight into human development, with hopefully many more to come in the future."
(Figure 1. Detection of somatic mutations acquired in early human embryogenesis )
(Figure 2. Unequal contributions of early embryonic cells to adult somatic tissues )
2017.03.23 View 8365
First Mutations in Human Life Discovered
The earliest mutations of human life have been observed by research team led by the Wellcome Trust Sanger Institute and their collaborators. Analyzing genomes from adult cells, the scientists could look back in time to reveal how each embryo developed.
Research team of the Sanger Institute including Professor Young Seok Ju of the Graduate School of Medical Science and Engineering at KAIST published an article of “Somatic Mutations Reveal Asymmetric Cellular Dynamics in the Early Human Embryo” in Nature on March 22.
The study shows that from the two-cell stage of the human embryo, one of these cells becomes more dominant than the other and leads to a higher proportion of the adult body.
A longstanding question for researchers has been what happens in the very early human development as this has proved impossible to study directly. Now, researchers have analyzed the whole genome sequences of blood samples (collected from 279 individuals with breast cancer) and discovered 163 mutations that occurred very early in the embryonic development of those people.
Once identified, the researchers used mutations from the first, second and third divisions of the fertilized egg to calculate which proportion of adult cells resulted from each of the first two cells in the embryo. They found that these first two cells contribute differently to the whole body. One cell gives rise to about 70 percent of the adult body tissues, whereas the other cell has a more minor contribution, leading to about 30percent of the tissues. This skewed contribution continues for some cells in the second and third generation too.
Originally pinpointed in normal blood cells from cancer patients, the researchers then looked for these mutations in cancer samples that had been surgically removed from the patients during treatment. Unlike normal tissues composed of multiple somatic cell clones, a cancer develops from one mutant cell. Therefore, each proposed embryonic mutation should either be present in all of the cancer cells in a tumor, or none of them. This proved to be the case, and by using these cancer samples, the researchers were able to validate that the mutations had originated during early development.
Dr. Young Seok Ju, first author from the Wellcome Trust Sanger Institute and KAIST, said: "This is the first time that anyone has seen where mutations arise in the very early human development. It is like finding a needle in a haystack. There are just a handful of these mutations, compared with millions of inherited genetic variations, and finding them allowed us to track what happened during embryogenesis."
Dr. Inigo Martincorena, from the Sanger Institute, said: "Having identified the mutations, we were able to use statistical analysis to better understand cell dynamics during embryo development. We determined the relative contribution of the first embryonic cells to the adult blood cell pool and found one dominant cell - that led to 70 percent of the blood cells - and one minor cell. We also sequenced normal lymph and breast cells, and the results suggested that the dominant cell also contributes to these other tissues at a similar level. This opens an unprecedented window into the earliest stages of human development."
During this study, the researchers were also able to measure the rate of mutation in early human development for the first time, up to three generations of cell division. Previous researchers had estimated one mutation per cell division, but this study measured three mutations for each cell doubling, in every daughter cell.
Mutations during the development of the embryo occur by two processes - known as mutational signatures 1 and 5. These mutations are fairly randomly distributed through the genome, and the vast majority of them will not affect the developing embryo. However, a mutation that occurs in an important gene can lead to disease such as developmental disorders.
Professor Sir Mike Stratton, lead author on the paper and Director of the Sanger Institute, said: "This is a significant step forward in widening the range of biological insights that can be extracted using genome sequences and mutations. Essentially, the mutations are archaeological traces of embryonic development left in our adult tissues, so if we can find and interpret them, we can understand human embryology better. This is just one early insight into human development, with hopefully many more to come in the future."
(Figure 1. Detection of somatic mutations acquired in early human embryogenesis )
(Figure 2. Unequal contributions of early embryonic cells to adult somatic tissues )
2017.03.23 View 8365