E.coli
-
 'Fingerprint' Machine Learning Technique Identifies Different Bacteria in Seconds
A synergistic combination of surface-enhanced Raman spectroscopy and deep learning serves as an effective platform for separation-free detection of bacteria in arbitrary media
Bacterial identification can take hours and often longer, precious time when diagnosing infections and selecting appropriate treatments. There may be a quicker, more accurate process according to researchers at KAIST. By teaching a deep learning algorithm to identify the “fingerprint” spectra of the molecular components of various bacteria, the researchers could classify various bacteria in different media with accuracies of up to 98%.
Their results were made available online on Jan. 18 in Biosensors and Bioelectronics, ahead of publication in the journal’s April issue.
Bacteria-induced illnesses, those caused by direct bacterial infection or by exposure to bacterial toxins, can induce painful symptoms and even lead to death, so the rapid detection of bacteria is crucial to prevent the intake of contaminated foods and to diagnose infections from clinical samples, such as urine. “By using surface-enhanced Raman spectroscopy (SERS) analysis boosted with a newly proposed deep learning model, we demonstrated a markedly simple, fast, and effective route to classify the signals of two common bacteria and their resident media without any separation procedures,” said Professor Sungho Jo from the School of Computing.
Raman spectroscopy sends light through a sample to see how it scatters. The results reveal structural information about the sample — the spectral fingerprint — allowing researchers to identify its molecules. The surface-enhanced version places sample cells on noble metal nanostructures that help amplify the sample’s signals.
However, it is challenging to obtain consistent and clear spectra of bacteria due to numerous overlapping peak sources, such as proteins in cell walls. “Moreover, strong signals of surrounding media are also enhanced to overwhelm target signals, requiring time-consuming and tedious bacterial separation steps,” said Professor Yeon Sik Jung from the Department of Materials Science and Engineering.
To parse through the noisy signals, the researchers implemented an artificial intelligence method called deep learning that can hierarchically extract certain features of the spectral information to classify data. They specifically designed their model, named the dual-branch wide-kernel network (DualWKNet), to efficiently learn the correlation between spectral features. Such an ability is critical for analyzing one-dimensional spectral data, according to Professor Jo.
“Despite having interfering signals or noise from the media, which make the general shapes of different bacterial spectra and their residing media signals look similar, high classification accuracies of bacterial types and their media were achieved,” Professor Jo said, explaining that DualWKNet allowed the team to identify key peaks in each class that were almost indiscernible in individual spectra, enhancing the classification accuracies. “Ultimately, with the use of DualWKNet replacing the bacteria and media separation steps, our method dramatically reduces analysis time.”
The researchers plan to use their platform to study more bacteria and media types, using the information to build a training data library of various bacterial types in additional media to reduce the collection and detection times for new samples.
“We developed a meaningful universal platform for rapid bacterial detection with the collaboration between SERS and deep learning,” Professor Jo said. “We hope to extend the use of our deep learning-based SERS analysis platform to detect numerous types of bacteria in additional media that are important for food or clinical analysis, such as blood.”
The National R&D Program, through a National Research Foundation of Korea grant funded by the Ministry of Science and ICT, supported this research.
-PublicationEojin Rho, Minjoon Kim, Seunghee H. Cho, Bongjae Choi, Hyungjoon Park, Hanhwi Jang, Yeon Sik Jung, Sungho Jo, “Separation-free bacterial identification in arbitrary media via deepneural network-based SERS analysis,” Biosensors and Bioelectronics online January 18, 2022 (doi.org/10.1016/j.bios.2022.113991)
-ProfileProfessor Yeon Sik JungDepartment of Materials Science and EngineeringKAIST
Professor Sungho JoSchool of ComputingKAIST
2022.03.04 View 23132
'Fingerprint' Machine Learning Technique Identifies Different Bacteria in Seconds
A synergistic combination of surface-enhanced Raman spectroscopy and deep learning serves as an effective platform for separation-free detection of bacteria in arbitrary media
Bacterial identification can take hours and often longer, precious time when diagnosing infections and selecting appropriate treatments. There may be a quicker, more accurate process according to researchers at KAIST. By teaching a deep learning algorithm to identify the “fingerprint” spectra of the molecular components of various bacteria, the researchers could classify various bacteria in different media with accuracies of up to 98%.
Their results were made available online on Jan. 18 in Biosensors and Bioelectronics, ahead of publication in the journal’s April issue.
Bacteria-induced illnesses, those caused by direct bacterial infection or by exposure to bacterial toxins, can induce painful symptoms and even lead to death, so the rapid detection of bacteria is crucial to prevent the intake of contaminated foods and to diagnose infections from clinical samples, such as urine. “By using surface-enhanced Raman spectroscopy (SERS) analysis boosted with a newly proposed deep learning model, we demonstrated a markedly simple, fast, and effective route to classify the signals of two common bacteria and their resident media without any separation procedures,” said Professor Sungho Jo from the School of Computing.
Raman spectroscopy sends light through a sample to see how it scatters. The results reveal structural information about the sample — the spectral fingerprint — allowing researchers to identify its molecules. The surface-enhanced version places sample cells on noble metal nanostructures that help amplify the sample’s signals.
However, it is challenging to obtain consistent and clear spectra of bacteria due to numerous overlapping peak sources, such as proteins in cell walls. “Moreover, strong signals of surrounding media are also enhanced to overwhelm target signals, requiring time-consuming and tedious bacterial separation steps,” said Professor Yeon Sik Jung from the Department of Materials Science and Engineering.
To parse through the noisy signals, the researchers implemented an artificial intelligence method called deep learning that can hierarchically extract certain features of the spectral information to classify data. They specifically designed their model, named the dual-branch wide-kernel network (DualWKNet), to efficiently learn the correlation between spectral features. Such an ability is critical for analyzing one-dimensional spectral data, according to Professor Jo.
“Despite having interfering signals or noise from the media, which make the general shapes of different bacterial spectra and their residing media signals look similar, high classification accuracies of bacterial types and their media were achieved,” Professor Jo said, explaining that DualWKNet allowed the team to identify key peaks in each class that were almost indiscernible in individual spectra, enhancing the classification accuracies. “Ultimately, with the use of DualWKNet replacing the bacteria and media separation steps, our method dramatically reduces analysis time.”
The researchers plan to use their platform to study more bacteria and media types, using the information to build a training data library of various bacterial types in additional media to reduce the collection and detection times for new samples.
“We developed a meaningful universal platform for rapid bacterial detection with the collaboration between SERS and deep learning,” Professor Jo said. “We hope to extend the use of our deep learning-based SERS analysis platform to detect numerous types of bacteria in additional media that are important for food or clinical analysis, such as blood.”
The National R&D Program, through a National Research Foundation of Korea grant funded by the Ministry of Science and ICT, supported this research.
-PublicationEojin Rho, Minjoon Kim, Seunghee H. Cho, Bongjae Choi, Hyungjoon Park, Hanhwi Jang, Yeon Sik Jung, Sungho Jo, “Separation-free bacterial identification in arbitrary media via deepneural network-based SERS analysis,” Biosensors and Bioelectronics online January 18, 2022 (doi.org/10.1016/j.bios.2022.113991)
-ProfileProfessor Yeon Sik JungDepartment of Materials Science and EngineeringKAIST
Professor Sungho JoSchool of ComputingKAIST
2022.03.04 View 23132 -
 DeepTFactor Predicts Transcription Factors
A deep learning-based tool predicts transcription factors using protein sequences as inputs
A joint research team from KAIST and UCSD has developed a deep neural network named DeepTFactor that predicts transcription factors from protein sequences. DeepTFactor will serve as a useful tool for understanding the regulatory systems of organisms, accelerating the use of deep learning for solving biological problems.
A transcription factor is a protein that specifically binds to DNA sequences to control the transcription initiation. Analyzing transcriptional regulation enables the understanding of how organisms control gene expression in response to genetic or environmental changes. In this regard, finding the transcription factor of an organism is the first step in the analysis of the transcriptional regulatory system of an organism.
Previously, transcription factors have been predicted by analyzing sequence homology with already characterized transcription factors or by data-driven approaches such as machine learning. Conventional machine learning models require a rigorous feature selection process that relies on domain expertise such as calculating the physicochemical properties of molecules or analyzing the homology of biological sequences. Meanwhile, deep learning can inherently learn latent features for the specific task.
A joint research team comprised of Ph.D. candidate Gi Bae Kim and Distinguished Professor Sang Yup Lee of the Department of Chemical and Biomolecular Engineering at KAIST, and Ye Gao and Professor Bernhard O. Palsson of the Department of Biochemical Engineering at UCSD reported a deep learning-based tool for the prediction of transcription factors. Their research paper “DeepTFactor: A deep learning-based tool for the prediction of transcription factors” was published online in PNAS.
Their article reports the development of DeepTFactor, a deep learning-based tool that predicts whether a given protein sequence is a transcription factor using three parallel convolutional neural networks. The joint research team predicted 332 transcription factors of Escherichia coli K-12 MG1655 using DeepTFactor and the performance of DeepTFactor by experimentally confirming the genome-wide binding sites of three predicted transcription factors (YqhC, YiaU, and YahB).
The joint research team further used a saliency method to understand the reasoning process of DeepTFactor. The researchers confirmed that even though information on the DNA binding domains of the transcription factor was not explicitly given the training process, DeepTFactor implicitly learned and used them for prediction. Unlike previous transcription factor prediction tools that were developed only for protein sequences of specific organisms, DeepTFactor is expected to be used in the analysis of the transcription systems of all organisms at a high level of performance.
Distinguished Professor Sang Yup Lee said, “DeepTFactor can be used to discover unknown transcription factors from numerous protein sequences that have not yet been characterized. It is expected that DeepTFactor will serve as an important tool for analyzing the regulatory systems of organisms of interest.”
This work was supported by the Technology Development Program to Solve Climate Changes on Systems Metabolic Engineering for Biorefineries from the Ministry of Science and ICT through the National Research Foundation of Korea.
-Publication
Gi Bae Kim, Ye Gao, Bernhard O. Palsson, and Sang Yup Lee. DeepTFactor: A deep learning-based tool for the prediction of transcription factors. (https://doi.org/10.1073/pnas202117118)
-Profile
Distinguished Professor Sang Yup Lee
leesy@kaist.ac.kr
Metabolic &Biomolecular Engineering National Research Laboratory
http://mbel.kaist.ac.kr
Department of Chemical and Biomolecular Engineering
KAIST
2021.01.05 View 10753
DeepTFactor Predicts Transcription Factors
A deep learning-based tool predicts transcription factors using protein sequences as inputs
A joint research team from KAIST and UCSD has developed a deep neural network named DeepTFactor that predicts transcription factors from protein sequences. DeepTFactor will serve as a useful tool for understanding the regulatory systems of organisms, accelerating the use of deep learning for solving biological problems.
A transcription factor is a protein that specifically binds to DNA sequences to control the transcription initiation. Analyzing transcriptional regulation enables the understanding of how organisms control gene expression in response to genetic or environmental changes. In this regard, finding the transcription factor of an organism is the first step in the analysis of the transcriptional regulatory system of an organism.
Previously, transcription factors have been predicted by analyzing sequence homology with already characterized transcription factors or by data-driven approaches such as machine learning. Conventional machine learning models require a rigorous feature selection process that relies on domain expertise such as calculating the physicochemical properties of molecules or analyzing the homology of biological sequences. Meanwhile, deep learning can inherently learn latent features for the specific task.
A joint research team comprised of Ph.D. candidate Gi Bae Kim and Distinguished Professor Sang Yup Lee of the Department of Chemical and Biomolecular Engineering at KAIST, and Ye Gao and Professor Bernhard O. Palsson of the Department of Biochemical Engineering at UCSD reported a deep learning-based tool for the prediction of transcription factors. Their research paper “DeepTFactor: A deep learning-based tool for the prediction of transcription factors” was published online in PNAS.
Their article reports the development of DeepTFactor, a deep learning-based tool that predicts whether a given protein sequence is a transcription factor using three parallel convolutional neural networks. The joint research team predicted 332 transcription factors of Escherichia coli K-12 MG1655 using DeepTFactor and the performance of DeepTFactor by experimentally confirming the genome-wide binding sites of three predicted transcription factors (YqhC, YiaU, and YahB).
The joint research team further used a saliency method to understand the reasoning process of DeepTFactor. The researchers confirmed that even though information on the DNA binding domains of the transcription factor was not explicitly given the training process, DeepTFactor implicitly learned and used them for prediction. Unlike previous transcription factor prediction tools that were developed only for protein sequences of specific organisms, DeepTFactor is expected to be used in the analysis of the transcription systems of all organisms at a high level of performance.
Distinguished Professor Sang Yup Lee said, “DeepTFactor can be used to discover unknown transcription factors from numerous protein sequences that have not yet been characterized. It is expected that DeepTFactor will serve as an important tool for analyzing the regulatory systems of organisms of interest.”
This work was supported by the Technology Development Program to Solve Climate Changes on Systems Metabolic Engineering for Biorefineries from the Ministry of Science and ICT through the National Research Foundation of Korea.
-Publication
Gi Bae Kim, Ye Gao, Bernhard O. Palsson, and Sang Yup Lee. DeepTFactor: A deep learning-based tool for the prediction of transcription factors. (https://doi.org/10.1073/pnas202117118)
-Profile
Distinguished Professor Sang Yup Lee
leesy@kaist.ac.kr
Metabolic &Biomolecular Engineering National Research Laboratory
http://mbel.kaist.ac.kr
Department of Chemical and Biomolecular Engineering
KAIST
2021.01.05 View 10753 -
 E. coli Engineered to Grow on CO₂ and Formic Acid as Sole Carbon Sources
- An E. coli strain that can grow to a relatively high cell density solely on CO₂ and formic acid was developed by employing metabolic engineering. -
Most biorefinery processes have relied on the use of biomass as a raw material for the production of chemicals and materials. Even though the use of CO₂ as a carbon source in biorefineries is desirable, it has not been possible to make common microbial strains such as E. coli grow on CO₂.
Now, a metabolic engineering research group at KAIST has developed a strategy to grow an E. coli strain to higher cell density solely on CO₂ and formic acid. Formic acid is a one carbon carboxylic acid, and can be easily produced from CO₂ using a variety of methods. Since it is easier to store and transport than CO₂, formic acid can be considered a good liquid-form alternative of CO₂.
With support from the C1 Gas Refinery R&D Center and the Ministry of Science and ICT, a research team led by Distinguished Professor Sang Yup Lee stepped up their work to develop an engineered E. coli strain capable of growing up to 11-fold higher cell density than those previously reported, using CO₂ and formic acid as sole carbon sources. This work was published in Nature Microbiology on September 28.
Despite the recent reports by several research groups on the development of E. coli strains capable of growing on CO₂ and formic acid, the maximum cell growth remained too low (optical density of around 1) and thus the production of chemicals from CO₂ and formic acid has been far from realized.
The team previously reported the reconstruction of the tetrahydrofolate cycle and reverse glycine cleavage pathway to construct an engineered E. coli strain that can sustain growth on CO₂ and formic acid. To further enhance the growth, the research team introduced the previously designed synthetic CO₂ and formic acid assimilation pathway, and two formate dehydrogenases.
Metabolic fluxes were also fine-tuned, the gluconeogenic flux enhanced, and the levels of cytochrome bo3 and bd-I ubiquinol oxidase for ATP generation were optimized. This engineered E. coli strain was able to grow to a relatively high OD600 of 7~11, showing promise as a platform strain growing solely on CO₂ and formic acid.
Professor Lee said, “We engineered E. coli that can grow to a higher cell density only using CO₂ and formic acid. We think that this is an important step forward, but this is not the end. The engineered strain we developed still needs further engineering so that it can grow faster to a much higher density.”
Professor Lee’s team is continuing to develop such a strain. “In the future, we would be delighted to see the production of chemicals from an engineered E. coli strain using CO₂ and formic acid as sole carbon sources,” he added.
-Profile:Distinguished Professor Sang Yup Leehttp://mbel.kaist.ac.krDepartment of Chemical and Biomolecular EngineeringKAIST
2020.09.29 View 12643
E. coli Engineered to Grow on CO₂ and Formic Acid as Sole Carbon Sources
- An E. coli strain that can grow to a relatively high cell density solely on CO₂ and formic acid was developed by employing metabolic engineering. -
Most biorefinery processes have relied on the use of biomass as a raw material for the production of chemicals and materials. Even though the use of CO₂ as a carbon source in biorefineries is desirable, it has not been possible to make common microbial strains such as E. coli grow on CO₂.
Now, a metabolic engineering research group at KAIST has developed a strategy to grow an E. coli strain to higher cell density solely on CO₂ and formic acid. Formic acid is a one carbon carboxylic acid, and can be easily produced from CO₂ using a variety of methods. Since it is easier to store and transport than CO₂, formic acid can be considered a good liquid-form alternative of CO₂.
With support from the C1 Gas Refinery R&D Center and the Ministry of Science and ICT, a research team led by Distinguished Professor Sang Yup Lee stepped up their work to develop an engineered E. coli strain capable of growing up to 11-fold higher cell density than those previously reported, using CO₂ and formic acid as sole carbon sources. This work was published in Nature Microbiology on September 28.
Despite the recent reports by several research groups on the development of E. coli strains capable of growing on CO₂ and formic acid, the maximum cell growth remained too low (optical density of around 1) and thus the production of chemicals from CO₂ and formic acid has been far from realized.
The team previously reported the reconstruction of the tetrahydrofolate cycle and reverse glycine cleavage pathway to construct an engineered E. coli strain that can sustain growth on CO₂ and formic acid. To further enhance the growth, the research team introduced the previously designed synthetic CO₂ and formic acid assimilation pathway, and two formate dehydrogenases.
Metabolic fluxes were also fine-tuned, the gluconeogenic flux enhanced, and the levels of cytochrome bo3 and bd-I ubiquinol oxidase for ATP generation were optimized. This engineered E. coli strain was able to grow to a relatively high OD600 of 7~11, showing promise as a platform strain growing solely on CO₂ and formic acid.
Professor Lee said, “We engineered E. coli that can grow to a higher cell density only using CO₂ and formic acid. We think that this is an important step forward, but this is not the end. The engineered strain we developed still needs further engineering so that it can grow faster to a much higher density.”
Professor Lee’s team is continuing to develop such a strain. “In the future, we would be delighted to see the production of chemicals from an engineered E. coli strain using CO₂ and formic acid as sole carbon sources,” he added.
-Profile:Distinguished Professor Sang Yup Leehttp://mbel.kaist.ac.krDepartment of Chemical and Biomolecular EngineeringKAIST
2020.09.29 View 12643 -
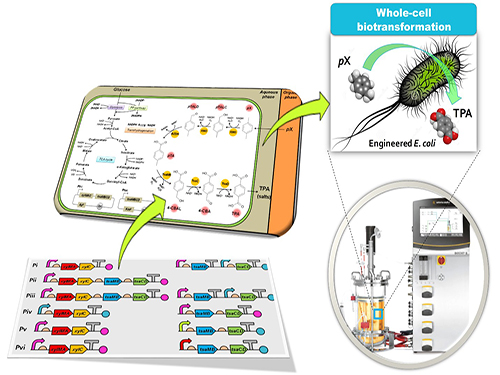 Bio-based p-Xylene Oxidation into Terephthalic Acid by Engineered E.coli
KAIST researchers have established an efficient biocatalytic system to produce terephthalic acid (TPA) from p-xylene (pX). It will allow this industrially important bulk chemical to be made available in a more environmentally-friendly manner.
The research team developed metabolically engineered Escherichia coli (E.coli) to biologically transform pX into TPA, a chemical necessary in the manufacturing of polyethylene terephthalate (PET). This biocatalysis system represents a greener and more efficient alternative to the traditional chemical methods for TPA production. This research, headed by Distinguished Professor Sang Yup Lee, was published in Nature Communications on May 31.
The research team utilized a metabolic engineering and synthetic biology approach to develop a recombinant microorganism that can oxidize pX into TPA using microbial fermentation. TPA is a globally important chemical commodity for manufacturing PET. It can be applied to manufacture plastic bottles, clothing fibers, films, and many other products. Currently, TPA is produced from pX oxidation through an industrially well-known chemical process (with a typical TPA yield of over 95 mol%), which shows, however, such drawbacks as intensive energy requirements at high temperatures and pressure, usage of heavy metal catalysts, and the unavoidable byproduct formation of 4-carboxybenzaldehyde.
The research team designed and constructed a synthetic metabolic pathway by incorporating the upper xylene degradation pathway of Pseudomonas putida F1 and the lower p-toluene sulfonate pathway of Comamonas testosteroni T-2, which successfully produced TPA from pX in small-scale cultures, with the formation of p-toluate (pTA) as the major byproduct. The team further optimized the pathway gene expression levels by using a synthetic biology toolkit, which gave the final engineered E. coli strain showing increased TPA production and the complete elimination of the byproduct.
Using this best-performing strain, the team designed an elegant two-phase (aqueous/organic) fermentation system for TPA production on a larger scale, where pX was supplied in the organic phase. Through a number of optimization steps, the team ultimately achieved production of 13.3 g TPA from 8.8 g pX, which represented an extraordinary yield of 97 mol%.
The team has developed a microbial biotechnology application which is reportedly the first successful example of the bio-based production of TPA from pX by the microbial fermentation of engineered E. coli. This bio-based TPA technology presents several advantages such as ambient reaction temperature and pressure, no use of heavy metals or other toxic chemicals, the removable of byproduct formation, and it is 100% environmentally compatible.
Professor Lee said, “We presented promising biotechnology for producing large amounts of the commodity chemical TPA, which is used for PET manufacturing, through metabolically engineered gut bacterium. Our research is meaningful in that it demonstrates the feasibility of the biotechnological production of bulk chemicals, and if reproducible when up-scaled, it will represent a breakthrough in hydrocarbon bioconversions.”
Ph.D. candidate Zi Wei Luo is the first author of this research (DOI:10.1038/ncomms15689).The research was supported by the Intelligent Synthetic Biology Center through the Global Frontier Project (2011-0031963) of the Ministry of Science, ICT & Future Planning through the National Research Foundation of Korea.
Figure: Biotransformation of pX into TPA by engineered E. coli.
This schematic diagram shows the overall conceptualization of how metabolically engineered E. coli produced TPA from pX. The engineered E. coli was developed through reconstituting a synthetic metabolic pathway for pX conversion to TPA and optimized for increased TPA yield and byproduct elimination. Two-phase partitioning fermentation system was developed for demonstrating the feasibility of large-scale production of TPA from pX using the engineered E. coli strains, where pX was supplied in the organic phase and TPA was produced in the aqueous phase.
2017.06.05 View 12858
Bio-based p-Xylene Oxidation into Terephthalic Acid by Engineered E.coli
KAIST researchers have established an efficient biocatalytic system to produce terephthalic acid (TPA) from p-xylene (pX). It will allow this industrially important bulk chemical to be made available in a more environmentally-friendly manner.
The research team developed metabolically engineered Escherichia coli (E.coli) to biologically transform pX into TPA, a chemical necessary in the manufacturing of polyethylene terephthalate (PET). This biocatalysis system represents a greener and more efficient alternative to the traditional chemical methods for TPA production. This research, headed by Distinguished Professor Sang Yup Lee, was published in Nature Communications on May 31.
The research team utilized a metabolic engineering and synthetic biology approach to develop a recombinant microorganism that can oxidize pX into TPA using microbial fermentation. TPA is a globally important chemical commodity for manufacturing PET. It can be applied to manufacture plastic bottles, clothing fibers, films, and many other products. Currently, TPA is produced from pX oxidation through an industrially well-known chemical process (with a typical TPA yield of over 95 mol%), which shows, however, such drawbacks as intensive energy requirements at high temperatures and pressure, usage of heavy metal catalysts, and the unavoidable byproduct formation of 4-carboxybenzaldehyde.
The research team designed and constructed a synthetic metabolic pathway by incorporating the upper xylene degradation pathway of Pseudomonas putida F1 and the lower p-toluene sulfonate pathway of Comamonas testosteroni T-2, which successfully produced TPA from pX in small-scale cultures, with the formation of p-toluate (pTA) as the major byproduct. The team further optimized the pathway gene expression levels by using a synthetic biology toolkit, which gave the final engineered E. coli strain showing increased TPA production and the complete elimination of the byproduct.
Using this best-performing strain, the team designed an elegant two-phase (aqueous/organic) fermentation system for TPA production on a larger scale, where pX was supplied in the organic phase. Through a number of optimization steps, the team ultimately achieved production of 13.3 g TPA from 8.8 g pX, which represented an extraordinary yield of 97 mol%.
The team has developed a microbial biotechnology application which is reportedly the first successful example of the bio-based production of TPA from pX by the microbial fermentation of engineered E. coli. This bio-based TPA technology presents several advantages such as ambient reaction temperature and pressure, no use of heavy metals or other toxic chemicals, the removable of byproduct formation, and it is 100% environmentally compatible.
Professor Lee said, “We presented promising biotechnology for producing large amounts of the commodity chemical TPA, which is used for PET manufacturing, through metabolically engineered gut bacterium. Our research is meaningful in that it demonstrates the feasibility of the biotechnological production of bulk chemicals, and if reproducible when up-scaled, it will represent a breakthrough in hydrocarbon bioconversions.”
Ph.D. candidate Zi Wei Luo is the first author of this research (DOI:10.1038/ncomms15689).The research was supported by the Intelligent Synthetic Biology Center through the Global Frontier Project (2011-0031963) of the Ministry of Science, ICT & Future Planning through the National Research Foundation of Korea.
Figure: Biotransformation of pX into TPA by engineered E. coli.
This schematic diagram shows the overall conceptualization of how metabolically engineered E. coli produced TPA from pX. The engineered E. coli was developed through reconstituting a synthetic metabolic pathway for pX conversion to TPA and optimized for increased TPA yield and byproduct elimination. Two-phase partitioning fermentation system was developed for demonstrating the feasibility of large-scale production of TPA from pX using the engineered E. coli strains, where pX was supplied in the organic phase and TPA was produced in the aqueous phase.
2017.06.05 View 12858