bio
-
 KAIST presents a fundamental technology to remove metastatic traits from lung cancer cells
KAIST (President Kwang Hyung Lee) announced on January 30th that a research team led by Professor Kwang-Hyun Cho from the Department of Bio and Brain Engineering succeeded in using systems biology research to change the properties of carcinogenic cells in the lungs and eliminate both drug resistance and their ability to proliferate out to other areas of the body.
As the incidences of cancer increase within aging populations, cancer has become the most lethal disease threatening healthy life. Fatality rates are especially high when early detection does not happen in time and metastasis has occurred in various organs. In order to resolve this problem, a series of attempts were made to remove or lower the ability of cancer cells to spread, but they resulted in cancer cells in the intermediate state becoming more unstable and even more malignant, which created serious treatment challenges.
Professor Kwang-Hyun Cho's research team simulated various cancer cell states in the Epithelial-to-Mesenchymal Transition (EMT) of lung cancer cells, between epithelial cells without metastatic ability and mesenchymal cells with metastatic ability. A mathematical model of molecular network was established, and key regulators that could reverse the state of invasive and drug resistant mesenchymal cells back to the epithelial state were discovered through computer simulation analysis and molecular cell experiments. In particular, this process succeeded in properly reverting the mesenchymal lung cancer cells to a state where they were sensitive to chemotherapy treatment while avoiding the unstable EMT hybrid cell state in the middle process, which had remained a difficult problem.
The results of this research, in which KAIST Ph.D. student Namhee Kim, Dr. Chae Young Hwang, Researcher Taeyoung Kim, and Ph.D. student Hyunjin Kim participated, were published as an online paper in the international journal “Cancer Research” published by the American Association for Cancer Research (AACR) on January 30th. (Paper title: A cell fate reprogramming strategy reverses epithelial-to-mesenchymal transition of lung cancer cells while avoiding hybrid states)
Cells in an EMT hybrid state, which are caused by incomplete transitions during the EMT process in cancer cells, have the characteristics of both epithelial cells and mesenchymal cells, and are known to have high drug resistance and metastatic potential by acquiring high stem cell capacity. In particular, EMT is further enhanced through factors such as transforming growth factor-beta (TGF-β) secreted from the tumor microenvironment (TME) and, as a result, various cell states with high plasticity appear. Due to the complexity of EMT, it has been very difficult to completely reverse the transitional process of the mesenchymal cancer cells to an epithelial cell state in which metastatic ability and drug resistance are eliminated while avoiding the EMT hybrid cell state with high metastatic ability and drug resistance.
Professor Kwang-Hyun Cho's research team established a mathematical model of the gene regulation network that governs the complex process of EMT, and then applied large-scale computer simulation analysis and complex system network control technology to identify and verify 'p53', 'SMAD4', and 'ERK1' and 'ERK 2' (collectively ERKs) through molecular cell experiments as the three key molecular targets that can transform lung cancer cells in the mesenchymal cell state, reversed back to an epithelial cell state that no longer demonstrates the ability to metastasize, while avoiding the EMT hybrid cell state.
In particular, by analyzing the molecular regulatory mechanism of the complex EMT process at the system level, the key pathways were identified that were linked to the positive feedback that plays an important role in completely returning cancer cells to an epithelial cell state in which metastatic ability and drug resistance are removed.
This discovery is significant in that it proved that mesenchymal cells can be reverted to the state of epithelial cells under conditions where TGF-β stimulation are present, like they are in the actual environment where cancer tissue forms in the human body.
Abnormal EMT in cancer cells leads to various malignant traits such as the migration and invasion of cancer cells, changes in responsiveness to chemotherapy treatment, enhanced stem cell function, and the dissemination of cancer. In particular, the acquisition of the metastatic ability of cancer cells is a key determinant factor for the prognosis of cancer patients. The EMT reversal technology in lung cancer cells developed in this research is a new anti-cancer treatment strategy that reprograms cancer cells to eliminate their high plasticity and metastatic potential and increase their responsiveness to chemotherapy.
Professor Kwang-Hyun Cho said, "By succeeding in reversing the state of lung cancer cells that acquired high metastatic traits and resistance to drugs and reverting them to a treatable epithelial cell state with renewed sensitivity to chemotherapy, the research findings propose a new strategy for treatments that can improve the prognosis of cancer patients.”
Professor Kwang-Hyun Cho's research team was the first to present the principle of reversal treatment to revert cancer cells to normal cells, following through with the announcement of the results of their study that reverted colon cancer cells to normal colon cells in January of 2020, and also presenting successful re-programming research where the most malignant basal type breast cancer cells turned into less-malignant luminal type breast cancer cells that were treatable with hormonal therapies in January of 2022. This latest research result is the third in the development of reversal technology where lung cancer cells that had acquired metastatic traits returned to a state in which their metastatic ability was removed and drug sensitivity was enhanced.
This research was carried out with support from the Ministry of Science and ICT and the National Research Foundation of Korea's Basic Research in Science & Engineering Program for Mid-Career Researchers.
< Figure 1. Construction of the mathematical model of the regulatory network to represent the EMT phenotype based on the interaction between various molecules related to EMT.
(A) Professor Kwang-Hyun Cho's research team investigated numerous literatures and databases related to complex EMT, and based on comparative analysis of cell line data showing epithelial and mesenchymal cell conditions, they extracted key signaling pathways related to EMT and built a mathematical model of regulatory network (B) By comparing the results of computer simulation analysis and the molecular cell experiments, it was verified how well the constructed mathematical model simulated the actual cellular phenomena. >
< Figure 2. Understanding of various EMT phenotypes through large-scale computer simulation analysis and complex system network control technology.
(A) Through computer simulation analysis and experiments, Professor Kwang-Hyun Cho's research team found that complete control of EMT is impossible with single-molecule control alone. In particular, through comparison of the relative stability of attractors, it was revealed that the cell state exhibiting EMT hybrid characteristics has unstable properties. (B), (C) Based on these results, Prof. Cho’s team identified two feedbacks (positive feedback consisting of Snail-miR-34 and ZEB1-miR-200) that play an important role in avoiding the EMT hybrid state that appeared in the TGF-β-ON state. It was found through computer simulation analysis that the two feedbacks restore relatively high stability when the excavated p53 and SMAD4 are regulated. In addition, molecular cell experiments demonstrated that the expression levels of E-cad and ZEB1, which are representative phenotypic markers of EMT, changed similarly to the expression profile in the epithelial cell state, despite the TGF-β-ON state. >
< Figure 3. Complex molecular network analysis and discovery of reprogramming molecular targets for intact elimination of EMT hybrid features.
(A) Controlling the expression of p53 and SMAD4 in lung cancer cell lines was expected to overcome drug resistance, but contrary to expectations, chemotherapy responsiveness was not restored. (B) Professor Kwang-Hyun Cho's research team additionally analyzed computer simulations, genome data, and experimental results and found that high expression levels of TWIST1 and EPCAM were related to drug resistance. (C) Prof. Cho’s team identified three key molecular targets: p53, SMAD4 and ERK1 & ERK2. (D), (E) Furthermore, they identified a key pathway that plays an important role in completely reversing into epithelial cells while avoiding EMT hybrid characteristics, and confirmed through network analysis and attractor analysis that high stability of the key pathway was restored when the proposed molecular target was controlled. >
< Figure 4. Verification through experiments with lung cancer cell lines.
When p53 was activated and SMAD4 and ERK1/2 were inhibited in lung cancer cell lines, (A), (B) E-cad protein expression increased and ZEB1 protein expression decreased, and (C) mesenchymal cell status including TWIST1 and EPCAM and gene expression of markers related to stem cell potential characteristics were completely inhibited. In addition, (D) it was confirmed that resistance to chemotherapy treatment was also overcome as the cell state was reversed by the regulated target. >
< Figure 5. A schematic representation of the research results.
Prof. Cho’s research team identified key molecular regulatory pathways to avoid high plasticity formed by abnormal EMT of cancer cells and reverse it to an epithelial cell state through systems biology research. From this analysis, a reprogramming molecular target that can reverse the state of mesenchymal cells with acquired invasiveness and drug resistance to the state of epithelial cells with restored drug responsiveness was discovered.
For lung cancer cells, when a drug that enhances the expression of p53, one of the molecular targets discovered, and inhibits the expression of SMAD4 and ERK1 & ERK2 is administered, the molecular network of genes in the state of mesenchymal cells is modified, eventually eliminating metastatic ability and it is reprogrammed to turn into epithelial cells without the resistance to chemotherapy treatments. >
2023.01.30 View 19322
KAIST presents a fundamental technology to remove metastatic traits from lung cancer cells
KAIST (President Kwang Hyung Lee) announced on January 30th that a research team led by Professor Kwang-Hyun Cho from the Department of Bio and Brain Engineering succeeded in using systems biology research to change the properties of carcinogenic cells in the lungs and eliminate both drug resistance and their ability to proliferate out to other areas of the body.
As the incidences of cancer increase within aging populations, cancer has become the most lethal disease threatening healthy life. Fatality rates are especially high when early detection does not happen in time and metastasis has occurred in various organs. In order to resolve this problem, a series of attempts were made to remove or lower the ability of cancer cells to spread, but they resulted in cancer cells in the intermediate state becoming more unstable and even more malignant, which created serious treatment challenges.
Professor Kwang-Hyun Cho's research team simulated various cancer cell states in the Epithelial-to-Mesenchymal Transition (EMT) of lung cancer cells, between epithelial cells without metastatic ability and mesenchymal cells with metastatic ability. A mathematical model of molecular network was established, and key regulators that could reverse the state of invasive and drug resistant mesenchymal cells back to the epithelial state were discovered through computer simulation analysis and molecular cell experiments. In particular, this process succeeded in properly reverting the mesenchymal lung cancer cells to a state where they were sensitive to chemotherapy treatment while avoiding the unstable EMT hybrid cell state in the middle process, which had remained a difficult problem.
The results of this research, in which KAIST Ph.D. student Namhee Kim, Dr. Chae Young Hwang, Researcher Taeyoung Kim, and Ph.D. student Hyunjin Kim participated, were published as an online paper in the international journal “Cancer Research” published by the American Association for Cancer Research (AACR) on January 30th. (Paper title: A cell fate reprogramming strategy reverses epithelial-to-mesenchymal transition of lung cancer cells while avoiding hybrid states)
Cells in an EMT hybrid state, which are caused by incomplete transitions during the EMT process in cancer cells, have the characteristics of both epithelial cells and mesenchymal cells, and are known to have high drug resistance and metastatic potential by acquiring high stem cell capacity. In particular, EMT is further enhanced through factors such as transforming growth factor-beta (TGF-β) secreted from the tumor microenvironment (TME) and, as a result, various cell states with high plasticity appear. Due to the complexity of EMT, it has been very difficult to completely reverse the transitional process of the mesenchymal cancer cells to an epithelial cell state in which metastatic ability and drug resistance are eliminated while avoiding the EMT hybrid cell state with high metastatic ability and drug resistance.
Professor Kwang-Hyun Cho's research team established a mathematical model of the gene regulation network that governs the complex process of EMT, and then applied large-scale computer simulation analysis and complex system network control technology to identify and verify 'p53', 'SMAD4', and 'ERK1' and 'ERK 2' (collectively ERKs) through molecular cell experiments as the three key molecular targets that can transform lung cancer cells in the mesenchymal cell state, reversed back to an epithelial cell state that no longer demonstrates the ability to metastasize, while avoiding the EMT hybrid cell state.
In particular, by analyzing the molecular regulatory mechanism of the complex EMT process at the system level, the key pathways were identified that were linked to the positive feedback that plays an important role in completely returning cancer cells to an epithelial cell state in which metastatic ability and drug resistance are removed.
This discovery is significant in that it proved that mesenchymal cells can be reverted to the state of epithelial cells under conditions where TGF-β stimulation are present, like they are in the actual environment where cancer tissue forms in the human body.
Abnormal EMT in cancer cells leads to various malignant traits such as the migration and invasion of cancer cells, changes in responsiveness to chemotherapy treatment, enhanced stem cell function, and the dissemination of cancer. In particular, the acquisition of the metastatic ability of cancer cells is a key determinant factor for the prognosis of cancer patients. The EMT reversal technology in lung cancer cells developed in this research is a new anti-cancer treatment strategy that reprograms cancer cells to eliminate their high plasticity and metastatic potential and increase their responsiveness to chemotherapy.
Professor Kwang-Hyun Cho said, "By succeeding in reversing the state of lung cancer cells that acquired high metastatic traits and resistance to drugs and reverting them to a treatable epithelial cell state with renewed sensitivity to chemotherapy, the research findings propose a new strategy for treatments that can improve the prognosis of cancer patients.”
Professor Kwang-Hyun Cho's research team was the first to present the principle of reversal treatment to revert cancer cells to normal cells, following through with the announcement of the results of their study that reverted colon cancer cells to normal colon cells in January of 2020, and also presenting successful re-programming research where the most malignant basal type breast cancer cells turned into less-malignant luminal type breast cancer cells that were treatable with hormonal therapies in January of 2022. This latest research result is the third in the development of reversal technology where lung cancer cells that had acquired metastatic traits returned to a state in which their metastatic ability was removed and drug sensitivity was enhanced.
This research was carried out with support from the Ministry of Science and ICT and the National Research Foundation of Korea's Basic Research in Science & Engineering Program for Mid-Career Researchers.
< Figure 1. Construction of the mathematical model of the regulatory network to represent the EMT phenotype based on the interaction between various molecules related to EMT.
(A) Professor Kwang-Hyun Cho's research team investigated numerous literatures and databases related to complex EMT, and based on comparative analysis of cell line data showing epithelial and mesenchymal cell conditions, they extracted key signaling pathways related to EMT and built a mathematical model of regulatory network (B) By comparing the results of computer simulation analysis and the molecular cell experiments, it was verified how well the constructed mathematical model simulated the actual cellular phenomena. >
< Figure 2. Understanding of various EMT phenotypes through large-scale computer simulation analysis and complex system network control technology.
(A) Through computer simulation analysis and experiments, Professor Kwang-Hyun Cho's research team found that complete control of EMT is impossible with single-molecule control alone. In particular, through comparison of the relative stability of attractors, it was revealed that the cell state exhibiting EMT hybrid characteristics has unstable properties. (B), (C) Based on these results, Prof. Cho’s team identified two feedbacks (positive feedback consisting of Snail-miR-34 and ZEB1-miR-200) that play an important role in avoiding the EMT hybrid state that appeared in the TGF-β-ON state. It was found through computer simulation analysis that the two feedbacks restore relatively high stability when the excavated p53 and SMAD4 are regulated. In addition, molecular cell experiments demonstrated that the expression levels of E-cad and ZEB1, which are representative phenotypic markers of EMT, changed similarly to the expression profile in the epithelial cell state, despite the TGF-β-ON state. >
< Figure 3. Complex molecular network analysis and discovery of reprogramming molecular targets for intact elimination of EMT hybrid features.
(A) Controlling the expression of p53 and SMAD4 in lung cancer cell lines was expected to overcome drug resistance, but contrary to expectations, chemotherapy responsiveness was not restored. (B) Professor Kwang-Hyun Cho's research team additionally analyzed computer simulations, genome data, and experimental results and found that high expression levels of TWIST1 and EPCAM were related to drug resistance. (C) Prof. Cho’s team identified three key molecular targets: p53, SMAD4 and ERK1 & ERK2. (D), (E) Furthermore, they identified a key pathway that plays an important role in completely reversing into epithelial cells while avoiding EMT hybrid characteristics, and confirmed through network analysis and attractor analysis that high stability of the key pathway was restored when the proposed molecular target was controlled. >
< Figure 4. Verification through experiments with lung cancer cell lines.
When p53 was activated and SMAD4 and ERK1/2 were inhibited in lung cancer cell lines, (A), (B) E-cad protein expression increased and ZEB1 protein expression decreased, and (C) mesenchymal cell status including TWIST1 and EPCAM and gene expression of markers related to stem cell potential characteristics were completely inhibited. In addition, (D) it was confirmed that resistance to chemotherapy treatment was also overcome as the cell state was reversed by the regulated target. >
< Figure 5. A schematic representation of the research results.
Prof. Cho’s research team identified key molecular regulatory pathways to avoid high plasticity formed by abnormal EMT of cancer cells and reverse it to an epithelial cell state through systems biology research. From this analysis, a reprogramming molecular target that can reverse the state of mesenchymal cells with acquired invasiveness and drug resistance to the state of epithelial cells with restored drug responsiveness was discovered.
For lung cancer cells, when a drug that enhances the expression of p53, one of the molecular targets discovered, and inhibits the expression of SMAD4 and ERK1 & ERK2 is administered, the molecular network of genes in the state of mesenchymal cells is modified, eventually eliminating metastatic ability and it is reprogrammed to turn into epithelial cells without the resistance to chemotherapy treatments. >
2023.01.30 View 19322 -
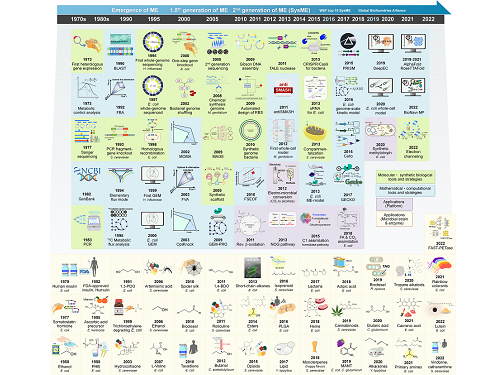 Overview of the 30-year history of metabolic engineering
< Distinguished Professor Sang Yup Lee from the Department of Chemical and Biomolecular Engineering at KAIST >
A research team comprised of Gi Bae Kim, Dr. So Young Choi, Dr. In Jin Cho, Da-Hee Ahn, and Distinguished Professor Sang Yup Lee from the Department of Chemical and Biomolecular Engineering at KAIST reported the 30-year history of metabolic engineering, highlighting examples of recent progress in the field and contributions to sustainability and health. Their paper “Metabolic engineering for sustainability and health” was published online in the 40th anniversary special issue of Trends in Biotechnology on January 10, 2023.
Metabolic engineering, a discipline of engineering that modifies cell phenotypes through molecular and genetic-level manipulations to improve cellular activities, has been studied since the early 1990s, and has progressed significantly over the past 30 years. In particular, metabolic engineering has enabled the engineering of microorganisms for the development of microbial cell factories capable of efficiently producing chemicals and materials as well as degrading recalcitrant contaminants.
This review article revisited how metabolic engineering has advanced over the past 30 years, from the advent of genetic engineering techniques such as recombinant DNA technologies to recent breakthroughs in systems metabolic engineering and data science aided by artificial intelligence. The research team highlighted momentous events and achievements in metabolic engineering, providing both trends and future directions in the field. Metabolic engineering’s contributions to bio-based sustainable chemicals and clean energy, health, and bioremediation were also reviewed. Finally, the research team shared their perspectives on the future challenges impacting metabolic engineering than must be overcome in order to achieve advancements in sustainability and health.
Distinguished Professor Sang Yup Lee said, “Replacing fossil resource-based chemical processes with bio-based sustainable processes for the production of chemicals, fuels, and materials using metabolic engineering has become our essential task for the future. By looking back on the 30+ years of metabolic engineering, we aimed to highlight the contributions of metabolic engineering to achieve sustainability and good health.” He added, “Metabolic engineering will play an increasingly important role as a key solution to the climate crisis, environmental pollution, food and energy shortages, and health problems in aging societies.”
< Figure: Metabolic Engineering Timeline >
2023.01.25 View 11745
Overview of the 30-year history of metabolic engineering
< Distinguished Professor Sang Yup Lee from the Department of Chemical and Biomolecular Engineering at KAIST >
A research team comprised of Gi Bae Kim, Dr. So Young Choi, Dr. In Jin Cho, Da-Hee Ahn, and Distinguished Professor Sang Yup Lee from the Department of Chemical and Biomolecular Engineering at KAIST reported the 30-year history of metabolic engineering, highlighting examples of recent progress in the field and contributions to sustainability and health. Their paper “Metabolic engineering for sustainability and health” was published online in the 40th anniversary special issue of Trends in Biotechnology on January 10, 2023.
Metabolic engineering, a discipline of engineering that modifies cell phenotypes through molecular and genetic-level manipulations to improve cellular activities, has been studied since the early 1990s, and has progressed significantly over the past 30 years. In particular, metabolic engineering has enabled the engineering of microorganisms for the development of microbial cell factories capable of efficiently producing chemicals and materials as well as degrading recalcitrant contaminants.
This review article revisited how metabolic engineering has advanced over the past 30 years, from the advent of genetic engineering techniques such as recombinant DNA technologies to recent breakthroughs in systems metabolic engineering and data science aided by artificial intelligence. The research team highlighted momentous events and achievements in metabolic engineering, providing both trends and future directions in the field. Metabolic engineering’s contributions to bio-based sustainable chemicals and clean energy, health, and bioremediation were also reviewed. Finally, the research team shared their perspectives on the future challenges impacting metabolic engineering than must be overcome in order to achieve advancements in sustainability and health.
Distinguished Professor Sang Yup Lee said, “Replacing fossil resource-based chemical processes with bio-based sustainable processes for the production of chemicals, fuels, and materials using metabolic engineering has become our essential task for the future. By looking back on the 30+ years of metabolic engineering, we aimed to highlight the contributions of metabolic engineering to achieve sustainability and good health.” He added, “Metabolic engineering will play an increasingly important role as a key solution to the climate crisis, environmental pollution, food and energy shortages, and health problems in aging societies.”
< Figure: Metabolic Engineering Timeline >
2023.01.25 View 11745 -
 Establishing a novel strategy to tackle Huntington’s disease
A platform to take on the Huntington’s disease via an innovative approach established by KAIST’s researchers through international collaboration with scientists in the Netherlands, France, and Sweden.
Through an international joint research effort involving ProQR Therapeutics of the Netherlands, Université Grenoble Alpes of France, and KTH Royal Institute of Technology of Sweden, Professor Ji-Soon Song's research team in the Department of Biological Sciences and KAIST Institute for BioCentury of KAIST, established a noble strategy to treat Huntington's disease. The new works showed that the protein converted from disease form to its disease-free form maintains its original function, providing new roadblocks to approach Huntington’s disease.
This research, titled, “A pathogenic-proteolysis resistant huntingtin isoform induced by an antisense oligonucleotide maintains huntingtin function”, co-authored by Hyeongju Kim, was published in the online edition of 'Journal of Clinical Investigation Insight' on August 9, 2022.
Huntington's disease is a dominantly inherited neurodegenerative disease and is caused by a mutation in a protein called ‘huntingtin’, which adds a distinctive feature of an expanded stretch of glutamine amino acids called polyglutamine to the protein. It is estimated that one in every 10,000 have Huntington's disease in United States. The patients would suffer a decade of regression before death, and, thus far, there is no known cure for the disease.
The cleavage near the stretched polyglutamine in mutated huntingtin is known to be the cause of the Huntington’s disease. However, as huntingtin protein is required for the development and normal function of the brain, it is critical to specifically eliminate the disease-causing protein while maintaining the ones that are still normally functioning.
The research team showed that huntingtin delta 12, the converted form of huntingtin that is resistant to developing cleavages at the ends of the protein, the known cause of the Huntington’s disease (HD), alleviated the disease’s symptoms while maintaining the functions of normal huntingtin.
Figure. Huntington's disease resistance huntingtin protein induced by antisense oligonucleotide (AON) is resistant to Caspase-6 cleavage, therefore, does not cause Huntington’s disease while maintaining normal functions of huntingtin.
The research was welcomed as it is sure to fuel innovate strategies to tackle Huntington’s disease without altering the essential function of huntingtin.
This work was supported by a Global Research Lab grant from the National Research Foundation of Korea (NRF) and by a EUREKA Eurostars 2 grant from European Union Horizon 2020.
2022.09.02 View 8158
Establishing a novel strategy to tackle Huntington’s disease
A platform to take on the Huntington’s disease via an innovative approach established by KAIST’s researchers through international collaboration with scientists in the Netherlands, France, and Sweden.
Through an international joint research effort involving ProQR Therapeutics of the Netherlands, Université Grenoble Alpes of France, and KTH Royal Institute of Technology of Sweden, Professor Ji-Soon Song's research team in the Department of Biological Sciences and KAIST Institute for BioCentury of KAIST, established a noble strategy to treat Huntington's disease. The new works showed that the protein converted from disease form to its disease-free form maintains its original function, providing new roadblocks to approach Huntington’s disease.
This research, titled, “A pathogenic-proteolysis resistant huntingtin isoform induced by an antisense oligonucleotide maintains huntingtin function”, co-authored by Hyeongju Kim, was published in the online edition of 'Journal of Clinical Investigation Insight' on August 9, 2022.
Huntington's disease is a dominantly inherited neurodegenerative disease and is caused by a mutation in a protein called ‘huntingtin’, which adds a distinctive feature of an expanded stretch of glutamine amino acids called polyglutamine to the protein. It is estimated that one in every 10,000 have Huntington's disease in United States. The patients would suffer a decade of regression before death, and, thus far, there is no known cure for the disease.
The cleavage near the stretched polyglutamine in mutated huntingtin is known to be the cause of the Huntington’s disease. However, as huntingtin protein is required for the development and normal function of the brain, it is critical to specifically eliminate the disease-causing protein while maintaining the ones that are still normally functioning.
The research team showed that huntingtin delta 12, the converted form of huntingtin that is resistant to developing cleavages at the ends of the protein, the known cause of the Huntington’s disease (HD), alleviated the disease’s symptoms while maintaining the functions of normal huntingtin.
Figure. Huntington's disease resistance huntingtin protein induced by antisense oligonucleotide (AON) is resistant to Caspase-6 cleavage, therefore, does not cause Huntington’s disease while maintaining normal functions of huntingtin.
The research was welcomed as it is sure to fuel innovate strategies to tackle Huntington’s disease without altering the essential function of huntingtin.
This work was supported by a Global Research Lab grant from the National Research Foundation of Korea (NRF) and by a EUREKA Eurostars 2 grant from European Union Horizon 2020.
2022.09.02 View 8158 -
 Phage resistant Escherichia coli strains developed to reduce fermentation failure
A genome engineering-based systematic strategy for developing phage resistant Escherichia coli strains has been successfully developed through the collaborative efforts of a team led by Professor Sang Yup Lee, Professor Shi Chen, and Professor Lianrong Wang. This study by Xuan Zou et al. was published in Nature Communications in August 2022 and featured in Nature Communications Editors’ Highlights. The collaboration by the School of Pharmaceutical Sciences at Wuhan University, the First Affiliated Hospital of Shenzhen University, and the KAIST Department of Chemical and Biomolecular Engineering has made an important advance in the metabolic engineering and fermentation industry as it solves a big problem of phage infection causing fermentation failure.
Systems metabolic engineering is a highly interdisciplinary field that has made the development of microbial cell factories to produce various bioproducts including chemicals, fuels, and materials possible in a sustainable and environmentally friendly way, mitigating the impact of worldwide resource depletion and climate change. Escherichia coli is one of the most important chassis microbial strains, given its wide applications in the bio-based production of a diverse range of chemicals and materials. With the development of tools and strategies for systems metabolic engineering using E. coli, a highly optimized and well-characterized cell factory will play a crucial role in converting cheap and readily available raw materials into products of great economic and industrial value.
However, the consistent problem of phage contamination in fermentation imposes a devastating impact on host cells and threatens the productivity of bacterial bioprocesses in biotechnology facilities, which can lead to widespread fermentation failure and immeasurable economic loss. Host-controlled defense systems can be developed into effective genetic engineering solutions to address bacteriophage contamination in industrial-scale fermentation; however, most of the resistance mechanisms only narrowly restrict phages and their effect on phage contamination will be limited.
There have been attempts to develop diverse abilities/systems for environmental adaptation or antiviral defense. The team’s collaborative efforts developed a new type II single-stranded DNA phosphorothioation (Ssp) defense system derived from E. coli 3234/A, which can be used in multiple industrial E. coli strains (e.g., E. coli K-12, B and W) to provide broad protection against various types of dsDNA coliphages. Furthermore, they developed a systematic genome engineering strategy involving the simultaneous genomic integration of the Ssp defense module and mutations in components that are essential to the phage life cycle. This strategy can be used to transform E. coli hosts that are highly susceptible to phage attack into strains with powerful restriction effects on the tested bacteriophages. This endows hosts with strong resistance against a wide spectrum of phage infections without affecting bacterial growth and normal physiological function. More importantly, the resulting engineered phage-resistant strains maintained the capabilities of producing the desired chemicals and recombinant proteins even under high levels of phage cocktail challenge, which provides crucial protection against phage attacks.
This is a major step forward, as it provides a systematic solution for engineering phage-resistant bacterial strains, especially industrial bioproduction strains, to protect cells from a wide range of bacteriophages. Considering the functionality of this engineering strategy with diverse E. coli strains, the strategy reported in this study can be widely extended to other bacterial species and industrial applications, which will be of great interest to researchers in academia and industry alike.
Fig. A schematic model of the systematic strategy for engineering phage-sensitive industrial E. coli strains into strains with broad antiphage activities. Through the simultaneous genomic integration of a DNA phosphorothioation-based Ssp defense module and mutations of components essential for the phage life cycle, the engineered E. coli strains show strong resistance against diverse phages tested and maintain the capabilities of producing example recombinant proteins, even under high levels of phage cocktail challenge.
2022.08.23 View 15467
Phage resistant Escherichia coli strains developed to reduce fermentation failure
A genome engineering-based systematic strategy for developing phage resistant Escherichia coli strains has been successfully developed through the collaborative efforts of a team led by Professor Sang Yup Lee, Professor Shi Chen, and Professor Lianrong Wang. This study by Xuan Zou et al. was published in Nature Communications in August 2022 and featured in Nature Communications Editors’ Highlights. The collaboration by the School of Pharmaceutical Sciences at Wuhan University, the First Affiliated Hospital of Shenzhen University, and the KAIST Department of Chemical and Biomolecular Engineering has made an important advance in the metabolic engineering and fermentation industry as it solves a big problem of phage infection causing fermentation failure.
Systems metabolic engineering is a highly interdisciplinary field that has made the development of microbial cell factories to produce various bioproducts including chemicals, fuels, and materials possible in a sustainable and environmentally friendly way, mitigating the impact of worldwide resource depletion and climate change. Escherichia coli is one of the most important chassis microbial strains, given its wide applications in the bio-based production of a diverse range of chemicals and materials. With the development of tools and strategies for systems metabolic engineering using E. coli, a highly optimized and well-characterized cell factory will play a crucial role in converting cheap and readily available raw materials into products of great economic and industrial value.
However, the consistent problem of phage contamination in fermentation imposes a devastating impact on host cells and threatens the productivity of bacterial bioprocesses in biotechnology facilities, which can lead to widespread fermentation failure and immeasurable economic loss. Host-controlled defense systems can be developed into effective genetic engineering solutions to address bacteriophage contamination in industrial-scale fermentation; however, most of the resistance mechanisms only narrowly restrict phages and their effect on phage contamination will be limited.
There have been attempts to develop diverse abilities/systems for environmental adaptation or antiviral defense. The team’s collaborative efforts developed a new type II single-stranded DNA phosphorothioation (Ssp) defense system derived from E. coli 3234/A, which can be used in multiple industrial E. coli strains (e.g., E. coli K-12, B and W) to provide broad protection against various types of dsDNA coliphages. Furthermore, they developed a systematic genome engineering strategy involving the simultaneous genomic integration of the Ssp defense module and mutations in components that are essential to the phage life cycle. This strategy can be used to transform E. coli hosts that are highly susceptible to phage attack into strains with powerful restriction effects on the tested bacteriophages. This endows hosts with strong resistance against a wide spectrum of phage infections without affecting bacterial growth and normal physiological function. More importantly, the resulting engineered phage-resistant strains maintained the capabilities of producing the desired chemicals and recombinant proteins even under high levels of phage cocktail challenge, which provides crucial protection against phage attacks.
This is a major step forward, as it provides a systematic solution for engineering phage-resistant bacterial strains, especially industrial bioproduction strains, to protect cells from a wide range of bacteriophages. Considering the functionality of this engineering strategy with diverse E. coli strains, the strategy reported in this study can be widely extended to other bacterial species and industrial applications, which will be of great interest to researchers in academia and industry alike.
Fig. A schematic model of the systematic strategy for engineering phage-sensitive industrial E. coli strains into strains with broad antiphage activities. Through the simultaneous genomic integration of a DNA phosphorothioation-based Ssp defense module and mutations of components essential for the phage life cycle, the engineered E. coli strains show strong resistance against diverse phages tested and maintain the capabilities of producing example recombinant proteins, even under high levels of phage cocktail challenge.
2022.08.23 View 15467 -
 Interactive Map of Metabolical Synthesis of Chemicals
An interactive map that compiled the chemicals produced by biological, chemical and combined reactions has been distributed on the web
- A team led by Distinguished Professor Sang Yup Lee of the Department of Chemical and Biomolecular Engineering, organized and distributed an all-inclusive listing of chemical substances that can be synthesized using microorganisms
- It is expected to be used by researchers around the world as it enables easy assessment of the synthetic pathway through the web.
A research team comprised of Woo Dae Jang, Gi Bae Kim, and Distinguished Professor Sang Yup Lee of the Department of Chemical and Biomolecular Engineering at KAIST reported an interactive metabolic map of bio-based chemicals. Their research paper “An interactive metabolic map of bio-based chemicals” was published online in Trends in Biotechnology on August 10, 2022.
As a response to rapid climate change and environmental pollution, research on the production of petrochemical products using microorganisms is receiving attention as a sustainable alternative to existing methods of productions. In order to synthesize various chemical substances, materials, and fuel using microorganisms, it is necessary to first construct the biosynthetic pathway toward desired product by exploration and discovery and introduce them into microorganisms. In addition, in order to efficiently synthesize various chemical substances, it is sometimes necessary to employ chemical methods along with bioengineering methods using microorganisms at the same time. For the production of non-native chemicals, novel pathways are designed by recruiting enzymes from heterologous sources or employing enzymes designed though rational engineering, directed evolution, or ab initio design.
The research team had completed a map of chemicals which compiled all available pathways of biological and/or chemical reactions that lead to the production of various bio-based chemicals back in 2019 and published the map in Nature Catalysis. The map was distributed in the form of a poster to industries and academia so that the synthesis paths of bio-based chemicals could be checked at a glance.
The research team has expanded the bio-based chemicals map this time in the form of an interactive map on the web so that anyone with internet access can quickly explore efficient paths to synthesize desired products. The web-based map provides interactive visual tools to allow interactive visualization, exploration, and analysis of complex networks of biological and/or chemical reactions toward the desired products. In addition, the reported paper also discusses the production of natural compounds that are used for diverse purposes such as food and medicine, which will help designing novel pathways through similar approaches or by exploiting the promiscuity of enzymes described in the map. The published bio-based chemicals map is also available at http://systemsbiotech.co.kr.
The co-first authors, Dr. Woo Dae Jang and Ph.D. student Gi Bae Kim, said, “We conducted this study to address the demand for updating the previously distributed chemicals map and enhancing its versatility.” “The map is expected to be utilized in a variety of research and in efforts to set strategies and prospects for chemical production incorporating bio and chemical methods that are detailed in the map.”
Distinguished Professor Sang Yup Lee said, “The interactive bio-based chemicals map is expected to help design and optimization of the metabolic pathways for the biosynthesis of target chemicals together with the strategies of chemical conversions, serving as a blueprint for developing further ideas on the production of desired chemicals through biological and/or chemical reactions.”
The interactive metabolic map of bio-based chemicals.
2022.08.11 View 16796
Interactive Map of Metabolical Synthesis of Chemicals
An interactive map that compiled the chemicals produced by biological, chemical and combined reactions has been distributed on the web
- A team led by Distinguished Professor Sang Yup Lee of the Department of Chemical and Biomolecular Engineering, organized and distributed an all-inclusive listing of chemical substances that can be synthesized using microorganisms
- It is expected to be used by researchers around the world as it enables easy assessment of the synthetic pathway through the web.
A research team comprised of Woo Dae Jang, Gi Bae Kim, and Distinguished Professor Sang Yup Lee of the Department of Chemical and Biomolecular Engineering at KAIST reported an interactive metabolic map of bio-based chemicals. Their research paper “An interactive metabolic map of bio-based chemicals” was published online in Trends in Biotechnology on August 10, 2022.
As a response to rapid climate change and environmental pollution, research on the production of petrochemical products using microorganisms is receiving attention as a sustainable alternative to existing methods of productions. In order to synthesize various chemical substances, materials, and fuel using microorganisms, it is necessary to first construct the biosynthetic pathway toward desired product by exploration and discovery and introduce them into microorganisms. In addition, in order to efficiently synthesize various chemical substances, it is sometimes necessary to employ chemical methods along with bioengineering methods using microorganisms at the same time. For the production of non-native chemicals, novel pathways are designed by recruiting enzymes from heterologous sources or employing enzymes designed though rational engineering, directed evolution, or ab initio design.
The research team had completed a map of chemicals which compiled all available pathways of biological and/or chemical reactions that lead to the production of various bio-based chemicals back in 2019 and published the map in Nature Catalysis. The map was distributed in the form of a poster to industries and academia so that the synthesis paths of bio-based chemicals could be checked at a glance.
The research team has expanded the bio-based chemicals map this time in the form of an interactive map on the web so that anyone with internet access can quickly explore efficient paths to synthesize desired products. The web-based map provides interactive visual tools to allow interactive visualization, exploration, and analysis of complex networks of biological and/or chemical reactions toward the desired products. In addition, the reported paper also discusses the production of natural compounds that are used for diverse purposes such as food and medicine, which will help designing novel pathways through similar approaches or by exploiting the promiscuity of enzymes described in the map. The published bio-based chemicals map is also available at http://systemsbiotech.co.kr.
The co-first authors, Dr. Woo Dae Jang and Ph.D. student Gi Bae Kim, said, “We conducted this study to address the demand for updating the previously distributed chemicals map and enhancing its versatility.” “The map is expected to be utilized in a variety of research and in efforts to set strategies and prospects for chemical production incorporating bio and chemical methods that are detailed in the map.”
Distinguished Professor Sang Yup Lee said, “The interactive bio-based chemicals map is expected to help design and optimization of the metabolic pathways for the biosynthesis of target chemicals together with the strategies of chemical conversions, serving as a blueprint for developing further ideas on the production of desired chemicals through biological and/or chemical reactions.”
The interactive metabolic map of bio-based chemicals.
2022.08.11 View 16796 -
 A New Therapeutic Drug for Alzheimer’s Disease without Inflammatory Side Effects
Although Aduhelm, a monoclonal antibody targeting amyloid beta (Aβ), recently became the first US FDA approved drug for Alzheimer’s disease (AD) based on its ability to decrease Aβ plaque burden in AD patients, its effect on cognitive improvement is still controversial. Moreover, about 40% of the patients treated with this antibody experienced serious side effects including cerebral edemas (ARIA-E) and hemorrhages (ARIA-H) that are likely related to inflammatory responses in the brain when the Aβ antibody binds Fc receptors (FCR) of immune cells such as microglia and macrophages.
These inflammatory side effects can cause neuronal cell death and synapse elimination by activated microglia, and even have the potential to exacerbate cognitive impairment in AD patients. Thus, current Aβ antibody-based immunotherapy holds the inherent risk of doing more harm than good due to their inflammatory side effects.
To overcome these problems, a team of researchers at KAIST in South Korea has developed a novel fusion protein drug, αAβ-Gas6, which efficiently eliminates Aβ via an entirely different mechanism than Aβ antibody-based immunotherapy. In a mouse model of AD, αAβ-Gas6 not only removed Aβ with higher potency, but also circumvented the neurotoxic inflammatory side effects associated with conventional antibody treatments.
Their findings were published on August 4 in Nature Medicine.
Schematic of a chimeric Gas6 fusion protein. A single chain variable fragment (scFv) of
an Amyloid β (Aβ)-targeting monoclonal antibody is fused with a truncated receptor binding
domain of Gas6, a bridging molecule for the clearance of dead cells via TAM (TYRO3, AXL,
and MERTK) receptors, which are expressed by microglia and astrocytes.
“FcR activation by Aβ targeting antibodies induces microglia-mediated Aβ phagocytosis, but it also produces inflammatory signals, inevitably damaging brain tissues,” said paper authors Chan Hyuk Kim and Won-Suk Chung, associate professors in the Department of Biological Sciences at KAIST.
“Therefore, we utilized efferocytosis, a cellular process by which dead cells are removed by phagocytes as an alternative pathway for the clearance of Aβ in the brain,” Prof. Kim and Chung said. “Efferocytosis is accompanied by anti-inflammatory responses to maintain tissue homeostasis. To exploit this process, we engineered Gas6, a soluble adaptor protein that mediates efferocytosis via TAM phagocytic receptors in such a way that its target specificity was redirected from dead cells to Aβ plaques.”
The professors and their team demonstrated that the resulting αAβ-Gas6 induced Aβ engulfment by activating not only microglial but also astrocytic phagocytosis since TAM phagocytic receptors are highly expressed by these two major phagocytes in the brain. Importantly, αAβ-Gas6 promoted the robust uptake of Aβ without showing any signs of inflammation and neurotoxicity, which contrasts sharply with the treatment using an Aβ monoclonal antibody. Moreover, they showed that αAβ-Gas6 substantially reduced excessive synapse elimination by microglia, consequently leading to better behavioral rescues in AD model mice.
“By using a mouse model of cerebral amyloid angiopathy (CAA), a cerebrovascular disorder caused by the deposition of Aβ within the walls of the brain’s blood vessels, we also showed that the intrathecal administration of Gas6 fusion protein significantly eliminated cerebrovascular amyloids, along with a reduction of microhemorrhages. These data demonstrate that aAb-Gas6 is a potent therapeutic agent in eliminating Aβ without exacerbating CAA-related microhemorrhages.”
The resulting αAβ-Gas6 clears Aβ oligomers and fibrils without causing neurotoxicity (a-b, neurons: red, and fragmented axons: yellow) and proinflammatory responses (c, TNF release), which are conversely exacerbated by the treatment of an Aβ-targeting monoclonal antibody (Aducanumab).
Professors Kim and Chung noted, “We believe our approach can be a breakthrough in treating AD without causing inflammatory side effects and synapse loss. Our approach holds promise as a novel therapeutic platform that is applicable to more than AD. By modifying the target-specificity of the fusion protein, the Gas6-fusion protein can be applied to various neurological disorders as well as autoimmune diseases affected by toxic molecules that should be removed without causing inflammatory responses.”
The number and total area of Aβ plaques (Thioflavin-T, green) were significantly reduced in αAβ-Gas6-treated AD mouse brains compared to Aducanumab-treated ones (a, b). The cognitive functions of AD model mice were significantly rescued by αAβ-Gas6 treatment, whereas Aducanumab-treated AD mice showed partial rescue in these cognitive tests (c-e).
Professors Kim and Chung founded “Illimis Therapeutics” based on this strategy of designing chimeric Gas6 fusion proteins that would remove toxic aggregates from the nervous system. Through this company, they are planning to further develop various Gas6-fusion proteins not only for Ab but also for Tau to treat AD symptoms.
This work was supported by KAIST and the Korea Health Technology R&D Project that was administered by the Korea Health Industry Development Institute (KHIDI) and the Korea Dementia Research Center (KDRC) funded by the Ministry of Health & Welfare (MOHW) and the Ministry of Science and ICT (MSIT), and KAIST.
Other contributors include Hyuncheol Jung and Se Young Lee, Sungjoon Lim, Hyeong Ryeol Choi, Yeseong Choi, Minjin Kim, Segi Kim, the Department of Biological Sciences, and the Korea Advanced Institute of Science and Technology (KAIST).
To receive more up-to-date information on this new development, follow “Illimis Therapeutics” on twitter @Illimistx.
2022.08.05 View 13648
A New Therapeutic Drug for Alzheimer’s Disease without Inflammatory Side Effects
Although Aduhelm, a monoclonal antibody targeting amyloid beta (Aβ), recently became the first US FDA approved drug for Alzheimer’s disease (AD) based on its ability to decrease Aβ plaque burden in AD patients, its effect on cognitive improvement is still controversial. Moreover, about 40% of the patients treated with this antibody experienced serious side effects including cerebral edemas (ARIA-E) and hemorrhages (ARIA-H) that are likely related to inflammatory responses in the brain when the Aβ antibody binds Fc receptors (FCR) of immune cells such as microglia and macrophages.
These inflammatory side effects can cause neuronal cell death and synapse elimination by activated microglia, and even have the potential to exacerbate cognitive impairment in AD patients. Thus, current Aβ antibody-based immunotherapy holds the inherent risk of doing more harm than good due to their inflammatory side effects.
To overcome these problems, a team of researchers at KAIST in South Korea has developed a novel fusion protein drug, αAβ-Gas6, which efficiently eliminates Aβ via an entirely different mechanism than Aβ antibody-based immunotherapy. In a mouse model of AD, αAβ-Gas6 not only removed Aβ with higher potency, but also circumvented the neurotoxic inflammatory side effects associated with conventional antibody treatments.
Their findings were published on August 4 in Nature Medicine.
Schematic of a chimeric Gas6 fusion protein. A single chain variable fragment (scFv) of
an Amyloid β (Aβ)-targeting monoclonal antibody is fused with a truncated receptor binding
domain of Gas6, a bridging molecule for the clearance of dead cells via TAM (TYRO3, AXL,
and MERTK) receptors, which are expressed by microglia and astrocytes.
“FcR activation by Aβ targeting antibodies induces microglia-mediated Aβ phagocytosis, but it also produces inflammatory signals, inevitably damaging brain tissues,” said paper authors Chan Hyuk Kim and Won-Suk Chung, associate professors in the Department of Biological Sciences at KAIST.
“Therefore, we utilized efferocytosis, a cellular process by which dead cells are removed by phagocytes as an alternative pathway for the clearance of Aβ in the brain,” Prof. Kim and Chung said. “Efferocytosis is accompanied by anti-inflammatory responses to maintain tissue homeostasis. To exploit this process, we engineered Gas6, a soluble adaptor protein that mediates efferocytosis via TAM phagocytic receptors in such a way that its target specificity was redirected from dead cells to Aβ plaques.”
The professors and their team demonstrated that the resulting αAβ-Gas6 induced Aβ engulfment by activating not only microglial but also astrocytic phagocytosis since TAM phagocytic receptors are highly expressed by these two major phagocytes in the brain. Importantly, αAβ-Gas6 promoted the robust uptake of Aβ without showing any signs of inflammation and neurotoxicity, which contrasts sharply with the treatment using an Aβ monoclonal antibody. Moreover, they showed that αAβ-Gas6 substantially reduced excessive synapse elimination by microglia, consequently leading to better behavioral rescues in AD model mice.
“By using a mouse model of cerebral amyloid angiopathy (CAA), a cerebrovascular disorder caused by the deposition of Aβ within the walls of the brain’s blood vessels, we also showed that the intrathecal administration of Gas6 fusion protein significantly eliminated cerebrovascular amyloids, along with a reduction of microhemorrhages. These data demonstrate that aAb-Gas6 is a potent therapeutic agent in eliminating Aβ without exacerbating CAA-related microhemorrhages.”
The resulting αAβ-Gas6 clears Aβ oligomers and fibrils without causing neurotoxicity (a-b, neurons: red, and fragmented axons: yellow) and proinflammatory responses (c, TNF release), which are conversely exacerbated by the treatment of an Aβ-targeting monoclonal antibody (Aducanumab).
Professors Kim and Chung noted, “We believe our approach can be a breakthrough in treating AD without causing inflammatory side effects and synapse loss. Our approach holds promise as a novel therapeutic platform that is applicable to more than AD. By modifying the target-specificity of the fusion protein, the Gas6-fusion protein can be applied to various neurological disorders as well as autoimmune diseases affected by toxic molecules that should be removed without causing inflammatory responses.”
The number and total area of Aβ plaques (Thioflavin-T, green) were significantly reduced in αAβ-Gas6-treated AD mouse brains compared to Aducanumab-treated ones (a, b). The cognitive functions of AD model mice were significantly rescued by αAβ-Gas6 treatment, whereas Aducanumab-treated AD mice showed partial rescue in these cognitive tests (c-e).
Professors Kim and Chung founded “Illimis Therapeutics” based on this strategy of designing chimeric Gas6 fusion proteins that would remove toxic aggregates from the nervous system. Through this company, they are planning to further develop various Gas6-fusion proteins not only for Ab but also for Tau to treat AD symptoms.
This work was supported by KAIST and the Korea Health Technology R&D Project that was administered by the Korea Health Industry Development Institute (KHIDI) and the Korea Dementia Research Center (KDRC) funded by the Ministry of Health & Welfare (MOHW) and the Ministry of Science and ICT (MSIT), and KAIST.
Other contributors include Hyuncheol Jung and Se Young Lee, Sungjoon Lim, Hyeong Ryeol Choi, Yeseong Choi, Minjin Kim, Segi Kim, the Department of Biological Sciences, and the Korea Advanced Institute of Science and Technology (KAIST).
To receive more up-to-date information on this new development, follow “Illimis Therapeutics” on twitter @Illimistx.
2022.08.05 View 13648 -
 Metabolically Engineered Bacterium Produces Lutein
A research group at KAIST has engineered a bacterial strain capable of producing lutein. The research team applied systems metabolic engineering strategies, including substrate channeling and electron channeling, to enhance the production of lutein in an engineered Escherichia coli strain. The strategies will be also useful for the efficient production of other industrially important natural products used in the food, pharmaceutical, and cosmetic industries.
Figure: Systems metabolic engineering was employed to construct and optimize the metabolic pathways for lutein production, and substrate channeling and electron channeling strategies were additionally employed to increase the production of the lutein with high productivity.
Lutein is classified as a xanthophyll chemical that is abundant in egg yolk, fruits, and vegetables. It protects the eye from oxidative damage from radiation and reduces the risk of eye diseases including macular degeneration and cataracts. Commercialized products featuring lutein are derived from the extracts of the marigold flower, which is known to harbor abundant amounts of lutein. However, the drawback of lutein production from nature is that it takes a long time to grow and harvest marigold flowers. Furthermore, it requires additional physical and chemical-based extractions with a low yield, which makes it economically unfeasible in terms of productivity. The high cost and low yield of these bioprocesses has made it difficult to readily meet the demand for lutein.
These challenges inspired the metabolic engineers at KAIST, including researchers Dr. Seon Young Park, Ph.D. Candidate Hyunmin Eun, and Distinguished Professor Sang Yup Lee from the Department of Chemical and Biomolecular Engineering. The team’s study entitled “Metabolic engineering of Escherichia coli with electron channeling for the production of natural products” was published in Nature Catalysis on August 5, 2022.
This research details the ability to produce lutein from E. coli with a high yield using a cheap carbon source, glycerol, via systems metabolic engineering. The research group focused on solving the bottlenecks of the biosynthetic pathway for lutein production constructed within an individual cell. First, using systems metabolic engineering, which is an integrated technology to engineer the metabolism of a microorganism, lutein was produced when the lutein biosynthesis pathway was introduced, albeit in very small amounts.
To improve the productivity of lutein production, the bottleneck enzymes within the metabolic pathway were first identified. It turned out that metabolic reactions that involve a promiscuous enzyme, an enzyme that is involved in two or more metabolic reactions, and electron-requiring cytochrome P450 enzymes are the main bottleneck steps of the pathway inhibiting lutein biosynthesis.
To overcome these challenges, substrate channeling, a strategy to artificially recruit enzymes in physical proximity within the cell in order to increase the local concentrations of substrates that can be converted into products, was employed to channel more metabolic flux towards the target chemical while reducing the formation of unwanted byproducts.
Furthermore, electron channeling, a strategy similar to substrate channeling but differing in terms of increasing the local concentrations of electrons required for oxidoreduction reactions mediated by P450 and its reductase partners, was applied to further streamline the metabolic flux towards lutein biosynthesis, which led to the highest titer of lutein production achieved in a bacterial host ever reported. The same electron channeling strategy was successfully applied for the production of other natural products including nootkatone and apigenin in E. coli, showcasing the general applicability of the strategy in the research field.
“It is expected that this microbial cell factory-based production of lutein will be able to replace the current plant extraction-based process,” said Dr. Seon Young Park, the first author of the paper. She explained that another important point of the research is that integrated metabolic engineering strategies developed from this study can be generally applicable for the efficient production of other natural products useful as pharmaceuticals or nutraceuticals.
“As maintaining good health in an aging society is becoming increasingly important, we expect that the technology and strategies developed here will play pivotal roles in producing other valuable natural products of medical or nutritional importance,” explained Distinguished Professor Sang Yup Lee.
This work was supported by the Cooperative Research Program for Agriculture Science & Technology Development funded by the Rural Development Administration of Korea, with further support from the Development of Next-generation Biorefinery Platform Technologies for Leading Bio-based Chemicals Industry Project and by the Development of Platform Technologies of Microbial Cell Factories for the Next-generation Biorefineries Project of the National Research Foundation funded by the Ministry of Science and ICT of Korea.
2022.08.05 View 11346
Metabolically Engineered Bacterium Produces Lutein
A research group at KAIST has engineered a bacterial strain capable of producing lutein. The research team applied systems metabolic engineering strategies, including substrate channeling and electron channeling, to enhance the production of lutein in an engineered Escherichia coli strain. The strategies will be also useful for the efficient production of other industrially important natural products used in the food, pharmaceutical, and cosmetic industries.
Figure: Systems metabolic engineering was employed to construct and optimize the metabolic pathways for lutein production, and substrate channeling and electron channeling strategies were additionally employed to increase the production of the lutein with high productivity.
Lutein is classified as a xanthophyll chemical that is abundant in egg yolk, fruits, and vegetables. It protects the eye from oxidative damage from radiation and reduces the risk of eye diseases including macular degeneration and cataracts. Commercialized products featuring lutein are derived from the extracts of the marigold flower, which is known to harbor abundant amounts of lutein. However, the drawback of lutein production from nature is that it takes a long time to grow and harvest marigold flowers. Furthermore, it requires additional physical and chemical-based extractions with a low yield, which makes it economically unfeasible in terms of productivity. The high cost and low yield of these bioprocesses has made it difficult to readily meet the demand for lutein.
These challenges inspired the metabolic engineers at KAIST, including researchers Dr. Seon Young Park, Ph.D. Candidate Hyunmin Eun, and Distinguished Professor Sang Yup Lee from the Department of Chemical and Biomolecular Engineering. The team’s study entitled “Metabolic engineering of Escherichia coli with electron channeling for the production of natural products” was published in Nature Catalysis on August 5, 2022.
This research details the ability to produce lutein from E. coli with a high yield using a cheap carbon source, glycerol, via systems metabolic engineering. The research group focused on solving the bottlenecks of the biosynthetic pathway for lutein production constructed within an individual cell. First, using systems metabolic engineering, which is an integrated technology to engineer the metabolism of a microorganism, lutein was produced when the lutein biosynthesis pathway was introduced, albeit in very small amounts.
To improve the productivity of lutein production, the bottleneck enzymes within the metabolic pathway were first identified. It turned out that metabolic reactions that involve a promiscuous enzyme, an enzyme that is involved in two or more metabolic reactions, and electron-requiring cytochrome P450 enzymes are the main bottleneck steps of the pathway inhibiting lutein biosynthesis.
To overcome these challenges, substrate channeling, a strategy to artificially recruit enzymes in physical proximity within the cell in order to increase the local concentrations of substrates that can be converted into products, was employed to channel more metabolic flux towards the target chemical while reducing the formation of unwanted byproducts.
Furthermore, electron channeling, a strategy similar to substrate channeling but differing in terms of increasing the local concentrations of electrons required for oxidoreduction reactions mediated by P450 and its reductase partners, was applied to further streamline the metabolic flux towards lutein biosynthesis, which led to the highest titer of lutein production achieved in a bacterial host ever reported. The same electron channeling strategy was successfully applied for the production of other natural products including nootkatone and apigenin in E. coli, showcasing the general applicability of the strategy in the research field.
“It is expected that this microbial cell factory-based production of lutein will be able to replace the current plant extraction-based process,” said Dr. Seon Young Park, the first author of the paper. She explained that another important point of the research is that integrated metabolic engineering strategies developed from this study can be generally applicable for the efficient production of other natural products useful as pharmaceuticals or nutraceuticals.
“As maintaining good health in an aging society is becoming increasingly important, we expect that the technology and strategies developed here will play pivotal roles in producing other valuable natural products of medical or nutritional importance,” explained Distinguished Professor Sang Yup Lee.
This work was supported by the Cooperative Research Program for Agriculture Science & Technology Development funded by the Rural Development Administration of Korea, with further support from the Development of Next-generation Biorefinery Platform Technologies for Leading Bio-based Chemicals Industry Project and by the Development of Platform Technologies of Microbial Cell Factories for the Next-generation Biorefineries Project of the National Research Foundation funded by the Ministry of Science and ICT of Korea.
2022.08.05 View 11346 -
 VP Sang Yup Lee Receives Honorary Doctorate from DTU
Vice President for Research, Distinguished Professor Sang Yup Lee at the Department of Chemical & Biomolecular Engineering, was awarded an honorary doctorate from the Technical University of Denmark (DTU) during the DTU Commemoration Day 2022 on April 29. The event drew distinguished guests, students, and faculty including HRH The Crown Prince Frederik Andre Henrik Christian and DTU President Anders Bjarklev.
Professor Lee was recognized for his exceptional scholarship in the field of systems metabolic engineering, which led to the development of microcell factories capable of producing a wide range of fuels, chemicals, materials, and natural compounds, many for the first time.
Professor Lee said in his acceptance speech that KAIST’s continued partnership with DTU in the field of biotechnology will lead to significant contributions in the global efforts to respond to climate change and promote green growth.
DTU CPO and CSO Dina Petronovic Nielson, who heads DTU Biosustain, also lauded Professor Lee saying, “It is not only a great honor for Professor Lee to be induced at DTU but also great honor for DTU to have him.”
Professor Lee also gave commemorative lectures at DTU Biosustain in Lingby and the Bio Innovation Research Institute at the Novo Nordisk Foundation in Copenhagen while in Denmark.
DTU, one of the leading science and technology universities in Europe, has been awarding honorary doctorates since 1921, including to Nobel laureate in chemistry Professor Frances Arnold at Caltech. Professor Lee is the first Korean to receive an honorary doctorate from DTU.
2022.05.03 View 12008
VP Sang Yup Lee Receives Honorary Doctorate from DTU
Vice President for Research, Distinguished Professor Sang Yup Lee at the Department of Chemical & Biomolecular Engineering, was awarded an honorary doctorate from the Technical University of Denmark (DTU) during the DTU Commemoration Day 2022 on April 29. The event drew distinguished guests, students, and faculty including HRH The Crown Prince Frederik Andre Henrik Christian and DTU President Anders Bjarklev.
Professor Lee was recognized for his exceptional scholarship in the field of systems metabolic engineering, which led to the development of microcell factories capable of producing a wide range of fuels, chemicals, materials, and natural compounds, many for the first time.
Professor Lee said in his acceptance speech that KAIST’s continued partnership with DTU in the field of biotechnology will lead to significant contributions in the global efforts to respond to climate change and promote green growth.
DTU CPO and CSO Dina Petronovic Nielson, who heads DTU Biosustain, also lauded Professor Lee saying, “It is not only a great honor for Professor Lee to be induced at DTU but also great honor for DTU to have him.”
Professor Lee also gave commemorative lectures at DTU Biosustain in Lingby and the Bio Innovation Research Institute at the Novo Nordisk Foundation in Copenhagen while in Denmark.
DTU, one of the leading science and technology universities in Europe, has been awarding honorary doctorates since 1921, including to Nobel laureate in chemistry Professor Frances Arnold at Caltech. Professor Lee is the first Korean to receive an honorary doctorate from DTU.
2022.05.03 View 12008 -
 Decoding Brain Signals to Control a Robotic Arm
Advanced brain-machine interface system successfully interprets arm movement directions from neural signals in the brain
Researchers have developed a mind-reading system for decoding neural signals from the brain during arm movement. The method, described in the journal Applied Soft Computing, can be used by a person to control a robotic arm through a brain-machine interface (BMI).
A BMI is a device that translates nerve signals into commands to control a machine, such as a computer or a robotic limb. There are two main techniques for monitoring neural signals in BMIs: electroencephalography (EEG) and electrocorticography (ECoG).
The EEG exhibits signals from electrodes on the surface of the scalp and is widely employed because it is non-invasive, relatively cheap, safe and easy to use. However, the EEG has low spatial resolution and detects irrelevant neural signals, which makes it difficult to interpret the intentions of individuals from the EEG.
On the other hand, the ECoG is an invasive method that involves placing electrodes directly on the surface of the cerebral cortex below the scalp. Compared with the EEG, the ECoG can monitor neural signals with much higher spatial resolution and less background noise. However, this technique has several drawbacks.
“The ECoG is primarily used to find potential sources of epileptic seizures, meaning the electrodes are placed in different locations for different patients and may not be in the optimal regions of the brain for detecting sensory and movement signals,” explained Professor Jaeseung Jeong, a brain scientist at KAIST. “This inconsistency makes it difficult to decode brain signals to predict movements.”
To overcome these problems, Professor Jeong’s team developed a new method for decoding ECoG neural signals during arm movement. The system is based on a machine-learning system for analysing and predicting neural signals called an ‘echo-state network’ and a mathematical probability model called the Gaussian distribution.
In the study, the researchers recorded ECoG signals from four individuals with epilepsy while they were performing a reach-and-grasp task. Because the ECoG electrodes were placed according to the potential sources of each patient’s epileptic seizures, only 22% to 44% of the electrodes were located in the regions of the brain responsible for controlling movement.
During the movement task, the participants were given visual cues, either by placing a real tennis ball in front of them, or via a virtual reality headset showing a clip of a human arm reaching forward in first-person view. They were asked to reach forward, grasp an object, then return their hand and release the object, while wearing motion sensors on their wrists and fingers. In a second task, they were instructed to imagine reaching forward without moving their arms.
The researchers monitored the signals from the ECoG electrodes during real and imaginary arm movements, and tested whether the new system could predict the direction of this movement from the neural signals. They found that the novel decoder successfully classified arm movements in 24 directions in three-dimensional space, both in the real and virtual tasks, and that the results were at least five times more accurate than chance. They also used a computer simulation to show that the novel ECoG decoder could control the movements of a robotic arm.
Overall, the results suggest that the new machine learning-based BCI system successfully used ECoG signals to interpret the direction of the intended movements. The next steps will be to improve the accuracy and efficiency of the decoder. In the future, it could be used in a real-time BMI device to help people with movement or sensory impairments.
This research was supported by the KAIST Global Singularity Research Program of 2021, Brain Research Program of the National Research Foundation of Korea funded by the Ministry of Science, ICT, and Future Planning, and the Basic Science Research Program through the National Research Foundation of Korea funded by the Ministry of Education.
-PublicationHoon-Hee Kim, Jaeseung Jeong, “An electrocorticographic decoder for arm movement for brain-machine interface using an echo state network and Gaussian readout,” Applied SoftComputing online December 31, 2021 (doi.org/10.1016/j.asoc.2021.108393)
-ProfileProfessor Jaeseung JeongDepartment of Bio and Brain EngineeringCollege of EngineeringKAIST
2022.03.18 View 13633
Decoding Brain Signals to Control a Robotic Arm
Advanced brain-machine interface system successfully interprets arm movement directions from neural signals in the brain
Researchers have developed a mind-reading system for decoding neural signals from the brain during arm movement. The method, described in the journal Applied Soft Computing, can be used by a person to control a robotic arm through a brain-machine interface (BMI).
A BMI is a device that translates nerve signals into commands to control a machine, such as a computer or a robotic limb. There are two main techniques for monitoring neural signals in BMIs: electroencephalography (EEG) and electrocorticography (ECoG).
The EEG exhibits signals from electrodes on the surface of the scalp and is widely employed because it is non-invasive, relatively cheap, safe and easy to use. However, the EEG has low spatial resolution and detects irrelevant neural signals, which makes it difficult to interpret the intentions of individuals from the EEG.
On the other hand, the ECoG is an invasive method that involves placing electrodes directly on the surface of the cerebral cortex below the scalp. Compared with the EEG, the ECoG can monitor neural signals with much higher spatial resolution and less background noise. However, this technique has several drawbacks.
“The ECoG is primarily used to find potential sources of epileptic seizures, meaning the electrodes are placed in different locations for different patients and may not be in the optimal regions of the brain for detecting sensory and movement signals,” explained Professor Jaeseung Jeong, a brain scientist at KAIST. “This inconsistency makes it difficult to decode brain signals to predict movements.”
To overcome these problems, Professor Jeong’s team developed a new method for decoding ECoG neural signals during arm movement. The system is based on a machine-learning system for analysing and predicting neural signals called an ‘echo-state network’ and a mathematical probability model called the Gaussian distribution.
In the study, the researchers recorded ECoG signals from four individuals with epilepsy while they were performing a reach-and-grasp task. Because the ECoG electrodes were placed according to the potential sources of each patient’s epileptic seizures, only 22% to 44% of the electrodes were located in the regions of the brain responsible for controlling movement.
During the movement task, the participants were given visual cues, either by placing a real tennis ball in front of them, or via a virtual reality headset showing a clip of a human arm reaching forward in first-person view. They were asked to reach forward, grasp an object, then return their hand and release the object, while wearing motion sensors on their wrists and fingers. In a second task, they were instructed to imagine reaching forward without moving their arms.
The researchers monitored the signals from the ECoG electrodes during real and imaginary arm movements, and tested whether the new system could predict the direction of this movement from the neural signals. They found that the novel decoder successfully classified arm movements in 24 directions in three-dimensional space, both in the real and virtual tasks, and that the results were at least five times more accurate than chance. They also used a computer simulation to show that the novel ECoG decoder could control the movements of a robotic arm.
Overall, the results suggest that the new machine learning-based BCI system successfully used ECoG signals to interpret the direction of the intended movements. The next steps will be to improve the accuracy and efficiency of the decoder. In the future, it could be used in a real-time BMI device to help people with movement or sensory impairments.
This research was supported by the KAIST Global Singularity Research Program of 2021, Brain Research Program of the National Research Foundation of Korea funded by the Ministry of Science, ICT, and Future Planning, and the Basic Science Research Program through the National Research Foundation of Korea funded by the Ministry of Education.
-PublicationHoon-Hee Kim, Jaeseung Jeong, “An electrocorticographic decoder for arm movement for brain-machine interface using an echo state network and Gaussian readout,” Applied SoftComputing online December 31, 2021 (doi.org/10.1016/j.asoc.2021.108393)
-ProfileProfessor Jaeseung JeongDepartment of Bio and Brain EngineeringCollege of EngineeringKAIST
2022.03.18 View 13633 -
 Five Projects Ranked in the Top 100 for National R&D Excellence
Five KAIST research projects were selected as the 2021 Top 100 for National R&D Excellence by the Ministry of Science and ICT and the Korea Institute of Science & Technology Evaluation and Planning.
The five projects are:-The development of E. coli that proliferates with only formic acid and carbon dioxide by Distinguished Professor Sang Yup Lee from the Department of Chemical and Biomolecular Engineering
-An original reverse aging technology that restores an old human skin cell into a younger one by Professor Kwang-Hyun Cho from the Department of Bio and Brain Engineering-The development of next-generation high-efficiency perovskite-silicon tandem solar cells by Professor Byungha Shin from the Department of Materials Science and Engineering-Research on the effects of ultrafine dust in the atmosphere has on energy consumption by Professor Jiyong Eom from the School of Business and Technology Management-Research on a molecular trigger that controls the phase transformation of bio materials by Professor Myungchul Kim from the Department of Bio and Brain Engineering
Started in 2006, an Evaluation Committee composed of experts in industries, universities, and research institutes has made the preliminary selections of the most outstanding research projects based on their significance as a scientific and technological development and their socioeconomic effects. The finalists went through an open public evaluation. The final 100 studies are from six fields: 18 from mechanics & materials, 26 from biology & marine sciences, 19 from ICT & electronics, 10 from interdisciplinary research, and nine from natural science and infrastructure.
The selected 100 studies will receive a certificate and an award plaque from the minister of MSIT as well as additional points for business and institutional evaluations according to appropriate regulations, and the selected researchers will be strongly recommended as candidates for national meritorious awards.
In particular, to help the 100 selected research projects become more accessible for the general public, their main contents will be provided in a free e-book ‘The Top 100 for National R&D Excellence of 2021’ that will be available from online booksellers.
2022.02.17 View 12010
Five Projects Ranked in the Top 100 for National R&D Excellence
Five KAIST research projects were selected as the 2021 Top 100 for National R&D Excellence by the Ministry of Science and ICT and the Korea Institute of Science & Technology Evaluation and Planning.
The five projects are:-The development of E. coli that proliferates with only formic acid and carbon dioxide by Distinguished Professor Sang Yup Lee from the Department of Chemical and Biomolecular Engineering
-An original reverse aging technology that restores an old human skin cell into a younger one by Professor Kwang-Hyun Cho from the Department of Bio and Brain Engineering-The development of next-generation high-efficiency perovskite-silicon tandem solar cells by Professor Byungha Shin from the Department of Materials Science and Engineering-Research on the effects of ultrafine dust in the atmosphere has on energy consumption by Professor Jiyong Eom from the School of Business and Technology Management-Research on a molecular trigger that controls the phase transformation of bio materials by Professor Myungchul Kim from the Department of Bio and Brain Engineering
Started in 2006, an Evaluation Committee composed of experts in industries, universities, and research institutes has made the preliminary selections of the most outstanding research projects based on their significance as a scientific and technological development and their socioeconomic effects. The finalists went through an open public evaluation. The final 100 studies are from six fields: 18 from mechanics & materials, 26 from biology & marine sciences, 19 from ICT & electronics, 10 from interdisciplinary research, and nine from natural science and infrastructure.
The selected 100 studies will receive a certificate and an award plaque from the minister of MSIT as well as additional points for business and institutional evaluations according to appropriate regulations, and the selected researchers will be strongly recommended as candidates for national meritorious awards.
In particular, to help the 100 selected research projects become more accessible for the general public, their main contents will be provided in a free e-book ‘The Top 100 for National R&D Excellence of 2021’ that will be available from online booksellers.
2022.02.17 View 12010 -
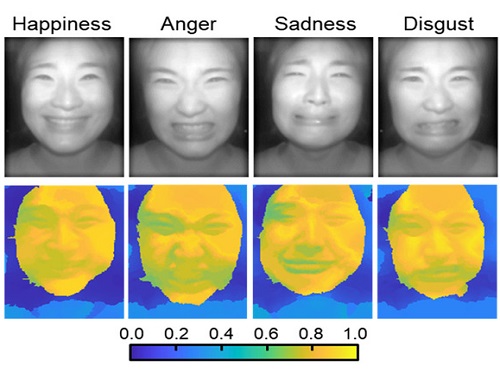 AI Light-Field Camera Reads 3D Facial Expressions
Machine-learned, light-field camera reads facial expressions from high-contrast illumination invariant 3D facial images
A joint research team led by Professors Ki-Hun Jeong and Doheon Lee from the KAIST Department of Bio and Brain Engineering reported the development of a technique for facial expression detection by merging near-infrared light-field camera techniques with artificial intelligence (AI) technology.
Unlike a conventional camera, the light-field camera contains micro-lens arrays in front of the image sensor, which makes the camera small enough to fit into a smart phone, while allowing it to acquire the spatial and directional information of the light with a single shot. The technique has received attention as it can reconstruct images in a variety of ways including multi-views, refocusing, and 3D image acquisition, giving rise to many potential applications.
However, the optical crosstalk between shadows caused by external light sources in the environment and the micro-lens has limited existing light-field cameras from being able to provide accurate image contrast and 3D reconstruction.
The joint research team applied a vertical-cavity surface-emitting laser (VCSEL) in the near-IR range to stabilize the accuracy of 3D image reconstruction that previously depended on environmental light. When an external light source is shone on a face at 0-, 30-, and 60-degree angles, the light field camera reduces 54% of image reconstruction errors. Additionally, by inserting a light-absorbing layer for visible and near-IR wavelengths between the micro-lens arrays, the team could minimize optical crosstalk while increasing the image contrast by 2.1 times.
Through this technique, the team could overcome the limitations of existing light-field cameras and was able to develop their NIR-based light-field camera (NIR-LFC), optimized for the 3D image reconstruction of facial expressions. Using the NIR-LFC, the team acquired high-quality 3D reconstruction images of facial expressions expressing various emotions regardless of the lighting conditions of the surrounding environment.
The facial expressions in the acquired 3D images were distinguished through machine learning with an average of 85% accuracy – a statistically significant figure compared to when 2D images were used. Furthermore, by calculating the interdependency of distance information that varies with facial expression in 3D images, the team could identify the information a light-field camera utilizes to distinguish human expressions.
Professor Ki-Hun Jeong said, “The sub-miniature light-field camera developed by the research team has the potential to become the new platform to quantitatively analyze the facial expressions and emotions of humans.” To highlight the significance of this research, he added, “It could be applied in various fields including mobile healthcare, field diagnosis, social cognition, and human-machine interactions.”
This research was published in Advanced Intelligent Systems online on December 16, under the title, “Machine-Learned Light-field Camera that Reads Facial Expression from High-Contrast and Illumination Invariant 3D Facial Images.” This research was funded by the Ministry of Science and ICT and the Ministry of Trade, Industry and Energy.
-Publication“Machine-learned light-field camera that reads fascial expression from high-contrast and illumination invariant 3D facial images,” Sang-In Bae, Sangyeon Lee, Jae-Myeong Kwon, Hyun-Kyung Kim. Kyung-Won Jang, Doheon Lee, Ki-Hun Jeong, Advanced Intelligent Systems, December 16, 2021 (doi.org/10.1002/aisy.202100182)
ProfileProfessor Ki-Hun JeongBiophotonic LaboratoryDepartment of Bio and Brain EngineeringKAIST
Professor Doheon LeeDepartment of Bio and Brain EngineeringKAIST
2022.01.21 View 13958
AI Light-Field Camera Reads 3D Facial Expressions
Machine-learned, light-field camera reads facial expressions from high-contrast illumination invariant 3D facial images
A joint research team led by Professors Ki-Hun Jeong and Doheon Lee from the KAIST Department of Bio and Brain Engineering reported the development of a technique for facial expression detection by merging near-infrared light-field camera techniques with artificial intelligence (AI) technology.
Unlike a conventional camera, the light-field camera contains micro-lens arrays in front of the image sensor, which makes the camera small enough to fit into a smart phone, while allowing it to acquire the spatial and directional information of the light with a single shot. The technique has received attention as it can reconstruct images in a variety of ways including multi-views, refocusing, and 3D image acquisition, giving rise to many potential applications.
However, the optical crosstalk between shadows caused by external light sources in the environment and the micro-lens has limited existing light-field cameras from being able to provide accurate image contrast and 3D reconstruction.
The joint research team applied a vertical-cavity surface-emitting laser (VCSEL) in the near-IR range to stabilize the accuracy of 3D image reconstruction that previously depended on environmental light. When an external light source is shone on a face at 0-, 30-, and 60-degree angles, the light field camera reduces 54% of image reconstruction errors. Additionally, by inserting a light-absorbing layer for visible and near-IR wavelengths between the micro-lens arrays, the team could minimize optical crosstalk while increasing the image contrast by 2.1 times.
Through this technique, the team could overcome the limitations of existing light-field cameras and was able to develop their NIR-based light-field camera (NIR-LFC), optimized for the 3D image reconstruction of facial expressions. Using the NIR-LFC, the team acquired high-quality 3D reconstruction images of facial expressions expressing various emotions regardless of the lighting conditions of the surrounding environment.
The facial expressions in the acquired 3D images were distinguished through machine learning with an average of 85% accuracy – a statistically significant figure compared to when 2D images were used. Furthermore, by calculating the interdependency of distance information that varies with facial expression in 3D images, the team could identify the information a light-field camera utilizes to distinguish human expressions.
Professor Ki-Hun Jeong said, “The sub-miniature light-field camera developed by the research team has the potential to become the new platform to quantitatively analyze the facial expressions and emotions of humans.” To highlight the significance of this research, he added, “It could be applied in various fields including mobile healthcare, field diagnosis, social cognition, and human-machine interactions.”
This research was published in Advanced Intelligent Systems online on December 16, under the title, “Machine-Learned Light-field Camera that Reads Facial Expression from High-Contrast and Illumination Invariant 3D Facial Images.” This research was funded by the Ministry of Science and ICT and the Ministry of Trade, Industry and Energy.
-Publication“Machine-learned light-field camera that reads fascial expression from high-contrast and illumination invariant 3D facial images,” Sang-In Bae, Sangyeon Lee, Jae-Myeong Kwon, Hyun-Kyung Kim. Kyung-Won Jang, Doheon Lee, Ki-Hun Jeong, Advanced Intelligent Systems, December 16, 2021 (doi.org/10.1002/aisy.202100182)
ProfileProfessor Ki-Hun JeongBiophotonic LaboratoryDepartment of Bio and Brain EngineeringKAIST
Professor Doheon LeeDepartment of Bio and Brain EngineeringKAIST
2022.01.21 View 13958 -
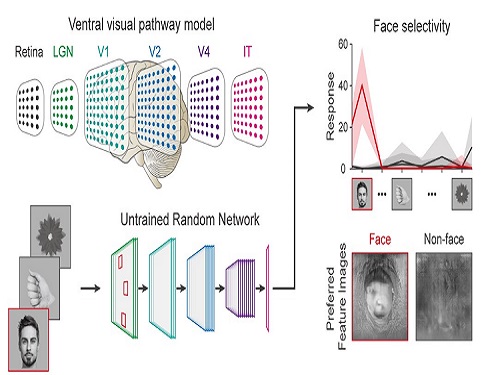 Face Detection in Untrained Deep Neural Networks
A KAIST team shows that primitive visual selectivity of faces can arise spontaneously in completely untrained deep neural networks
Researchers have found that higher visual cognitive functions can arise spontaneously in untrained neural networks. A KAIST research team led by Professor Se-Bum Paik from the Department of Bio and Brain Engineering has shown that visual selectivity of facial images can arise even in completely untrained deep neural networks.
This new finding has provided revelatory insights into mechanisms underlying the development of cognitive functions in both biological and artificial neural networks, also making a significant impact on our understanding of the origin of early brain functions before sensory experiences.
The study published in Nature Communications on December 16 demonstrates that neuronal activities selective to facial images are observed in randomly initialized deep neural networks in the complete absence of learning, and that they show the characteristics of those observed in biological brains.
The ability to identify and recognize faces is a crucial function for social behavior, and this ability is thought to originate from neuronal tuning at the single or multi-neuronal level. Neurons that selectively respond to faces are observed in young animals of various species, and this raises intense debate whether face-selective neurons can arise innately in the brain or if they require visual experience.
Using a model neural network that captures properties of the ventral stream of the visual cortex, the research team found that face-selectivity can emerge spontaneously from random feedforward wirings in untrained deep neural networks. The team showed that the character of this innate face-selectivity is comparable to that observed with face-selective neurons in the brain, and that this spontaneous neuronal tuning for faces enables the network to perform face detection tasks.
These results imply a possible scenario in which the random feedforward connections that develop in early, untrained networks may be sufficient for initializing primitive visual cognitive functions.
Professor Paik said, “Our findings suggest that innate cognitive functions can emerge spontaneously from the statistical complexity embedded in the hierarchical feedforward projection circuitry, even in the complete absence of learning”.
He continued, “Our results provide a broad conceptual advance as well as advanced insight into the mechanisms underlying the development of innate functions in both biological and artificial neural networks, which may unravel the mystery of the generation and evolution of intelligence.” This work was supported by the National Research Foundation of Korea (NRF) and by the KAIST singularity research project.
-PublicationSeungdae Baek, Min Song, Jaeson Jang, Gwangsu Kim, and Se-Bum Baik, “Face detection in untrained deep neural network,” Nature Communications 12, 7328 on Dec.16, 2021
(https://doi.org/10.1038/s41467-021-27606-9)
-ProfileProfessor Se-Bum PaikVisual System and Neural Network LaboratoryProgram of Brain and Cognitive EngineeringDepartment of Bio and Brain EngineeringCollege of EngineeringKAIST
2021.12.21 View 11410
Face Detection in Untrained Deep Neural Networks
A KAIST team shows that primitive visual selectivity of faces can arise spontaneously in completely untrained deep neural networks
Researchers have found that higher visual cognitive functions can arise spontaneously in untrained neural networks. A KAIST research team led by Professor Se-Bum Paik from the Department of Bio and Brain Engineering has shown that visual selectivity of facial images can arise even in completely untrained deep neural networks.
This new finding has provided revelatory insights into mechanisms underlying the development of cognitive functions in both biological and artificial neural networks, also making a significant impact on our understanding of the origin of early brain functions before sensory experiences.
The study published in Nature Communications on December 16 demonstrates that neuronal activities selective to facial images are observed in randomly initialized deep neural networks in the complete absence of learning, and that they show the characteristics of those observed in biological brains.
The ability to identify and recognize faces is a crucial function for social behavior, and this ability is thought to originate from neuronal tuning at the single or multi-neuronal level. Neurons that selectively respond to faces are observed in young animals of various species, and this raises intense debate whether face-selective neurons can arise innately in the brain or if they require visual experience.
Using a model neural network that captures properties of the ventral stream of the visual cortex, the research team found that face-selectivity can emerge spontaneously from random feedforward wirings in untrained deep neural networks. The team showed that the character of this innate face-selectivity is comparable to that observed with face-selective neurons in the brain, and that this spontaneous neuronal tuning for faces enables the network to perform face detection tasks.
These results imply a possible scenario in which the random feedforward connections that develop in early, untrained networks may be sufficient for initializing primitive visual cognitive functions.
Professor Paik said, “Our findings suggest that innate cognitive functions can emerge spontaneously from the statistical complexity embedded in the hierarchical feedforward projection circuitry, even in the complete absence of learning”.
He continued, “Our results provide a broad conceptual advance as well as advanced insight into the mechanisms underlying the development of innate functions in both biological and artificial neural networks, which may unravel the mystery of the generation and evolution of intelligence.” This work was supported by the National Research Foundation of Korea (NRF) and by the KAIST singularity research project.
-PublicationSeungdae Baek, Min Song, Jaeson Jang, Gwangsu Kim, and Se-Bum Baik, “Face detection in untrained deep neural network,” Nature Communications 12, 7328 on Dec.16, 2021
(https://doi.org/10.1038/s41467-021-27606-9)
-ProfileProfessor Se-Bum PaikVisual System and Neural Network LaboratoryProgram of Brain and Cognitive EngineeringDepartment of Bio and Brain EngineeringCollege of EngineeringKAIST
2021.12.21 View 11410