Science
-
 Scientists re-writes FDA-recommended equation to improve estimation of drug-drug interaction
Drugs absorbed into the body are metabolized and thus removed by enzymes from several organs like the liver. How fast a drug is cleared out of the system can be affected by other drugs that are taken together because added substance can increase the amount of enzyme secretion in the body. This dramatically decreases the concentration of a drug, reducing its efficacy, often leading to the failure of having any effect at all. Therefore, accurately predicting the clearance rate in the presence of drug-drug interaction* is critical in the process of drug prescription and development of a new drug in order to ensure its efficacy and/or to avoid unwanted side-effects.
*Drug-drug interaction: In terms of metabolism, drug-drug interaction is a phenomenon in which one drug changes the metabolism of another drug to promote or inhibit its excretion from the body when two or more drugs are taken together. As a result, it increases the toxicity of medicines or causes loss of efficacy.
Since it is practically impossible to evaluate all interactions between new drug candidates and all marketed drugs during the development process, the FDA recommends indirect evaluation of drug interactions using a formula suggested in their guidance, first published in 1997, revised in January of 2020, in order to evaluate drug interactions and minimize side effects of having to use more than one type of drugs at once.
The formula relies on the 110-year-old Michaelis-Menten (MM) model, which has a fundamental limit of making a very broad and groundless assumption on the part of the presence of the enzymes that metabolizes the drug. While MM equation has been one of the most widely known equations in biochemistry used in more than 220,000 published papers, the MM equation is accurate only when the concentration of the enzyme that metabolizes the drug is almost non-existent, causing the accuracy of the equation highly unsatisfactory – only 38 percent of the predictions had less than two-fold errors.
“To make up for the gap, researcher resorted to plugging in scientifically unjustified constants into the equation,” Professor Jung-woo Chae of Chungnam National University College of Pharmacy said. “This is comparable to having to have the epicyclic orbits introduced to explain the motion of the planets back in the days in order to explain the now-defunct Ptolemaic theory, because it was 'THE' theory back then.”
< (From left) Ph.D. student Yun Min Song (KAIST, co-first authors), Professor Sang Kyum Kim (Chungnam National University, co-corresponding author), Jae Kyoung Kim, CI (KAIST, co-corresponding author), Professor Jung-woo Chae (Chungnam National University, co-corresponding author), Ph.D. students Quyen Thi Tran and Ngoc-Anh Thi Vu (Chungnam National University, co-first authors) >
A joint research team composed of mathematicians from the Biomedical Mathematics Group within the Institute for Basic Science (IBS) and the Korea Advanced Institute of Science and Technology (KAIST) and pharmacological scientists from the Chungnam National University reported that they identified the major causes of the FDA-recommended equation’s inaccuracies and presented a solution.
When estimating the gut bioavailability (Fg), which is the key parameter of the equation, the fraction absorbed from the gut lumen (Fa) is usually assumed to be 1. However, many experiments have shown that Fa is less than 1, obviously since it can’t be expected that all of the orally taken drugs to be completely absorbed by the intestines. To solve this problem, the research team used an “estimated Fa” value based on factors such as the drug’s transit time, intestine radius, and permeability values and used it to re-calculate Fg.
Also, taking a different approach from the MM equation, the team used an alternative model they derived in a previous study back in 2020, which can more accurately predict the drug metabolism rate regardless of the enzyme concentration. Combining these changes, the modified equation with re-calculated Fg had a dramatically increased accuracy of the resulting estimate. The existing FDA formula predicted drug interactions within a 2-fold margin of error at the rate of 38%, whereas the accuracy rate of the revised formula reached 80%.
“Such drastic improvement in drug-drug interaction prediction accuracy is expected to make great contribution to increasing the success rate of new drug development and drug efficacy in clinical trials. As the results of this study were published in one of the top clinical pharmacology journal, it is expected that the FDA guidance will be revised according to the results of this study.” said Professor Sang Kyum Kim from Chungnam National University College of Pharmacy.
Furthermore, this study highlights the importance of collaborative research between research groups in vastly different disciplines, in a field that is as dynamic as drug interactions.
“Thanks to the collaborative research between mathematics and pharmacy, we were able to recify the formula that we have accepted to be the right answer for so long to finally grasp on the leads toward healthier life for mankind.,” said Professor Jae Kyung Kim. He continued, “I hope seeing a ‘K-formula’ entered into the US FDA guidance one day.”
The results of this study were published in the online edition of Clinical Pharmacology and Therapeutics (IF 7.051), an authoritative journal in the field of clinical pharmacology, on December 15, 2022 (Korean time).
Thesis Title: Beyond the Michaelis-Menten: Accurate Prediction of Drug Interactions through Cytochrome P450 3A4 Induction (doi: 10.1002/cpt.2824)
< Figure 1. The formula proposed by the FDA guidance for predicting drug-drug interactions (top) and the formula newly derived by the researchers (bottom). AUCR (the ratio of substrate area under the plasma concentration-time curve) represents the rate of change in drug concentration due to drug interactions. The research team more than doubled the accuracy of drug interaction prediction compared to the existing formula. >
< Figure 2. Existing FDA formulas tend to underestimate the extent of drug-drug interactions (gray dots) than the actual measured values. On the other hand, the newly derived equation (red dot) has a prediction rate that is within the error range of 2 times (0.5 to 2 times) of the measured value, and is more than twice as high as the existing equation. The solid line in the figure represents the predicted value that matches the measured value. The dotted line represents the predicted value with an error of 0.5 to 2 times. >
For further information or to request media assistance, please contact Jae Kyoung Kim at Biomedical Mathematics Group, Institute for Basic Science (IBS) (jaekkim@ibs.re.kr) or William I. Suh at the IBS Communications Team (willisuh@ibs.re.kr).
- About the Institute for Basic Science (IBS)
IBS was founded in 2011 by the government of the Republic of Korea with the sole purpose of driving forward the development of basic science in South Korea. IBS has 4 research institutes and 33 research centers as of January 2023. There are eleven physics, three mathematics, five chemistry, nine life science, two earth science, and three interdisciplinary research centers.
2023.01.18 View 14490
Scientists re-writes FDA-recommended equation to improve estimation of drug-drug interaction
Drugs absorbed into the body are metabolized and thus removed by enzymes from several organs like the liver. How fast a drug is cleared out of the system can be affected by other drugs that are taken together because added substance can increase the amount of enzyme secretion in the body. This dramatically decreases the concentration of a drug, reducing its efficacy, often leading to the failure of having any effect at all. Therefore, accurately predicting the clearance rate in the presence of drug-drug interaction* is critical in the process of drug prescription and development of a new drug in order to ensure its efficacy and/or to avoid unwanted side-effects.
*Drug-drug interaction: In terms of metabolism, drug-drug interaction is a phenomenon in which one drug changes the metabolism of another drug to promote or inhibit its excretion from the body when two or more drugs are taken together. As a result, it increases the toxicity of medicines or causes loss of efficacy.
Since it is practically impossible to evaluate all interactions between new drug candidates and all marketed drugs during the development process, the FDA recommends indirect evaluation of drug interactions using a formula suggested in their guidance, first published in 1997, revised in January of 2020, in order to evaluate drug interactions and minimize side effects of having to use more than one type of drugs at once.
The formula relies on the 110-year-old Michaelis-Menten (MM) model, which has a fundamental limit of making a very broad and groundless assumption on the part of the presence of the enzymes that metabolizes the drug. While MM equation has been one of the most widely known equations in biochemistry used in more than 220,000 published papers, the MM equation is accurate only when the concentration of the enzyme that metabolizes the drug is almost non-existent, causing the accuracy of the equation highly unsatisfactory – only 38 percent of the predictions had less than two-fold errors.
“To make up for the gap, researcher resorted to plugging in scientifically unjustified constants into the equation,” Professor Jung-woo Chae of Chungnam National University College of Pharmacy said. “This is comparable to having to have the epicyclic orbits introduced to explain the motion of the planets back in the days in order to explain the now-defunct Ptolemaic theory, because it was 'THE' theory back then.”
< (From left) Ph.D. student Yun Min Song (KAIST, co-first authors), Professor Sang Kyum Kim (Chungnam National University, co-corresponding author), Jae Kyoung Kim, CI (KAIST, co-corresponding author), Professor Jung-woo Chae (Chungnam National University, co-corresponding author), Ph.D. students Quyen Thi Tran and Ngoc-Anh Thi Vu (Chungnam National University, co-first authors) >
A joint research team composed of mathematicians from the Biomedical Mathematics Group within the Institute for Basic Science (IBS) and the Korea Advanced Institute of Science and Technology (KAIST) and pharmacological scientists from the Chungnam National University reported that they identified the major causes of the FDA-recommended equation’s inaccuracies and presented a solution.
When estimating the gut bioavailability (Fg), which is the key parameter of the equation, the fraction absorbed from the gut lumen (Fa) is usually assumed to be 1. However, many experiments have shown that Fa is less than 1, obviously since it can’t be expected that all of the orally taken drugs to be completely absorbed by the intestines. To solve this problem, the research team used an “estimated Fa” value based on factors such as the drug’s transit time, intestine radius, and permeability values and used it to re-calculate Fg.
Also, taking a different approach from the MM equation, the team used an alternative model they derived in a previous study back in 2020, which can more accurately predict the drug metabolism rate regardless of the enzyme concentration. Combining these changes, the modified equation with re-calculated Fg had a dramatically increased accuracy of the resulting estimate. The existing FDA formula predicted drug interactions within a 2-fold margin of error at the rate of 38%, whereas the accuracy rate of the revised formula reached 80%.
“Such drastic improvement in drug-drug interaction prediction accuracy is expected to make great contribution to increasing the success rate of new drug development and drug efficacy in clinical trials. As the results of this study were published in one of the top clinical pharmacology journal, it is expected that the FDA guidance will be revised according to the results of this study.” said Professor Sang Kyum Kim from Chungnam National University College of Pharmacy.
Furthermore, this study highlights the importance of collaborative research between research groups in vastly different disciplines, in a field that is as dynamic as drug interactions.
“Thanks to the collaborative research between mathematics and pharmacy, we were able to recify the formula that we have accepted to be the right answer for so long to finally grasp on the leads toward healthier life for mankind.,” said Professor Jae Kyung Kim. He continued, “I hope seeing a ‘K-formula’ entered into the US FDA guidance one day.”
The results of this study were published in the online edition of Clinical Pharmacology and Therapeutics (IF 7.051), an authoritative journal in the field of clinical pharmacology, on December 15, 2022 (Korean time).
Thesis Title: Beyond the Michaelis-Menten: Accurate Prediction of Drug Interactions through Cytochrome P450 3A4 Induction (doi: 10.1002/cpt.2824)
< Figure 1. The formula proposed by the FDA guidance for predicting drug-drug interactions (top) and the formula newly derived by the researchers (bottom). AUCR (the ratio of substrate area under the plasma concentration-time curve) represents the rate of change in drug concentration due to drug interactions. The research team more than doubled the accuracy of drug interaction prediction compared to the existing formula. >
< Figure 2. Existing FDA formulas tend to underestimate the extent of drug-drug interactions (gray dots) than the actual measured values. On the other hand, the newly derived equation (red dot) has a prediction rate that is within the error range of 2 times (0.5 to 2 times) of the measured value, and is more than twice as high as the existing equation. The solid line in the figure represents the predicted value that matches the measured value. The dotted line represents the predicted value with an error of 0.5 to 2 times. >
For further information or to request media assistance, please contact Jae Kyoung Kim at Biomedical Mathematics Group, Institute for Basic Science (IBS) (jaekkim@ibs.re.kr) or William I. Suh at the IBS Communications Team (willisuh@ibs.re.kr).
- About the Institute for Basic Science (IBS)
IBS was founded in 2011 by the government of the Republic of Korea with the sole purpose of driving forward the development of basic science in South Korea. IBS has 4 research institutes and 33 research centers as of January 2023. There are eleven physics, three mathematics, five chemistry, nine life science, two earth science, and three interdisciplinary research centers.
2023.01.18 View 14490 -
 KAIST Offers Hope to Musicians with Dystonia
< Photo 1. Conductor and Pianist João Carlos Martins before the Recital at the Carnegie Hall preparing with his bionic gloves >
KAIST’s neuroscientist and professor, Dr. Daesoo Kim attended the “Conference for Musicians with Dystonia” supported by the World Health Organization (WHO) and the Carnegie Hall concert of legendary pianist João Carlos Martins, who is also a dystonia patient, to announce his team’s recent advancements toward finding a cure for dystonia.
On November 19, 2022, a “miracle concert” was held in Carnegie Hall. João Carlos Martins was a renowned world-class pianist in the 70s and 80s, but he had to put an end to his musical career due to focal dystonia in his fingers. But in 2020, he began using a bionic glove developed by industrial designer Ubiratã Bizarro Costa and after years of hard work he was back in Carnegie Hall as an 82-year-old man.
During the concert, he conducted the NOVUS NY orchestra in a performance of Bach, and later even played the piano himself. In particular, between his performances, he gave shout-outs to scientists studying dystonia including KAIST Professor Daesoo Kim, asking them to continue working towards curing rare diseases for musicians.
< Photo 2. Professor Daesoo Kim with Conductor and Pianist João Carlos Martins >
Musician’s dystonia affects 1-3% of musicians around the world and musicians make up approximately 5% of the total number of dystonia patients. Musicians who are no longer able to practice music due to the disease often experience stress and depression, which may even lead to suicide in extreme cases. Musicians are known to be particularly prone to such diseases due to excessive practice regimens, perfectionism, and even genetics. Currently, botulinum toxin (Botox) is used to suppress abnormal muscles, but muscle function suppression ultimately means that the musician is no longer able to play the instrument. João Carlos Martins himself underwent several Botox procedures and three brain surgeries, but saw no therapeutic results. This is why a new treatment was necessary.
Professor Daesoo Kim’s research team at KAIST took note of the fact that abnormal muscle tension is caused by excessive stress, and developed NT-1, a treatment that blocks the development of the symptoms of dystonia from the brain, allowing patients to use their muscles as they normally would. The research team published their findings in Science Advances in 2021, and João Carlos Martins invited Professor Daesoo Kim to the UN conference and his concert after reading this paper.
< Photo 3. Professor Daesoo Kim (3rd from the left) photographed with other guests at the recital including Dr. Dévora Kestel, the Director of the Mental Health and Substance Use at WHO, sharing the center with Conductor and Pianist João Carlos Martins >
During the UN conference held the day prior to the Carnegie Hall concert, Dr. Dévora Kestel, Director of the Mental Health and Substance Use at WHO, said, “Although dystonia is not as well-known, it is a common disease around the world, and needs our society’s attention and the devotion of many researchers.” Professor Daesoo Kim said, “NT-1 is a drug that blocks the cause of dystonia in the brain, and will allow musicians to continue practicing music. We aim to attain clinical approval in Korea by 2024.”
NT-1 is currently under development by NeuroTobe, a faculty-led start-up company at KAIST, headed by Professor Daesoo Kim as the CEO. The synthesis of the drug for clinical testing has been successfully completed, and it has shown excellent efficacy and safety through various rounds of animal testing. Unlike Botox, which takes a few days to show its therapeutic effects after receiving the procedure from a hospital, NT-1 shows its therapeutic effects within an hour after taking it. As a so-called “edible Botox”, it is expected to help treat various muscular diseases and ailments.
2022.12.27 View 12299
KAIST Offers Hope to Musicians with Dystonia
< Photo 1. Conductor and Pianist João Carlos Martins before the Recital at the Carnegie Hall preparing with his bionic gloves >
KAIST’s neuroscientist and professor, Dr. Daesoo Kim attended the “Conference for Musicians with Dystonia” supported by the World Health Organization (WHO) and the Carnegie Hall concert of legendary pianist João Carlos Martins, who is also a dystonia patient, to announce his team’s recent advancements toward finding a cure for dystonia.
On November 19, 2022, a “miracle concert” was held in Carnegie Hall. João Carlos Martins was a renowned world-class pianist in the 70s and 80s, but he had to put an end to his musical career due to focal dystonia in his fingers. But in 2020, he began using a bionic glove developed by industrial designer Ubiratã Bizarro Costa and after years of hard work he was back in Carnegie Hall as an 82-year-old man.
During the concert, he conducted the NOVUS NY orchestra in a performance of Bach, and later even played the piano himself. In particular, between his performances, he gave shout-outs to scientists studying dystonia including KAIST Professor Daesoo Kim, asking them to continue working towards curing rare diseases for musicians.
< Photo 2. Professor Daesoo Kim with Conductor and Pianist João Carlos Martins >
Musician’s dystonia affects 1-3% of musicians around the world and musicians make up approximately 5% of the total number of dystonia patients. Musicians who are no longer able to practice music due to the disease often experience stress and depression, which may even lead to suicide in extreme cases. Musicians are known to be particularly prone to such diseases due to excessive practice regimens, perfectionism, and even genetics. Currently, botulinum toxin (Botox) is used to suppress abnormal muscles, but muscle function suppression ultimately means that the musician is no longer able to play the instrument. João Carlos Martins himself underwent several Botox procedures and three brain surgeries, but saw no therapeutic results. This is why a new treatment was necessary.
Professor Daesoo Kim’s research team at KAIST took note of the fact that abnormal muscle tension is caused by excessive stress, and developed NT-1, a treatment that blocks the development of the symptoms of dystonia from the brain, allowing patients to use their muscles as they normally would. The research team published their findings in Science Advances in 2021, and João Carlos Martins invited Professor Daesoo Kim to the UN conference and his concert after reading this paper.
< Photo 3. Professor Daesoo Kim (3rd from the left) photographed with other guests at the recital including Dr. Dévora Kestel, the Director of the Mental Health and Substance Use at WHO, sharing the center with Conductor and Pianist João Carlos Martins >
During the UN conference held the day prior to the Carnegie Hall concert, Dr. Dévora Kestel, Director of the Mental Health and Substance Use at WHO, said, “Although dystonia is not as well-known, it is a common disease around the world, and needs our society’s attention and the devotion of many researchers.” Professor Daesoo Kim said, “NT-1 is a drug that blocks the cause of dystonia in the brain, and will allow musicians to continue practicing music. We aim to attain clinical approval in Korea by 2024.”
NT-1 is currently under development by NeuroTobe, a faculty-led start-up company at KAIST, headed by Professor Daesoo Kim as the CEO. The synthesis of the drug for clinical testing has been successfully completed, and it has shown excellent efficacy and safety through various rounds of animal testing. Unlike Botox, which takes a few days to show its therapeutic effects after receiving the procedure from a hospital, NT-1 shows its therapeutic effects within an hour after taking it. As a so-called “edible Botox”, it is expected to help treat various muscular diseases and ailments.
2022.12.27 View 12299 -
 KAIST develops biocompatible adhesive applicable to hair transplants
Aside from being used as a new medical adhesive, the new material can be applied to developing a new method of hair transplants, which cannot be repeated multiple times using current method of implanting the wholly intact follicles into the skin.
Medical adhesives are materials that can be applied to various uses such as wound healing, hemostasis, vascular anastomosis, and tissue engineering, and is expected to contribute greatly to the development of minimally invasive surgery and organ transplants. However, adhesives with high adhesion, low toxicity, and capable of decomposing in the body are rare. Adhesives based on natural proteins, such as fibrin and collagen, have high biocompatibility but insufficient adhesive strength. Synthetic polymer adhesives based on urethane or acrylic have greater adhesion but do not decompose well and may cause an inflammatory reaction in the body.
A joint research team led by Professor Myungeun Seo and Professor Haeshin Lee from the KAIST Department of Chemistry developed a bio-friendly adhesive from biocompatible polymers using tannic acid, the source of astringency in wine.
The research team focused on tannic acid, a natural polyphenolic product. Tannic acid is a polyphenol present in large amounts in fruit peels, nuts, and cacao. It has a high affinity and coating ability on other substances, and we sense the astringent taste in wine when tannic acid sticks to the surface of our tongue. When tannic acid is mixed with hydrophilic polymers, they form coacervates, or small droplets of jelly-like fluids that sink. If the polymers used are biocompatible, the mixture can be applied as a medical adhesive with low toxicity. However, coacervates are fundamentally fluid-like and cannot withstand large forces, which limits their adhesive capabilities. Thus, while research to utilize it as an adhesive has been actively discussed, a biodegradable material exhibiting strong adhesion due to its high shear strength has not yet been developed.
The research team figured out a way to enhance adhesion by mixing two biocompatible FDA-approved polymers, polyethylene glycol (PEG) and polylactic acid (PLA). While PEG, which is used widely in eyedrops and cream, is hydrophilic, PLA, a well-known bioplastic derived from lactic acid, is insoluble in water. The team combined the two into a block copolymer, which forms hydrophilic PLA aggregates in water with PEG blocks surrounding them. A coacervate created by mixing the micelles and tannic acid would behave like a solid due to the hard PLA components, and show an elastic modulus improved by a thousand times compared to PEG, enabling it to withstand much greater force as an adhesive.
Figure 1. (Above) Principle of biodegradable adhesive made by mixing poly(ethylene glycol)-poly(lactic acid) diblock copolymer and tannic acid in water. Yellow coacervate is precipitated through hydrogen bonding between the block copolymer micelles and tannic acid, and exhibits adhesion. After heat treatment, hydrogen bonds are rearranged to further improve adhesion. (Bottom) Adhesion comparison. Compared to using poly(ethylene glycol) polymer (d), it can support 10 times more weight when using block copolymer (e) and 60 times more weight after heat treatment (f). The indicated G' values represent the elastic modulus of the material.
Furthermore, the research team observed that the material’s mechanical properties can be improved by over a hundred times through a heating and cooling process that is used to heat-treat metals. They also discovered that this is due to the enforced interactions between micelle and tannic acid arrays.
The research team used the fact that the material shows minimal irritation to the skin and decomposes well in the body to demonstrate its possible application as an adhesive for hair transplantation through an animal experiment. Professor Haeshin Lee, who has pioneered various application fields including medical adhesives, hemostatic agents, and browning shampoo, focused on the adhesive capacities and low toxicity of polyphenols like tannic acid, and now looks forward to it improving the limitations of current hair transplant methods, which still involve follicle transfer and are difficult to be repeated multiple times.
Figure 2. (a) Overview of a hair transplantation method using a biodegradable adhesive (right) compared to a conventional hair transplantation method (left) that transplants hair containing hair follicles. After applying an adhesive to the tip of the hair, it is fixed to the skin by implanting it through a subcutaneous injection, and repeated treatment is possible. (b) Initial animal test results. One day after 15 hair transplantation, 12 strands of hair remain. If you pull the 3 strands of hair, you can see that the whole body is pulled up, indicating that it is firmly implanted into the skin. All strands of hair applied without the new adhesive material fell off, and in the case of adhesive without heat treatment, the efficiency was 1/7.
This research was conducted by first co-authors Dr. Jongmin Park (currently a senior researcher at the Korea Research Institute of Chemical Technology) from Professor Myeongeun Seo’s team and Dr. Eunsook Park from Professor Haeshin Lee’s team in the KAIST Department of Chemistry, and through joint research with the teams led by Professor Hyungjun Kim from the KAIST Department of Chemistry and Professor Siyoung Choi from the Department of Chemical and Biomolecular Engineering. The research was published online on August 22 in the international journal Au (JACS Au) under the title Biodegradable Block Copolymer-Tannic Acid Glue.
This study was funded by the Support Research Under Protection Project of the National Research Foundation (NRF), Leading Research Center Support Project (Research Center for Multiscale Chiral Structure), Biodegradable Plastics Commercialization and Demonstration Project by the Ministry of Trade and Industry, and institutional funding from the Korea Research Institute of Chemical Technology.
2022.10.07 View 12619
KAIST develops biocompatible adhesive applicable to hair transplants
Aside from being used as a new medical adhesive, the new material can be applied to developing a new method of hair transplants, which cannot be repeated multiple times using current method of implanting the wholly intact follicles into the skin.
Medical adhesives are materials that can be applied to various uses such as wound healing, hemostasis, vascular anastomosis, and tissue engineering, and is expected to contribute greatly to the development of minimally invasive surgery and organ transplants. However, adhesives with high adhesion, low toxicity, and capable of decomposing in the body are rare. Adhesives based on natural proteins, such as fibrin and collagen, have high biocompatibility but insufficient adhesive strength. Synthetic polymer adhesives based on urethane or acrylic have greater adhesion but do not decompose well and may cause an inflammatory reaction in the body.
A joint research team led by Professor Myungeun Seo and Professor Haeshin Lee from the KAIST Department of Chemistry developed a bio-friendly adhesive from biocompatible polymers using tannic acid, the source of astringency in wine.
The research team focused on tannic acid, a natural polyphenolic product. Tannic acid is a polyphenol present in large amounts in fruit peels, nuts, and cacao. It has a high affinity and coating ability on other substances, and we sense the astringent taste in wine when tannic acid sticks to the surface of our tongue. When tannic acid is mixed with hydrophilic polymers, they form coacervates, or small droplets of jelly-like fluids that sink. If the polymers used are biocompatible, the mixture can be applied as a medical adhesive with low toxicity. However, coacervates are fundamentally fluid-like and cannot withstand large forces, which limits their adhesive capabilities. Thus, while research to utilize it as an adhesive has been actively discussed, a biodegradable material exhibiting strong adhesion due to its high shear strength has not yet been developed.
The research team figured out a way to enhance adhesion by mixing two biocompatible FDA-approved polymers, polyethylene glycol (PEG) and polylactic acid (PLA). While PEG, which is used widely in eyedrops and cream, is hydrophilic, PLA, a well-known bioplastic derived from lactic acid, is insoluble in water. The team combined the two into a block copolymer, which forms hydrophilic PLA aggregates in water with PEG blocks surrounding them. A coacervate created by mixing the micelles and tannic acid would behave like a solid due to the hard PLA components, and show an elastic modulus improved by a thousand times compared to PEG, enabling it to withstand much greater force as an adhesive.
Figure 1. (Above) Principle of biodegradable adhesive made by mixing poly(ethylene glycol)-poly(lactic acid) diblock copolymer and tannic acid in water. Yellow coacervate is precipitated through hydrogen bonding between the block copolymer micelles and tannic acid, and exhibits adhesion. After heat treatment, hydrogen bonds are rearranged to further improve adhesion. (Bottom) Adhesion comparison. Compared to using poly(ethylene glycol) polymer (d), it can support 10 times more weight when using block copolymer (e) and 60 times more weight after heat treatment (f). The indicated G' values represent the elastic modulus of the material.
Furthermore, the research team observed that the material’s mechanical properties can be improved by over a hundred times through a heating and cooling process that is used to heat-treat metals. They also discovered that this is due to the enforced interactions between micelle and tannic acid arrays.
The research team used the fact that the material shows minimal irritation to the skin and decomposes well in the body to demonstrate its possible application as an adhesive for hair transplantation through an animal experiment. Professor Haeshin Lee, who has pioneered various application fields including medical adhesives, hemostatic agents, and browning shampoo, focused on the adhesive capacities and low toxicity of polyphenols like tannic acid, and now looks forward to it improving the limitations of current hair transplant methods, which still involve follicle transfer and are difficult to be repeated multiple times.
Figure 2. (a) Overview of a hair transplantation method using a biodegradable adhesive (right) compared to a conventional hair transplantation method (left) that transplants hair containing hair follicles. After applying an adhesive to the tip of the hair, it is fixed to the skin by implanting it through a subcutaneous injection, and repeated treatment is possible. (b) Initial animal test results. One day after 15 hair transplantation, 12 strands of hair remain. If you pull the 3 strands of hair, you can see that the whole body is pulled up, indicating that it is firmly implanted into the skin. All strands of hair applied without the new adhesive material fell off, and in the case of adhesive without heat treatment, the efficiency was 1/7.
This research was conducted by first co-authors Dr. Jongmin Park (currently a senior researcher at the Korea Research Institute of Chemical Technology) from Professor Myeongeun Seo’s team and Dr. Eunsook Park from Professor Haeshin Lee’s team in the KAIST Department of Chemistry, and through joint research with the teams led by Professor Hyungjun Kim from the KAIST Department of Chemistry and Professor Siyoung Choi from the Department of Chemical and Biomolecular Engineering. The research was published online on August 22 in the international journal Au (JACS Au) under the title Biodegradable Block Copolymer-Tannic Acid Glue.
This study was funded by the Support Research Under Protection Project of the National Research Foundation (NRF), Leading Research Center Support Project (Research Center for Multiscale Chiral Structure), Biodegradable Plastics Commercialization and Demonstration Project by the Ministry of Trade and Industry, and institutional funding from the Korea Research Institute of Chemical Technology.
2022.10.07 View 12619 -
 Establishing a novel strategy to tackle Huntington’s disease
A platform to take on the Huntington’s disease via an innovative approach established by KAIST’s researchers through international collaboration with scientists in the Netherlands, France, and Sweden.
Through an international joint research effort involving ProQR Therapeutics of the Netherlands, Université Grenoble Alpes of France, and KTH Royal Institute of Technology of Sweden, Professor Ji-Soon Song's research team in the Department of Biological Sciences and KAIST Institute for BioCentury of KAIST, established a noble strategy to treat Huntington's disease. The new works showed that the protein converted from disease form to its disease-free form maintains its original function, providing new roadblocks to approach Huntington’s disease.
This research, titled, “A pathogenic-proteolysis resistant huntingtin isoform induced by an antisense oligonucleotide maintains huntingtin function”, co-authored by Hyeongju Kim, was published in the online edition of 'Journal of Clinical Investigation Insight' on August 9, 2022.
Huntington's disease is a dominantly inherited neurodegenerative disease and is caused by a mutation in a protein called ‘huntingtin’, which adds a distinctive feature of an expanded stretch of glutamine amino acids called polyglutamine to the protein. It is estimated that one in every 10,000 have Huntington's disease in United States. The patients would suffer a decade of regression before death, and, thus far, there is no known cure for the disease.
The cleavage near the stretched polyglutamine in mutated huntingtin is known to be the cause of the Huntington’s disease. However, as huntingtin protein is required for the development and normal function of the brain, it is critical to specifically eliminate the disease-causing protein while maintaining the ones that are still normally functioning.
The research team showed that huntingtin delta 12, the converted form of huntingtin that is resistant to developing cleavages at the ends of the protein, the known cause of the Huntington’s disease (HD), alleviated the disease’s symptoms while maintaining the functions of normal huntingtin.
Figure. Huntington's disease resistance huntingtin protein induced by antisense oligonucleotide (AON) is resistant to Caspase-6 cleavage, therefore, does not cause Huntington’s disease while maintaining normal functions of huntingtin.
The research was welcomed as it is sure to fuel innovate strategies to tackle Huntington’s disease without altering the essential function of huntingtin.
This work was supported by a Global Research Lab grant from the National Research Foundation of Korea (NRF) and by a EUREKA Eurostars 2 grant from European Union Horizon 2020.
2022.09.02 View 8038
Establishing a novel strategy to tackle Huntington’s disease
A platform to take on the Huntington’s disease via an innovative approach established by KAIST’s researchers through international collaboration with scientists in the Netherlands, France, and Sweden.
Through an international joint research effort involving ProQR Therapeutics of the Netherlands, Université Grenoble Alpes of France, and KTH Royal Institute of Technology of Sweden, Professor Ji-Soon Song's research team in the Department of Biological Sciences and KAIST Institute for BioCentury of KAIST, established a noble strategy to treat Huntington's disease. The new works showed that the protein converted from disease form to its disease-free form maintains its original function, providing new roadblocks to approach Huntington’s disease.
This research, titled, “A pathogenic-proteolysis resistant huntingtin isoform induced by an antisense oligonucleotide maintains huntingtin function”, co-authored by Hyeongju Kim, was published in the online edition of 'Journal of Clinical Investigation Insight' on August 9, 2022.
Huntington's disease is a dominantly inherited neurodegenerative disease and is caused by a mutation in a protein called ‘huntingtin’, which adds a distinctive feature of an expanded stretch of glutamine amino acids called polyglutamine to the protein. It is estimated that one in every 10,000 have Huntington's disease in United States. The patients would suffer a decade of regression before death, and, thus far, there is no known cure for the disease.
The cleavage near the stretched polyglutamine in mutated huntingtin is known to be the cause of the Huntington’s disease. However, as huntingtin protein is required for the development and normal function of the brain, it is critical to specifically eliminate the disease-causing protein while maintaining the ones that are still normally functioning.
The research team showed that huntingtin delta 12, the converted form of huntingtin that is resistant to developing cleavages at the ends of the protein, the known cause of the Huntington’s disease (HD), alleviated the disease’s symptoms while maintaining the functions of normal huntingtin.
Figure. Huntington's disease resistance huntingtin protein induced by antisense oligonucleotide (AON) is resistant to Caspase-6 cleavage, therefore, does not cause Huntington’s disease while maintaining normal functions of huntingtin.
The research was welcomed as it is sure to fuel innovate strategies to tackle Huntington’s disease without altering the essential function of huntingtin.
This work was supported by a Global Research Lab grant from the National Research Foundation of Korea (NRF) and by a EUREKA Eurostars 2 grant from European Union Horizon 2020.
2022.09.02 View 8038 -
 A New Therapeutic Drug for Alzheimer’s Disease without Inflammatory Side Effects
Although Aduhelm, a monoclonal antibody targeting amyloid beta (Aβ), recently became the first US FDA approved drug for Alzheimer’s disease (AD) based on its ability to decrease Aβ plaque burden in AD patients, its effect on cognitive improvement is still controversial. Moreover, about 40% of the patients treated with this antibody experienced serious side effects including cerebral edemas (ARIA-E) and hemorrhages (ARIA-H) that are likely related to inflammatory responses in the brain when the Aβ antibody binds Fc receptors (FCR) of immune cells such as microglia and macrophages.
These inflammatory side effects can cause neuronal cell death and synapse elimination by activated microglia, and even have the potential to exacerbate cognitive impairment in AD patients. Thus, current Aβ antibody-based immunotherapy holds the inherent risk of doing more harm than good due to their inflammatory side effects.
To overcome these problems, a team of researchers at KAIST in South Korea has developed a novel fusion protein drug, αAβ-Gas6, which efficiently eliminates Aβ via an entirely different mechanism than Aβ antibody-based immunotherapy. In a mouse model of AD, αAβ-Gas6 not only removed Aβ with higher potency, but also circumvented the neurotoxic inflammatory side effects associated with conventional antibody treatments.
Their findings were published on August 4 in Nature Medicine.
Schematic of a chimeric Gas6 fusion protein. A single chain variable fragment (scFv) of
an Amyloid β (Aβ)-targeting monoclonal antibody is fused with a truncated receptor binding
domain of Gas6, a bridging molecule for the clearance of dead cells via TAM (TYRO3, AXL,
and MERTK) receptors, which are expressed by microglia and astrocytes.
“FcR activation by Aβ targeting antibodies induces microglia-mediated Aβ phagocytosis, but it also produces inflammatory signals, inevitably damaging brain tissues,” said paper authors Chan Hyuk Kim and Won-Suk Chung, associate professors in the Department of Biological Sciences at KAIST.
“Therefore, we utilized efferocytosis, a cellular process by which dead cells are removed by phagocytes as an alternative pathway for the clearance of Aβ in the brain,” Prof. Kim and Chung said. “Efferocytosis is accompanied by anti-inflammatory responses to maintain tissue homeostasis. To exploit this process, we engineered Gas6, a soluble adaptor protein that mediates efferocytosis via TAM phagocytic receptors in such a way that its target specificity was redirected from dead cells to Aβ plaques.”
The professors and their team demonstrated that the resulting αAβ-Gas6 induced Aβ engulfment by activating not only microglial but also astrocytic phagocytosis since TAM phagocytic receptors are highly expressed by these two major phagocytes in the brain. Importantly, αAβ-Gas6 promoted the robust uptake of Aβ without showing any signs of inflammation and neurotoxicity, which contrasts sharply with the treatment using an Aβ monoclonal antibody. Moreover, they showed that αAβ-Gas6 substantially reduced excessive synapse elimination by microglia, consequently leading to better behavioral rescues in AD model mice.
“By using a mouse model of cerebral amyloid angiopathy (CAA), a cerebrovascular disorder caused by the deposition of Aβ within the walls of the brain’s blood vessels, we also showed that the intrathecal administration of Gas6 fusion protein significantly eliminated cerebrovascular amyloids, along with a reduction of microhemorrhages. These data demonstrate that aAb-Gas6 is a potent therapeutic agent in eliminating Aβ without exacerbating CAA-related microhemorrhages.”
The resulting αAβ-Gas6 clears Aβ oligomers and fibrils without causing neurotoxicity (a-b, neurons: red, and fragmented axons: yellow) and proinflammatory responses (c, TNF release), which are conversely exacerbated by the treatment of an Aβ-targeting monoclonal antibody (Aducanumab).
Professors Kim and Chung noted, “We believe our approach can be a breakthrough in treating AD without causing inflammatory side effects and synapse loss. Our approach holds promise as a novel therapeutic platform that is applicable to more than AD. By modifying the target-specificity of the fusion protein, the Gas6-fusion protein can be applied to various neurological disorders as well as autoimmune diseases affected by toxic molecules that should be removed without causing inflammatory responses.”
The number and total area of Aβ plaques (Thioflavin-T, green) were significantly reduced in αAβ-Gas6-treated AD mouse brains compared to Aducanumab-treated ones (a, b). The cognitive functions of AD model mice were significantly rescued by αAβ-Gas6 treatment, whereas Aducanumab-treated AD mice showed partial rescue in these cognitive tests (c-e).
Professors Kim and Chung founded “Illimis Therapeutics” based on this strategy of designing chimeric Gas6 fusion proteins that would remove toxic aggregates from the nervous system. Through this company, they are planning to further develop various Gas6-fusion proteins not only for Ab but also for Tau to treat AD symptoms.
This work was supported by KAIST and the Korea Health Technology R&D Project that was administered by the Korea Health Industry Development Institute (KHIDI) and the Korea Dementia Research Center (KDRC) funded by the Ministry of Health & Welfare (MOHW) and the Ministry of Science and ICT (MSIT), and KAIST.
Other contributors include Hyuncheol Jung and Se Young Lee, Sungjoon Lim, Hyeong Ryeol Choi, Yeseong Choi, Minjin Kim, Segi Kim, the Department of Biological Sciences, and the Korea Advanced Institute of Science and Technology (KAIST).
To receive more up-to-date information on this new development, follow “Illimis Therapeutics” on twitter @Illimistx.
2022.08.05 View 13441
A New Therapeutic Drug for Alzheimer’s Disease without Inflammatory Side Effects
Although Aduhelm, a monoclonal antibody targeting amyloid beta (Aβ), recently became the first US FDA approved drug for Alzheimer’s disease (AD) based on its ability to decrease Aβ plaque burden in AD patients, its effect on cognitive improvement is still controversial. Moreover, about 40% of the patients treated with this antibody experienced serious side effects including cerebral edemas (ARIA-E) and hemorrhages (ARIA-H) that are likely related to inflammatory responses in the brain when the Aβ antibody binds Fc receptors (FCR) of immune cells such as microglia and macrophages.
These inflammatory side effects can cause neuronal cell death and synapse elimination by activated microglia, and even have the potential to exacerbate cognitive impairment in AD patients. Thus, current Aβ antibody-based immunotherapy holds the inherent risk of doing more harm than good due to their inflammatory side effects.
To overcome these problems, a team of researchers at KAIST in South Korea has developed a novel fusion protein drug, αAβ-Gas6, which efficiently eliminates Aβ via an entirely different mechanism than Aβ antibody-based immunotherapy. In a mouse model of AD, αAβ-Gas6 not only removed Aβ with higher potency, but also circumvented the neurotoxic inflammatory side effects associated with conventional antibody treatments.
Their findings were published on August 4 in Nature Medicine.
Schematic of a chimeric Gas6 fusion protein. A single chain variable fragment (scFv) of
an Amyloid β (Aβ)-targeting monoclonal antibody is fused with a truncated receptor binding
domain of Gas6, a bridging molecule for the clearance of dead cells via TAM (TYRO3, AXL,
and MERTK) receptors, which are expressed by microglia and astrocytes.
“FcR activation by Aβ targeting antibodies induces microglia-mediated Aβ phagocytosis, but it also produces inflammatory signals, inevitably damaging brain tissues,” said paper authors Chan Hyuk Kim and Won-Suk Chung, associate professors in the Department of Biological Sciences at KAIST.
“Therefore, we utilized efferocytosis, a cellular process by which dead cells are removed by phagocytes as an alternative pathway for the clearance of Aβ in the brain,” Prof. Kim and Chung said. “Efferocytosis is accompanied by anti-inflammatory responses to maintain tissue homeostasis. To exploit this process, we engineered Gas6, a soluble adaptor protein that mediates efferocytosis via TAM phagocytic receptors in such a way that its target specificity was redirected from dead cells to Aβ plaques.”
The professors and their team demonstrated that the resulting αAβ-Gas6 induced Aβ engulfment by activating not only microglial but also astrocytic phagocytosis since TAM phagocytic receptors are highly expressed by these two major phagocytes in the brain. Importantly, αAβ-Gas6 promoted the robust uptake of Aβ without showing any signs of inflammation and neurotoxicity, which contrasts sharply with the treatment using an Aβ monoclonal antibody. Moreover, they showed that αAβ-Gas6 substantially reduced excessive synapse elimination by microglia, consequently leading to better behavioral rescues in AD model mice.
“By using a mouse model of cerebral amyloid angiopathy (CAA), a cerebrovascular disorder caused by the deposition of Aβ within the walls of the brain’s blood vessels, we also showed that the intrathecal administration of Gas6 fusion protein significantly eliminated cerebrovascular amyloids, along with a reduction of microhemorrhages. These data demonstrate that aAb-Gas6 is a potent therapeutic agent in eliminating Aβ without exacerbating CAA-related microhemorrhages.”
The resulting αAβ-Gas6 clears Aβ oligomers and fibrils without causing neurotoxicity (a-b, neurons: red, and fragmented axons: yellow) and proinflammatory responses (c, TNF release), which are conversely exacerbated by the treatment of an Aβ-targeting monoclonal antibody (Aducanumab).
Professors Kim and Chung noted, “We believe our approach can be a breakthrough in treating AD without causing inflammatory side effects and synapse loss. Our approach holds promise as a novel therapeutic platform that is applicable to more than AD. By modifying the target-specificity of the fusion protein, the Gas6-fusion protein can be applied to various neurological disorders as well as autoimmune diseases affected by toxic molecules that should be removed without causing inflammatory responses.”
The number and total area of Aβ plaques (Thioflavin-T, green) were significantly reduced in αAβ-Gas6-treated AD mouse brains compared to Aducanumab-treated ones (a, b). The cognitive functions of AD model mice were significantly rescued by αAβ-Gas6 treatment, whereas Aducanumab-treated AD mice showed partial rescue in these cognitive tests (c-e).
Professors Kim and Chung founded “Illimis Therapeutics” based on this strategy of designing chimeric Gas6 fusion proteins that would remove toxic aggregates from the nervous system. Through this company, they are planning to further develop various Gas6-fusion proteins not only for Ab but also for Tau to treat AD symptoms.
This work was supported by KAIST and the Korea Health Technology R&D Project that was administered by the Korea Health Industry Development Institute (KHIDI) and the Korea Dementia Research Center (KDRC) funded by the Ministry of Health & Welfare (MOHW) and the Ministry of Science and ICT (MSIT), and KAIST.
Other contributors include Hyuncheol Jung and Se Young Lee, Sungjoon Lim, Hyeong Ryeol Choi, Yeseong Choi, Minjin Kim, Segi Kim, the Department of Biological Sciences, and the Korea Advanced Institute of Science and Technology (KAIST).
To receive more up-to-date information on this new development, follow “Illimis Therapeutics” on twitter @Illimistx.
2022.08.05 View 13441 -
 KAIST Research Team Proves How a Neurotransmitter may be the Key in Controlling Alzheimer’s Toxicity
With nearly 50 million dementia patients worldwide, and Alzheimers’s disease is the most common neurodegenerative disease. Its main symptom is the impairment of general cognitive abilities, including the ability to speak or to remember. The importance of finding a cure is widely understood with increasingly aging population and the life expectancy being ever-extended. However, even the cause of the grim disease is yet to be given a clear definition.
A KAIST research team in the Department of Chemistry led by professor Mi Hee Lim took on a lead to discovered a new role for somatostatin, a protein-based neurotransmitter, in reducing the toxicity caused in the pathogenic mechanism taken towards development of Alzheimer’s disease. The study was published in the July issue of Nature Chemistry under the title, “Conformational and functional changes of the native neuropeptide somatostatin occur in the presence of copper and amyloid-β”.
According to the amyloid hypothesis, the abnormal deposition of Aβ proteins causes death of neuronal cells. While Aβ agglomerations make up most of the aged plaques through fibrosis, in recent studies, high concentrations of transitional metal were found in the plaques from Alzheimer’s patients.
This suggests a close interaction between metallic ions and Aβ, which accelerates the fibrosis of proteins. Copper in particular is a redox-activating transition metal that can produce large amounts of oxygen and cause serious oxidative stress on cell organelles. Aβ proteins and transition metals can closely interact with neurotransmitters at synapses, but the direct effects of such abnormalities on the structure and function of neurotransmitters are yet to be understood.
Figure 1. Functional shift of somatostatin (SST) by factors in the pathogenesis of Alzheimer's disease.
Figure 2. Somatostatin’s loss-of-function as neurotransmitter. a. Schematic diagram of SST auto-aggregation due to Alzheimer's pathological factors. b. SST’s aggregation by copper ions. c. Coordination-prediction structure and N-terminal folding of copper-SST. d. Inhibition of SST receptor binding specificity by metals.
In their research, Professor Lim’s team discovered that when somatostatin, the protein-based neurotransmitter, is met with copper, Aβ, and metal-Aβ complexes, self-aggregates and ceases to perform its innate function of transmitting neural signals, but begins to attenuate the toxicity and agglomeration of metal-Aβ complexes.
Figure 3. Gain-of-function of somatostatin (SST) in the dementia setting. a. Prediction of docking of SST and amyloid beta. b. SST making metal-amyloid beta aggregates into an amorphous form. c. Cytotoxic mitigation effect of SST. d. SST mitigating the interaction between amyloid beta protein with the cell membrane.
This research, by Dr. Jiyeon Han et al. from the KAIST Department of Chemistry, revealed the coordination structure between copper and somatostatin at a molecular level through which it suggested the agglomeration mechanism, and discovered the effects of somatostatin on Aβ agglomeration path depending on the presence or absence of metals. The team has further confirmed somatostatin’s receptor binding, interactions with cell membranes, and effects on cell toxicity for the first time to receive international attention.
Professor Mi Hee Lim said, “This research has great significance in having discovered a new role of neurotransmitters in the pathogenesis of Alzheimer’s disease.” “We expect this research to contribute to defining the pathogenic network of neurodegenerative diseases caused by aging, and to the development of future biomarkers and medicine,” she added.
This research was conducted jointly by Professor Seung-Hee Lee’s team of KAIST Department of Biological Sciences, Professor Kiyoung Park’s Team of KAIST Department of Chemistry, and Professor Yulong Li’s team of Peking University.
The research was funded by Basic Science Research Program of the National Research Foundation of Korea and KAIST.
For more information about the research team, visit the website: https://sites.google.com/site/miheelimlab/1-professor-mi-hee-lim.
2022.07.29 View 15892
KAIST Research Team Proves How a Neurotransmitter may be the Key in Controlling Alzheimer’s Toxicity
With nearly 50 million dementia patients worldwide, and Alzheimers’s disease is the most common neurodegenerative disease. Its main symptom is the impairment of general cognitive abilities, including the ability to speak or to remember. The importance of finding a cure is widely understood with increasingly aging population and the life expectancy being ever-extended. However, even the cause of the grim disease is yet to be given a clear definition.
A KAIST research team in the Department of Chemistry led by professor Mi Hee Lim took on a lead to discovered a new role for somatostatin, a protein-based neurotransmitter, in reducing the toxicity caused in the pathogenic mechanism taken towards development of Alzheimer’s disease. The study was published in the July issue of Nature Chemistry under the title, “Conformational and functional changes of the native neuropeptide somatostatin occur in the presence of copper and amyloid-β”.
According to the amyloid hypothesis, the abnormal deposition of Aβ proteins causes death of neuronal cells. While Aβ agglomerations make up most of the aged plaques through fibrosis, in recent studies, high concentrations of transitional metal were found in the plaques from Alzheimer’s patients.
This suggests a close interaction between metallic ions and Aβ, which accelerates the fibrosis of proteins. Copper in particular is a redox-activating transition metal that can produce large amounts of oxygen and cause serious oxidative stress on cell organelles. Aβ proteins and transition metals can closely interact with neurotransmitters at synapses, but the direct effects of such abnormalities on the structure and function of neurotransmitters are yet to be understood.
Figure 1. Functional shift of somatostatin (SST) by factors in the pathogenesis of Alzheimer's disease.
Figure 2. Somatostatin’s loss-of-function as neurotransmitter. a. Schematic diagram of SST auto-aggregation due to Alzheimer's pathological factors. b. SST’s aggregation by copper ions. c. Coordination-prediction structure and N-terminal folding of copper-SST. d. Inhibition of SST receptor binding specificity by metals.
In their research, Professor Lim’s team discovered that when somatostatin, the protein-based neurotransmitter, is met with copper, Aβ, and metal-Aβ complexes, self-aggregates and ceases to perform its innate function of transmitting neural signals, but begins to attenuate the toxicity and agglomeration of metal-Aβ complexes.
Figure 3. Gain-of-function of somatostatin (SST) in the dementia setting. a. Prediction of docking of SST and amyloid beta. b. SST making metal-amyloid beta aggregates into an amorphous form. c. Cytotoxic mitigation effect of SST. d. SST mitigating the interaction between amyloid beta protein with the cell membrane.
This research, by Dr. Jiyeon Han et al. from the KAIST Department of Chemistry, revealed the coordination structure between copper and somatostatin at a molecular level through which it suggested the agglomeration mechanism, and discovered the effects of somatostatin on Aβ agglomeration path depending on the presence or absence of metals. The team has further confirmed somatostatin’s receptor binding, interactions with cell membranes, and effects on cell toxicity for the first time to receive international attention.
Professor Mi Hee Lim said, “This research has great significance in having discovered a new role of neurotransmitters in the pathogenesis of Alzheimer’s disease.” “We expect this research to contribute to defining the pathogenic network of neurodegenerative diseases caused by aging, and to the development of future biomarkers and medicine,” she added.
This research was conducted jointly by Professor Seung-Hee Lee’s team of KAIST Department of Biological Sciences, Professor Kiyoung Park’s Team of KAIST Department of Chemistry, and Professor Yulong Li’s team of Peking University.
The research was funded by Basic Science Research Program of the National Research Foundation of Korea and KAIST.
For more information about the research team, visit the website: https://sites.google.com/site/miheelimlab/1-professor-mi-hee-lim.
2022.07.29 View 15892 -
 Success in Real-Time Observation of the Formation Process of Topological Solitons, a Core Technology for Next-Generation Information Transfer
< From left) Geonhyeong Park (Ph.D. Candidate), Yun-Seok Choi (Ph.D.), Professor Dong Ki Yoon, and Changjae Lee (Ph.D. Candidate) of the Department of Chemistry >
Professor Dong Ki Yoon's research team in the Department of Chemistry at KAIST announced on the 11th that they have succeeded in controlling the formation of topological solitons in a regular, large-area manner through the self-assembly of chiral liquid crystal materials and observing their formation process in real-time.
A soliton refers to a phenomenon where a specific wave persists without dissipating through interaction with its surroundings. In particular, even when a wave is transmitted over long distances, it retains its unique information until it reaches the desired destination. Therefore, in today's digital society, which is susceptible to hacking, solitons are highly anticipated to be the core of future communication due to their inherent high stability. Furthermore, topological solitons created using organic liquid crystal molecules are expected to be utilized as next-generation anti-counterfeiting devices and memory elements due to their unique spin directionality.
Professor Yoon's team specifically revealed the formation process of topological solitons in this study, which had not been observable in real-time under mild conditions such as room temperature until now. This was made possible by using self-assembling chiral liquid crystal materials in a confined space created by air pillars.
This research, in which Geonhyeong Park (Ph.D. Candidate, Department of Chemistry) and Dr. Ahram Suh participated as co-first authors, and Dr. Yun-Seok Choi and Changjae Lee (Ph.D. Candidate) from the same group also participated, was published online in the international journal 'Advanced Materials' on June 5th and is scheduled to be featured as the back cover of the July issue. (Paper title: "Fabrication of Arrays of Topological Solitons in Patterned Chiral Liquid Crystals for Real-Time Observation of Morphogenesis")
< Figure 1. Schematic diagram of the research>
< Figure 2. Real-time observation of topological soliton formation using liquid crystals>
In this study, Professor Yoon's team implemented topological soliton structures at approximately 30 degrees Celsius, similar to room temperature, using chiral (asymmetric) liquid crystal materials instead of the conventional liquid crystal molecules widely used as core materials in liquid crystal displays (LCDs). Generally, complex equipment is required to control the formation of topological solitons, and their formation time is very short, which has hindered research into their formation process until now.
To achieve regular formation and control of topological solitons formed by chiral liquid crystal molecules, Professor Yoon's team precisely controlled a combination of vertical alignment layers, which can orient molecules vertically, and air pillars. Specifically, they prepared concave patterns based on circular silicon material, several micrometers (one-millionth of a meter) in size, coated with a vertical alignment layer, and a glass substrate. By adjusting the gap to several micrometers and injecting chiral liquid crystal material, air pillars were spontaneously formed on the concave patterns. Subsequently, the liquid crystal molecules were vertically aligned on all substrates, inevitably causing regular distortions between the substrates, and between the substrate and the air pillars, thus developing a system where chiral molecular structures, i.e., topological solitons, could be formed.
The key to the formation and control of topological solitons lies in controlling the thermal phase transition to occur regularly as desired when cooling from the isotropic phase temperature (approximately 40 degrees Celsius) to the liquid crystal phase temperature (approximately 30 degrees Celsius), where the liquid crystal material near the air pillars is cooler than the liquid crystal material between the glass substrate and the silicon patterned parts. This is consistent with the everyday wisdom of eating steamed eggs from a 'Ttukbaegi' (earthen pot) by starting from the relatively cooler part exposed to the air (near the air pillars) rather than the hot pot part (silicon or glass substrate part).
Through real-time analysis, the research team elucidated that topological defects are formed by the naturally formed air pillars through controlled thermal phase transition, and topological solitons are formed only at the locations of these defects. This analysis technique has the potential for application in various fields, including the interpretation of topological soliton formation found in other physical phenomena such as skyrmion particles in electromagnetism.
< Figure 3. Snapshots during the formation process of regularly arranged topological solitons>
Professor Dong Ki Yoon stated, "General topological solitons are known to be highly stable, capable only of generation or annihilation. Through the results of this research, we can understand the formation process of solitons in more detail, and they can be used as spintronics application technology, considered a next-generation semiconductor device for storing and recording information."
This research was conducted in collaboration with Professor Ivan Smalyukh's laboratory at the University of Colorado, Department of Physics, and was supported by the Multiscale Chiral Structures Research Center and strategic projects of the National Research Foundation of Korea under the Ministry of Science and ICT.
2022.07.11 View 488
Success in Real-Time Observation of the Formation Process of Topological Solitons, a Core Technology for Next-Generation Information Transfer
< From left) Geonhyeong Park (Ph.D. Candidate), Yun-Seok Choi (Ph.D.), Professor Dong Ki Yoon, and Changjae Lee (Ph.D. Candidate) of the Department of Chemistry >
Professor Dong Ki Yoon's research team in the Department of Chemistry at KAIST announced on the 11th that they have succeeded in controlling the formation of topological solitons in a regular, large-area manner through the self-assembly of chiral liquid crystal materials and observing their formation process in real-time.
A soliton refers to a phenomenon where a specific wave persists without dissipating through interaction with its surroundings. In particular, even when a wave is transmitted over long distances, it retains its unique information until it reaches the desired destination. Therefore, in today's digital society, which is susceptible to hacking, solitons are highly anticipated to be the core of future communication due to their inherent high stability. Furthermore, topological solitons created using organic liquid crystal molecules are expected to be utilized as next-generation anti-counterfeiting devices and memory elements due to their unique spin directionality.
Professor Yoon's team specifically revealed the formation process of topological solitons in this study, which had not been observable in real-time under mild conditions such as room temperature until now. This was made possible by using self-assembling chiral liquid crystal materials in a confined space created by air pillars.
This research, in which Geonhyeong Park (Ph.D. Candidate, Department of Chemistry) and Dr. Ahram Suh participated as co-first authors, and Dr. Yun-Seok Choi and Changjae Lee (Ph.D. Candidate) from the same group also participated, was published online in the international journal 'Advanced Materials' on June 5th and is scheduled to be featured as the back cover of the July issue. (Paper title: "Fabrication of Arrays of Topological Solitons in Patterned Chiral Liquid Crystals for Real-Time Observation of Morphogenesis")
< Figure 1. Schematic diagram of the research>
< Figure 2. Real-time observation of topological soliton formation using liquid crystals>
In this study, Professor Yoon's team implemented topological soliton structures at approximately 30 degrees Celsius, similar to room temperature, using chiral (asymmetric) liquid crystal materials instead of the conventional liquid crystal molecules widely used as core materials in liquid crystal displays (LCDs). Generally, complex equipment is required to control the formation of topological solitons, and their formation time is very short, which has hindered research into their formation process until now.
To achieve regular formation and control of topological solitons formed by chiral liquid crystal molecules, Professor Yoon's team precisely controlled a combination of vertical alignment layers, which can orient molecules vertically, and air pillars. Specifically, they prepared concave patterns based on circular silicon material, several micrometers (one-millionth of a meter) in size, coated with a vertical alignment layer, and a glass substrate. By adjusting the gap to several micrometers and injecting chiral liquid crystal material, air pillars were spontaneously formed on the concave patterns. Subsequently, the liquid crystal molecules were vertically aligned on all substrates, inevitably causing regular distortions between the substrates, and between the substrate and the air pillars, thus developing a system where chiral molecular structures, i.e., topological solitons, could be formed.
The key to the formation and control of topological solitons lies in controlling the thermal phase transition to occur regularly as desired when cooling from the isotropic phase temperature (approximately 40 degrees Celsius) to the liquid crystal phase temperature (approximately 30 degrees Celsius), where the liquid crystal material near the air pillars is cooler than the liquid crystal material between the glass substrate and the silicon patterned parts. This is consistent with the everyday wisdom of eating steamed eggs from a 'Ttukbaegi' (earthen pot) by starting from the relatively cooler part exposed to the air (near the air pillars) rather than the hot pot part (silicon or glass substrate part).
Through real-time analysis, the research team elucidated that topological defects are formed by the naturally formed air pillars through controlled thermal phase transition, and topological solitons are formed only at the locations of these defects. This analysis technique has the potential for application in various fields, including the interpretation of topological soliton formation found in other physical phenomena such as skyrmion particles in electromagnetism.
< Figure 3. Snapshots during the formation process of regularly arranged topological solitons>
Professor Dong Ki Yoon stated, "General topological solitons are known to be highly stable, capable only of generation or annihilation. Through the results of this research, we can understand the formation process of solitons in more detail, and they can be used as spintronics application technology, considered a next-generation semiconductor device for storing and recording information."
This research was conducted in collaboration with Professor Ivan Smalyukh's laboratory at the University of Colorado, Department of Physics, and was supported by the Multiscale Chiral Structures Research Center and strategic projects of the National Research Foundation of Korea under the Ministry of Science and ICT.
2022.07.11 View 488 -
 New Polymer Mesophase Structure Discovered
Bilayer-folded lamellar mesophase induced by random polymer sequence
Polymers, large molecules made up of repeating smaller molecules called monomers, are found in nearly everything we use in our day-to-day lives. Polymers can be natural or created synthetically. Natural polymers, also called biopolymers, include DNA, proteins, and materials like silk, gelatin, and collagen. Synthetic polymers make up many different kinds of materials, including plastic, that are used in constructing everything from toys to industrial fiber cables to brake pads.
As polymers are formed through a process called polymerization, the monomers are connected through a chain. As the chain develops, the structure of the polymer determines its unique physical and chemical properties. Researchers are continually studying polymers, how they form, how they are structured, and how they develop these unique properties. By understanding this information, scientists can develop new uses for polymers and create new materials that can be used in a wide variety of industries.
In a paper published in Nature Communications on May 4, researchers describe a new structure found in an aqueous solution of an amphiphilic copolymer, called a bilayer-folded lamellar mesophase, that has been discovered through a random copolymer sequence.
“A new mesophase is an important discovery as it shows a new way for molecules to self-organize,” said Professor Myungeun Seo at the Department of Chemistry at KAIST. “We were particularly thrilled to identify this bilayer-folded lamellar phase because pure bilayer membranes are difficult to fold thermodynamically.”
Researchers think that this mesophase structure comes from the sequence of the monomers within the copolymer. The way the different monomers arrange themselves in the chain that makes up a copolymer is important and can have implications for what the copolymer can do. Many copolymers are random, which means that their structure relies on how the monomers interact with each other. In this case, the interaction between the hydrophobic monomers associates the copolymer chains to conceal the hydrophobic domain from water. As the structure gets more complex, researchers have found that a visible order develops so that monomers can be matched up with the right pair.
“While we tend to think random means disorder, here we showed that a periodic order can spontaneously arise from the random copolymer sequence based on their collective behavior,” said Professor Seo. “We believe this comes from the sequence matching problem: finding a perfectly complementary pair for a long sequence is nearly impossible.”
This is what creates the unique structure of this newly discovered mesophase. The copolymer spontaneously folds and creates a multilamellar structure that is separated by water. A multilamellar structure refers to plate-like folds and the folded layers stack on top of each other. The resulting mesophase is birefringent, meaning light refracts through it, it is similar to liquid crystalline, and viscoelastic, which means that it is both viscous and elastic at the same time.
Looking ahead, researchers hope to learn more about this new mesophase and figure out how to control the outcome. Once more is understood about the mesophase and how it is formed, it’s possible that new mesophases could be discovered as more sequences are researched. “One of the obvious questions for us is how to control the folding frequency and adjust the folded height, which we are currently working to address. Ultimately, we want to understand how different multinary sequences can associate with another to create order and apply the knowledge to develop new materials,” said Professor Seo. The National Research Foundation, the Ministry of Education, and the Ministry of Science and ICT of Korea funded this research.
-PublicationMinjoong Shin, Hayeon Kim, Geonhyeong Park, Jongmin Park, Hyungju Ahn, Dong Ki Yoon, Eunji Lee, Myungeun Seo, “Bilayer-folded lamellar mesophase induced by random polymersequence,” May 4, 2022, Nature Communications (https://doi.org/10.1038/s41467-022-30122-z)
-ProfileProfessor Myungeun SeoMacromolecular Materials Chemistry Lab (https://nanopsg.kaist.ac.kr/)Department of ChemistryCollege of Natural SciencesKAIST
2022.06.17 View 9846
New Polymer Mesophase Structure Discovered
Bilayer-folded lamellar mesophase induced by random polymer sequence
Polymers, large molecules made up of repeating smaller molecules called monomers, are found in nearly everything we use in our day-to-day lives. Polymers can be natural or created synthetically. Natural polymers, also called biopolymers, include DNA, proteins, and materials like silk, gelatin, and collagen. Synthetic polymers make up many different kinds of materials, including plastic, that are used in constructing everything from toys to industrial fiber cables to brake pads.
As polymers are formed through a process called polymerization, the monomers are connected through a chain. As the chain develops, the structure of the polymer determines its unique physical and chemical properties. Researchers are continually studying polymers, how they form, how they are structured, and how they develop these unique properties. By understanding this information, scientists can develop new uses for polymers and create new materials that can be used in a wide variety of industries.
In a paper published in Nature Communications on May 4, researchers describe a new structure found in an aqueous solution of an amphiphilic copolymer, called a bilayer-folded lamellar mesophase, that has been discovered through a random copolymer sequence.
“A new mesophase is an important discovery as it shows a new way for molecules to self-organize,” said Professor Myungeun Seo at the Department of Chemistry at KAIST. “We were particularly thrilled to identify this bilayer-folded lamellar phase because pure bilayer membranes are difficult to fold thermodynamically.”
Researchers think that this mesophase structure comes from the sequence of the monomers within the copolymer. The way the different monomers arrange themselves in the chain that makes up a copolymer is important and can have implications for what the copolymer can do. Many copolymers are random, which means that their structure relies on how the monomers interact with each other. In this case, the interaction between the hydrophobic monomers associates the copolymer chains to conceal the hydrophobic domain from water. As the structure gets more complex, researchers have found that a visible order develops so that monomers can be matched up with the right pair.
“While we tend to think random means disorder, here we showed that a periodic order can spontaneously arise from the random copolymer sequence based on their collective behavior,” said Professor Seo. “We believe this comes from the sequence matching problem: finding a perfectly complementary pair for a long sequence is nearly impossible.”
This is what creates the unique structure of this newly discovered mesophase. The copolymer spontaneously folds and creates a multilamellar structure that is separated by water. A multilamellar structure refers to plate-like folds and the folded layers stack on top of each other. The resulting mesophase is birefringent, meaning light refracts through it, it is similar to liquid crystalline, and viscoelastic, which means that it is both viscous and elastic at the same time.
Looking ahead, researchers hope to learn more about this new mesophase and figure out how to control the outcome. Once more is understood about the mesophase and how it is formed, it’s possible that new mesophases could be discovered as more sequences are researched. “One of the obvious questions for us is how to control the folding frequency and adjust the folded height, which we are currently working to address. Ultimately, we want to understand how different multinary sequences can associate with another to create order and apply the knowledge to develop new materials,” said Professor Seo. The National Research Foundation, the Ministry of Education, and the Ministry of Science and ICT of Korea funded this research.
-PublicationMinjoong Shin, Hayeon Kim, Geonhyeong Park, Jongmin Park, Hyungju Ahn, Dong Ki Yoon, Eunji Lee, Myungeun Seo, “Bilayer-folded lamellar mesophase induced by random polymersequence,” May 4, 2022, Nature Communications (https://doi.org/10.1038/s41467-022-30122-z)
-ProfileProfessor Myungeun SeoMacromolecular Materials Chemistry Lab (https://nanopsg.kaist.ac.kr/)Department of ChemistryCollege of Natural SciencesKAIST
2022.06.17 View 9846 -
 Distinguished Professor Sukbok Chang Named the 2022 Ho-Am Laureate
Distinguished Professor Sukbok Chang from the Department of Chemistry was named the awardee of the Ho-Am Prize in the fields of chemistry and life sciences. The award has recognized the most distinguished scholars, individuals, and organizations in physics and mathematics, chemistry and life sciences, engineering, medicine, arts, and community service in honor of the late founder of Samsung Group Byong-Chul Lee, whose penname is Ho-Am. The awards ceremony will be held on May 31 and awardees will receive 300 million KRW in prize money.
Professor Chang became the fourth KAIST Ho-Am laureate following Distinguished Professor Sang Yup Lee in engineering in 2014, Distinguished Professor Jun Ho Oh in engineering in 2016, and Distinguished Professor Gou Young Koh in medicine in 2018.
Professor Chang is a renowned chemist who has made pioneering research in the area of transition metal catalysis for organic transformations. Professor Chang is also one of the Highly Cited Researchers who rank in the top 1% of citations by field and publication year in the Web of Science citation index. He has made the list seven years in a row from 2016.
Professor Chang has developed a range of new and impactful C-H bond functionalization reactions. By using his approaches, value-added molecules can be readily produced from chemical feedstocks, representatively hydrocarbons and (hetero)arenes. His research team elucidated fundamental key mechanistic aspects in the course of the essential C-H bond activation process of unreactive starting materials. He was able to utilize the obtained mechanistic understanding for the subsequent catalyst design to develop more efficient and highly (stereo)selective catalytic reactions.
Among the numerous contributions he made, the design of new mechanistic approaches toward metal nitrenoid transfers are of especially high impact to the chemical community. Indeed, a series of important transition metal catalyst systems were developed by Professor Chang to enable the direct and selective C-H amidation of unreactive organic compounds, thereby producing aminated compounds that have important applicability in synthetic, medicinal, and materials science. He has also pioneered in the area of asymmetric C-H amination chemistry by creatively devising various types of chiral transition metal catalyst systems, and his team proved for the first time that chiral lactam compounds can be obtained at an excellent level of stereoselectivity.
Another significant contribution of Professor. Chang was the introduction of dioxazolones as a robust but highly reactive source of acyl nitrenoids for the catalytic C-H amidation reactions, and this reagent is now broadly utilized in synthetic chemistry worldwide.
Professor Chang also leads a research group in the Center for Catalytic Hydrocarbon Functionalizations at the Institute for Basic Science.
2022.04.06 View 8901
Distinguished Professor Sukbok Chang Named the 2022 Ho-Am Laureate
Distinguished Professor Sukbok Chang from the Department of Chemistry was named the awardee of the Ho-Am Prize in the fields of chemistry and life sciences. The award has recognized the most distinguished scholars, individuals, and organizations in physics and mathematics, chemistry and life sciences, engineering, medicine, arts, and community service in honor of the late founder of Samsung Group Byong-Chul Lee, whose penname is Ho-Am. The awards ceremony will be held on May 31 and awardees will receive 300 million KRW in prize money.
Professor Chang became the fourth KAIST Ho-Am laureate following Distinguished Professor Sang Yup Lee in engineering in 2014, Distinguished Professor Jun Ho Oh in engineering in 2016, and Distinguished Professor Gou Young Koh in medicine in 2018.
Professor Chang is a renowned chemist who has made pioneering research in the area of transition metal catalysis for organic transformations. Professor Chang is also one of the Highly Cited Researchers who rank in the top 1% of citations by field and publication year in the Web of Science citation index. He has made the list seven years in a row from 2016.
Professor Chang has developed a range of new and impactful C-H bond functionalization reactions. By using his approaches, value-added molecules can be readily produced from chemical feedstocks, representatively hydrocarbons and (hetero)arenes. His research team elucidated fundamental key mechanistic aspects in the course of the essential C-H bond activation process of unreactive starting materials. He was able to utilize the obtained mechanistic understanding for the subsequent catalyst design to develop more efficient and highly (stereo)selective catalytic reactions.
Among the numerous contributions he made, the design of new mechanistic approaches toward metal nitrenoid transfers are of especially high impact to the chemical community. Indeed, a series of important transition metal catalyst systems were developed by Professor Chang to enable the direct and selective C-H amidation of unreactive organic compounds, thereby producing aminated compounds that have important applicability in synthetic, medicinal, and materials science. He has also pioneered in the area of asymmetric C-H amination chemistry by creatively devising various types of chiral transition metal catalyst systems, and his team proved for the first time that chiral lactam compounds can be obtained at an excellent level of stereoselectivity.
Another significant contribution of Professor. Chang was the introduction of dioxazolones as a robust but highly reactive source of acyl nitrenoids for the catalytic C-H amidation reactions, and this reagent is now broadly utilized in synthetic chemistry worldwide.
Professor Chang also leads a research group in the Center for Catalytic Hydrocarbon Functionalizations at the Institute for Basic Science.
2022.04.06 View 8901 -
 Mathematicians Identify a Key Source of Cell-to-Cell Variability in Cell Signaling
Systematic inferences identify a major source of heterogeneity in cell signaling dynamics
Why do genetically identical cells respond differently to the same external stimuli, such as antibiotics? This long-standing mystery has been solved by KAIST and IBS mathematicians who have developed a new framework for analyzing cell responses to some stimuli. The team found that the cell-to-cell variability in antibiotic stress response increases as the effective length of the cell signaling pathway (i.e., the number of rate-limiting steps) increases. This finding could identify more effective chemotherapies to overcome the fractional killing of cancer cells caused by cell-to-cell variability.
Cells in the human body contain signal transduction systems that respond to various external stimuli such as antibiotics and changes in osmotic pressure. When an external stimulus is detected, various biochemical reactions occur sequentially. This leads to the expression of relevant genes, allowing the cells to respond to the perturbed external environment. Furthermore, signal transduction leads to a drug response (e.g., antibiotic resistance genes are expressed when antibiotic drugs are given).
However, even when the same external stimuli are detected, the responses of individual cells are greatly heterogeneous. This leads to the emergence of persister cells that are highly resistant to drugs. To identify potential sources of this cell-to cell variability, many studies have been conducted. However, most of the intermediate signal transduction reactions are unobservable with current experimental techniques.
A group of researchers including Dae Wook Kim and Hyukpyo Hong and led by Professor Jae Kyoung Kim from the KAIST Department of Mathematical Sciences and IBS Biomedical Mathematics Group solved the mystery by exploiting queueing theory and Bayesian inference methodology. They proposed a queueing process that describes the signal transduction system in cells. Based on this, they developed Bayesian inference computational software using MBI (the Moment-based Bayesian Inference method). This enables the analysis of the signal transduction system without a direct observation of the intermediate steps. This study was published in Science Advances.
By analyzing experimental data from Escherichia coli using MBI, the research team found that cell-to-cell variability increases as the number of rate-limiting steps in the signaling pathway increases.
The rate-limiting steps denote the slowest steps (i.e., bottlenecks) in sequential biochemical reaction steps composing cell signaling pathways and thus dominates most of the signaling time. As the number of the rate-limiting steps increases, the intensity of the transduced signal becomes greatly heterogeneous even in a population of genetically identical cells. This finding is expected to provide a new paradigm for studying the heterogeneous antibiotic resistance of cells, which is a big challenge in cancer medicine.
Professor Kim said, “As a mathematician, I am excited to help advance the understanding of cell-to-cell variability in response to external stimuli. I hope this finding facilitates the development of more effective chemotherapies.”
This work was supported by the Samsung Science and Technology Foundation, the National Research Foundation of Korea, and the Institute for Basic Science.
-Publication:Dae Wook Kim, Hyukpyo Hong, and Jae Kyoung Kim (2022) “Systematic inference identifies a major source of heterogeneity in cell signaling dynamics: the rate-limiting step number,”Science Advances March 18, 2022 (DOI: 10.1126/sciadv.abl4598)
-Profile:Professor Jae Kyoung Kimhttp://mathsci.kaist.ac.kr/~jaekkim
jaekkim@kaist.ac.kr@umichkim on TwitterDepartment of Mathematical SciencesKAIST
2022.03.29 View 11405
Mathematicians Identify a Key Source of Cell-to-Cell Variability in Cell Signaling
Systematic inferences identify a major source of heterogeneity in cell signaling dynamics
Why do genetically identical cells respond differently to the same external stimuli, such as antibiotics? This long-standing mystery has been solved by KAIST and IBS mathematicians who have developed a new framework for analyzing cell responses to some stimuli. The team found that the cell-to-cell variability in antibiotic stress response increases as the effective length of the cell signaling pathway (i.e., the number of rate-limiting steps) increases. This finding could identify more effective chemotherapies to overcome the fractional killing of cancer cells caused by cell-to-cell variability.
Cells in the human body contain signal transduction systems that respond to various external stimuli such as antibiotics and changes in osmotic pressure. When an external stimulus is detected, various biochemical reactions occur sequentially. This leads to the expression of relevant genes, allowing the cells to respond to the perturbed external environment. Furthermore, signal transduction leads to a drug response (e.g., antibiotic resistance genes are expressed when antibiotic drugs are given).
However, even when the same external stimuli are detected, the responses of individual cells are greatly heterogeneous. This leads to the emergence of persister cells that are highly resistant to drugs. To identify potential sources of this cell-to cell variability, many studies have been conducted. However, most of the intermediate signal transduction reactions are unobservable with current experimental techniques.
A group of researchers including Dae Wook Kim and Hyukpyo Hong and led by Professor Jae Kyoung Kim from the KAIST Department of Mathematical Sciences and IBS Biomedical Mathematics Group solved the mystery by exploiting queueing theory and Bayesian inference methodology. They proposed a queueing process that describes the signal transduction system in cells. Based on this, they developed Bayesian inference computational software using MBI (the Moment-based Bayesian Inference method). This enables the analysis of the signal transduction system without a direct observation of the intermediate steps. This study was published in Science Advances.
By analyzing experimental data from Escherichia coli using MBI, the research team found that cell-to-cell variability increases as the number of rate-limiting steps in the signaling pathway increases.
The rate-limiting steps denote the slowest steps (i.e., bottlenecks) in sequential biochemical reaction steps composing cell signaling pathways and thus dominates most of the signaling time. As the number of the rate-limiting steps increases, the intensity of the transduced signal becomes greatly heterogeneous even in a population of genetically identical cells. This finding is expected to provide a new paradigm for studying the heterogeneous antibiotic resistance of cells, which is a big challenge in cancer medicine.
Professor Kim said, “As a mathematician, I am excited to help advance the understanding of cell-to-cell variability in response to external stimuli. I hope this finding facilitates the development of more effective chemotherapies.”
This work was supported by the Samsung Science and Technology Foundation, the National Research Foundation of Korea, and the Institute for Basic Science.
-Publication:Dae Wook Kim, Hyukpyo Hong, and Jae Kyoung Kim (2022) “Systematic inference identifies a major source of heterogeneity in cell signaling dynamics: the rate-limiting step number,”Science Advances March 18, 2022 (DOI: 10.1126/sciadv.abl4598)
-Profile:Professor Jae Kyoung Kimhttp://mathsci.kaist.ac.kr/~jaekkim
jaekkim@kaist.ac.kr@umichkim on TwitterDepartment of Mathematical SciencesKAIST
2022.03.29 View 11405 -
 Tomographic Measurement of Dielectric Tensors
Dielectric tensor tomography allows the direct measurement of the 3D dielectric tensors of optically anisotropic structures
A research team reported the direct measurement of dielectric tensors of anisotropic structures including the spatial variations of principal refractive indices and directors. The group also demonstrated quantitative tomographic measurements of various nematic liquid-crystal structures and their fast 3D nonequilibrium dynamics using a 3D label-free tomographic method. The method was described in Nature Materials.
Light-matter interactions are described by the dielectric tensor. Despite their importance in basic science and applications, it has not been possible to measure 3D dielectric tensors directly. The main challenge was due to the vectorial nature of light scattering from a 3D anisotropic structure. Previous approaches only addressed 3D anisotropic information indirectly and were limited to two-dimensional, qualitative, strict sample conditions or assumptions.
The research team developed a method enabling the tomographic reconstruction of 3D dielectric tensors without any preparation or assumptions. A sample is illuminated with a laser beam with various angles and circularly polarization states. Then, the light fields scattered from a sample are holographically measured and converted into vectorial diffraction components. Finally, by inversely solving a vectorial wave equation, the 3D dielectric tensor is reconstructed.
Professor YongKeun Park said, “There were a greater number of unknowns in direct measuring than with the conventional approach. We applied our approach to measure additional holographic images by slightly tilting the incident angle.”
He said that the slightly tilted illumination provides an additional orthogonal polarization, which makes the underdetermined problem become the determined problem. “Although scattered fields are dependent on the illumination angle, the Fourier differentiation theorem enables the extraction of the same dielectric tensor for the slightly tilted illumination,” Professor Park added.
His team’s method was validated by reconstructing well-known liquid crystal (LC) structures, including the twisted nematic, hybrid aligned nematic, radial, and bipolar configurations. Furthermore, the research team demonstrated the experimental measurements of the non-equilibrium dynamics of annihilating, nucleating, and merging LC droplets, and the LC polymer network with repeating 3D topological defects.
“This is the first experimental measurement of non-equilibrium dynamics and 3D topological defects in LC structures in a label-free manner. Our method enables the exploration of inaccessible nematic structures and interactions in non-equilibrium dynamics,” first author Dr. Seungwoo Shin explained.
-PublicationSeungwoo Shin, Jonghee Eun, Sang Seok Lee, Changjae Lee, Herve Hugonnet, Dong Ki Yoon, Shin-Hyun Kim, Jongwoo Jeong, YongKeun Park, “Tomographic Measurement ofDielectric Tensors at Optical Frequency,” Nature Materials March 02, 2022 (https://doi.org/10/1038/s41563-022-01202-8)
-ProfileProfessor YongKeun ParkBiomedical Optics Laboratory (http://bmol.kaist.ac.kr)Department of PhysicsCollege of Natural SciencesKAIST
2022.03.22 View 9472
Tomographic Measurement of Dielectric Tensors
Dielectric tensor tomography allows the direct measurement of the 3D dielectric tensors of optically anisotropic structures
A research team reported the direct measurement of dielectric tensors of anisotropic structures including the spatial variations of principal refractive indices and directors. The group also demonstrated quantitative tomographic measurements of various nematic liquid-crystal structures and their fast 3D nonequilibrium dynamics using a 3D label-free tomographic method. The method was described in Nature Materials.
Light-matter interactions are described by the dielectric tensor. Despite their importance in basic science and applications, it has not been possible to measure 3D dielectric tensors directly. The main challenge was due to the vectorial nature of light scattering from a 3D anisotropic structure. Previous approaches only addressed 3D anisotropic information indirectly and were limited to two-dimensional, qualitative, strict sample conditions or assumptions.
The research team developed a method enabling the tomographic reconstruction of 3D dielectric tensors without any preparation or assumptions. A sample is illuminated with a laser beam with various angles and circularly polarization states. Then, the light fields scattered from a sample are holographically measured and converted into vectorial diffraction components. Finally, by inversely solving a vectorial wave equation, the 3D dielectric tensor is reconstructed.
Professor YongKeun Park said, “There were a greater number of unknowns in direct measuring than with the conventional approach. We applied our approach to measure additional holographic images by slightly tilting the incident angle.”
He said that the slightly tilted illumination provides an additional orthogonal polarization, which makes the underdetermined problem become the determined problem. “Although scattered fields are dependent on the illumination angle, the Fourier differentiation theorem enables the extraction of the same dielectric tensor for the slightly tilted illumination,” Professor Park added.
His team’s method was validated by reconstructing well-known liquid crystal (LC) structures, including the twisted nematic, hybrid aligned nematic, radial, and bipolar configurations. Furthermore, the research team demonstrated the experimental measurements of the non-equilibrium dynamics of annihilating, nucleating, and merging LC droplets, and the LC polymer network with repeating 3D topological defects.
“This is the first experimental measurement of non-equilibrium dynamics and 3D topological defects in LC structures in a label-free manner. Our method enables the exploration of inaccessible nematic structures and interactions in non-equilibrium dynamics,” first author Dr. Seungwoo Shin explained.
-PublicationSeungwoo Shin, Jonghee Eun, Sang Seok Lee, Changjae Lee, Herve Hugonnet, Dong Ki Yoon, Shin-Hyun Kim, Jongwoo Jeong, YongKeun Park, “Tomographic Measurement ofDielectric Tensors at Optical Frequency,” Nature Materials March 02, 2022 (https://doi.org/10/1038/s41563-022-01202-8)
-ProfileProfessor YongKeun ParkBiomedical Optics Laboratory (http://bmol.kaist.ac.kr)Department of PhysicsCollege of Natural SciencesKAIST
2022.03.22 View 9472 -
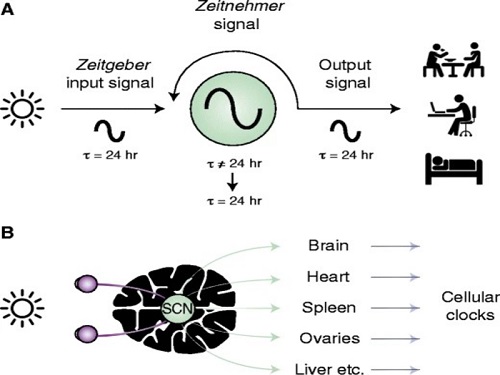 Scientist Discover How Circadian Rhythm Can Be Both Strong and Flexible
Study reveals that master and slave oscillators function via different molecular mechanisms
From tiny fruit flies to human beings, all animals on Earth maintain their daily rhythms based on their internal circadian clock. The circadian clock enables organisms to undergo rhythmic changes in behavior and physiology based on a 24-hour circadian cycle. For example, our own biological clock tells our brain to release melatonin, a sleep-inducing hormone, at night time.
The discovery of the molecular mechanism of the circadian clock was bestowed the Nobel Prize in Physiology or Medicine 2017. From what we know, no one centralized clock is responsible for our circadian cycles. Instead, it operates in a hierarchical network where there are “master pacemaker” and “slave oscillator”.
The master pacemaker receives various input signals from the environment such as light. The master then drives the slave oscillator that regulates various outputs such as sleep, feeding, and metabolism. Despite the different roles of the pacemaker neurons, they are known to share common molecular mechanisms that are well conserved in all lifeforms. For example, interlocked systems of multiple transcriptional-translational feedback loops (TTFLs) composed of core clock proteins have been deeply studied in fruit flies.
However, there is still much that we need to learn about our own biological clock. The hierarchically-organized nature of master and slave clock neurons leads to a prevailing belief that they share an identical molecular clockwork. At the same time, the different roles they serve in regulating bodily rhythms also raise the question of whether they might function under different molecular clockworks.
Research team led by Professor Kim Jae Kyoung from the Department of Mathematical Sciences, a chief investigator at the Biomedical Mathematics Group at the Institute for Basic Science, used a combination of mathematical and experimental approaches using fruit flies to answer this question. The team found that the master clock and the slave clock operate via different molecular mechanisms.
In both master and slave neurons of fruit flies, a circadian rhythm-related protein called PER is produced and degraded at different rates depending on the time of the day. Previously, the team found that the master clock neuron (sLNvs) and the slave clock neuron (DN1ps) have different profiles of PER in wild-type and Clk-Δ mutant Drosophila. This hinted that there might be a potential difference in molecular clockworks between the master and slave clock neurons.
However, due to the complexity of the molecular clockwork, it was challenging to identify the source of such differences. Thus, the team developed a mathematical model describing the molecular clockworks of the master and slave clocks. Then, all possible molecular differences between the master and slave clock neurons were systematically investigated by using computer simulations. The model predicted that PER is more efficiently produced and then rapidly degraded in the master clock compared to the slave clock neurons. This prediction was then confirmed by the follow-up experiments using animal.
Then, why do the master clock neurons have such different molecular properties from the slave clock neurons? To answer this question, the research team again used the combination of mathematical model simulation and experiments. It was found that the faster rate of synthesis of PER in the master clock neurons allows them to generate synchronized rhythms with a high level of amplitude. Generation of such a strong rhythm with high amplitude is critical to delivering clear signals to slave clock neurons.
However, such strong rhythms would typically be unfavorable when it comes to adapting to environmental changes. These include natural causes such as different daylight hours across summer and winter seasons, up to more extreme artificial cases such as jet lag that occurs after international travel. Thanks to the distinct property of the master clock neurons, it is able to undergo phase dispersion when the standard light-dark cycle is disrupted, drastically reducing the level of PER. The master clock neurons can then easily adapt to the new diurnal cycle. Our master pacemaker’s plasticity explains how we can quickly adjust to the new time zones after international flights after just a brief period of jet lag.
It is hoped that the findings of this study can have future clinical implications when it comes to treating various disorders that affect our circadian rhythm. Professor Kim notes, “When the circadian clock loses its robustness and flexibility, the circadian rhythms sleep disorders can occur. As this study identifies the molecular mechanism that generates robustness and flexibility of the circadian clock, it can facilitate the identification of the cause of and treatment strategy for the circadian rhythm sleep disorders.” This work was supported by the Human Frontier Science Program.
-PublicationEui Min Jeong, Miri Kwon, Eunjoo Cho, Sang Hyuk Lee, Hyun Kim, Eun Young Kim, and Jae Kyoung Kim, “Systematic modeling-driven experiments identify distinct molecularclockworks underlying hierarchically organized pacemaker neurons,” February 22, 2022, Proceedings of the National Academy of Sciences of the United States of America
-ProfileProfessor Jae Kyoung KimDepartment of Mathematical SciencesKAIST
2022.02.23 View 11386
Scientist Discover How Circadian Rhythm Can Be Both Strong and Flexible
Study reveals that master and slave oscillators function via different molecular mechanisms
From tiny fruit flies to human beings, all animals on Earth maintain their daily rhythms based on their internal circadian clock. The circadian clock enables organisms to undergo rhythmic changes in behavior and physiology based on a 24-hour circadian cycle. For example, our own biological clock tells our brain to release melatonin, a sleep-inducing hormone, at night time.
The discovery of the molecular mechanism of the circadian clock was bestowed the Nobel Prize in Physiology or Medicine 2017. From what we know, no one centralized clock is responsible for our circadian cycles. Instead, it operates in a hierarchical network where there are “master pacemaker” and “slave oscillator”.
The master pacemaker receives various input signals from the environment such as light. The master then drives the slave oscillator that regulates various outputs such as sleep, feeding, and metabolism. Despite the different roles of the pacemaker neurons, they are known to share common molecular mechanisms that are well conserved in all lifeforms. For example, interlocked systems of multiple transcriptional-translational feedback loops (TTFLs) composed of core clock proteins have been deeply studied in fruit flies.
However, there is still much that we need to learn about our own biological clock. The hierarchically-organized nature of master and slave clock neurons leads to a prevailing belief that they share an identical molecular clockwork. At the same time, the different roles they serve in regulating bodily rhythms also raise the question of whether they might function under different molecular clockworks.
Research team led by Professor Kim Jae Kyoung from the Department of Mathematical Sciences, a chief investigator at the Biomedical Mathematics Group at the Institute for Basic Science, used a combination of mathematical and experimental approaches using fruit flies to answer this question. The team found that the master clock and the slave clock operate via different molecular mechanisms.
In both master and slave neurons of fruit flies, a circadian rhythm-related protein called PER is produced and degraded at different rates depending on the time of the day. Previously, the team found that the master clock neuron (sLNvs) and the slave clock neuron (DN1ps) have different profiles of PER in wild-type and Clk-Δ mutant Drosophila. This hinted that there might be a potential difference in molecular clockworks between the master and slave clock neurons.
However, due to the complexity of the molecular clockwork, it was challenging to identify the source of such differences. Thus, the team developed a mathematical model describing the molecular clockworks of the master and slave clocks. Then, all possible molecular differences between the master and slave clock neurons were systematically investigated by using computer simulations. The model predicted that PER is more efficiently produced and then rapidly degraded in the master clock compared to the slave clock neurons. This prediction was then confirmed by the follow-up experiments using animal.
Then, why do the master clock neurons have such different molecular properties from the slave clock neurons? To answer this question, the research team again used the combination of mathematical model simulation and experiments. It was found that the faster rate of synthesis of PER in the master clock neurons allows them to generate synchronized rhythms with a high level of amplitude. Generation of such a strong rhythm with high amplitude is critical to delivering clear signals to slave clock neurons.
However, such strong rhythms would typically be unfavorable when it comes to adapting to environmental changes. These include natural causes such as different daylight hours across summer and winter seasons, up to more extreme artificial cases such as jet lag that occurs after international travel. Thanks to the distinct property of the master clock neurons, it is able to undergo phase dispersion when the standard light-dark cycle is disrupted, drastically reducing the level of PER. The master clock neurons can then easily adapt to the new diurnal cycle. Our master pacemaker’s plasticity explains how we can quickly adjust to the new time zones after international flights after just a brief period of jet lag.
It is hoped that the findings of this study can have future clinical implications when it comes to treating various disorders that affect our circadian rhythm. Professor Kim notes, “When the circadian clock loses its robustness and flexibility, the circadian rhythms sleep disorders can occur. As this study identifies the molecular mechanism that generates robustness and flexibility of the circadian clock, it can facilitate the identification of the cause of and treatment strategy for the circadian rhythm sleep disorders.” This work was supported by the Human Frontier Science Program.
-PublicationEui Min Jeong, Miri Kwon, Eunjoo Cho, Sang Hyuk Lee, Hyun Kim, Eun Young Kim, and Jae Kyoung Kim, “Systematic modeling-driven experiments identify distinct molecularclockworks underlying hierarchically organized pacemaker neurons,” February 22, 2022, Proceedings of the National Academy of Sciences of the United States of America
-ProfileProfessor Jae Kyoung KimDepartment of Mathematical SciencesKAIST
2022.02.23 View 11386