Science
-
 KAIST Proposes a New Way to Circumvent a Long-time Frustration in Neural Computing
The human brain begins learning through spontaneous random activities even before it receives sensory information from the external world. The technology developed by the KAIST research team enables much faster and more accurate learning when exposed to actual data by pre-learning random information in a brain-mimicking artificial neural network, and is expected to be a breakthrough in the development of brain-based artificial intelligence and neuromorphic computing technology in the future.
KAIST (President Kwang-Hyung Lee) announced on the 16th of December that Professor Se-Bum Paik 's research team in the Department of Brain Cognitive Sciences solved the weight transport problem*, a long-standing challenge in neural network learning, and through this, explained the principles that enable resource-efficient learning in biological brain neural networks.
*Weight transport problem: This is the biggest obstacle to the development of artificial intelligence that mimics the biological brain. It is the fundamental reason why large-scale memory and computational work are required in the learning of general artificial neural networks, unlike biological brains.
Over the past several decades, the development of artificial intelligence has been based on error backpropagation learning proposed by Geoffery Hinton, who won the Nobel Prize in Physics this year. However, error backpropagation learning was thought to be impossible in biological brains because it requires the unrealistic assumption that individual neurons must know all the connected information across multiple layers in order to calculate the error signal for learning.
< Figure 1. Illustration depicting the method of random noise training and its effects >
This difficult problem, called the weight transport problem, was raised by Francis Crick, who won the Nobel Prize in Physiology or Medicine for the discovery of the structure of DNA, after the error backpropagation learning was proposed by Hinton in 1986. Since then, it has been considered the reason why the operating principles of natural neural networks and artificial neural networks will forever be fundamentally different.
At the borderline of artificial intelligence and neuroscience, researchers including Hinton have continued to attempt to create biologically plausible models that can implement the learning principles of the brain by solving the weight transport problem.
In 2016, a joint research team from Oxford University and DeepMind in the UK first proposed the concept of error backpropagation learning being possible without weight transport, drawing attention from the academic world. However, biologically plausible error backpropagation learning without weight transport was inefficient, with slow learning speeds and low accuracy, making it difficult to apply in reality.
KAIST research team noted that the biological brain begins learning through internal spontaneous random neural activity even before experiencing external sensory experiences. To mimic this, the research team pre-trained a biologically plausible neural network without weight transport with meaningless random information (random noise).
As a result, they showed that the symmetry of the forward and backward neural cell connections of the neural network, which is an essential condition for error backpropagation learning, can be created. In other words, learning without weight transport is possible through random pre-training.
< Figure 2. Illustration depicting the meta-learning effect of random noise training >
The research team revealed that learning random information before learning actual data has the property of meta-learning, which is ‘learning how to learn.’ It was shown that neural networks that pre-learned random noise perform much faster and more accurate learning when exposed to actual data, and can achieve high learning efficiency without weight transport.
< Figure 3. Illustration depicting research on understanding the brain's operating principles through artificial neural networks >
Professor Se-Bum Paik said, “It breaks the conventional understanding of existing machine learning that only data learning is important, and provides a new perspective that focuses on the neuroscience principles of creating appropriate conditions before learning,” and added, “It is significant in that it solves important problems in artificial neural network learning through clues from developmental neuroscience, and at the same time provides insight into the brain’s learning principles through artificial neural network models.”
This study, in which Jeonghwan Cheon, a Master’s candidate of KAIST Department of Brain and Cognitive Sciences participated as the first author and Professor Sang Wan Lee of the same department as a co-author, was presented at the 38th Neural Information Processing Systems (NeurIPS), the world's top artificial intelligence conference, on December 14th in Vancouver, Canada. (Paper title: Pretraining with random noise for fast and robust learning without weight transport)
This study was conducted with the support of the National Research Foundation of Korea's Basic Research Program in Science and Engineering, the Information and Communications Technology Planning and Evaluation Institute's Talent Development Program, and the KAIST Singularity Professor Program.
2024.12.16 View 7212
KAIST Proposes a New Way to Circumvent a Long-time Frustration in Neural Computing
The human brain begins learning through spontaneous random activities even before it receives sensory information from the external world. The technology developed by the KAIST research team enables much faster and more accurate learning when exposed to actual data by pre-learning random information in a brain-mimicking artificial neural network, and is expected to be a breakthrough in the development of brain-based artificial intelligence and neuromorphic computing technology in the future.
KAIST (President Kwang-Hyung Lee) announced on the 16th of December that Professor Se-Bum Paik 's research team in the Department of Brain Cognitive Sciences solved the weight transport problem*, a long-standing challenge in neural network learning, and through this, explained the principles that enable resource-efficient learning in biological brain neural networks.
*Weight transport problem: This is the biggest obstacle to the development of artificial intelligence that mimics the biological brain. It is the fundamental reason why large-scale memory and computational work are required in the learning of general artificial neural networks, unlike biological brains.
Over the past several decades, the development of artificial intelligence has been based on error backpropagation learning proposed by Geoffery Hinton, who won the Nobel Prize in Physics this year. However, error backpropagation learning was thought to be impossible in biological brains because it requires the unrealistic assumption that individual neurons must know all the connected information across multiple layers in order to calculate the error signal for learning.
< Figure 1. Illustration depicting the method of random noise training and its effects >
This difficult problem, called the weight transport problem, was raised by Francis Crick, who won the Nobel Prize in Physiology or Medicine for the discovery of the structure of DNA, after the error backpropagation learning was proposed by Hinton in 1986. Since then, it has been considered the reason why the operating principles of natural neural networks and artificial neural networks will forever be fundamentally different.
At the borderline of artificial intelligence and neuroscience, researchers including Hinton have continued to attempt to create biologically plausible models that can implement the learning principles of the brain by solving the weight transport problem.
In 2016, a joint research team from Oxford University and DeepMind in the UK first proposed the concept of error backpropagation learning being possible without weight transport, drawing attention from the academic world. However, biologically plausible error backpropagation learning without weight transport was inefficient, with slow learning speeds and low accuracy, making it difficult to apply in reality.
KAIST research team noted that the biological brain begins learning through internal spontaneous random neural activity even before experiencing external sensory experiences. To mimic this, the research team pre-trained a biologically plausible neural network without weight transport with meaningless random information (random noise).
As a result, they showed that the symmetry of the forward and backward neural cell connections of the neural network, which is an essential condition for error backpropagation learning, can be created. In other words, learning without weight transport is possible through random pre-training.
< Figure 2. Illustration depicting the meta-learning effect of random noise training >
The research team revealed that learning random information before learning actual data has the property of meta-learning, which is ‘learning how to learn.’ It was shown that neural networks that pre-learned random noise perform much faster and more accurate learning when exposed to actual data, and can achieve high learning efficiency without weight transport.
< Figure 3. Illustration depicting research on understanding the brain's operating principles through artificial neural networks >
Professor Se-Bum Paik said, “It breaks the conventional understanding of existing machine learning that only data learning is important, and provides a new perspective that focuses on the neuroscience principles of creating appropriate conditions before learning,” and added, “It is significant in that it solves important problems in artificial neural network learning through clues from developmental neuroscience, and at the same time provides insight into the brain’s learning principles through artificial neural network models.”
This study, in which Jeonghwan Cheon, a Master’s candidate of KAIST Department of Brain and Cognitive Sciences participated as the first author and Professor Sang Wan Lee of the same department as a co-author, was presented at the 38th Neural Information Processing Systems (NeurIPS), the world's top artificial intelligence conference, on December 14th in Vancouver, Canada. (Paper title: Pretraining with random noise for fast and robust learning without weight transport)
This study was conducted with the support of the National Research Foundation of Korea's Basic Research Program in Science and Engineering, the Information and Communications Technology Planning and Evaluation Institute's Talent Development Program, and the KAIST Singularity Professor Program.
2024.12.16 View 7212 -
 KAIST Unveils New Possibilities for Treating Intractable Brain Tumors
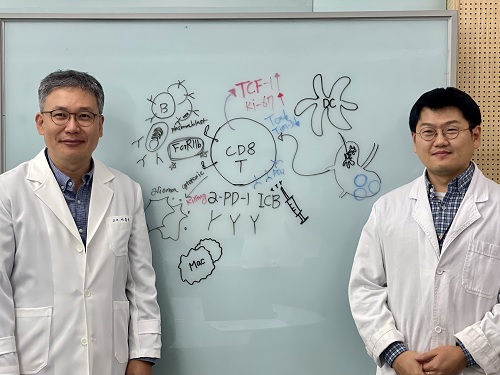
< Photo 1. (From left) Professor Heung Kyu Lee, KAIST Department of Biological Sciences, and Dr. Keun Bon Ku >
Immunotherapy, which enhances the immune system's T cell response to eliminate cancer cells, has emerged as a key approach in cancer treatment. However, in the case of glioblastoma, an aggressive and treatment-resistant brain tumor, numerous clinical trials have failed to confirm their efficacy. Korean researchers have recently analyzed the mechanisms that cause T cell exhaustion, which is characterized by a loss of function or a weakened response following prolonged exposure to antigens in such intractable cancers, identifying key control factors in T cell activation and clarifying the mechanisms that enhance therapeutic effectiveness.
KAIST (represented by President Kwang Hyung Lee) announced on the 6th of November that Professor Heung Kyu Lee’s team from the Department of Biological Sciences, in collaboration with the Korea Research Institute of Chemical Technology (represented by President Young Kuk Lee), has confirmed improved survival rates in a glioblastoma mouse model. By removing the inhibitory Fc gamma receptor (FcγRIIB), the research team was able to restore the responsiveness of cytotoxic T cells to immune checkpoint inhibitors, leading to enhanced anticancer activity.
The research team examined the effect of FcγRIIB, an inhibitory receptor recently found in cytotoxic T cells, on tumor-infiltrating T cells and the therapeutic effectiveness of the anti-PD-1 immune checkpoint inhibitor.
< Figure 1. Study results on improved survival rate due to increased antitumor activity of anti-PD-1 treatment in inhibitory Fc gamma receptor(Fcgr2b) ablation mice with murine glioblastoma. >
Their findings showed that deleting FcγRIIB induced the increase of tumor antigen-specific memory T cells, which helps to suppress exhaustion, enhances stem-like qualities, and reactivates T cell-mediated antitumor immunity, particularly in response to anti-PD-1 treatment. Furthermore, FcγRIIB deletion led to an increase in antigen-specific memory T cells that maintained continuous infiltration into the tumor tissue.
This study presents a new therapeutic target for tumors unresponsive to immune checkpoint inhibitors and demonstrates that combining FcγRIIB inhibition with anti-PD-1 treatment can produce synergistic effects, potentially improving therapeutic outcomes for tumors like glioblastoma, which typically show resistance to anti-PD-1 therapy.
< Figure 2. Overview of the study on the enhanced response to anti-PD-1 therapy for glioblastoma brain tumors upon deletion of the inhibitory Fc gamma receptor (FcγRIIB) in tumor microenvironment. When the inhibitory Fc gamma receptor (FcγRIIB) of cytotoxic T cells is deleted, an increase in tumor-specific memory T cells (Ttsms) was observed. In addition, this T cell subset is identified as originating from the tumor-draining lymph nodes(TdLNs) and leads to persistent infiltration into the tumor tissue. Anti-PD-1 therapy leads to an increased anti-tumor immune response via Ttsms, which is confirmed by increased tumor cell toxicity and increased cell division and decreased cell de-migration indices. Ultimately, the increased cytotoxic T cell immune response leads to an increase in the survival rate of glioblastoma. >
Professor Heung Kyu Lee explained, "This study offers a way to overcome clinical failures in treating brain tumors with immune checkpoint therapy and opens possibilities for broader applications to other intractable cancers. It also highlights the potential of utilizing cytotoxic T cells for tumor cell therapy."
The study, led by Dr. Keun Bon Ku of KAIST (currently a senior researcher at the Korea Research Institute of Chemical Technology's Center for Infectious Disease Diagnosis and Prevention), along with Chae Won Kim, Yumin Kim, Byeong Hoon Kang, Jeongwoo La, In Kang, Won Hyung Park, Stephen Ahn, and Sung Ki Lee, was published online on October 26 in the Journal for ImmunoTherapy of Cancer, an international journal in tumor immunology and therapy from the Society for Immunotherapy of Cancer. (Paper title: “Inhibitory Fcγ receptor deletion enhances CD8 T cell stemness increasing anti-PD-1 therapy responsiveness against glioblastoma,” http://dx.doi.org/10.1136/jitc-2024-009449).
This research received support from the National Research Foundation of Korea, the Bio & Medical Technology Development Program, and the Samsung Science & Technology Foundation.
2024.11.15 View 5258
KAIST Unveils New Possibilities for Treating Intractable Brain Tumors
< Photo 1. (From left) Professor Heung Kyu Lee, KAIST Department of Biological Sciences, and Dr. Keun Bon Ku >
Immunotherapy, which enhances the immune system's T cell response to eliminate cancer cells, has emerged as a key approach in cancer treatment. However, in the case of glioblastoma, an aggressive and treatment-resistant brain tumor, numerous clinical trials have failed to confirm their efficacy. Korean researchers have recently analyzed the mechanisms that cause T cell exhaustion, which is characterized by a loss of function or a weakened response following prolonged exposure to antigens in such intractable cancers, identifying key control factors in T cell activation and clarifying the mechanisms that enhance therapeutic effectiveness.
KAIST (represented by President Kwang Hyung Lee) announced on the 6th of November that Professor Heung Kyu Lee’s team from the Department of Biological Sciences, in collaboration with the Korea Research Institute of Chemical Technology (represented by President Young Kuk Lee), has confirmed improved survival rates in a glioblastoma mouse model. By removing the inhibitory Fc gamma receptor (FcγRIIB), the research team was able to restore the responsiveness of cytotoxic T cells to immune checkpoint inhibitors, leading to enhanced anticancer activity.
The research team examined the effect of FcγRIIB, an inhibitory receptor recently found in cytotoxic T cells, on tumor-infiltrating T cells and the therapeutic effectiveness of the anti-PD-1 immune checkpoint inhibitor.
< Figure 1. Study results on improved survival rate due to increased antitumor activity of anti-PD-1 treatment in inhibitory Fc gamma receptor(Fcgr2b) ablation mice with murine glioblastoma. >
Their findings showed that deleting FcγRIIB induced the increase of tumor antigen-specific memory T cells, which helps to suppress exhaustion, enhances stem-like qualities, and reactivates T cell-mediated antitumor immunity, particularly in response to anti-PD-1 treatment. Furthermore, FcγRIIB deletion led to an increase in antigen-specific memory T cells that maintained continuous infiltration into the tumor tissue.
This study presents a new therapeutic target for tumors unresponsive to immune checkpoint inhibitors and demonstrates that combining FcγRIIB inhibition with anti-PD-1 treatment can produce synergistic effects, potentially improving therapeutic outcomes for tumors like glioblastoma, which typically show resistance to anti-PD-1 therapy.
< Figure 2. Overview of the study on the enhanced response to anti-PD-1 therapy for glioblastoma brain tumors upon deletion of the inhibitory Fc gamma receptor (FcγRIIB) in tumor microenvironment. When the inhibitory Fc gamma receptor (FcγRIIB) of cytotoxic T cells is deleted, an increase in tumor-specific memory T cells (Ttsms) was observed. In addition, this T cell subset is identified as originating from the tumor-draining lymph nodes(TdLNs) and leads to persistent infiltration into the tumor tissue. Anti-PD-1 therapy leads to an increased anti-tumor immune response via Ttsms, which is confirmed by increased tumor cell toxicity and increased cell division and decreased cell de-migration indices. Ultimately, the increased cytotoxic T cell immune response leads to an increase in the survival rate of glioblastoma. >
Professor Heung Kyu Lee explained, "This study offers a way to overcome clinical failures in treating brain tumors with immune checkpoint therapy and opens possibilities for broader applications to other intractable cancers. It also highlights the potential of utilizing cytotoxic T cells for tumor cell therapy."
The study, led by Dr. Keun Bon Ku of KAIST (currently a senior researcher at the Korea Research Institute of Chemical Technology's Center for Infectious Disease Diagnosis and Prevention), along with Chae Won Kim, Yumin Kim, Byeong Hoon Kang, Jeongwoo La, In Kang, Won Hyung Park, Stephen Ahn, and Sung Ki Lee, was published online on October 26 in the Journal for ImmunoTherapy of Cancer, an international journal in tumor immunology and therapy from the Society for Immunotherapy of Cancer. (Paper title: “Inhibitory Fcγ receptor deletion enhances CD8 T cell stemness increasing anti-PD-1 therapy responsiveness against glioblastoma,” http://dx.doi.org/10.1136/jitc-2024-009449).
This research received support from the National Research Foundation of Korea, the Bio & Medical Technology Development Program, and the Samsung Science & Technology Foundation.
2024.11.15 View 5258 -
 KAIST Succeeds in the Real-time Observation of Organoids using Holotomography
Organoids, which are 3D miniature organs that mimic the structure and function of human organs, play an essential role in disease research and drug development. A Korean research team has overcome the limitations of existing imaging technologies, succeeding in the real-time, high-resolution observation of living organoids.
KAIST (represented by President Kwang Hyung Lee) announced on the 14th of October that Professor YongKeun Park’s research team from the Department of Physics, in collaboration with the Genome Editing Research Center (Director Bon-Kyoung Koo) of the Institute for Basic Science (IBS President Do-Young Noh) and Tomocube Inc., has developed an imaging technology using holotomography to observe live, small intestinal organoids in real time at a high resolution.
Existing imaging techniques have struggled to observe living organoids in high resolution over extended periods and often required additional treatments like fluorescent staining.
< Figure 1. Overview of the low-coherence HT workflow. Using holotomography, 3D morphological restoration and quantitative analysis of organoids can be performed. In order to improve the limited field of view, which is a limitation of the microscope, our research team utilized a large-area field of view combination algorithm and made a 3D restoration by acquiring multi-focus holographic images for 3D measurements. After that, the organoids were compartmentalized to divide the parts necessary for analysis and quantitatively evaluated the protein concentration measurable from the refractive index and the survival rate of the organoids. >
The research team introduced holotomography technology to address these issues, which provides high-resolution images without the need for fluorescent staining and allows for the long-term observation of dynamic changes in real time without causing cell damage.
The team validated this technology using small intestinal organoids from experimental mice and were able to observe various cell structures inside the organoids in detail. They also captured dynamic changes such as growth processes, cell division, and cell death in real time using holotomography.
Additionally, the technology allowed for the precise analysis of the organoids' responses to drug treatments, verifying the survival of the cells.
The researchers believe that this breakthrough will open new horizons in organoid research, enabling the greater utilization of organoids in drug development, personalized medicine, and regenerative medicine.
Future research is expected to more accurately replicate the in vivo environment of organoids, contributing significantly to a more detailed understanding of various life phenomena at the cellular level through more precise 3D imaging.
< Figure 2. Real-time organoid morphology analysis. Using holotomography, it is possible to observe the lumen and villus development process of intestinal organoids in real time, which was difficult to observe with a conventional microscope. In addition, various information about intestinal organoids can be obtained by quantifying the size and protein amount of intestinal organoids through image analysis. >
Dr. Mahn Jae Lee, a graduate of KAIST's Graduate School of Medical Science and Engineering, currently at Chungnam National University Hospital and the first author of the paper, commented, "This research represents a new imaging technology that surpasses previous limitations and is expected to make a major contribution to disease modeling, personalized treatments, and drug development research using organoids."
The research results were published online in the international journal Experimental & Molecular Medicine on October 1, 2024, and the technology has been recognized for its applicability in various fields of life sciences. (Paper title: “Long-term three-dimensional high-resolution imaging of live unlabeled small intestinal organoids via low-coherence holotomography”)
This research was supported by the National Research Foundation of Korea, KAIST Institutes, and the Institute for Basic Science.
2024.10.14 View 5088
KAIST Succeeds in the Real-time Observation of Organoids using Holotomography
Organoids, which are 3D miniature organs that mimic the structure and function of human organs, play an essential role in disease research and drug development. A Korean research team has overcome the limitations of existing imaging technologies, succeeding in the real-time, high-resolution observation of living organoids.
KAIST (represented by President Kwang Hyung Lee) announced on the 14th of October that Professor YongKeun Park’s research team from the Department of Physics, in collaboration with the Genome Editing Research Center (Director Bon-Kyoung Koo) of the Institute for Basic Science (IBS President Do-Young Noh) and Tomocube Inc., has developed an imaging technology using holotomography to observe live, small intestinal organoids in real time at a high resolution.
Existing imaging techniques have struggled to observe living organoids in high resolution over extended periods and often required additional treatments like fluorescent staining.
< Figure 1. Overview of the low-coherence HT workflow. Using holotomography, 3D morphological restoration and quantitative analysis of organoids can be performed. In order to improve the limited field of view, which is a limitation of the microscope, our research team utilized a large-area field of view combination algorithm and made a 3D restoration by acquiring multi-focus holographic images for 3D measurements. After that, the organoids were compartmentalized to divide the parts necessary for analysis and quantitatively evaluated the protein concentration measurable from the refractive index and the survival rate of the organoids. >
The research team introduced holotomography technology to address these issues, which provides high-resolution images without the need for fluorescent staining and allows for the long-term observation of dynamic changes in real time without causing cell damage.
The team validated this technology using small intestinal organoids from experimental mice and were able to observe various cell structures inside the organoids in detail. They also captured dynamic changes such as growth processes, cell division, and cell death in real time using holotomography.
Additionally, the technology allowed for the precise analysis of the organoids' responses to drug treatments, verifying the survival of the cells.
The researchers believe that this breakthrough will open new horizons in organoid research, enabling the greater utilization of organoids in drug development, personalized medicine, and regenerative medicine.
Future research is expected to more accurately replicate the in vivo environment of organoids, contributing significantly to a more detailed understanding of various life phenomena at the cellular level through more precise 3D imaging.
< Figure 2. Real-time organoid morphology analysis. Using holotomography, it is possible to observe the lumen and villus development process of intestinal organoids in real time, which was difficult to observe with a conventional microscope. In addition, various information about intestinal organoids can be obtained by quantifying the size and protein amount of intestinal organoids through image analysis. >
Dr. Mahn Jae Lee, a graduate of KAIST's Graduate School of Medical Science and Engineering, currently at Chungnam National University Hospital and the first author of the paper, commented, "This research represents a new imaging technology that surpasses previous limitations and is expected to make a major contribution to disease modeling, personalized treatments, and drug development research using organoids."
The research results were published online in the international journal Experimental & Molecular Medicine on October 1, 2024, and the technology has been recognized for its applicability in various fields of life sciences. (Paper title: “Long-term three-dimensional high-resolution imaging of live unlabeled small intestinal organoids via low-coherence holotomography”)
This research was supported by the National Research Foundation of Korea, KAIST Institutes, and the Institute for Basic Science.
2024.10.14 View 5088 -
 KAIST Changes the Paradigm of Drug Discovery with World's First Atomic Editing
In pioneering drug development, the new technology that enables the easy and rapid editing of key atoms responsible for drug efficacy has been regarded as a fundamental and "dream" technology, revolutionizing the process of discovering potential drug candidates. KAIST researchers have become the first in the world to successfully develop single-atom editing technology that maximizes drug efficacy.
On October 8th, KAIST (represented by President Kwang-Hyung Lee) announced that Professor Yoonsu Park’s research team from the Department of Chemistry successfully developed technology that enables the easy editing and correction of oxygen atoms in furan compounds into nitrogen atoms, directly converting them into pyrrole frameworks, which are widely used in pharmaceuticals.
< Image. Conceptual image illustrating the main idea of the research >
This research was published in the prestigious scientific journal Science on October 3rd under the title "Photocatalytic Furan-to-Pyrrole Conversion."
Many drugs have complex chemical structures, but their efficacy is often determined by a single critical atom. Atoms like oxygen and nitrogen play a central role in enhancing the pharmacological effects of these drugs, particularly against viruses.
This phenomenon, where the introduction of specific atoms into a drug molecule dramatically affects its efficacy, is known as the "Single Atom Effect." In leading-edge drug development, discovering atoms that maximize drug efficacy is key.
However, evaluating the Single Atom Effect has traditionally required multi-step, costly synthesis processes, as it has been difficult to selectively edit single atoms within stable ring structures containing oxygen or nitrogen.
Professor Park’s team overcame this challenge by introducing a photocatalyst that uses light energy. They developed a photocatalyst that acts as a “molecular scissor,” freely cutting and attaching five-membered rings, enabling single-atom editing at room temperature and atmospheric pressure—a world first.
The team discovered a new reaction mechanism in which the excited molecular scissor removes oxygen from furan via single-electron oxidation and then sequentially adds a nitrogen atom.
Donghyeon Kim and Jaehyun You, the study's first authors and candidates in KAIST’s integrated master's and doctoral program in the Department of Chemistry, explained that this technique offers high versatility by utilizing light energy to replace harsh conditions. They further noted that the technology enables selective editing, even when applied to complex natural products or pharmaceuticals. Professor Yoonsu Park, who led the research, remarked, "This breakthrough, which allows for the selective editing of five-membered organic ring structures, will open new doors for building libraries of drug candidates, a key challenge in pharmaceuticals. I hope this foundational technology will be used to revolutionize the drug development process."
The significance of this research was highlighted in the Perspective section of Science, a feature where a peer scientist of prominence outside of the project group provides commentary on an impactful research.
This research was supported by the National Research Foundation of Korea’s Creative Research Program, the Cross-Generation Collaborative Lab Project at KAIST, and the POSCO Science Fellowship of the POSCO TJ Park Foundation.
2024.10.11 View 5592
KAIST Changes the Paradigm of Drug Discovery with World's First Atomic Editing
In pioneering drug development, the new technology that enables the easy and rapid editing of key atoms responsible for drug efficacy has been regarded as a fundamental and "dream" technology, revolutionizing the process of discovering potential drug candidates. KAIST researchers have become the first in the world to successfully develop single-atom editing technology that maximizes drug efficacy.
On October 8th, KAIST (represented by President Kwang-Hyung Lee) announced that Professor Yoonsu Park’s research team from the Department of Chemistry successfully developed technology that enables the easy editing and correction of oxygen atoms in furan compounds into nitrogen atoms, directly converting them into pyrrole frameworks, which are widely used in pharmaceuticals.
< Image. Conceptual image illustrating the main idea of the research >
This research was published in the prestigious scientific journal Science on October 3rd under the title "Photocatalytic Furan-to-Pyrrole Conversion."
Many drugs have complex chemical structures, but their efficacy is often determined by a single critical atom. Atoms like oxygen and nitrogen play a central role in enhancing the pharmacological effects of these drugs, particularly against viruses.
This phenomenon, where the introduction of specific atoms into a drug molecule dramatically affects its efficacy, is known as the "Single Atom Effect." In leading-edge drug development, discovering atoms that maximize drug efficacy is key.
However, evaluating the Single Atom Effect has traditionally required multi-step, costly synthesis processes, as it has been difficult to selectively edit single atoms within stable ring structures containing oxygen or nitrogen.
Professor Park’s team overcame this challenge by introducing a photocatalyst that uses light energy. They developed a photocatalyst that acts as a “molecular scissor,” freely cutting and attaching five-membered rings, enabling single-atom editing at room temperature and atmospheric pressure—a world first.
The team discovered a new reaction mechanism in which the excited molecular scissor removes oxygen from furan via single-electron oxidation and then sequentially adds a nitrogen atom.
Donghyeon Kim and Jaehyun You, the study's first authors and candidates in KAIST’s integrated master's and doctoral program in the Department of Chemistry, explained that this technique offers high versatility by utilizing light energy to replace harsh conditions. They further noted that the technology enables selective editing, even when applied to complex natural products or pharmaceuticals. Professor Yoonsu Park, who led the research, remarked, "This breakthrough, which allows for the selective editing of five-membered organic ring structures, will open new doors for building libraries of drug candidates, a key challenge in pharmaceuticals. I hope this foundational technology will be used to revolutionize the drug development process."
The significance of this research was highlighted in the Perspective section of Science, a feature where a peer scientist of prominence outside of the project group provides commentary on an impactful research.
This research was supported by the National Research Foundation of Korea’s Creative Research Program, the Cross-Generation Collaborative Lab Project at KAIST, and the POSCO Science Fellowship of the POSCO TJ Park Foundation.
2024.10.11 View 5592 -
 KAIST presents strategies for Holotomography in advanced bio research
Measuring and analyzing three-dimensional (3D) images of live cells and tissues is considered crucial in advanced fields of biology and medicine. Organoids, which are 3D structures that mimic organs, are particular examples that significantly benefits 3D live imaging. Organoids provide effective alternatives to animal testing in the drug development processes, and can rapidly determine personalized medicine. On the other hand, active researches are ongoing to utilize organoids for organ replacement.
< Figure 1. Schematic illustration of holotomography compared to X-ray CT. Similar to CT, they share the commonality of measuring the optical properties of an unlabeled specimen in three dimensions. Instead of X-rays, holotomography irradiates light in the visible range, and provides refractive index measurements of transparent specimens rather than absorptivity. While CT obtains three-dimensional information only through mechanical rotation of the irradiating light, holotomography can replace this by applying wavefront control technology in the visible range. >
Organelle-level observation of 3D biological specimens such as organoids and stem cell colonies without staining or preprocessing holds significant implications for both innovating basic research and bioindustrial applications related to regenerative medicine and bioindustrial applications.
Holotomography (HT) is a 3D optical microscopy that implements 3D reconstruction analogous to that of X-ray computed tomography (CT). Although HT and CT share a similar theoretical background, HT facilitates high-resolution examination inside cells and tissues, instead of the human body. HT obtains 3D images of cells and tissues at the organelle level without chemical or genetic labeling, thus overcomes various challenges of existing methods in bio research and industry. Its potential is highlighted in research fields where sample physiology must not be disrupted, such as regenerative medicine, personalized medicine, and infertility treatment.
< Figure 2. Label-free 3D imaging of diverse live cells. Time-lapse image of Hep3B cells illustrating subcellular morphology changes upon H2O2 treatment, followed by cellular recovery after returning to the regular cell culture medium. >
This paper introduces the advantages and broad applicability of HT to biomedical researchers, while presenting an overview of principles and future technical challenges to optical researchers. It showcases various cases of applying HT in studies such as 3D biology, regenerative medicine, and cancer research, as well as suggesting future optical development. Also, it categorizes HT based on the light source, to describe the principles, limitations, and improvements of each category in detail. Particularly, the paper addresses strategies for deepening cell and organoid studies by introducing artificial intelligence (AI) to HT.
Due to its potential to drive advanced bioindustry, HT is attracting interest and investment from universities and corporates worldwide. The KAIST research team has been leading this international field by developing core technologies and carrying out key application researches throughout the last decade.
< Figure 3. Various types of cells and organelles that make up the imaging barrier of a living intestinal organoid can be observed using holotomography. >
This paper, co-authored by Dr. Geon Kim from KAIST Research Center for Natural Sciences, Professor Ki-Jun Yoon's team from the Department of Biological Sciences, Director Bon-Kyoung Koo's team from the Institute for Basic Science (IBS) Center for Genome Engineering, and Dr. Seongsoo Lee's team from the Korea Basic Science Institute (KBSI), was published in 'Nature Reviews Methods Primers' on the 25th of July. This research was supported by the Leader Grant and Basic Science Research Program of the National Research Foundation, the Hologram Core Technology Development Grant of the Ministry of Science and ICT, the Nano and Material Technology Development Project, and the Health and Medical R&D Project of the Ministry of Health and Welfare.
2024.07.30 View 5598
KAIST presents strategies for Holotomography in advanced bio research
Measuring and analyzing three-dimensional (3D) images of live cells and tissues is considered crucial in advanced fields of biology and medicine. Organoids, which are 3D structures that mimic organs, are particular examples that significantly benefits 3D live imaging. Organoids provide effective alternatives to animal testing in the drug development processes, and can rapidly determine personalized medicine. On the other hand, active researches are ongoing to utilize organoids for organ replacement.
< Figure 1. Schematic illustration of holotomography compared to X-ray CT. Similar to CT, they share the commonality of measuring the optical properties of an unlabeled specimen in three dimensions. Instead of X-rays, holotomography irradiates light in the visible range, and provides refractive index measurements of transparent specimens rather than absorptivity. While CT obtains three-dimensional information only through mechanical rotation of the irradiating light, holotomography can replace this by applying wavefront control technology in the visible range. >
Organelle-level observation of 3D biological specimens such as organoids and stem cell colonies without staining or preprocessing holds significant implications for both innovating basic research and bioindustrial applications related to regenerative medicine and bioindustrial applications.
Holotomography (HT) is a 3D optical microscopy that implements 3D reconstruction analogous to that of X-ray computed tomography (CT). Although HT and CT share a similar theoretical background, HT facilitates high-resolution examination inside cells and tissues, instead of the human body. HT obtains 3D images of cells and tissues at the organelle level without chemical or genetic labeling, thus overcomes various challenges of existing methods in bio research and industry. Its potential is highlighted in research fields where sample physiology must not be disrupted, such as regenerative medicine, personalized medicine, and infertility treatment.
< Figure 2. Label-free 3D imaging of diverse live cells. Time-lapse image of Hep3B cells illustrating subcellular morphology changes upon H2O2 treatment, followed by cellular recovery after returning to the regular cell culture medium. >
This paper introduces the advantages and broad applicability of HT to biomedical researchers, while presenting an overview of principles and future technical challenges to optical researchers. It showcases various cases of applying HT in studies such as 3D biology, regenerative medicine, and cancer research, as well as suggesting future optical development. Also, it categorizes HT based on the light source, to describe the principles, limitations, and improvements of each category in detail. Particularly, the paper addresses strategies for deepening cell and organoid studies by introducing artificial intelligence (AI) to HT.
Due to its potential to drive advanced bioindustry, HT is attracting interest and investment from universities and corporates worldwide. The KAIST research team has been leading this international field by developing core technologies and carrying out key application researches throughout the last decade.
< Figure 3. Various types of cells and organelles that make up the imaging barrier of a living intestinal organoid can be observed using holotomography. >
This paper, co-authored by Dr. Geon Kim from KAIST Research Center for Natural Sciences, Professor Ki-Jun Yoon's team from the Department of Biological Sciences, Director Bon-Kyoung Koo's team from the Institute for Basic Science (IBS) Center for Genome Engineering, and Dr. Seongsoo Lee's team from the Korea Basic Science Institute (KBSI), was published in 'Nature Reviews Methods Primers' on the 25th of July. This research was supported by the Leader Grant and Basic Science Research Program of the National Research Foundation, the Hologram Core Technology Development Grant of the Ministry of Science and ICT, the Nano and Material Technology Development Project, and the Health and Medical R&D Project of the Ministry of Health and Welfare.
2024.07.30 View 5598 -
 Unraveling Mitochondrial DNA Mutations in Human Cells
Throughout our lifetime, cells accumulate DNA mutations, which contribute to genetic diversity, or “mosaicism”, among cells. These genomic mutations are pivotal for the aging process and the onset of various diseases, including cancer. Mitochondria, essential cellular organelles involved in energy metabolism and apoptosis, possess their own DNA, which are susceptible to mutations. However, studies on mtDNA mutations and mosaicism have been limited due to a variety of technical challenges.
Genomic scientists from KAIST have revealed the genetic mosaicism characterized by variations in mitochondrial DNA (mtDNA) across normal human cells. This study provides fundamental insights into understanding human aging and disease onset mechanisms.
The study, “Mitochondrial DNA mosaicism in normal human somatic cells,” was published in Nature Genetics on July 22. It was led by graduate student Jisong An under the supervision of Professor Young Seok Ju from the Graduate School of Medical Science and Engineering.
Researchers from Seoul National University College of Medicine, Yonsei University College of Medicine, Korea University College of Medicine, Washington University School of Medicine National Cancer Center, Seoul National University Hospital, Gangnam Severance Hospital and KAIST faculty startup company Inocras Inc. also participated in this study.
< Figure 1. a. Flow of experiment. b. Schematic diagram illustrating the origin and dynamics of mtDNA alterations across a lifetime. >
The study involved a bioinformatic analysis of whole-genome sequences from 2,096 single cells obtained from normal human colorectal epithelial tissue, fibroblasts, and blood collected from 31 individuals. The study highlights an average of three significant mtDNA differences between cells, with approximately 6% of these variations confirmed to be inherited as heteroplasmy from the mother.
Moreover, mutations significantly increased during tumorigenesis, with some mutations contributing to instability in mitochondrial RNA. Based on these findings, the study illustrates a computational model that comprehensively elucidates the evolution of mitochondria from embryonic development to aging and tumorigenesis.
This study systematically reveals the mechanisms behind mitochondrial DNA mosaicism in normal human cells, establishing a crucial foundation for understanding the impact of mtDNA on aging and disease onset.
Professor Ju remarked, “By systematically utilizing whole-genome big data, we can illuminate previously unknown phenomena in life sciences.” He emphasized the significance of the study, adding, “For the first time, we have established a method to systematically understand mitochondrial DNA changes occurring during human embryonic development, aging, and cancer development.”
This work was supported by the National Research Foundation of Korea and the Suh Kyungbae Foundation.
2024.07.24 View 5406
Unraveling Mitochondrial DNA Mutations in Human Cells
Throughout our lifetime, cells accumulate DNA mutations, which contribute to genetic diversity, or “mosaicism”, among cells. These genomic mutations are pivotal for the aging process and the onset of various diseases, including cancer. Mitochondria, essential cellular organelles involved in energy metabolism and apoptosis, possess their own DNA, which are susceptible to mutations. However, studies on mtDNA mutations and mosaicism have been limited due to a variety of technical challenges.
Genomic scientists from KAIST have revealed the genetic mosaicism characterized by variations in mitochondrial DNA (mtDNA) across normal human cells. This study provides fundamental insights into understanding human aging and disease onset mechanisms.
The study, “Mitochondrial DNA mosaicism in normal human somatic cells,” was published in Nature Genetics on July 22. It was led by graduate student Jisong An under the supervision of Professor Young Seok Ju from the Graduate School of Medical Science and Engineering.
Researchers from Seoul National University College of Medicine, Yonsei University College of Medicine, Korea University College of Medicine, Washington University School of Medicine National Cancer Center, Seoul National University Hospital, Gangnam Severance Hospital and KAIST faculty startup company Inocras Inc. also participated in this study.
< Figure 1. a. Flow of experiment. b. Schematic diagram illustrating the origin and dynamics of mtDNA alterations across a lifetime. >
The study involved a bioinformatic analysis of whole-genome sequences from 2,096 single cells obtained from normal human colorectal epithelial tissue, fibroblasts, and blood collected from 31 individuals. The study highlights an average of three significant mtDNA differences between cells, with approximately 6% of these variations confirmed to be inherited as heteroplasmy from the mother.
Moreover, mutations significantly increased during tumorigenesis, with some mutations contributing to instability in mitochondrial RNA. Based on these findings, the study illustrates a computational model that comprehensively elucidates the evolution of mitochondria from embryonic development to aging and tumorigenesis.
This study systematically reveals the mechanisms behind mitochondrial DNA mosaicism in normal human cells, establishing a crucial foundation for understanding the impact of mtDNA on aging and disease onset.
Professor Ju remarked, “By systematically utilizing whole-genome big data, we can illuminate previously unknown phenomena in life sciences.” He emphasized the significance of the study, adding, “For the first time, we have established a method to systematically understand mitochondrial DNA changes occurring during human embryonic development, aging, and cancer development.”
This work was supported by the National Research Foundation of Korea and the Suh Kyungbae Foundation.
2024.07.24 View 5406 -
 A 20-year-old puzzle solved: KAIST research team reveals the 'three-dimensional vortex' of zero-dimensional ferroelectrics
Materials that can maintain a magnetized state by themselves without an external magnetic field (i.e., permanent magnets) are called ferromagnets. Ferroelectrics can be thought of as the electric counterpart to ferromagnets, as they maintain a polarized state without an external electric field. It is well-known that ferromagnets lose their magnetic properties when reduced to nano sizes below a certain threshold. What happens when ferroelectrics are similarly made extremely small in all directions (i.e., into a zero-dimensional structure such as nanoparticles) has been a topic of controversy for a long time.
< (From left) Professor Yongsoo Yang, the corresponding author, and Chaehwa Jeong, the first author studying in the integrated master’s and doctoral program, of the KAIST Department of Physics >
The research team led by Dr. Yongsoo Yang from the Department of Physics at KAIST has, for the first time, experimentally clarified the three-dimensional, vortex-shaped polarization distribution inside ferroelectric nanoparticles through international collaborative research with POSTECH, SNU, KBSI, LBNL and University of Arkansas.
About 20 years ago, Prof. Laurent Bellaiche (currently at University of Arkansas) and his colleagues theoretically predicted that a unique form of polarization distribution, arranged in a toroidal vortex shape, could occur inside ferroelectric nanodots. They also suggested that if this vortex distribution could be properly controlled, it could be applied to ultra-high-density memory devices with capacities over 10,000 times greater than existing ones. However, experimental clarification had not been achieved due to the difficulty of measuring the three-dimensional polarization distribution within ferroelectric nanostructures.
The research team at KAIST successfully solved this 20-year-old challenge by implementing a technique called atomic electron tomography. This technique works by acquiring atomic-resolution transmission electron microscope images of the nanomaterials from multiple tilt angles, and then reconstructing them back into three-dimensional structures using advanced reconstruction algorithms. Electron tomography can be understood as essentially the same method with the CT scans used in hospitals to view internal organs in three dimensions; the KAIST team adapted it uniquely for nanomaterials, utilizing an electron microscope at the single-atom level.
< Figure 1. Three-dimensional polarization distribution of BaTiO3 nanoparticles revealed by atomic electron tomography. >(Left) Schematic of the electron tomography technique, which involves acquiring transmission electron microscope images at multiple tilt angles and reconstructing them into 3D atomic structures.(Center) Experimentally determined three-dimensional polarization distribution inside a BaTiO3 nanoparticle via atomic electron tomography. A vortex-like structure is clearly visible near the bottom (blue dot).(Right) A two-dimensional cross-section of the polarization distribution, thinly sliced at the center of the vortex, with the color and arrows together indicating the direction of the polarization. A distinct vortex structure can be observed.
Using atomic electron tomography, the team completely measured the positions of cation atoms inside barium titanate (BaTiO3) nanoparticles, a well-known ferroelectric material, in three dimensions. From the precisely determined 3D atomic arrangements, they were able to further calculate the internal three-dimensional polarization distribution at the single-atom level. The analysis of the polarization distribution revealed, for the first time experimentally, that topological polarization orderings including vortices, anti-vortices, skyrmions, and a Bloch point occur inside the 0-dimensional ferroelectrics, as theoretically predicted 20 years ago. Furthermore, it was also found that the number of internal vortices can be controlled depending on their sizes.
Prof. Sergey Prosandeev and Prof. Bellaiche (who proposed with other co-workers the polar vortex ordering theoretically 20 years ago), joined this collaboration and further proved that the vortex distribution results obtained from experiments are consistent with theoretical calculations.
By controlling the number and orientation of these polarization distributions, it is expected that this can be utilized into next-generation high-density memory device that can store more than 10,000 times the amount of information in the same-sized device compared to existing ones.
Dr. Yang, who led the research, explained the significance of the results: “This result suggests that controlling the size and shape of ferroelectrics alone, without needing to tune the substrate or surrounding environmental effects such as epitaxial strain, can manipulate ferroelectric vortices or other topological orderings at the nano-scale. Further research could then be applied to the development of next-generation ultra-high-density memory.”
This research, with Chaehwa Jeong from the Department of Physics at KAIST as the first author, was published online in Nature Communications on May 8th (Title: Revealing the Three-Dimensional Arrangement of Polar Topology in Nanoparticles).
The study was mainly supported by the National Research Foundation of Korea (NRF) Grants funded by the Korean Government (MSIT).
2024.05.31 View 7649
A 20-year-old puzzle solved: KAIST research team reveals the 'three-dimensional vortex' of zero-dimensional ferroelectrics
Materials that can maintain a magnetized state by themselves without an external magnetic field (i.e., permanent magnets) are called ferromagnets. Ferroelectrics can be thought of as the electric counterpart to ferromagnets, as they maintain a polarized state without an external electric field. It is well-known that ferromagnets lose their magnetic properties when reduced to nano sizes below a certain threshold. What happens when ferroelectrics are similarly made extremely small in all directions (i.e., into a zero-dimensional structure such as nanoparticles) has been a topic of controversy for a long time.
< (From left) Professor Yongsoo Yang, the corresponding author, and Chaehwa Jeong, the first author studying in the integrated master’s and doctoral program, of the KAIST Department of Physics >
The research team led by Dr. Yongsoo Yang from the Department of Physics at KAIST has, for the first time, experimentally clarified the three-dimensional, vortex-shaped polarization distribution inside ferroelectric nanoparticles through international collaborative research with POSTECH, SNU, KBSI, LBNL and University of Arkansas.
About 20 years ago, Prof. Laurent Bellaiche (currently at University of Arkansas) and his colleagues theoretically predicted that a unique form of polarization distribution, arranged in a toroidal vortex shape, could occur inside ferroelectric nanodots. They also suggested that if this vortex distribution could be properly controlled, it could be applied to ultra-high-density memory devices with capacities over 10,000 times greater than existing ones. However, experimental clarification had not been achieved due to the difficulty of measuring the three-dimensional polarization distribution within ferroelectric nanostructures.
The research team at KAIST successfully solved this 20-year-old challenge by implementing a technique called atomic electron tomography. This technique works by acquiring atomic-resolution transmission electron microscope images of the nanomaterials from multiple tilt angles, and then reconstructing them back into three-dimensional structures using advanced reconstruction algorithms. Electron tomography can be understood as essentially the same method with the CT scans used in hospitals to view internal organs in three dimensions; the KAIST team adapted it uniquely for nanomaterials, utilizing an electron microscope at the single-atom level.
< Figure 1. Three-dimensional polarization distribution of BaTiO3 nanoparticles revealed by atomic electron tomography. >(Left) Schematic of the electron tomography technique, which involves acquiring transmission electron microscope images at multiple tilt angles and reconstructing them into 3D atomic structures.(Center) Experimentally determined three-dimensional polarization distribution inside a BaTiO3 nanoparticle via atomic electron tomography. A vortex-like structure is clearly visible near the bottom (blue dot).(Right) A two-dimensional cross-section of the polarization distribution, thinly sliced at the center of the vortex, with the color and arrows together indicating the direction of the polarization. A distinct vortex structure can be observed.
Using atomic electron tomography, the team completely measured the positions of cation atoms inside barium titanate (BaTiO3) nanoparticles, a well-known ferroelectric material, in three dimensions. From the precisely determined 3D atomic arrangements, they were able to further calculate the internal three-dimensional polarization distribution at the single-atom level. The analysis of the polarization distribution revealed, for the first time experimentally, that topological polarization orderings including vortices, anti-vortices, skyrmions, and a Bloch point occur inside the 0-dimensional ferroelectrics, as theoretically predicted 20 years ago. Furthermore, it was also found that the number of internal vortices can be controlled depending on their sizes.
Prof. Sergey Prosandeev and Prof. Bellaiche (who proposed with other co-workers the polar vortex ordering theoretically 20 years ago), joined this collaboration and further proved that the vortex distribution results obtained from experiments are consistent with theoretical calculations.
By controlling the number and orientation of these polarization distributions, it is expected that this can be utilized into next-generation high-density memory device that can store more than 10,000 times the amount of information in the same-sized device compared to existing ones.
Dr. Yang, who led the research, explained the significance of the results: “This result suggests that controlling the size and shape of ferroelectrics alone, without needing to tune the substrate or surrounding environmental effects such as epitaxial strain, can manipulate ferroelectric vortices or other topological orderings at the nano-scale. Further research could then be applied to the development of next-generation ultra-high-density memory.”
This research, with Chaehwa Jeong from the Department of Physics at KAIST as the first author, was published online in Nature Communications on May 8th (Title: Revealing the Three-Dimensional Arrangement of Polar Topology in Nanoparticles).
The study was mainly supported by the National Research Foundation of Korea (NRF) Grants funded by the Korean Government (MSIT).
2024.05.31 View 7649 -
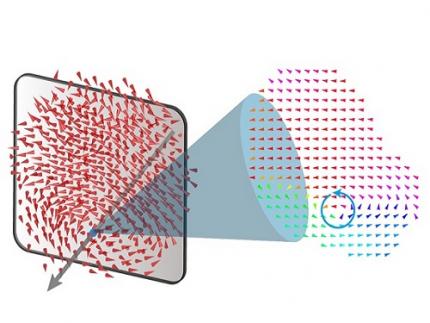
 Revolutionary 'scLENS' Unveiled to Decode Complex Single-Cell Genomic Data
Unlocking biological information from complex single-cell genomic data has just become easier and more precise, thanks to the innovative 'scLENS' tool developed by the Biomedical Mathematics Group within the IBS Center for Mathematical and Computational Sciences led by Chief Investigator Jae Kyoung Kim, who is also a professor at KAIST. This new finding represents a significant leap forward in the field of single-cell transcriptomics.
Single-cell genomic analysis is an advanced technique that measures gene expression at the individual cell level, revealing cellular changes and interactions that are not observable with traditional genomic analysis methods. When applied to cancer tissues, this analysis can delineate the composition of diverse cell types within a tumor, providing insights into how cancer progresses and identifying key genes involved during each stage of progression.
Despite the immense potential of single-cell genomic analysis, handling the vast amount of data that it generates has always been challenging. The amount of data covers the expression of tens of thousands of genes across hundreds to thousands of individual cells. This not only results in large datasets but also introduces noise-related distortions, which arise in part due to current measurement limitations.
< Figure 1. Overview of scLENS (single-cell Low-dimensional embedding using the effective Noise Subtract) >
(Left) Current dimensionality reduction methods for scRNA-seq data involve conventional data preprocessing steps, such as log normalization, followed by manual selection of signals from the scaled data. However, this study reveals that the high levels of sparsity and variability in scRNA-seq data can lead to signal distortion during the data preprocessing, compromising the accuracy of downstream analyses.
(Right) To address this issue, the researchers integrated L2 normalization into the conventional preprocessing pipeline, effectively mitigating signal distortion. Moreover, they developed a novel signal detection algorithm that eliminates the need for user intervention by leveraging random matrix theory-based noise filtering and signal robustness testing. By incorporating these techniques, scLENS enables accurate and automated analysis of scRNA-seq data, overcoming the limitations of existing dimensionality reduction methods.
Corresponding author Jae Kyoung Kim highlighted, “There has been a remarkable advancement in experimental technologies for analyzing single-cell transcriptomes over the past decade. However, due to limitations in data analysis methods, there has been a struggle to fully utilize valuable data obtained through extensive cost and time."
Researchers have developed numerous analysis methods over the years to discern biological signals from this noise. However, the accuracy of these methods has been less than satisfactory. A critical issue is that determining signal and noise thresholds often depends on subjective decisions from the users.
The newly developed scLENS tool harnesses Random Matrix Theory and Signal robustness test to automatically differentiate signals from noise without relying on subjective user input.
First author Hyun Kim stated, "Previously, users had to arbitrarily decide the threshold for signal and noise, which compromised the reproducibility of analysis results and introduced subjectivity. scLENS eliminates this problem by automatically detecting signals using only the inherent structure of the data."
During the development of scLENS, researchers identified the fundamental reasons for inaccuracies in existing analysis methods. They found that commonly used data preprocessing methods distort both biological signals and noise. The new preprocessing approach that scLENS offers is free from such distortions.
By resolving issues related to noise threshold determined by subjective user choice and signal distortion in conventional data preprocessing, scLENS significantly outperforms existing methods in accuracy. Additionally, scLENS automates the laborious process of signal dimension selection, allowing researchers to extract biological signals conveniently and automatically.
CI Kim added, "scLENS solves major issues in single-cell transcriptome data analysis, substantially improving the accuracy and efficiency throughout the analysis process. This is a prime example of how fundamental mathematical theories can drive innovation in life sciences research, allowing researchers to more quickly and accurately answer biological questions and uncover secrets of life that were previously hidden."
This research was published in the international journal 'Nature Communications' on April 27.
Terminology
* Single-cell RNA sequencing (scRNA-seq): A technique used to measure gene expression levels in individual cells, providing insights into cell heterogeneity and rare cell types.
* Dimensionality reduction: A method to reduce the number of features or variables in a dataset while preserving the most important information, making data analysis more manageable and interpretable.
* Random matrix theory: A mathematical framework used to model and analyze the properties of large, random matrices, which can be applied to filter out noise in high-dimensional data.
* Signal robustness test: Among the signals, this test selects signals that are robust to the slight perturbation in data because real biological signals should be invariant for such slight modification in the data.
2024.05.09 View 6098
Revolutionary 'scLENS' Unveiled to Decode Complex Single-Cell Genomic Data
Unlocking biological information from complex single-cell genomic data has just become easier and more precise, thanks to the innovative 'scLENS' tool developed by the Biomedical Mathematics Group within the IBS Center for Mathematical and Computational Sciences led by Chief Investigator Jae Kyoung Kim, who is also a professor at KAIST. This new finding represents a significant leap forward in the field of single-cell transcriptomics.
Single-cell genomic analysis is an advanced technique that measures gene expression at the individual cell level, revealing cellular changes and interactions that are not observable with traditional genomic analysis methods. When applied to cancer tissues, this analysis can delineate the composition of diverse cell types within a tumor, providing insights into how cancer progresses and identifying key genes involved during each stage of progression.
Despite the immense potential of single-cell genomic analysis, handling the vast amount of data that it generates has always been challenging. The amount of data covers the expression of tens of thousands of genes across hundreds to thousands of individual cells. This not only results in large datasets but also introduces noise-related distortions, which arise in part due to current measurement limitations.
< Figure 1. Overview of scLENS (single-cell Low-dimensional embedding using the effective Noise Subtract) >
(Left) Current dimensionality reduction methods for scRNA-seq data involve conventional data preprocessing steps, such as log normalization, followed by manual selection of signals from the scaled data. However, this study reveals that the high levels of sparsity and variability in scRNA-seq data can lead to signal distortion during the data preprocessing, compromising the accuracy of downstream analyses.
(Right) To address this issue, the researchers integrated L2 normalization into the conventional preprocessing pipeline, effectively mitigating signal distortion. Moreover, they developed a novel signal detection algorithm that eliminates the need for user intervention by leveraging random matrix theory-based noise filtering and signal robustness testing. By incorporating these techniques, scLENS enables accurate and automated analysis of scRNA-seq data, overcoming the limitations of existing dimensionality reduction methods.
Corresponding author Jae Kyoung Kim highlighted, “There has been a remarkable advancement in experimental technologies for analyzing single-cell transcriptomes over the past decade. However, due to limitations in data analysis methods, there has been a struggle to fully utilize valuable data obtained through extensive cost and time."
Researchers have developed numerous analysis methods over the years to discern biological signals from this noise. However, the accuracy of these methods has been less than satisfactory. A critical issue is that determining signal and noise thresholds often depends on subjective decisions from the users.
The newly developed scLENS tool harnesses Random Matrix Theory and Signal robustness test to automatically differentiate signals from noise without relying on subjective user input.
First author Hyun Kim stated, "Previously, users had to arbitrarily decide the threshold for signal and noise, which compromised the reproducibility of analysis results and introduced subjectivity. scLENS eliminates this problem by automatically detecting signals using only the inherent structure of the data."
During the development of scLENS, researchers identified the fundamental reasons for inaccuracies in existing analysis methods. They found that commonly used data preprocessing methods distort both biological signals and noise. The new preprocessing approach that scLENS offers is free from such distortions.
By resolving issues related to noise threshold determined by subjective user choice and signal distortion in conventional data preprocessing, scLENS significantly outperforms existing methods in accuracy. Additionally, scLENS automates the laborious process of signal dimension selection, allowing researchers to extract biological signals conveniently and automatically.
CI Kim added, "scLENS solves major issues in single-cell transcriptome data analysis, substantially improving the accuracy and efficiency throughout the analysis process. This is a prime example of how fundamental mathematical theories can drive innovation in life sciences research, allowing researchers to more quickly and accurately answer biological questions and uncover secrets of life that were previously hidden."
This research was published in the international journal 'Nature Communications' on April 27.
Terminology
* Single-cell RNA sequencing (scRNA-seq): A technique used to measure gene expression levels in individual cells, providing insights into cell heterogeneity and rare cell types.
* Dimensionality reduction: A method to reduce the number of features or variables in a dataset while preserving the most important information, making data analysis more manageable and interpretable.
* Random matrix theory: A mathematical framework used to model and analyze the properties of large, random matrices, which can be applied to filter out noise in high-dimensional data.
* Signal robustness test: Among the signals, this test selects signals that are robust to the slight perturbation in data because real biological signals should be invariant for such slight modification in the data.
2024.05.09 View 6098 -
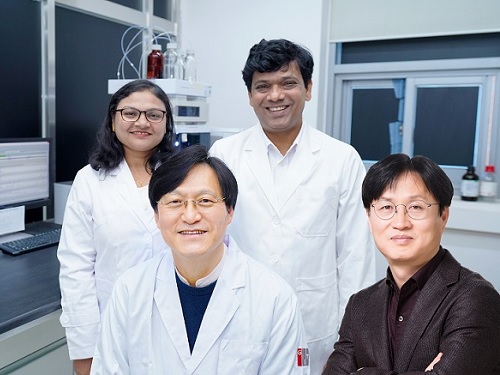 A Korean research team develops a new clinical candidate for fatty liver disease
A team of Korean researchers have succeeded in developing a new drug candidate for the treatment of non-alcoholic fatty liver disease (NAFLD) acting on peripheral tissues. To date, there has not been an optimal treatment for non-alcoholic steatohepatitis (NASH), and this discovery is expected to set the grounds for the development of new drugs that can safely suppress both liver fat accumulation and liver fibrosis at the same time.
A joint research team led by Professor Jin Hee Ahn from Gwangju Institute of Science and Technology (GIST) and Professor Hail Kim from the KAIST Graduate School of Medical Science and Engineering developed a new chemical that can suppress disease-specific protein (HTR2A) through years of basic research. The team also revealed to have verified its efficacy and safety through preclinical tests (animal tests) at JD Bioscience Inc., a start-up company founded by Professor Ahn.
Although NAFLD has a prevalence rate as high as 20-30%, and about 5% of the global adult population suffers from NASH, there are no commercial drugs targeting them to date. NAFLD is a chronic disease that starts from the fatty liver and progresses into steatohepatitis, fibrosis, cirrhosis, and liver cancer. The mortality rate of patients increases with accompanied cardiovascular diseases and liver-related complications, and appropriate treatment in the early stage is hence necessary.
< Figure 1. Strategy and history of 5HT2A antagonists. Library and rational design for the development of compound 11c as a potent 5HT2A antagonist. Previous research efforts were discontinued due to limited oral absorption and safety. A therapeutic candidate to overcome this problem was identified and phase 1 clinical trials are currently in progress. >
The new synthetic chemical developed by the joint GIST-KAIST research is an innovative drug candidate that shows therapeutic effects on NASH based on a dual action mechanism that inhibits the accumulation of fat in the liver and liver fibrosis by suppressing the serotonin receptor protein 5HT2A.
The research team confirmed its therapeutic effects in animal models for NAFLD and NASH, in which hepatic steatosis and liver fibrosis* caused by fat accumulation in the liver were suppressed simultaneously by 50-70%.
*fibrosis: stiffening of parts of the liver, also used as a major indicator to track the prognosis of steatosis
The research team explained that the material was designed with optimal polarity and lipid affinity to minimize its permeability across the blood-brain barrier. It therefore does not affect the brain, and causes little side effects in the central nervous system (CNS) such as depression and suicidal ideations, while demonstrating excellent inhibition on its target protein present in tissues outside brain (IC50* = 14 nM). The team also demonstrated its superior efficacy in improving liver fibrosis when compared to similar drugs in the phase 3 clinical trial.
*IC50 (half maximal inhibitory concentration): the concentration at which a chemical suppresses 50% of a particular biological function
< Figure 2. GM-60106 (11c)'s effect on obesity: When GM-60106 was administered to an obese animal model (mice) for 2 months, body weight, body fat mass, and blood sugar were significantly reduced (a-d). In addition, the steatohepatitis level (NAFLD Activity Score) and the expression of genes of the treated mice involved in adipogenesis along with blood/liver fat decreased (e-h) >
Based on the pharmacological data obtained through preclinical trials, the team evaluated the effects of the drug on 88 healthy adults as part of their phase 1 clinical trial, where the side effects and the safe dosage of a drug are tested against healthy adults. Results showed no serious side effects and a good level of drug safety.
In addition, a preliminary efficacy evaluation on eight adults with steatohepatitis is currently underway.
Professor Jin Hee Ahn said, “The aim of this research is to develop a treatment for NASH with little side effects and guaranteed safety by developing a new target. The developed chemical is currently going through phase 1 of the global clinical trial in Australia through JD Bioscience Inc., a bio venture company for innovative drug development.” he added, “The candidate material the research team is currently developing shows not only a high level of safety and preventative effects by suppressing fat accumulation in the liver, but also a direct therapeutic effect on liver fibrosis. This is a strength that distinguishes our material from other competing drugs.”
< Figure 3. Efficacy of GM-60106 (11c) on liver fibrosis: When GM-60106 was administered to a steatohepatitis model (mice) for 3 months, the expression of genes associated with tissue fibrosis was significantly reduced (b-c). As a result of a detailed analysis of the tissues of the animal model, it was confirmed that the rate of tissue fibrosis was reduced and the expression rate of genes related to tissue fibrosis and inflammation was also significantly reduced (e-h). >
Professor Hail Kim from KAIST said, “Until now, this disease did not have a method of treatment other than weight control, and there has been no attempt to develop a drug that can be used for non-obese patients.” He added, “Through this research, we look forward to the development of various treatment techniques targeting a range of metabolic diseases including NASH that do not affect the weight of the patient.”
This study, conducted together by the research teams led by Professor Ahn from GIST and Professor Kim from KAIST, as well as the research team from JD Bioscience Inc., was supported by the Ministry of Science and ICT, and the National New Drug Development Project. The results of this research were published by Nature Communications on January 20.
The team also presented the results of their clinical study on the candidate material coded GM-60106 targeting metabolic abnormality-related MASH* at NASH-TAG Conference 2024, which was held in Utah for three days starting on January 4, which was selected as an excellent abstract.
*MASH (Metabolic Dysfunction-Associated Steatohepatitis): new replacement term for NASH
2024.02.21 View 9571
A Korean research team develops a new clinical candidate for fatty liver disease
A team of Korean researchers have succeeded in developing a new drug candidate for the treatment of non-alcoholic fatty liver disease (NAFLD) acting on peripheral tissues. To date, there has not been an optimal treatment for non-alcoholic steatohepatitis (NASH), and this discovery is expected to set the grounds for the development of new drugs that can safely suppress both liver fat accumulation and liver fibrosis at the same time.
A joint research team led by Professor Jin Hee Ahn from Gwangju Institute of Science and Technology (GIST) and Professor Hail Kim from the KAIST Graduate School of Medical Science and Engineering developed a new chemical that can suppress disease-specific protein (HTR2A) through years of basic research. The team also revealed to have verified its efficacy and safety through preclinical tests (animal tests) at JD Bioscience Inc., a start-up company founded by Professor Ahn.
Although NAFLD has a prevalence rate as high as 20-30%, and about 5% of the global adult population suffers from NASH, there are no commercial drugs targeting them to date. NAFLD is a chronic disease that starts from the fatty liver and progresses into steatohepatitis, fibrosis, cirrhosis, and liver cancer. The mortality rate of patients increases with accompanied cardiovascular diseases and liver-related complications, and appropriate treatment in the early stage is hence necessary.
< Figure 1. Strategy and history of 5HT2A antagonists. Library and rational design for the development of compound 11c as a potent 5HT2A antagonist. Previous research efforts were discontinued due to limited oral absorption and safety. A therapeutic candidate to overcome this problem was identified and phase 1 clinical trials are currently in progress. >
The new synthetic chemical developed by the joint GIST-KAIST research is an innovative drug candidate that shows therapeutic effects on NASH based on a dual action mechanism that inhibits the accumulation of fat in the liver and liver fibrosis by suppressing the serotonin receptor protein 5HT2A.
The research team confirmed its therapeutic effects in animal models for NAFLD and NASH, in which hepatic steatosis and liver fibrosis* caused by fat accumulation in the liver were suppressed simultaneously by 50-70%.
*fibrosis: stiffening of parts of the liver, also used as a major indicator to track the prognosis of steatosis
The research team explained that the material was designed with optimal polarity and lipid affinity to minimize its permeability across the blood-brain barrier. It therefore does not affect the brain, and causes little side effects in the central nervous system (CNS) such as depression and suicidal ideations, while demonstrating excellent inhibition on its target protein present in tissues outside brain (IC50* = 14 nM). The team also demonstrated its superior efficacy in improving liver fibrosis when compared to similar drugs in the phase 3 clinical trial.
*IC50 (half maximal inhibitory concentration): the concentration at which a chemical suppresses 50% of a particular biological function
< Figure 2. GM-60106 (11c)'s effect on obesity: When GM-60106 was administered to an obese animal model (mice) for 2 months, body weight, body fat mass, and blood sugar were significantly reduced (a-d). In addition, the steatohepatitis level (NAFLD Activity Score) and the expression of genes of the treated mice involved in adipogenesis along with blood/liver fat decreased (e-h) >
Based on the pharmacological data obtained through preclinical trials, the team evaluated the effects of the drug on 88 healthy adults as part of their phase 1 clinical trial, where the side effects and the safe dosage of a drug are tested against healthy adults. Results showed no serious side effects and a good level of drug safety.
In addition, a preliminary efficacy evaluation on eight adults with steatohepatitis is currently underway.
Professor Jin Hee Ahn said, “The aim of this research is to develop a treatment for NASH with little side effects and guaranteed safety by developing a new target. The developed chemical is currently going through phase 1 of the global clinical trial in Australia through JD Bioscience Inc., a bio venture company for innovative drug development.” he added, “The candidate material the research team is currently developing shows not only a high level of safety and preventative effects by suppressing fat accumulation in the liver, but also a direct therapeutic effect on liver fibrosis. This is a strength that distinguishes our material from other competing drugs.”
< Figure 3. Efficacy of GM-60106 (11c) on liver fibrosis: When GM-60106 was administered to a steatohepatitis model (mice) for 3 months, the expression of genes associated with tissue fibrosis was significantly reduced (b-c). As a result of a detailed analysis of the tissues of the animal model, it was confirmed that the rate of tissue fibrosis was reduced and the expression rate of genes related to tissue fibrosis and inflammation was also significantly reduced (e-h). >
Professor Hail Kim from KAIST said, “Until now, this disease did not have a method of treatment other than weight control, and there has been no attempt to develop a drug that can be used for non-obese patients.” He added, “Through this research, we look forward to the development of various treatment techniques targeting a range of metabolic diseases including NASH that do not affect the weight of the patient.”
This study, conducted together by the research teams led by Professor Ahn from GIST and Professor Kim from KAIST, as well as the research team from JD Bioscience Inc., was supported by the Ministry of Science and ICT, and the National New Drug Development Project. The results of this research were published by Nature Communications on January 20.
The team also presented the results of their clinical study on the candidate material coded GM-60106 targeting metabolic abnormality-related MASH* at NASH-TAG Conference 2024, which was held in Utah for three days starting on January 4, which was selected as an excellent abstract.
*MASH (Metabolic Dysfunction-Associated Steatohepatitis): new replacement term for NASH
2024.02.21 View 9571 -
 KAIST Research Team Breaks Down Musical Instincts with AI
Music, often referred to as the universal language, is known to be a common component in all cultures. Then, could ‘musical instinct’ be something that is shared to some degree despite the extensive environmental differences amongst cultures?
On January 16, a KAIST research team led by Professor Hawoong Jung from the Department of Physics announced to have identified the principle by which musical instincts emerge from the human brain without special learning using an artificial neural network model.
Previously, many researchers have attempted to identify the similarities and differences between the music that exist in various different cultures, and tried to understand the origin of the universality. A paper published in Science in 2019 had revealed that music is produced in all ethnographically distinct cultures, and that similar forms of beats and tunes are used. Neuroscientist have also previously found out that a specific part of the human brain, namely the auditory cortex, is responsible for processing musical information.
Professor Jung’s team used an artificial neural network model to show that cognitive functions for music forms spontaneously as a result of processing auditory information received from nature, without being taught music. The research team utilized AudioSet, a large-scale collection of sound data provided by Google, and taught the artificial neural network to learn the various sounds. Interestingly, the research team discovered that certain neurons within the network model would respond selectively to music. In other words, they observed the spontaneous generation of neurons that reacted minimally to various other sounds like those of animals, nature, or machines, but showed high levels of response to various forms of music including both instrumental and vocal.
The neurons in the artificial neural network model showed similar reactive behaviours to those in the auditory cortex of a real brain. For example, artificial neurons responded less to the sound of music that was cropped into short intervals and were rearranged. This indicates that the spontaneously-generated music-selective neurons encode the temporal structure of music. This property was not limited to a specific genre of music, but emerged across 25 different genres including classic, pop, rock, jazz, and electronic.
< Figure 1. Illustration of the musicality of the brain and artificial neural network (created with DALL·E3 AI based on the paper content) >
Furthermore, suppressing the activity of the music-selective neurons was found to greatly impede the cognitive accuracy for other natural sounds. That is to say, the neural function that processes musical information helps process other sounds, and that ‘musical ability’ may be an instinct formed as a result of an evolutionary adaptation acquired to better process sounds from nature.
Professor Hawoong Jung, who advised the research, said, “The results of our study imply that evolutionary pressure has contributed to forming the universal basis for processing musical information in various cultures.” As for the significance of the research, he explained, “We look forward for this artificially built model with human-like musicality to become an original model for various applications including AI music generation, musical therapy, and for research in musical cognition.” He also commented on its limitations, adding, “This research however does not take into consideration the developmental process that follows the learning of music, and it must be noted that this is a study on the foundation of processing musical information in early development.”
< Figure 2. The artificial neural network that learned to recognize non-musical natural sounds in the cyber space distinguishes between music and non-music. >
This research, conducted by first author Dr. Gwangsu Kim of the KAIST Department of Physics (current affiliation: MIT Department of Brain and Cognitive Sciences) and Dr. Dong-Kyum Kim (current affiliation: IBS) was published in Nature Communications under the title, “Spontaneous emergence of rudimentary music detectors in deep neural networks”.
This research was supported by the National Research Foundation of Korea.
2024.01.23 View 7798
KAIST Research Team Breaks Down Musical Instincts with AI
Music, often referred to as the universal language, is known to be a common component in all cultures. Then, could ‘musical instinct’ be something that is shared to some degree despite the extensive environmental differences amongst cultures?
On January 16, a KAIST research team led by Professor Hawoong Jung from the Department of Physics announced to have identified the principle by which musical instincts emerge from the human brain without special learning using an artificial neural network model.
Previously, many researchers have attempted to identify the similarities and differences between the music that exist in various different cultures, and tried to understand the origin of the universality. A paper published in Science in 2019 had revealed that music is produced in all ethnographically distinct cultures, and that similar forms of beats and tunes are used. Neuroscientist have also previously found out that a specific part of the human brain, namely the auditory cortex, is responsible for processing musical information.
Professor Jung’s team used an artificial neural network model to show that cognitive functions for music forms spontaneously as a result of processing auditory information received from nature, without being taught music. The research team utilized AudioSet, a large-scale collection of sound data provided by Google, and taught the artificial neural network to learn the various sounds. Interestingly, the research team discovered that certain neurons within the network model would respond selectively to music. In other words, they observed the spontaneous generation of neurons that reacted minimally to various other sounds like those of animals, nature, or machines, but showed high levels of response to various forms of music including both instrumental and vocal.
The neurons in the artificial neural network model showed similar reactive behaviours to those in the auditory cortex of a real brain. For example, artificial neurons responded less to the sound of music that was cropped into short intervals and were rearranged. This indicates that the spontaneously-generated music-selective neurons encode the temporal structure of music. This property was not limited to a specific genre of music, but emerged across 25 different genres including classic, pop, rock, jazz, and electronic.
< Figure 1. Illustration of the musicality of the brain and artificial neural network (created with DALL·E3 AI based on the paper content) >
Furthermore, suppressing the activity of the music-selective neurons was found to greatly impede the cognitive accuracy for other natural sounds. That is to say, the neural function that processes musical information helps process other sounds, and that ‘musical ability’ may be an instinct formed as a result of an evolutionary adaptation acquired to better process sounds from nature.
Professor Hawoong Jung, who advised the research, said, “The results of our study imply that evolutionary pressure has contributed to forming the universal basis for processing musical information in various cultures.” As for the significance of the research, he explained, “We look forward for this artificially built model with human-like musicality to become an original model for various applications including AI music generation, musical therapy, and for research in musical cognition.” He also commented on its limitations, adding, “This research however does not take into consideration the developmental process that follows the learning of music, and it must be noted that this is a study on the foundation of processing musical information in early development.”
< Figure 2. The artificial neural network that learned to recognize non-musical natural sounds in the cyber space distinguishes between music and non-music. >
This research, conducted by first author Dr. Gwangsu Kim of the KAIST Department of Physics (current affiliation: MIT Department of Brain and Cognitive Sciences) and Dr. Dong-Kyum Kim (current affiliation: IBS) was published in Nature Communications under the title, “Spontaneous emergence of rudimentary music detectors in deep neural networks”.
This research was supported by the National Research Foundation of Korea.
2024.01.23 View 7798 -
 A KAIST Research Team Observes the Processes of Memory and Cognition in Real Time
The human brain contains approximately 86 billion neurons and 600 trillion synapses that exchange signals between the neurons to help us control the various functions of the brain including cognition, emotion, and memory. Interestingly, the number of synapses decrease with age or as a result of diseases like Alzheimer’s, and research on synapses thus attracts a lot of attention. However, limitations have existed in observing the dynamics of synapse structures in real time.
On January 9, a joint research team led by Professor Won Do Heo from the KAIST Department of Biological Sciences, Professor Hyung-Bae Kwon from Johns Hopkins School of Medicine, and Professor Sangkyu Lee from the Institute for Basic Science (IBS) revealed that they have developed the world’s first technique to allow a real-time observation of synapse formation, extinction, and alterations.
Professor Heo’s team conjugated dimerization-dependent fluorescent proteins (ddFP) to synapses in order to observe the process in which synapses create connections between neurons in real time. The team named this technique SynapShot, by combining the words ‘synapse’ and snapshot’, and successfully tracked and observed the live formation and extinction processes of synapses as well as their dynamic changes.
< Figure 1. To observe dynamically changing synapses, dimerization-dependent fluorescent protein (ddFP) was expressed to observe flourescent signals upon synapse formation as ddFP enables fluorescence detection through reversible binding to pre- and postsynaptic terminals. >
Through a joint research project, the teams led by Professor Heo and Professor Sangkyu Lee at IBS together designed a SynapShot with green and red fluorescence, and were able to easily distinguish the synapse connecting two different neurons. Additionally, by combining an optogenetic technique that can control the function of a molecule using light, the team was able to observe the changes in the synapses while simultaneously inducing certain functions of the neurons using light.
Through more joint research with the team led by Professor Hyung-Bae Kwon at the Johns Hopkins School of Medicine, Professor Heo’s team induced several situations on live mice, including visual discrimination training, exercise, and anaesthesia, and used SynapShot to observe the changes in the synapses during each situation in real time. The observations revealed that each synapse could change fairly quickly and dynamically. This was the first-ever case in which the changes in synapses were observed in a live mammal.
< Figure 2. Microscopic photos observed through changes of the flourescence of the synapse sensor (SynapShot) by cultivating the neurons of an experimental rat and expressing the SynapShot. The changes in the synapse that is created when the pre- and post-synaptic terminals come into contact and the synapse that disappears after a certain period of time are measured by the fluorescence of the SynapShot. >
Professor Heo said, “Our group developed SynapShot through a collaboration with domestic and international research teams, and have opened up the possibility for first-hand live observations of the quick and dynamic changes of synapses, which was previously difficult to do. We expect this technique to revolutionize research methodology in the neurological field, and play an important role in brightening the future of brain science.”
This research, conducted by co-first authors Seungkyu Son (Ph.D. candidate), Jinsu Lee (Ph.D. candidate) and Dr. Kanghoon Jung from Johns Hopkins, was published in the online edition of Nature Methods on January 8 under the title “Real-time visualization of structural dynamics of synapses in live cells in vivo”, and will be printed in the February volume.
< Figure 3. Simultaneous use of green-SynapShot and red-SynapShot to distinguish and observe synapses with one post-terminal and different pre-terminals. >
< Figure 4. Dimer-dependent fluorescent protein (ddFP) exists as a green fluorescent protein as well as a red fluorescent protein, and can be applied together with blue light-activated optogenetic technology. After activating Tropomyosin receptor kinase B (TrkB) by blue light using optogenetic technology, the strengthening of synaptic connections through signals of brain-derived neurotrophic factor is observed using red-SynapShot. >
< Figure 5. Micrographs showing real-time changing synapses in the visual cortex of mice trained through visual training using in vivo imaging techniques such as two-photon microscopy as well as at the cellular level. >
This research was supported by Mid-Sized Research Funds and the Singularity Project from KAIST, and by IBS.
2024.01.18 View 7325
A KAIST Research Team Observes the Processes of Memory and Cognition in Real Time
The human brain contains approximately 86 billion neurons and 600 trillion synapses that exchange signals between the neurons to help us control the various functions of the brain including cognition, emotion, and memory. Interestingly, the number of synapses decrease with age or as a result of diseases like Alzheimer’s, and research on synapses thus attracts a lot of attention. However, limitations have existed in observing the dynamics of synapse structures in real time.
On January 9, a joint research team led by Professor Won Do Heo from the KAIST Department of Biological Sciences, Professor Hyung-Bae Kwon from Johns Hopkins School of Medicine, and Professor Sangkyu Lee from the Institute for Basic Science (IBS) revealed that they have developed the world’s first technique to allow a real-time observation of synapse formation, extinction, and alterations.
Professor Heo’s team conjugated dimerization-dependent fluorescent proteins (ddFP) to synapses in order to observe the process in which synapses create connections between neurons in real time. The team named this technique SynapShot, by combining the words ‘synapse’ and snapshot’, and successfully tracked and observed the live formation and extinction processes of synapses as well as their dynamic changes.
< Figure 1. To observe dynamically changing synapses, dimerization-dependent fluorescent protein (ddFP) was expressed to observe flourescent signals upon synapse formation as ddFP enables fluorescence detection through reversible binding to pre- and postsynaptic terminals. >
Through a joint research project, the teams led by Professor Heo and Professor Sangkyu Lee at IBS together designed a SynapShot with green and red fluorescence, and were able to easily distinguish the synapse connecting two different neurons. Additionally, by combining an optogenetic technique that can control the function of a molecule using light, the team was able to observe the changes in the synapses while simultaneously inducing certain functions of the neurons using light.
Through more joint research with the team led by Professor Hyung-Bae Kwon at the Johns Hopkins School of Medicine, Professor Heo’s team induced several situations on live mice, including visual discrimination training, exercise, and anaesthesia, and used SynapShot to observe the changes in the synapses during each situation in real time. The observations revealed that each synapse could change fairly quickly and dynamically. This was the first-ever case in which the changes in synapses were observed in a live mammal.
< Figure 2. Microscopic photos observed through changes of the flourescence of the synapse sensor (SynapShot) by cultivating the neurons of an experimental rat and expressing the SynapShot. The changes in the synapse that is created when the pre- and post-synaptic terminals come into contact and the synapse that disappears after a certain period of time are measured by the fluorescence of the SynapShot. >
Professor Heo said, “Our group developed SynapShot through a collaboration with domestic and international research teams, and have opened up the possibility for first-hand live observations of the quick and dynamic changes of synapses, which was previously difficult to do. We expect this technique to revolutionize research methodology in the neurological field, and play an important role in brightening the future of brain science.”
This research, conducted by co-first authors Seungkyu Son (Ph.D. candidate), Jinsu Lee (Ph.D. candidate) and Dr. Kanghoon Jung from Johns Hopkins, was published in the online edition of Nature Methods on January 8 under the title “Real-time visualization of structural dynamics of synapses in live cells in vivo”, and will be printed in the February volume.
< Figure 3. Simultaneous use of green-SynapShot and red-SynapShot to distinguish and observe synapses with one post-terminal and different pre-terminals. >
< Figure 4. Dimer-dependent fluorescent protein (ddFP) exists as a green fluorescent protein as well as a red fluorescent protein, and can be applied together with blue light-activated optogenetic technology. After activating Tropomyosin receptor kinase B (TrkB) by blue light using optogenetic technology, the strengthening of synaptic connections through signals of brain-derived neurotrophic factor is observed using red-SynapShot. >
< Figure 5. Micrographs showing real-time changing synapses in the visual cortex of mice trained through visual training using in vivo imaging techniques such as two-photon microscopy as well as at the cellular level. >
This research was supported by Mid-Sized Research Funds and the Singularity Project from KAIST, and by IBS.
2024.01.18 View 7325 -
 The Relentless Rain: East Asia's Recent Floods and What Lies Beneath
In just a month's time, East Asia witnessed torrential downpours that would usually span an entire season. Japan, battered by three times its usual monthly rainfall, faced landslides and flooding that claimed over 200 lives. Meanwhile, South Korea grappled with its longest monsoon in seven years, leaving more than 40 individuals dead or missing. But these events, as harrowing as they sound, are more than just weather anomalies. They're telltale signs, symptoms of a larger malaise that has gripped our planet.
Diving deep into these rain-soaked mysteries, a recently published paper in the journal Science Advances offers a fresh perspective. Led by a research team at the Korea Advanced Institute of Science and Technology (KAIST), Korea, the research unpacks the influence of human-induced climate changes on the East Asia Summer Monsoon frontal system.
Heavy summer rain has a significant impact on agriculture and industry, and can be said to be one of the greatest threats to human society by causing disasters such as floods and landslides, affecting the local ecosystem. It has been reported from all over the world that the intensity of summer heavy rain has changed over the past few decades. However, summer rain in East Asia is caused by various forms such as typhoons, extratropical cyclones, and fronts, and efforts to study heavy frontal rain, which account for more than 40% of summer rainfall, is still insufficient. In addition, because heavy rain is also influenced by natural fluctuations or coincidences in the climate system, it is not yet known to what extent warming due to human activities affects the intensity of heavy frontal precipitation.
An international joint research team consisting of eight institutions from Korea, the United States, and Japan, including KAIST, Tokyo University, Tokyo Institute of Technology, Chonnam National University, GIST, and Utah State University, confirmed the intensity of heavy rain caused by the weather fronts in East Asia using observation data for the past 60 years and found that the coast of southeastern China. It was found that the intensity of heavy rain increased by about 17% across the Korean Peninsula and Japan. To investigate the cause of these changes, the research team used the Earth Metaverse experiment, which simulated Earth with and without greenhouse gas emissions due to human activities, and found that heavy rain intensity was strengthened by about 6% due to greenhouse gas emissions, and the changes discovered were has succeeded for the first time in the world in showing that warming cannot be explained without the effects of human activities.
< Figure 1. (Left) Observed difference in frontal rainfall intensity (FRI) from the later (1991–2015) to the earlier periods (1958–1982) (Right) Visualization of the impact of anthropogenic warming on the intensity of heavy frontal rain analyzed using the Earth Metaverse experiment. >
"It's not just about connecting the dots," said Moon, the first author of the paper, "it's about seeing the larger pattern. Our data analysis reveals a clear and intensified trend in East Asia's frontal rainfall, one that's intertwined with human actions and increasingly harmful for lives and property."
One of the intriguing finds from the study is the mechanics behind this intensification. The team found increased moisture transport due to a warmer climate, which, when coupled with the strengthening of a gigantic weather system called the West North Pacific Subtropical High, results in enhanced frontal rainfall. It’s akin to the climate dialing up the volume on rain events. As the atmosphere warms, it holds more moisture, leading to heavier downpours when conditions are right.
Nobuyuki Utsumi, another voice from the team, makes the science accessible for all, saying, "Monsoon rain isn't just rain anymore. The frequency, the intensity, it's changing. And our actions, our carbon footprint, are casting a larger shadow than we anticipated."
While the devastating news of floods fills headlines, Professor Simon Wang of Utah State University, reminds us of the underlying importance of their study. "It's like reading a detective novel. To solve the mystery of these floods, one has to trace them back to their roots. This work hints at a future where such intense rain events aren't the exception but might become an everyday reality."
Hyungjun Kim, the principal investigator of the team throws in a note of caution, "Understanding this is just the first step. Predicting and preparing for these extremes is the real challenge. Every amplified rainfall event is a message from the future, urging us to adapt." So far, predicting rainfall intensity and locations remains a challenging task for even the most sophisticated weather models.
< Figure 2. Comparison of rates of change in Anthropocene fingerprints. The horizontal axis shows the long-term change slope of the Anthropocene fingerprint signal (1958 to 2015). Shows the probability distribution of slopes extracted from the non-warming experiment (blue) and the warming experiment (red). The vertical solid lines are the slope of the Anthropocene fingerprint signal extracted from observational data. >
The researchers say, “Facing climate change, the narrative of this new study is more than mere numbers and data. It's a story of our planet, our actions, and the rain-soaked repercussions we're beginning to face. As we mop up the aftermath of another flood, research like Moon's beckons us to look deeper, understand better, and act wiser.”
< Figure 3. Comparison of water vapor convergence and rate of change of the western North Pacific high pressure. Shows the gradient of change in water vapor convergence (horizontal axis) and the Northwestern Pacific-East Asia pressure gradient (vertical axis) extracted from warming (red) and non-warming (blue) experiments. Shows the distribution of slope changes of the two indices during the period 1958 to 1982 (P1) and 1991 to 2015 (P2). >
The results of this study were published on November 24 in Science Advances. (Paper title: Anthropogenic warming induced intensification of summer monsoon frontal precipitation over East Asia)
This research was conducted with support from the National Research Foundation of Korea's Overseas Scientist Attraction Project (BP+) and the Anthropocene Research Center.
2023.12.05 View 6374
The Relentless Rain: East Asia's Recent Floods and What Lies Beneath
In just a month's time, East Asia witnessed torrential downpours that would usually span an entire season. Japan, battered by three times its usual monthly rainfall, faced landslides and flooding that claimed over 200 lives. Meanwhile, South Korea grappled with its longest monsoon in seven years, leaving more than 40 individuals dead or missing. But these events, as harrowing as they sound, are more than just weather anomalies. They're telltale signs, symptoms of a larger malaise that has gripped our planet.
Diving deep into these rain-soaked mysteries, a recently published paper in the journal Science Advances offers a fresh perspective. Led by a research team at the Korea Advanced Institute of Science and Technology (KAIST), Korea, the research unpacks the influence of human-induced climate changes on the East Asia Summer Monsoon frontal system.
Heavy summer rain has a significant impact on agriculture and industry, and can be said to be one of the greatest threats to human society by causing disasters such as floods and landslides, affecting the local ecosystem. It has been reported from all over the world that the intensity of summer heavy rain has changed over the past few decades. However, summer rain in East Asia is caused by various forms such as typhoons, extratropical cyclones, and fronts, and efforts to study heavy frontal rain, which account for more than 40% of summer rainfall, is still insufficient. In addition, because heavy rain is also influenced by natural fluctuations or coincidences in the climate system, it is not yet known to what extent warming due to human activities affects the intensity of heavy frontal precipitation.
An international joint research team consisting of eight institutions from Korea, the United States, and Japan, including KAIST, Tokyo University, Tokyo Institute of Technology, Chonnam National University, GIST, and Utah State University, confirmed the intensity of heavy rain caused by the weather fronts in East Asia using observation data for the past 60 years and found that the coast of southeastern China. It was found that the intensity of heavy rain increased by about 17% across the Korean Peninsula and Japan. To investigate the cause of these changes, the research team used the Earth Metaverse experiment, which simulated Earth with and without greenhouse gas emissions due to human activities, and found that heavy rain intensity was strengthened by about 6% due to greenhouse gas emissions, and the changes discovered were has succeeded for the first time in the world in showing that warming cannot be explained without the effects of human activities.
< Figure 1. (Left) Observed difference in frontal rainfall intensity (FRI) from the later (1991–2015) to the earlier periods (1958–1982) (Right) Visualization of the impact of anthropogenic warming on the intensity of heavy frontal rain analyzed using the Earth Metaverse experiment. >
"It's not just about connecting the dots," said Moon, the first author of the paper, "it's about seeing the larger pattern. Our data analysis reveals a clear and intensified trend in East Asia's frontal rainfall, one that's intertwined with human actions and increasingly harmful for lives and property."
One of the intriguing finds from the study is the mechanics behind this intensification. The team found increased moisture transport due to a warmer climate, which, when coupled with the strengthening of a gigantic weather system called the West North Pacific Subtropical High, results in enhanced frontal rainfall. It’s akin to the climate dialing up the volume on rain events. As the atmosphere warms, it holds more moisture, leading to heavier downpours when conditions are right.
Nobuyuki Utsumi, another voice from the team, makes the science accessible for all, saying, "Monsoon rain isn't just rain anymore. The frequency, the intensity, it's changing. And our actions, our carbon footprint, are casting a larger shadow than we anticipated."
While the devastating news of floods fills headlines, Professor Simon Wang of Utah State University, reminds us of the underlying importance of their study. "It's like reading a detective novel. To solve the mystery of these floods, one has to trace them back to their roots. This work hints at a future where such intense rain events aren't the exception but might become an everyday reality."
Hyungjun Kim, the principal investigator of the team throws in a note of caution, "Understanding this is just the first step. Predicting and preparing for these extremes is the real challenge. Every amplified rainfall event is a message from the future, urging us to adapt." So far, predicting rainfall intensity and locations remains a challenging task for even the most sophisticated weather models.
< Figure 2. Comparison of rates of change in Anthropocene fingerprints. The horizontal axis shows the long-term change slope of the Anthropocene fingerprint signal (1958 to 2015). Shows the probability distribution of slopes extracted from the non-warming experiment (blue) and the warming experiment (red). The vertical solid lines are the slope of the Anthropocene fingerprint signal extracted from observational data. >
The researchers say, “Facing climate change, the narrative of this new study is more than mere numbers and data. It's a story of our planet, our actions, and the rain-soaked repercussions we're beginning to face. As we mop up the aftermath of another flood, research like Moon's beckons us to look deeper, understand better, and act wiser.”
< Figure 3. Comparison of water vapor convergence and rate of change of the western North Pacific high pressure. Shows the gradient of change in water vapor convergence (horizontal axis) and the Northwestern Pacific-East Asia pressure gradient (vertical axis) extracted from warming (red) and non-warming (blue) experiments. Shows the distribution of slope changes of the two indices during the period 1958 to 1982 (P1) and 1991 to 2015 (P2). >
The results of this study were published on November 24 in Science Advances. (Paper title: Anthropogenic warming induced intensification of summer monsoon frontal precipitation over East Asia)
This research was conducted with support from the National Research Foundation of Korea's Overseas Scientist Attraction Project (BP+) and the Anthropocene Research Center.
2023.12.05 View 6374