ACT
-
 KAIST proposes alternatives to chemical factories through “iBridge”
- A computer simulation program “iBridge” was developed at KAIST that can put together microbial cell factories quickly and efficiently to produce cosmetics and food additives, and raw materials for nylons
- Eco-friendly and sustainable fermentation process to establish an alternative to chemical plants
As climate change and environmental concerns intensify, sustainable microbial cell factories garner significant attention as candidates to replace chemical plants. To develop microorganisms to be used in the microbial cell factories, it is crucial to modify their metabolic processes to induce efficient target chemical production by modulating its gene expressions. Yet, the challenge persists in determining which gene expressions to amplify and suppress, and the experimental verification of these modification targets is a time- and resource-intensive process even for experts. The challenges were addressed by a team of researchers at KAIST (President Kwang-Hyung Lee) led by Distinguished Professor Sang Yup Lee.
It was announced on the 9th by the school that a method for building a microbial factory at low cost, quickly and efficiently, was presented by a novel computer simulation program developed by the team under Professor Lee’s guidance, which is named “iBridge”. This innovative system is designed to predict gene targets to either overexpress or downregulate in the goal of producing a desired compound to enable the cost-effective and efficient construction of microbial cell factories specifically tailored for producing the chemical compound in demand from renewable biomass.
Systems metabolic engineering is a field of research and engineering pioneered by KAIST’s Distinguished Professor Sang Yup Lee that seeks to produce valuable compounds in industrial demands using microorganisms that are re-configured by a combination of methods including, but not limited to, metabolic engineering, synthetic biology, systems biology, and fermentation engineering.
In order to improve microorganisms’ capability to produce useful compounds, it is essential to delete, suppress, or overexpress microbial genes. However, it is difficult even for the experts to identify the gene targets to modify without experimental confirmations for each of them, which can take up immeasurable amount of time and resources.
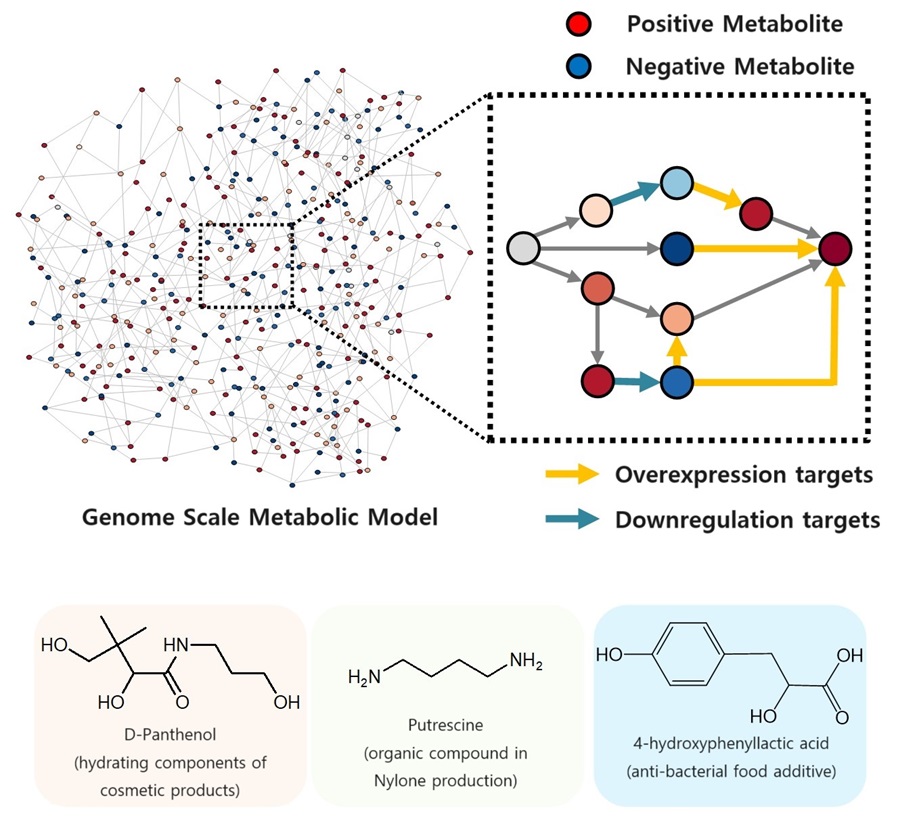
The newly developed iBridge identifies positive and negative metabolites within cells, which exert positive and/or negative impact on formation of the products, by calculating the sum of covariances of their outgoing (consuming) reaction fluxes for a target chemical. Subsequently, it pinpoints "bridge" reactions responsible for converting negative metabolites into positive ones as candidates for overexpression, while identifying the opposites as targets for downregulation.
The research team successfully utilized the iBridge simulation to establish E. coli microbial cell factories each capable of producing three of the compounds that are in high demands at a production capacity that has not been reported around the world. They developed E. coli strains that can each produce panthenol, a moisturizing agent found in many cosmetics, putrescine, which is one of the key components in nylon production, and 4-hydroxyphenyllactic acid, an anti-bacterial food additive. In addition to these three compounds, the study presents predictions for overexpression and suppression genes to construct microbial factories for 298 other industrially valuable compounds.
Dr. Youngjoon Lee, the co-first author of this paper from KAIST, emphasized the accelerated construction of various microbial factories the newly developed simulation enabled. He stated, "With the use of this simulation, multiple microbial cell factories have been established significantly faster than it would have been using the conventional methods. Microbial cell factories producing a wider range of valuable compounds can now be constructed quickly using this technology."
Professor Sang Yup Lee said, "Systems metabolic engineering is a crucial technology for addressing the current climate change issues." He added, "This simulation could significantly expedite the transition from resorting to conventional chemical factories to utilizing environmentally friendly microbial factories."
< Figure. Conceptual diagram of the flow of iBridge simulation >
The team’s work on iBridge is described in a paper titled "Genome-Wide Identification of Overexpression and Downregulation Gene Targets Based on the Sum of Covariances of the Outgoing Reaction Fluxes" written by Dr. Won Jun Kim, and Dr. Youngjoon Lee of the Bioprocess Research Center and Professors Hyun Uk Kim and Sang Yup Lee of the Department of Chemical and Biomolecular Engineering of KAIST. The paper was published via peer-review on the 6th of November on “Cell Systems” by Cell Press.
This research was conducted with the support from the Development of Platform Technologies of Microbial Cell Factories for the Next-generation Biorefineries Project (Project Leader: Distinguished Professor Sang Yup Lee, KAIST) and Development of Platform Technology for the Production of Novel Aromatic Bioplastic using Microbial Cell Factories Project (Project Leader: Research Professor So Young Choi, KAIST) of the Korean Ministry of Science and ICT.
2023.11.09 View 6546
KAIST proposes alternatives to chemical factories through “iBridge”
- A computer simulation program “iBridge” was developed at KAIST that can put together microbial cell factories quickly and efficiently to produce cosmetics and food additives, and raw materials for nylons
- Eco-friendly and sustainable fermentation process to establish an alternative to chemical plants
As climate change and environmental concerns intensify, sustainable microbial cell factories garner significant attention as candidates to replace chemical plants. To develop microorganisms to be used in the microbial cell factories, it is crucial to modify their metabolic processes to induce efficient target chemical production by modulating its gene expressions. Yet, the challenge persists in determining which gene expressions to amplify and suppress, and the experimental verification of these modification targets is a time- and resource-intensive process even for experts. The challenges were addressed by a team of researchers at KAIST (President Kwang-Hyung Lee) led by Distinguished Professor Sang Yup Lee.
It was announced on the 9th by the school that a method for building a microbial factory at low cost, quickly and efficiently, was presented by a novel computer simulation program developed by the team under Professor Lee’s guidance, which is named “iBridge”. This innovative system is designed to predict gene targets to either overexpress or downregulate in the goal of producing a desired compound to enable the cost-effective and efficient construction of microbial cell factories specifically tailored for producing the chemical compound in demand from renewable biomass.
Systems metabolic engineering is a field of research and engineering pioneered by KAIST’s Distinguished Professor Sang Yup Lee that seeks to produce valuable compounds in industrial demands using microorganisms that are re-configured by a combination of methods including, but not limited to, metabolic engineering, synthetic biology, systems biology, and fermentation engineering.
In order to improve microorganisms’ capability to produce useful compounds, it is essential to delete, suppress, or overexpress microbial genes. However, it is difficult even for the experts to identify the gene targets to modify without experimental confirmations for each of them, which can take up immeasurable amount of time and resources.
The newly developed iBridge identifies positive and negative metabolites within cells, which exert positive and/or negative impact on formation of the products, by calculating the sum of covariances of their outgoing (consuming) reaction fluxes for a target chemical. Subsequently, it pinpoints "bridge" reactions responsible for converting negative metabolites into positive ones as candidates for overexpression, while identifying the opposites as targets for downregulation.
The research team successfully utilized the iBridge simulation to establish E. coli microbial cell factories each capable of producing three of the compounds that are in high demands at a production capacity that has not been reported around the world. They developed E. coli strains that can each produce panthenol, a moisturizing agent found in many cosmetics, putrescine, which is one of the key components in nylon production, and 4-hydroxyphenyllactic acid, an anti-bacterial food additive. In addition to these three compounds, the study presents predictions for overexpression and suppression genes to construct microbial factories for 298 other industrially valuable compounds.
Dr. Youngjoon Lee, the co-first author of this paper from KAIST, emphasized the accelerated construction of various microbial factories the newly developed simulation enabled. He stated, "With the use of this simulation, multiple microbial cell factories have been established significantly faster than it would have been using the conventional methods. Microbial cell factories producing a wider range of valuable compounds can now be constructed quickly using this technology."
Professor Sang Yup Lee said, "Systems metabolic engineering is a crucial technology for addressing the current climate change issues." He added, "This simulation could significantly expedite the transition from resorting to conventional chemical factories to utilizing environmentally friendly microbial factories."
< Figure. Conceptual diagram of the flow of iBridge simulation >
The team’s work on iBridge is described in a paper titled "Genome-Wide Identification of Overexpression and Downregulation Gene Targets Based on the Sum of Covariances of the Outgoing Reaction Fluxes" written by Dr. Won Jun Kim, and Dr. Youngjoon Lee of the Bioprocess Research Center and Professors Hyun Uk Kim and Sang Yup Lee of the Department of Chemical and Biomolecular Engineering of KAIST. The paper was published via peer-review on the 6th of November on “Cell Systems” by Cell Press.
This research was conducted with the support from the Development of Platform Technologies of Microbial Cell Factories for the Next-generation Biorefineries Project (Project Leader: Distinguished Professor Sang Yup Lee, KAIST) and Development of Platform Technology for the Production of Novel Aromatic Bioplastic using Microbial Cell Factories Project (Project Leader: Research Professor So Young Choi, KAIST) of the Korean Ministry of Science and ICT.
2023.11.09 View 6546 -
 A KAIST Research Team Produces Eco-Friendly Nylon with Engineered Bacterium
With worsening climate change and environmental issues, in recent years, there has been increased interest in the eco-friendly production of polymers like nylon.
On August 10, Dr. Taehee Han from a KAIST research team led by Distinguished Professor Sang Yup Lee in the Department of Chemical and Biomolecular Engineering revealed the successful development of a microbial strain that produces valerolactam, a monomer of nylon-5.
Valerolactam is an important monomer that constitutes nylon-5 and nylon-6,5. Nylon is the oldest synthetic polymer, and nylon-5 is one of its derivatives composed of monomers with five carbons, while nylon-5,6 is composed of two types of monomers with either five or six carbons. They not only have excellent processability, but are also light and tough, which allows them to be applied in a wide range of industrial sectors including clothing, badminton rackets, fishing nets, tents, and gear parts. Monomers are materials that can be built into polymers, and synthetic processes are what connects them into a polymer.
The chemical production of valerolactam, however, is based on petrochemistry, where extreme reaction conditions are required and toxic waste is produced. To solve these problems, efforts are being made to develop environmentally friendly and highly efficient microbial cell factories for lactam production. Systems metabolic engineering, a key strategy for effective microbial strain development, is a research field pioneered by Professor Sang Yup Lee.
Professor Lee’s team used metabolic engineering, a technique for manipulating microbial metabolic pathways, to construct a synthetic metabolic pathway for valerolactam production in Corynebacteriam glutamicum, a bacterium commonly used for amino acid production. With this, they successfully developed a microbial strain that utilizes biomass-derived glucose as a carbon source to produce high-value valerolactam.
In 2017, the team suggested a novel method that metabolically manipulates Escherichia coli to produce valerolactam. However, there were several limitations at the time including low producibility and the generation of harmful byproducts.
< Figure 1. Schematic graphical representation of the development of microorganisms that produce valerolactam, a nylon-5 monomer >
In this research, the team improved valerolactam producibility and incorporated an additional systems metabolic strategy to the developed microbial strain while eliminating the harmful byproducts. By removing the gene involved in the production of the main byproduct and through gene screening, the team successfully converted 5-aminovaleric acid, a byproduct and a precursor, into valerolactam.
Furthermore, by employing a strategy where the 5-aminovaleric acid-converting gene is inserted multiple times into the genome, the team strengthened the metabolic flux for valerolactam production. As a result, they reached a world-record concentration of 76.1 g/L, which is 6.17 times greater than what was previously reported.
This study was published in Metabolic Engineering on July 12, under the title, “Metabolic engineering of Corynebacterium glutamicum for the high-level production of valerolactam, a nylon-5 monomer”.
Dr. Taehee Han, the first author of the paper, said, “The significance of this research lies in our development of an environmentally friendly technology that efficiently produces monomer lactam for nylon production using microorganisms.” She added, “Through this technology, we will be able to take a step forward in replacing the petrochemical industry with a microorganism-based biopolymer industry.”
This work was supported by the “Development of Next-Generation Biofinery Platform Technologies for Leading Bio-based Chemicals Industry Project” funded by the Korean Ministry of Science and ICT.
2023.08.24 View 5163
A KAIST Research Team Produces Eco-Friendly Nylon with Engineered Bacterium
With worsening climate change and environmental issues, in recent years, there has been increased interest in the eco-friendly production of polymers like nylon.
On August 10, Dr. Taehee Han from a KAIST research team led by Distinguished Professor Sang Yup Lee in the Department of Chemical and Biomolecular Engineering revealed the successful development of a microbial strain that produces valerolactam, a monomer of nylon-5.
Valerolactam is an important monomer that constitutes nylon-5 and nylon-6,5. Nylon is the oldest synthetic polymer, and nylon-5 is one of its derivatives composed of monomers with five carbons, while nylon-5,6 is composed of two types of monomers with either five or six carbons. They not only have excellent processability, but are also light and tough, which allows them to be applied in a wide range of industrial sectors including clothing, badminton rackets, fishing nets, tents, and gear parts. Monomers are materials that can be built into polymers, and synthetic processes are what connects them into a polymer.
The chemical production of valerolactam, however, is based on petrochemistry, where extreme reaction conditions are required and toxic waste is produced. To solve these problems, efforts are being made to develop environmentally friendly and highly efficient microbial cell factories for lactam production. Systems metabolic engineering, a key strategy for effective microbial strain development, is a research field pioneered by Professor Sang Yup Lee.
Professor Lee’s team used metabolic engineering, a technique for manipulating microbial metabolic pathways, to construct a synthetic metabolic pathway for valerolactam production in Corynebacteriam glutamicum, a bacterium commonly used for amino acid production. With this, they successfully developed a microbial strain that utilizes biomass-derived glucose as a carbon source to produce high-value valerolactam.
In 2017, the team suggested a novel method that metabolically manipulates Escherichia coli to produce valerolactam. However, there were several limitations at the time including low producibility and the generation of harmful byproducts.
< Figure 1. Schematic graphical representation of the development of microorganisms that produce valerolactam, a nylon-5 monomer >
In this research, the team improved valerolactam producibility and incorporated an additional systems metabolic strategy to the developed microbial strain while eliminating the harmful byproducts. By removing the gene involved in the production of the main byproduct and through gene screening, the team successfully converted 5-aminovaleric acid, a byproduct and a precursor, into valerolactam.
Furthermore, by employing a strategy where the 5-aminovaleric acid-converting gene is inserted multiple times into the genome, the team strengthened the metabolic flux for valerolactam production. As a result, they reached a world-record concentration of 76.1 g/L, which is 6.17 times greater than what was previously reported.
This study was published in Metabolic Engineering on July 12, under the title, “Metabolic engineering of Corynebacterium glutamicum for the high-level production of valerolactam, a nylon-5 monomer”.
Dr. Taehee Han, the first author of the paper, said, “The significance of this research lies in our development of an environmentally friendly technology that efficiently produces monomer lactam for nylon production using microorganisms.” She added, “Through this technology, we will be able to take a step forward in replacing the petrochemical industry with a microorganism-based biopolymer industry.”
This work was supported by the “Development of Next-Generation Biofinery Platform Technologies for Leading Bio-based Chemicals Industry Project” funded by the Korean Ministry of Science and ICT.
2023.08.24 View 5163 -
 KAIST presents a microbial cell factory as a source of eco-friendly food and cosmetic coloring
Despite decades of global population growth, global food crisis seems to be at hand yet again because the food productivity is cut severely due to prolonged presence of abnormal weather from intensifying climate change and global food supply chain is deteriorated due to international conflicts such as wars exacerbating food shortages and nutritional inequality around the globe. At the same time, however, as awareness of the environment and sustainability rises, an increase in demand for more eco-friendly and high-quality food and beauty products is being observed not without a sense of irony. At a time like this, microorganisms are attracting attention as a key that can handle this couple of seemingly distant problems.
KAIST (President Kwang-Hyung Lee) announced on the 26th that Kyeong Rok Choi, a research professor of the Bioprocess Research Center and Sang Yup Lee, a Distinguished Professor of the Department of Chemical and Biomolecular Engineering, published a paper titled “Metabolic Engineering of Microorganisms for Food and Cosmetics Production” upon invitation by “Nature Reviews Bioengineering” to be published online published by Nature after peer review.
※ Paper title: Systems metabolic engineering of microorganisms for food and cosmetics production
※ Author information: Kyeong Rok Choi (first author) and Sang Yup Lee (corresponding author)
Systems metabolic engineering is a research field founded by Distinguished Professor Sang Yup Lee of KAIST to more effectively develop microbial cell factories, the core factor of the next-generation bio industry to replace the existing chemical industry that relies heavily on petroleum. By applying a systemic metabolic engineering strategy, the researchers have developed a number of high-performance microbial cell factories that produce a variety of food and cosmetic compounds including natural substances like heme and zinc protoporphyrin IX compounds which can improve the flavor and color of synthetic meat, lycopene and β-carotene which are functional natural pigments that can be widely used in food and cosmetics, and methyl anthranilate, a grape-derived compound widely used to impart grape flavor in food and beverage manufacturing.
In this paper written upon invitation by Nature, the research team covered remarkable cases of microbial cell factory that can produce amino acids, proteins, fats and fatty acids, vitamins, flavors, pigments, alcohols, functional compounds and other food additives used in various foods and cosmetics and the companies that have successfully commercialized these microbial-derived materials Furthermore, the paper organized and presents systems metabolic engineering strategies that can spur the development of industrial microbial cell factories that can produce more diverse food and cosmetic compounds in an eco-friendly way with economic feasibility.
< Figure 1. Examples of production of food and cosmetic compounds using microbial cell factories >
For example, by producing proteins or amino acids with high nutritional value through non-edible biomass used as animal feed or fertilizer through the microbial fermentation process, it will contribute to the increase in production and stable supply of food around the world. Furthermore, by contributing to developing more viable alternative meat, further reducing dependence on animal protein, it can also contribute to reducing greenhouse gases and environmental pollution generated through livestock breeding or fish farming.
In addition, vanillin or methyl anthranilate, which give off vanilla or grape flavor, are widely added to various foods, but natural products isolated and refined from plants are low in production and high in production cost, so in most cases, petrochemicals substances derived from vanillin and methylanthranilic acid are added to food. These materials can also be produced through an eco-friendly and human-friendly method by borrowing the power of microorganisms.
Ethical and resource problems that arise in producing compounds like Calmin (cochineal pigment), a coloring added to various cosmetics and foods such as red lipstick and strawberry-flavored milk, which must be extracted from cochineal insects that live only in certain cacti. and Hyaluronic acid, which is widely consumed as a health supplement, but is only present in omega-3 fatty acids extracted from shark or fish livers, can also be resolved when they can be produced in an eco-friendly way using microorganisms.
KAIST Research Professor Kyeong Rok Choi, the first author of this paper, said, “In addition to traditional fermented foods such as kimchi and yogurt, foods produced with the help of microorganisms like cocoa butter, a base ingredient for chocolate that can only be obtained from fermented cacao beans, and monosodium glutamate, a seasoning produced through microbial fermentation are already familiar to us”. “In the future, we will be able to acquire a wider variety of foods and cosmetics even more easily produced in an eco-friendly and sustainable way in our daily lives through microbial cell factories.” he added.
< Figure 2. Systems metabolic engineering strategy to improve metabolic flow in microbial cell factories >
Distinguished Professor Sang Yup Lee said, “It is engineers’ mission to make the world a better place utilizing science and technology.” and added, “Continuous advancement and active use of systems metabolic engineering will contribute greatly to easing and resolving the problems arising from both the food crisis and the climate change."
This research was carried out as a part of the “Development of Protein Production Technology from Inorganic Substances through Control of Microbial Metabolism System Project” (Project Leader: Kyeong Rok Choi, KAIST Research Professor) of the the Center for Agricultural Microorganism and Enzyme (Director Pahn-Shick Chang) supported by the Rural Development Administration and the “Development of Platform Technologies of Microbial Cell Factories for the Next-generation Biorefineries Project” (Project Leader: Sang Yup Lee, KAIST Distinguished Professor) of the Petroleum-Substitute Eco-friendly Chemical Technology Development Program supported by the Ministry of Science and ICT.
2023.07.28 View 7337
KAIST presents a microbial cell factory as a source of eco-friendly food and cosmetic coloring
Despite decades of global population growth, global food crisis seems to be at hand yet again because the food productivity is cut severely due to prolonged presence of abnormal weather from intensifying climate change and global food supply chain is deteriorated due to international conflicts such as wars exacerbating food shortages and nutritional inequality around the globe. At the same time, however, as awareness of the environment and sustainability rises, an increase in demand for more eco-friendly and high-quality food and beauty products is being observed not without a sense of irony. At a time like this, microorganisms are attracting attention as a key that can handle this couple of seemingly distant problems.
KAIST (President Kwang-Hyung Lee) announced on the 26th that Kyeong Rok Choi, a research professor of the Bioprocess Research Center and Sang Yup Lee, a Distinguished Professor of the Department of Chemical and Biomolecular Engineering, published a paper titled “Metabolic Engineering of Microorganisms for Food and Cosmetics Production” upon invitation by “Nature Reviews Bioengineering” to be published online published by Nature after peer review.
※ Paper title: Systems metabolic engineering of microorganisms for food and cosmetics production
※ Author information: Kyeong Rok Choi (first author) and Sang Yup Lee (corresponding author)
Systems metabolic engineering is a research field founded by Distinguished Professor Sang Yup Lee of KAIST to more effectively develop microbial cell factories, the core factor of the next-generation bio industry to replace the existing chemical industry that relies heavily on petroleum. By applying a systemic metabolic engineering strategy, the researchers have developed a number of high-performance microbial cell factories that produce a variety of food and cosmetic compounds including natural substances like heme and zinc protoporphyrin IX compounds which can improve the flavor and color of synthetic meat, lycopene and β-carotene which are functional natural pigments that can be widely used in food and cosmetics, and methyl anthranilate, a grape-derived compound widely used to impart grape flavor in food and beverage manufacturing.
In this paper written upon invitation by Nature, the research team covered remarkable cases of microbial cell factory that can produce amino acids, proteins, fats and fatty acids, vitamins, flavors, pigments, alcohols, functional compounds and other food additives used in various foods and cosmetics and the companies that have successfully commercialized these microbial-derived materials Furthermore, the paper organized and presents systems metabolic engineering strategies that can spur the development of industrial microbial cell factories that can produce more diverse food and cosmetic compounds in an eco-friendly way with economic feasibility.
< Figure 1. Examples of production of food and cosmetic compounds using microbial cell factories >
For example, by producing proteins or amino acids with high nutritional value through non-edible biomass used as animal feed or fertilizer through the microbial fermentation process, it will contribute to the increase in production and stable supply of food around the world. Furthermore, by contributing to developing more viable alternative meat, further reducing dependence on animal protein, it can also contribute to reducing greenhouse gases and environmental pollution generated through livestock breeding or fish farming.
In addition, vanillin or methyl anthranilate, which give off vanilla or grape flavor, are widely added to various foods, but natural products isolated and refined from plants are low in production and high in production cost, so in most cases, petrochemicals substances derived from vanillin and methylanthranilic acid are added to food. These materials can also be produced through an eco-friendly and human-friendly method by borrowing the power of microorganisms.
Ethical and resource problems that arise in producing compounds like Calmin (cochineal pigment), a coloring added to various cosmetics and foods such as red lipstick and strawberry-flavored milk, which must be extracted from cochineal insects that live only in certain cacti. and Hyaluronic acid, which is widely consumed as a health supplement, but is only present in omega-3 fatty acids extracted from shark or fish livers, can also be resolved when they can be produced in an eco-friendly way using microorganisms.
KAIST Research Professor Kyeong Rok Choi, the first author of this paper, said, “In addition to traditional fermented foods such as kimchi and yogurt, foods produced with the help of microorganisms like cocoa butter, a base ingredient for chocolate that can only be obtained from fermented cacao beans, and monosodium glutamate, a seasoning produced through microbial fermentation are already familiar to us”. “In the future, we will be able to acquire a wider variety of foods and cosmetics even more easily produced in an eco-friendly and sustainable way in our daily lives through microbial cell factories.” he added.
< Figure 2. Systems metabolic engineering strategy to improve metabolic flow in microbial cell factories >
Distinguished Professor Sang Yup Lee said, “It is engineers’ mission to make the world a better place utilizing science and technology.” and added, “Continuous advancement and active use of systems metabolic engineering will contribute greatly to easing and resolving the problems arising from both the food crisis and the climate change."
This research was carried out as a part of the “Development of Protein Production Technology from Inorganic Substances through Control of Microbial Metabolism System Project” (Project Leader: Kyeong Rok Choi, KAIST Research Professor) of the the Center for Agricultural Microorganism and Enzyme (Director Pahn-Shick Chang) supported by the Rural Development Administration and the “Development of Platform Technologies of Microbial Cell Factories for the Next-generation Biorefineries Project” (Project Leader: Sang Yup Lee, KAIST Distinguished Professor) of the Petroleum-Substitute Eco-friendly Chemical Technology Development Program supported by the Ministry of Science and ICT.
2023.07.28 View 7337 -
 2023 Global Entrepreneurship Summer School in Silicon Valley Successfully Concluded
< 2023 Silicon Valley Global Entrepreneurship Summer School Participants >
The 2023 KAIST Global Entrepreneurship Summer School (GESS) was successfully held. Co-hosted by the Center for Global Strategies and Planning (GSP) (Director Man-Sung Yim) and the Startup KAIST (Director Hyeonmin Bae), the 2023 KAIST GESS was the second one of the summer programs, repeating the Silicon Valley global entrepreneurship bootcamp of 2022 (2022 GESC), based on industry-academia collaboration. This program was designed to provide students with the opportunity to visit Silicon Valley, the global hub of entrepreneurship, and personally experience the Silicon Valley culture while developing human networks that would serve as a foundation for their overseas startup development.
A total of 20 participants were selected earlier this year, including potential KAISTian entrepreneurs and early-stage entrepreneurs from KAIST within one year of incorporation. In particular, a number of foreign students of various nationalities such as Vietnam, Azerbaijan, Honduras, Indonesia, Philippines, and Kazakhstan, increased significantly, demonstrating the enthusiasm for entrepreneurship across national boundaries along with the program's growing international status. This year's event was also open to 20 Impact MBA and Social Entrepreneur (SE) students from KAIST's College of Business for the Silicon Valley program.
For the past two months, the participants have trained on business model development and pitching at KAIST's main campus in Daejeon. From June 21st to the 30th, they visited the campuses of leading universities, such as, Stanford University, UC Santa Cruz, and UC Berkeley, as well as KOTRA Silicon Valley Trade Center (Manager Hyoung il Kim), and local alumni companies and Apple company to experience the global technology startups. The start-ups by KAIST alums including B Garage (CEO Aiden Kim), ImpriMed (CEO Sungwon Lim), Medic Life Sciences (CEO Kyuho Han), and VESSL AI (CEO Jaeman Ahn) participated in the program and gave lectures and company tours to inspire the participants to have passion to take on the entrepreneurial endeavors and challenges.
On the last day, the participants gave presentations on their team’s business items in front of local venture capitalists in Silicon Valley. After receiving continuous coaching from Silicon Valley's professional accelerators through remote video conferencing and face-to-face mentoring for the last two months, the participants developed their business models and presented their creative and innovative ideas, revealing their potential as future global entrepreneurs. At the final competition, Team Sparky that developed “Snoove” won the first prize. Snoove is a scientifically-proven mattress accessory that applies mild vibration to the mattress to aid users in achieving better sleep, a method previously used to soothe infants.
< GESS Pitching Day Presentation >
Kevin Choi from the Team Sparky said, "Seeing and experiencing the realities of entrepreneurship in Silicon Valley, a global startup scene, made me think about the importance of unlearning, challenging, and failing to be a global entrepreneur who contributes to our society." Man-Sung Yim, the Associate Vice President of the International Office, who organized the event added, "Through this experience, we expect KAIST students to grow to become global leaders who would create global values and enhance the international reputation of our university."
Meanwhile, the GSP and Startup KAIST commented that they will to continue to develop the KAIST GESS program to foster prospective entrepreneurs who can compete in the global market based on the success of this program.
2023.07.05 View 9466
2023 Global Entrepreneurship Summer School in Silicon Valley Successfully Concluded
< 2023 Silicon Valley Global Entrepreneurship Summer School Participants >
The 2023 KAIST Global Entrepreneurship Summer School (GESS) was successfully held. Co-hosted by the Center for Global Strategies and Planning (GSP) (Director Man-Sung Yim) and the Startup KAIST (Director Hyeonmin Bae), the 2023 KAIST GESS was the second one of the summer programs, repeating the Silicon Valley global entrepreneurship bootcamp of 2022 (2022 GESC), based on industry-academia collaboration. This program was designed to provide students with the opportunity to visit Silicon Valley, the global hub of entrepreneurship, and personally experience the Silicon Valley culture while developing human networks that would serve as a foundation for their overseas startup development.
A total of 20 participants were selected earlier this year, including potential KAISTian entrepreneurs and early-stage entrepreneurs from KAIST within one year of incorporation. In particular, a number of foreign students of various nationalities such as Vietnam, Azerbaijan, Honduras, Indonesia, Philippines, and Kazakhstan, increased significantly, demonstrating the enthusiasm for entrepreneurship across national boundaries along with the program's growing international status. This year's event was also open to 20 Impact MBA and Social Entrepreneur (SE) students from KAIST's College of Business for the Silicon Valley program.
For the past two months, the participants have trained on business model development and pitching at KAIST's main campus in Daejeon. From June 21st to the 30th, they visited the campuses of leading universities, such as, Stanford University, UC Santa Cruz, and UC Berkeley, as well as KOTRA Silicon Valley Trade Center (Manager Hyoung il Kim), and local alumni companies and Apple company to experience the global technology startups. The start-ups by KAIST alums including B Garage (CEO Aiden Kim), ImpriMed (CEO Sungwon Lim), Medic Life Sciences (CEO Kyuho Han), and VESSL AI (CEO Jaeman Ahn) participated in the program and gave lectures and company tours to inspire the participants to have passion to take on the entrepreneurial endeavors and challenges.
On the last day, the participants gave presentations on their team’s business items in front of local venture capitalists in Silicon Valley. After receiving continuous coaching from Silicon Valley's professional accelerators through remote video conferencing and face-to-face mentoring for the last two months, the participants developed their business models and presented their creative and innovative ideas, revealing their potential as future global entrepreneurs. At the final competition, Team Sparky that developed “Snoove” won the first prize. Snoove is a scientifically-proven mattress accessory that applies mild vibration to the mattress to aid users in achieving better sleep, a method previously used to soothe infants.
< GESS Pitching Day Presentation >
Kevin Choi from the Team Sparky said, "Seeing and experiencing the realities of entrepreneurship in Silicon Valley, a global startup scene, made me think about the importance of unlearning, challenging, and failing to be a global entrepreneur who contributes to our society." Man-Sung Yim, the Associate Vice President of the International Office, who organized the event added, "Through this experience, we expect KAIST students to grow to become global leaders who would create global values and enhance the international reputation of our university."
Meanwhile, the GSP and Startup KAIST commented that they will to continue to develop the KAIST GESS program to foster prospective entrepreneurs who can compete in the global market based on the success of this program.
2023.07.05 View 9466 -
 Synthetic sRNAs to knockdown genes in medical and industrial bacteria
Bacteria are intimately involved in our daily lives. These microorganisms have been used in human history for food such as cheese, yogurt, and wine, In more recent years, through metabolic engineering, microorganisms been used extensively as microbial cell factories to manufacture plastics, feed for livestock, dietary supplements, and drugs. However, in addition to these bacteria that are beneficial to human lives, pathogens such as Pneumonia, Salmonella, and Staphylococcus that cause various infectious diseases are also ubiquitously present. It is important to be able to metabolically control these beneficial industrial bacteria for high value-added chemicals production and to manipulate harmful pathogens to suppress its pathogenic traits.
KAIST (President Kwang Hyung Lee) announced on the 10th that a research team led by Distinguished Professor Sang Yup Lee of the Department of Biochemical Engineering has developed a new sRNA tool that can effectively inhibit target genes in various bacteria, including both Gram-negative and Gram-positive bacteria. The research results were published online on April 24 in Nature Communications.
※ Thesis title: Targeted and high-throughput gene knockdown in diverse bacteria using synthetic sRNAs
※ Author information : Jae Sung Cho (co-1st), Dongsoo Yang (co-1st), Cindy Pricilia Surya Prabowo (co-author), Mohammad Rifqi Ghiffary (co-author), Taehee Han (co-author), Kyeong Rok Choi (co-author), Cheon Woo Moon (co-author), Hengrui Zhou (co-author), Jae Yong Ryu (co-author), Hyun Uk Kim (co-author) and Sang Yup Lee (corresponding author).
sRNA is an effective tool for synthesizing and regulating target genes in E. coli, but it has been difficult to apply to industrially useful Gram-positive bacteria such as Bacillus subtilis and Corynebacterium in addition to Gram-negative bacteria such as E. coli.
To address this issue, a research team led by Distinguished Professor Lee Sang Yup Lee of the Department of Chemical and Biomolecular Engineering at KAIST developed a new sRNA platform that can effectively suppress target genes in various bacteria, including both Gram-negative and positive bacteria. The research team surveyed thousands of microbial-derived sRNA systems in the microbial database, and eventually designated the sRNA system derived from 'Bacillus subtilis' that showed the highest gene knockdown efficiency, and designated it as “Broad-Host-Range sRNA”, or BHR-sRNA.
A similar well-known system is the CRISPR interference (CRISPRi) system, which is a modified CRISPR system that knocks down gene expression by suppressing the gene transcription process. However, the Cas9 protein in the CRISPRi system has a very high molecular weight, and there have been reports growth inhibition in bacteria. The BHR-sRNA system developed in this study did not affect bacterial growth while showing similar gene knockdown efficiencies to CRISPRi.
< Figure 1. a) Schematic illustration demonstrating the mechanism of syntetic sRNA b) Phylogenetic tree of the 16 Gram-negative and Gram-positive bacterial species tested for gene knockdown by the BHR-sRNA system. >
To validate the versatility of the BHR-sRNA system, 16 different gram-negative and gram-positive bacteria were selected and tested, where the BHR-sRNA system worked successfully in 15 of them. In addition, it was demonstrated that the gene knockdown capability was more effective than that of the existing E. coli-based sRNA system in 10 bacteria. The BHR-sRNA system proved to be a universal tool capable of effectively inhibiting gene expression in various bacteria.
In order to address the problem of antibiotic-resistant pathogens that have recently become more serious, the BHR-sRNA was demonstrated to suppress the pathogenicity by suppressing the gene producing the virulence factor. By using BHR-sRNA, biofilm formation, one of the factors resulting in antibiotic resistance, was inhibited by 73% in Staphylococcus epidermidis a pathogen that can cause hospital-acquired infections. Antibiotic resistance was also weakened by 58% in the pneumonia causing bacteria Klebsiella pneumoniae. In addition, BHR-sRNA was applied to industrial bacteria to develop microbial cell factories to produce high value-added chemicals with better production performance. Notably, superior industrial strains were constructed with the aid of BHR-sRNA to produce the following chemicals: valerolactam, a raw material for polyamide polymers, methyl-anthranilate, a grape-flavor food additive, and indigoidine, a blue-toned natural dye.
The BHR-sRNA developed through this study will help expedite the commercialization of bioprocesses to produce high value-added compounds and materials such as artificial meat, jet fuel, health supplements, pharmaceuticals, and plastics. It is also anticipated that to help eradicating antibiotic-resistant pathogens in preparation for another upcoming pandemic. “In the past, we could only develop new tools for gene knockdown for each bacterium, but now we have developed a tool that works for a variety of bacteria” said Distinguished Professor Sang Yup Lee.
This work was supported by the Development of Next-generation Biorefinery Platform Technologies for Leading Bio-based Chemicals Industry Project and the Development of Platform Technologies of Microbial Cell Factories for the Next-generation Biorefineries Project from NRF supported by the Korean MSIT.
2023.05.10 View 6642
Synthetic sRNAs to knockdown genes in medical and industrial bacteria
Bacteria are intimately involved in our daily lives. These microorganisms have been used in human history for food such as cheese, yogurt, and wine, In more recent years, through metabolic engineering, microorganisms been used extensively as microbial cell factories to manufacture plastics, feed for livestock, dietary supplements, and drugs. However, in addition to these bacteria that are beneficial to human lives, pathogens such as Pneumonia, Salmonella, and Staphylococcus that cause various infectious diseases are also ubiquitously present. It is important to be able to metabolically control these beneficial industrial bacteria for high value-added chemicals production and to manipulate harmful pathogens to suppress its pathogenic traits.
KAIST (President Kwang Hyung Lee) announced on the 10th that a research team led by Distinguished Professor Sang Yup Lee of the Department of Biochemical Engineering has developed a new sRNA tool that can effectively inhibit target genes in various bacteria, including both Gram-negative and Gram-positive bacteria. The research results were published online on April 24 in Nature Communications.
※ Thesis title: Targeted and high-throughput gene knockdown in diverse bacteria using synthetic sRNAs
※ Author information : Jae Sung Cho (co-1st), Dongsoo Yang (co-1st), Cindy Pricilia Surya Prabowo (co-author), Mohammad Rifqi Ghiffary (co-author), Taehee Han (co-author), Kyeong Rok Choi (co-author), Cheon Woo Moon (co-author), Hengrui Zhou (co-author), Jae Yong Ryu (co-author), Hyun Uk Kim (co-author) and Sang Yup Lee (corresponding author).
sRNA is an effective tool for synthesizing and regulating target genes in E. coli, but it has been difficult to apply to industrially useful Gram-positive bacteria such as Bacillus subtilis and Corynebacterium in addition to Gram-negative bacteria such as E. coli.
To address this issue, a research team led by Distinguished Professor Lee Sang Yup Lee of the Department of Chemical and Biomolecular Engineering at KAIST developed a new sRNA platform that can effectively suppress target genes in various bacteria, including both Gram-negative and positive bacteria. The research team surveyed thousands of microbial-derived sRNA systems in the microbial database, and eventually designated the sRNA system derived from 'Bacillus subtilis' that showed the highest gene knockdown efficiency, and designated it as “Broad-Host-Range sRNA”, or BHR-sRNA.
A similar well-known system is the CRISPR interference (CRISPRi) system, which is a modified CRISPR system that knocks down gene expression by suppressing the gene transcription process. However, the Cas9 protein in the CRISPRi system has a very high molecular weight, and there have been reports growth inhibition in bacteria. The BHR-sRNA system developed in this study did not affect bacterial growth while showing similar gene knockdown efficiencies to CRISPRi.
< Figure 1. a) Schematic illustration demonstrating the mechanism of syntetic sRNA b) Phylogenetic tree of the 16 Gram-negative and Gram-positive bacterial species tested for gene knockdown by the BHR-sRNA system. >
To validate the versatility of the BHR-sRNA system, 16 different gram-negative and gram-positive bacteria were selected and tested, where the BHR-sRNA system worked successfully in 15 of them. In addition, it was demonstrated that the gene knockdown capability was more effective than that of the existing E. coli-based sRNA system in 10 bacteria. The BHR-sRNA system proved to be a universal tool capable of effectively inhibiting gene expression in various bacteria.
In order to address the problem of antibiotic-resistant pathogens that have recently become more serious, the BHR-sRNA was demonstrated to suppress the pathogenicity by suppressing the gene producing the virulence factor. By using BHR-sRNA, biofilm formation, one of the factors resulting in antibiotic resistance, was inhibited by 73% in Staphylococcus epidermidis a pathogen that can cause hospital-acquired infections. Antibiotic resistance was also weakened by 58% in the pneumonia causing bacteria Klebsiella pneumoniae. In addition, BHR-sRNA was applied to industrial bacteria to develop microbial cell factories to produce high value-added chemicals with better production performance. Notably, superior industrial strains were constructed with the aid of BHR-sRNA to produce the following chemicals: valerolactam, a raw material for polyamide polymers, methyl-anthranilate, a grape-flavor food additive, and indigoidine, a blue-toned natural dye.
The BHR-sRNA developed through this study will help expedite the commercialization of bioprocesses to produce high value-added compounds and materials such as artificial meat, jet fuel, health supplements, pharmaceuticals, and plastics. It is also anticipated that to help eradicating antibiotic-resistant pathogens in preparation for another upcoming pandemic. “In the past, we could only develop new tools for gene knockdown for each bacterium, but now we have developed a tool that works for a variety of bacteria” said Distinguished Professor Sang Yup Lee.
This work was supported by the Development of Next-generation Biorefinery Platform Technologies for Leading Bio-based Chemicals Industry Project and the Development of Platform Technologies of Microbial Cell Factories for the Next-generation Biorefineries Project from NRF supported by the Korean MSIT.
2023.05.10 View 6642 -
 KAIST leads AI-based analysis on drug-drug interactions involving Paxlovid
KAIST (President Kwang Hyung Lee) announced on the 16th that an advanced AI-based drug interaction prediction technology developed by the Distinguished Professor Sang Yup Lee's research team in the Department of Biochemical Engineering that analyzed the interaction between the PaxlovidTM ingredients that are used as COVID-19 treatment and other prescription drugs was published as a thesis. This paper was published in the online edition of 「Proceedings of the National Academy of Sciences of America」 (PNAS), an internationally renowned academic journal, on the 13th of March.
* Thesis Title: Computational prediction of interactions between Paxlovid and prescription drugs (Authored by Yeji Kim (KAIST, co-first author), Jae Yong Ryu (Duksung Women's University, co-first author), Hyun Uk Kim (KAIST, co-first author), and Sang Yup Lee (KAIST, corresponding author))
In this study, the research team developed DeepDDI2, an advanced version of DeepDDI, an AI-based drug interaction prediction model they developed in 2018. DeepDDI2 is able to compute for and process a total of 113 drug-drug interaction (DDI) types, more than the 86 DDI types covered by the existing DeepDDI.
The research team used DeepDDI2 to predict possible interactions between the ingredients (ritonavir, nirmatrelvir) of Paxlovid*, a COVID-19 treatment, and other prescription drugs. The research team said that while among COVID-19 patients, high-risk patients with chronic diseases such as high blood pressure and diabetes are likely to be taking other drugs, drug-drug interactions and adverse drug reactions for Paxlovid have not been sufficiently analyzed, yet. This study was pursued in light of seeing how continued usage of the drug may lead to serious and unwanted complications.
* Paxlovid: Paxlovid is a COVID-19 treatment developed by Pfizer, an American pharmaceutical company, and received emergency use approval (EUA) from the US Food and Drug Administration (FDA) in December 2021.
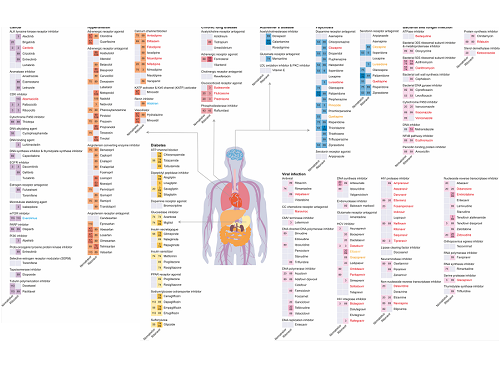
The research team used DeepDDI2 to predict how Paxrovid's components, ritonavir and nirmatrelvir, would interact with 2,248 prescription drugs. As a result of the prediction, ritonavir was predicted to interact with 1,403 prescription drugs and nirmatrelvir with 673 drugs.
Using the prediction results, the research team proposed alternative drugs with the same mechanism but low drug interaction potential for prescription drugs with high adverse drug events (ADEs). Accordingly, 124 alternative drugs that could reduce the possible adverse DDI with ritonavir and 239 alternative drugs for nirmatrelvir were identified.
Through this research achievement, it became possible to use an deep learning technology to accurately predict drug-drug interactions (DDIs), and this is expected to play an important role in the digital healthcare, precision medicine and pharmaceutical industries by providing useful information in the process of developing new drugs and making prescriptions.
Distinguished Professor Sang Yup Lee said, "The results of this study are meaningful at times like when we would have to resort to using drugs that are developed in a hurry in the face of an urgent situations like the COVID-19 pandemic, that it is now possible to identify and take necessary actions against adverse drug reactions caused by drug-drug interactions very quickly.”
This research was carried out with the support of the KAIST New-Deal Project for COVID-19 Science and Technology and the Bio·Medical Technology Development Project supported by the Ministry of Science and ICT.
Figure 1. Results of drug interaction prediction between Paxlovid ingredients and representative approved drugs using DeepDDI2
2023.03.16 View 7343
KAIST leads AI-based analysis on drug-drug interactions involving Paxlovid
KAIST (President Kwang Hyung Lee) announced on the 16th that an advanced AI-based drug interaction prediction technology developed by the Distinguished Professor Sang Yup Lee's research team in the Department of Biochemical Engineering that analyzed the interaction between the PaxlovidTM ingredients that are used as COVID-19 treatment and other prescription drugs was published as a thesis. This paper was published in the online edition of 「Proceedings of the National Academy of Sciences of America」 (PNAS), an internationally renowned academic journal, on the 13th of March.
* Thesis Title: Computational prediction of interactions between Paxlovid and prescription drugs (Authored by Yeji Kim (KAIST, co-first author), Jae Yong Ryu (Duksung Women's University, co-first author), Hyun Uk Kim (KAIST, co-first author), and Sang Yup Lee (KAIST, corresponding author))
In this study, the research team developed DeepDDI2, an advanced version of DeepDDI, an AI-based drug interaction prediction model they developed in 2018. DeepDDI2 is able to compute for and process a total of 113 drug-drug interaction (DDI) types, more than the 86 DDI types covered by the existing DeepDDI.
The research team used DeepDDI2 to predict possible interactions between the ingredients (ritonavir, nirmatrelvir) of Paxlovid*, a COVID-19 treatment, and other prescription drugs. The research team said that while among COVID-19 patients, high-risk patients with chronic diseases such as high blood pressure and diabetes are likely to be taking other drugs, drug-drug interactions and adverse drug reactions for Paxlovid have not been sufficiently analyzed, yet. This study was pursued in light of seeing how continued usage of the drug may lead to serious and unwanted complications.
* Paxlovid: Paxlovid is a COVID-19 treatment developed by Pfizer, an American pharmaceutical company, and received emergency use approval (EUA) from the US Food and Drug Administration (FDA) in December 2021.
The research team used DeepDDI2 to predict how Paxrovid's components, ritonavir and nirmatrelvir, would interact with 2,248 prescription drugs. As a result of the prediction, ritonavir was predicted to interact with 1,403 prescription drugs and nirmatrelvir with 673 drugs.
Using the prediction results, the research team proposed alternative drugs with the same mechanism but low drug interaction potential for prescription drugs with high adverse drug events (ADEs). Accordingly, 124 alternative drugs that could reduce the possible adverse DDI with ritonavir and 239 alternative drugs for nirmatrelvir were identified.
Through this research achievement, it became possible to use an deep learning technology to accurately predict drug-drug interactions (DDIs), and this is expected to play an important role in the digital healthcare, precision medicine and pharmaceutical industries by providing useful information in the process of developing new drugs and making prescriptions.
Distinguished Professor Sang Yup Lee said, "The results of this study are meaningful at times like when we would have to resort to using drugs that are developed in a hurry in the face of an urgent situations like the COVID-19 pandemic, that it is now possible to identify and take necessary actions against adverse drug reactions caused by drug-drug interactions very quickly.”
This research was carried out with the support of the KAIST New-Deal Project for COVID-19 Science and Technology and the Bio·Medical Technology Development Project supported by the Ministry of Science and ICT.
Figure 1. Results of drug interaction prediction between Paxlovid ingredients and representative approved drugs using DeepDDI2
2023.03.16 View 7343 -
 KAIST team develops smart immune system that can pin down on malignant tumors
A joint research team led by Professor Jung Kyoon Choi of the KAIST Department of Bio and Brain Engineering and Professor Jong-Eun Park of the KAIST Graduate School of Medical Science and Engineering (GSMSE) announced the development of the key technologies to treat cancers using smart immune cells designed based on AI and big data analysis. This technology is expected to be a next-generation immunotherapy that allows precision targeting of tumor cells by having the chimeric antigen receptors (CARs) operate through a logical circuit. Professor Hee Jung An of CHA Bundang Medical Center and Professor Hae-Ock Lee of the Catholic University of Korea also participated in this research to contribute joint effort.
Professor Jung Kyoon Choi’s team built a gene expression database from millions of cells, and used this to successfully develop and verify a deep-learning algorithm that could detect the differences in gene expression patterns between tumor cells and normal cells through a logical circuit. CAR immune cells that were fitted with the logic circuits discovered through this methodology could distinguish between tumorous and normal cells as a computer would, and therefore showed potentials to strike only on tumor cells accurately without causing unwanted side effects.
This research, conducted by co-first authors Dr. Joonha Kwon of the KAIST Department of Bio and Brain Engineering and Ph.D. candidate Junho Kang of KAIST GSMSE, was published by Nature Biotechnology on February 16, under the title Single-cell mapping of combinatorial target antigens for CAR switches using logic gates.
An area in cancer research where the most attempts and advances have been made in recent years is immunotherapy. This field of treatment, which utilizes the patient’s own immune system in order to overcome cancer, has several methods including immune checkpoint inhibitors, cancer vaccines and cellular treatments. Immune cells like CAR-T or CAR-NK equipped with chimera antigen receptors, in particular, can recognize cancer antigens and directly destroy cancer cells.
Starting with its success in blood cancer treatment, scientists have been trying to expand the application of CAR cell therapy to treat solid cancer. But there have been difficulties to develop CAR cells with effective killing abilities against solid cancer cells with minimized side effects. Accordingly, in recent years, the development of smarter CAR engineering technologies, i.e., computational logic gates such as AND, OR, and NOT, to effectively target cancer cells has been underway.
At this point in time, the research team built a large-scale database for cancer and normal cells to discover the exact genes that are expressed only from cancer cells at a single-cell level. The team followed this up by developing an AI algorithm that could search for a combination of genes that best distinguishes cancer cells from normal cells. This algorithm, in particular, has been used to find a logic circuit that can specifically target cancer cells through cell-level simulations of all gene combinations. CAR-T cells equipped with logic circuits discovered through this methodology are expected to distinguish cancerous cells from normal cells like computers, thereby minimizing side effects and maximizing the effects of chemotherapy.
Dr. Joonha Kwon, who is the first author of this paper, said, “this research suggests a new method that hasn’t been tried before. What’s particularly noteworthy is the process in which we found the optimal CAR cell circuit through simulations of millions of individual tumors and normal cells.” He added, “This is an innovative technology that can apply AI and computer logic circuits to immune cell engineering. It would contribute greatly to expanding CAR therapy, which is being successfully used for blood cancer, to solid cancers as well.”
This research was funded by the Original Technology Development Project and Research Program for Next Generation Applied Omic of the Korea Research Foundation.
Figure 1. A schematic diagram of manufacturing and administration process of CAR therapy and of cancer cell-specific dual targeting using CAR.
Figure 2. Deep learning (convolutional neural networks, CNNs) algorithm for selection of dual targets based on gene combination (left) and algorithm for calculating expressing cell fractions by gene combination according to logical circuit (right).
2023.03.09 View 8622
KAIST team develops smart immune system that can pin down on malignant tumors
A joint research team led by Professor Jung Kyoon Choi of the KAIST Department of Bio and Brain Engineering and Professor Jong-Eun Park of the KAIST Graduate School of Medical Science and Engineering (GSMSE) announced the development of the key technologies to treat cancers using smart immune cells designed based on AI and big data analysis. This technology is expected to be a next-generation immunotherapy that allows precision targeting of tumor cells by having the chimeric antigen receptors (CARs) operate through a logical circuit. Professor Hee Jung An of CHA Bundang Medical Center and Professor Hae-Ock Lee of the Catholic University of Korea also participated in this research to contribute joint effort.
Professor Jung Kyoon Choi’s team built a gene expression database from millions of cells, and used this to successfully develop and verify a deep-learning algorithm that could detect the differences in gene expression patterns between tumor cells and normal cells through a logical circuit. CAR immune cells that were fitted with the logic circuits discovered through this methodology could distinguish between tumorous and normal cells as a computer would, and therefore showed potentials to strike only on tumor cells accurately without causing unwanted side effects.
This research, conducted by co-first authors Dr. Joonha Kwon of the KAIST Department of Bio and Brain Engineering and Ph.D. candidate Junho Kang of KAIST GSMSE, was published by Nature Biotechnology on February 16, under the title Single-cell mapping of combinatorial target antigens for CAR switches using logic gates.
An area in cancer research where the most attempts and advances have been made in recent years is immunotherapy. This field of treatment, which utilizes the patient’s own immune system in order to overcome cancer, has several methods including immune checkpoint inhibitors, cancer vaccines and cellular treatments. Immune cells like CAR-T or CAR-NK equipped with chimera antigen receptors, in particular, can recognize cancer antigens and directly destroy cancer cells.
Starting with its success in blood cancer treatment, scientists have been trying to expand the application of CAR cell therapy to treat solid cancer. But there have been difficulties to develop CAR cells with effective killing abilities against solid cancer cells with minimized side effects. Accordingly, in recent years, the development of smarter CAR engineering technologies, i.e., computational logic gates such as AND, OR, and NOT, to effectively target cancer cells has been underway.
At this point in time, the research team built a large-scale database for cancer and normal cells to discover the exact genes that are expressed only from cancer cells at a single-cell level. The team followed this up by developing an AI algorithm that could search for a combination of genes that best distinguishes cancer cells from normal cells. This algorithm, in particular, has been used to find a logic circuit that can specifically target cancer cells through cell-level simulations of all gene combinations. CAR-T cells equipped with logic circuits discovered through this methodology are expected to distinguish cancerous cells from normal cells like computers, thereby minimizing side effects and maximizing the effects of chemotherapy.
Dr. Joonha Kwon, who is the first author of this paper, said, “this research suggests a new method that hasn’t been tried before. What’s particularly noteworthy is the process in which we found the optimal CAR cell circuit through simulations of millions of individual tumors and normal cells.” He added, “This is an innovative technology that can apply AI and computer logic circuits to immune cell engineering. It would contribute greatly to expanding CAR therapy, which is being successfully used for blood cancer, to solid cancers as well.”
This research was funded by the Original Technology Development Project and Research Program for Next Generation Applied Omic of the Korea Research Foundation.
Figure 1. A schematic diagram of manufacturing and administration process of CAR therapy and of cancer cell-specific dual targeting using CAR.
Figure 2. Deep learning (convolutional neural networks, CNNs) algorithm for selection of dual targets based on gene combination (left) and algorithm for calculating expressing cell fractions by gene combination according to logical circuit (right).
2023.03.09 View 8622 -
 KAIST presents a fundamental technology to remove metastatic traits from lung cancer cells
KAIST (President Kwang Hyung Lee) announced on January 30th that a research team led by Professor Kwang-Hyun Cho from the Department of Bio and Brain Engineering succeeded in using systems biology research to change the properties of carcinogenic cells in the lungs and eliminate both drug resistance and their ability to proliferate out to other areas of the body.
As the incidences of cancer increase within aging populations, cancer has become the most lethal disease threatening healthy life. Fatality rates are especially high when early detection does not happen in time and metastasis has occurred in various organs. In order to resolve this problem, a series of attempts were made to remove or lower the ability of cancer cells to spread, but they resulted in cancer cells in the intermediate state becoming more unstable and even more malignant, which created serious treatment challenges.
Professor Kwang-Hyun Cho's research team simulated various cancer cell states in the Epithelial-to-Mesenchymal Transition (EMT) of lung cancer cells, between epithelial cells without metastatic ability and mesenchymal cells with metastatic ability. A mathematical model of molecular network was established, and key regulators that could reverse the state of invasive and drug resistant mesenchymal cells back to the epithelial state were discovered through computer simulation analysis and molecular cell experiments. In particular, this process succeeded in properly reverting the mesenchymal lung cancer cells to a state where they were sensitive to chemotherapy treatment while avoiding the unstable EMT hybrid cell state in the middle process, which had remained a difficult problem.
The results of this research, in which KAIST Ph.D. student Namhee Kim, Dr. Chae Young Hwang, Researcher Taeyoung Kim, and Ph.D. student Hyunjin Kim participated, were published as an online paper in the international journal “Cancer Research” published by the American Association for Cancer Research (AACR) on January 30th. (Paper title: A cell fate reprogramming strategy reverses epithelial-to-mesenchymal transition of lung cancer cells while avoiding hybrid states)
Cells in an EMT hybrid state, which are caused by incomplete transitions during the EMT process in cancer cells, have the characteristics of both epithelial cells and mesenchymal cells, and are known to have high drug resistance and metastatic potential by acquiring high stem cell capacity. In particular, EMT is further enhanced through factors such as transforming growth factor-beta (TGF-β) secreted from the tumor microenvironment (TME) and, as a result, various cell states with high plasticity appear. Due to the complexity of EMT, it has been very difficult to completely reverse the transitional process of the mesenchymal cancer cells to an epithelial cell state in which metastatic ability and drug resistance are eliminated while avoiding the EMT hybrid cell state with high metastatic ability and drug resistance.
Professor Kwang-Hyun Cho's research team established a mathematical model of the gene regulation network that governs the complex process of EMT, and then applied large-scale computer simulation analysis and complex system network control technology to identify and verify 'p53', 'SMAD4', and 'ERK1' and 'ERK 2' (collectively ERKs) through molecular cell experiments as the three key molecular targets that can transform lung cancer cells in the mesenchymal cell state, reversed back to an epithelial cell state that no longer demonstrates the ability to metastasize, while avoiding the EMT hybrid cell state.
In particular, by analyzing the molecular regulatory mechanism of the complex EMT process at the system level, the key pathways were identified that were linked to the positive feedback that plays an important role in completely returning cancer cells to an epithelial cell state in which metastatic ability and drug resistance are removed.
This discovery is significant in that it proved that mesenchymal cells can be reverted to the state of epithelial cells under conditions where TGF-β stimulation are present, like they are in the actual environment where cancer tissue forms in the human body.
Abnormal EMT in cancer cells leads to various malignant traits such as the migration and invasion of cancer cells, changes in responsiveness to chemotherapy treatment, enhanced stem cell function, and the dissemination of cancer. In particular, the acquisition of the metastatic ability of cancer cells is a key determinant factor for the prognosis of cancer patients. The EMT reversal technology in lung cancer cells developed in this research is a new anti-cancer treatment strategy that reprograms cancer cells to eliminate their high plasticity and metastatic potential and increase their responsiveness to chemotherapy.
Professor Kwang-Hyun Cho said, "By succeeding in reversing the state of lung cancer cells that acquired high metastatic traits and resistance to drugs and reverting them to a treatable epithelial cell state with renewed sensitivity to chemotherapy, the research findings propose a new strategy for treatments that can improve the prognosis of cancer patients.”
Professor Kwang-Hyun Cho's research team was the first to present the principle of reversal treatment to revert cancer cells to normal cells, following through with the announcement of the results of their study that reverted colon cancer cells to normal colon cells in January of 2020, and also presenting successful re-programming research where the most malignant basal type breast cancer cells turned into less-malignant luminal type breast cancer cells that were treatable with hormonal therapies in January of 2022. This latest research result is the third in the development of reversal technology where lung cancer cells that had acquired metastatic traits returned to a state in which their metastatic ability was removed and drug sensitivity was enhanced.
This research was carried out with support from the Ministry of Science and ICT and the National Research Foundation of Korea's Basic Research in Science & Engineering Program for Mid-Career Researchers.
< Figure 1. Construction of the mathematical model of the regulatory network to represent the EMT phenotype based on the interaction between various molecules related to EMT.
(A) Professor Kwang-Hyun Cho's research team investigated numerous literatures and databases related to complex EMT, and based on comparative analysis of cell line data showing epithelial and mesenchymal cell conditions, they extracted key signaling pathways related to EMT and built a mathematical model of regulatory network (B) By comparing the results of computer simulation analysis and the molecular cell experiments, it was verified how well the constructed mathematical model simulated the actual cellular phenomena. >
< Figure 2. Understanding of various EMT phenotypes through large-scale computer simulation analysis and complex system network control technology.
(A) Through computer simulation analysis and experiments, Professor Kwang-Hyun Cho's research team found that complete control of EMT is impossible with single-molecule control alone. In particular, through comparison of the relative stability of attractors, it was revealed that the cell state exhibiting EMT hybrid characteristics has unstable properties. (B), (C) Based on these results, Prof. Cho’s team identified two feedbacks (positive feedback consisting of Snail-miR-34 and ZEB1-miR-200) that play an important role in avoiding the EMT hybrid state that appeared in the TGF-β-ON state. It was found through computer simulation analysis that the two feedbacks restore relatively high stability when the excavated p53 and SMAD4 are regulated. In addition, molecular cell experiments demonstrated that the expression levels of E-cad and ZEB1, which are representative phenotypic markers of EMT, changed similarly to the expression profile in the epithelial cell state, despite the TGF-β-ON state. >
< Figure 3. Complex molecular network analysis and discovery of reprogramming molecular targets for intact elimination of EMT hybrid features.
(A) Controlling the expression of p53 and SMAD4 in lung cancer cell lines was expected to overcome drug resistance, but contrary to expectations, chemotherapy responsiveness was not restored. (B) Professor Kwang-Hyun Cho's research team additionally analyzed computer simulations, genome data, and experimental results and found that high expression levels of TWIST1 and EPCAM were related to drug resistance. (C) Prof. Cho’s team identified three key molecular targets: p53, SMAD4 and ERK1 & ERK2. (D), (E) Furthermore, they identified a key pathway that plays an important role in completely reversing into epithelial cells while avoiding EMT hybrid characteristics, and confirmed through network analysis and attractor analysis that high stability of the key pathway was restored when the proposed molecular target was controlled. >
< Figure 4. Verification through experiments with lung cancer cell lines.
When p53 was activated and SMAD4 and ERK1/2 were inhibited in lung cancer cell lines, (A), (B) E-cad protein expression increased and ZEB1 protein expression decreased, and (C) mesenchymal cell status including TWIST1 and EPCAM and gene expression of markers related to stem cell potential characteristics were completely inhibited. In addition, (D) it was confirmed that resistance to chemotherapy treatment was also overcome as the cell state was reversed by the regulated target. >
< Figure 5. A schematic representation of the research results.
Prof. Cho’s research team identified key molecular regulatory pathways to avoid high plasticity formed by abnormal EMT of cancer cells and reverse it to an epithelial cell state through systems biology research. From this analysis, a reprogramming molecular target that can reverse the state of mesenchymal cells with acquired invasiveness and drug resistance to the state of epithelial cells with restored drug responsiveness was discovered.
For lung cancer cells, when a drug that enhances the expression of p53, one of the molecular targets discovered, and inhibits the expression of SMAD4 and ERK1 & ERK2 is administered, the molecular network of genes in the state of mesenchymal cells is modified, eventually eliminating metastatic ability and it is reprogrammed to turn into epithelial cells without the resistance to chemotherapy treatments. >
2023.01.30 View 15968
KAIST presents a fundamental technology to remove metastatic traits from lung cancer cells
KAIST (President Kwang Hyung Lee) announced on January 30th that a research team led by Professor Kwang-Hyun Cho from the Department of Bio and Brain Engineering succeeded in using systems biology research to change the properties of carcinogenic cells in the lungs and eliminate both drug resistance and their ability to proliferate out to other areas of the body.
As the incidences of cancer increase within aging populations, cancer has become the most lethal disease threatening healthy life. Fatality rates are especially high when early detection does not happen in time and metastasis has occurred in various organs. In order to resolve this problem, a series of attempts were made to remove or lower the ability of cancer cells to spread, but they resulted in cancer cells in the intermediate state becoming more unstable and even more malignant, which created serious treatment challenges.
Professor Kwang-Hyun Cho's research team simulated various cancer cell states in the Epithelial-to-Mesenchymal Transition (EMT) of lung cancer cells, between epithelial cells without metastatic ability and mesenchymal cells with metastatic ability. A mathematical model of molecular network was established, and key regulators that could reverse the state of invasive and drug resistant mesenchymal cells back to the epithelial state were discovered through computer simulation analysis and molecular cell experiments. In particular, this process succeeded in properly reverting the mesenchymal lung cancer cells to a state where they were sensitive to chemotherapy treatment while avoiding the unstable EMT hybrid cell state in the middle process, which had remained a difficult problem.
The results of this research, in which KAIST Ph.D. student Namhee Kim, Dr. Chae Young Hwang, Researcher Taeyoung Kim, and Ph.D. student Hyunjin Kim participated, were published as an online paper in the international journal “Cancer Research” published by the American Association for Cancer Research (AACR) on January 30th. (Paper title: A cell fate reprogramming strategy reverses epithelial-to-mesenchymal transition of lung cancer cells while avoiding hybrid states)
Cells in an EMT hybrid state, which are caused by incomplete transitions during the EMT process in cancer cells, have the characteristics of both epithelial cells and mesenchymal cells, and are known to have high drug resistance and metastatic potential by acquiring high stem cell capacity. In particular, EMT is further enhanced through factors such as transforming growth factor-beta (TGF-β) secreted from the tumor microenvironment (TME) and, as a result, various cell states with high plasticity appear. Due to the complexity of EMT, it has been very difficult to completely reverse the transitional process of the mesenchymal cancer cells to an epithelial cell state in which metastatic ability and drug resistance are eliminated while avoiding the EMT hybrid cell state with high metastatic ability and drug resistance.
Professor Kwang-Hyun Cho's research team established a mathematical model of the gene regulation network that governs the complex process of EMT, and then applied large-scale computer simulation analysis and complex system network control technology to identify and verify 'p53', 'SMAD4', and 'ERK1' and 'ERK 2' (collectively ERKs) through molecular cell experiments as the three key molecular targets that can transform lung cancer cells in the mesenchymal cell state, reversed back to an epithelial cell state that no longer demonstrates the ability to metastasize, while avoiding the EMT hybrid cell state.
In particular, by analyzing the molecular regulatory mechanism of the complex EMT process at the system level, the key pathways were identified that were linked to the positive feedback that plays an important role in completely returning cancer cells to an epithelial cell state in which metastatic ability and drug resistance are removed.
This discovery is significant in that it proved that mesenchymal cells can be reverted to the state of epithelial cells under conditions where TGF-β stimulation are present, like they are in the actual environment where cancer tissue forms in the human body.
Abnormal EMT in cancer cells leads to various malignant traits such as the migration and invasion of cancer cells, changes in responsiveness to chemotherapy treatment, enhanced stem cell function, and the dissemination of cancer. In particular, the acquisition of the metastatic ability of cancer cells is a key determinant factor for the prognosis of cancer patients. The EMT reversal technology in lung cancer cells developed in this research is a new anti-cancer treatment strategy that reprograms cancer cells to eliminate their high plasticity and metastatic potential and increase their responsiveness to chemotherapy.
Professor Kwang-Hyun Cho said, "By succeeding in reversing the state of lung cancer cells that acquired high metastatic traits and resistance to drugs and reverting them to a treatable epithelial cell state with renewed sensitivity to chemotherapy, the research findings propose a new strategy for treatments that can improve the prognosis of cancer patients.”
Professor Kwang-Hyun Cho's research team was the first to present the principle of reversal treatment to revert cancer cells to normal cells, following through with the announcement of the results of their study that reverted colon cancer cells to normal colon cells in January of 2020, and also presenting successful re-programming research where the most malignant basal type breast cancer cells turned into less-malignant luminal type breast cancer cells that were treatable with hormonal therapies in January of 2022. This latest research result is the third in the development of reversal technology where lung cancer cells that had acquired metastatic traits returned to a state in which their metastatic ability was removed and drug sensitivity was enhanced.
This research was carried out with support from the Ministry of Science and ICT and the National Research Foundation of Korea's Basic Research in Science & Engineering Program for Mid-Career Researchers.
< Figure 1. Construction of the mathematical model of the regulatory network to represent the EMT phenotype based on the interaction between various molecules related to EMT.
(A) Professor Kwang-Hyun Cho's research team investigated numerous literatures and databases related to complex EMT, and based on comparative analysis of cell line data showing epithelial and mesenchymal cell conditions, they extracted key signaling pathways related to EMT and built a mathematical model of regulatory network (B) By comparing the results of computer simulation analysis and the molecular cell experiments, it was verified how well the constructed mathematical model simulated the actual cellular phenomena. >
< Figure 2. Understanding of various EMT phenotypes through large-scale computer simulation analysis and complex system network control technology.
(A) Through computer simulation analysis and experiments, Professor Kwang-Hyun Cho's research team found that complete control of EMT is impossible with single-molecule control alone. In particular, through comparison of the relative stability of attractors, it was revealed that the cell state exhibiting EMT hybrid characteristics has unstable properties. (B), (C) Based on these results, Prof. Cho’s team identified two feedbacks (positive feedback consisting of Snail-miR-34 and ZEB1-miR-200) that play an important role in avoiding the EMT hybrid state that appeared in the TGF-β-ON state. It was found through computer simulation analysis that the two feedbacks restore relatively high stability when the excavated p53 and SMAD4 are regulated. In addition, molecular cell experiments demonstrated that the expression levels of E-cad and ZEB1, which are representative phenotypic markers of EMT, changed similarly to the expression profile in the epithelial cell state, despite the TGF-β-ON state. >
< Figure 3. Complex molecular network analysis and discovery of reprogramming molecular targets for intact elimination of EMT hybrid features.
(A) Controlling the expression of p53 and SMAD4 in lung cancer cell lines was expected to overcome drug resistance, but contrary to expectations, chemotherapy responsiveness was not restored. (B) Professor Kwang-Hyun Cho's research team additionally analyzed computer simulations, genome data, and experimental results and found that high expression levels of TWIST1 and EPCAM were related to drug resistance. (C) Prof. Cho’s team identified three key molecular targets: p53, SMAD4 and ERK1 & ERK2. (D), (E) Furthermore, they identified a key pathway that plays an important role in completely reversing into epithelial cells while avoiding EMT hybrid characteristics, and confirmed through network analysis and attractor analysis that high stability of the key pathway was restored when the proposed molecular target was controlled. >
< Figure 4. Verification through experiments with lung cancer cell lines.
When p53 was activated and SMAD4 and ERK1/2 were inhibited in lung cancer cell lines, (A), (B) E-cad protein expression increased and ZEB1 protein expression decreased, and (C) mesenchymal cell status including TWIST1 and EPCAM and gene expression of markers related to stem cell potential characteristics were completely inhibited. In addition, (D) it was confirmed that resistance to chemotherapy treatment was also overcome as the cell state was reversed by the regulated target. >
< Figure 5. A schematic representation of the research results.
Prof. Cho’s research team identified key molecular regulatory pathways to avoid high plasticity formed by abnormal EMT of cancer cells and reverse it to an epithelial cell state through systems biology research. From this analysis, a reprogramming molecular target that can reverse the state of mesenchymal cells with acquired invasiveness and drug resistance to the state of epithelial cells with restored drug responsiveness was discovered.
For lung cancer cells, when a drug that enhances the expression of p53, one of the molecular targets discovered, and inhibits the expression of SMAD4 and ERK1 & ERK2 is administered, the molecular network of genes in the state of mesenchymal cells is modified, eventually eliminating metastatic ability and it is reprogrammed to turn into epithelial cells without the resistance to chemotherapy treatments. >
2023.01.30 View 15968 -
 Overview of the 30-year history of metabolic engineering
< Distinguished Professor Sang Yup Lee from the Department of Chemical and Biomolecular Engineering at KAIST >
A research team comprised of Gi Bae Kim, Dr. So Young Choi, Dr. In Jin Cho, Da-Hee Ahn, and Distinguished Professor Sang Yup Lee from the Department of Chemical and Biomolecular Engineering at KAIST reported the 30-year history of metabolic engineering, highlighting examples of recent progress in the field and contributions to sustainability and health. Their paper “Metabolic engineering for sustainability and health” was published online in the 40th anniversary special issue of Trends in Biotechnology on January 10, 2023.
Metabolic engineering, a discipline of engineering that modifies cell phenotypes through molecular and genetic-level manipulations to improve cellular activities, has been studied since the early 1990s, and has progressed significantly over the past 30 years. In particular, metabolic engineering has enabled the engineering of microorganisms for the development of microbial cell factories capable of efficiently producing chemicals and materials as well as degrading recalcitrant contaminants.
This review article revisited how metabolic engineering has advanced over the past 30 years, from the advent of genetic engineering techniques such as recombinant DNA technologies to recent breakthroughs in systems metabolic engineering and data science aided by artificial intelligence. The research team highlighted momentous events and achievements in metabolic engineering, providing both trends and future directions in the field. Metabolic engineering’s contributions to bio-based sustainable chemicals and clean energy, health, and bioremediation were also reviewed. Finally, the research team shared their perspectives on the future challenges impacting metabolic engineering than must be overcome in order to achieve advancements in sustainability and health.
Distinguished Professor Sang Yup Lee said, “Replacing fossil resource-based chemical processes with bio-based sustainable processes for the production of chemicals, fuels, and materials using metabolic engineering has become our essential task for the future. By looking back on the 30+ years of metabolic engineering, we aimed to highlight the contributions of metabolic engineering to achieve sustainability and good health.” He added, “Metabolic engineering will play an increasingly important role as a key solution to the climate crisis, environmental pollution, food and energy shortages, and health problems in aging societies.”
< Figure: Metabolic Engineering Timeline >
2023.01.25 View 9518
Overview of the 30-year history of metabolic engineering
< Distinguished Professor Sang Yup Lee from the Department of Chemical and Biomolecular Engineering at KAIST >
A research team comprised of Gi Bae Kim, Dr. So Young Choi, Dr. In Jin Cho, Da-Hee Ahn, and Distinguished Professor Sang Yup Lee from the Department of Chemical and Biomolecular Engineering at KAIST reported the 30-year history of metabolic engineering, highlighting examples of recent progress in the field and contributions to sustainability and health. Their paper “Metabolic engineering for sustainability and health” was published online in the 40th anniversary special issue of Trends in Biotechnology on January 10, 2023.
Metabolic engineering, a discipline of engineering that modifies cell phenotypes through molecular and genetic-level manipulations to improve cellular activities, has been studied since the early 1990s, and has progressed significantly over the past 30 years. In particular, metabolic engineering has enabled the engineering of microorganisms for the development of microbial cell factories capable of efficiently producing chemicals and materials as well as degrading recalcitrant contaminants.
This review article revisited how metabolic engineering has advanced over the past 30 years, from the advent of genetic engineering techniques such as recombinant DNA technologies to recent breakthroughs in systems metabolic engineering and data science aided by artificial intelligence. The research team highlighted momentous events and achievements in metabolic engineering, providing both trends and future directions in the field. Metabolic engineering’s contributions to bio-based sustainable chemicals and clean energy, health, and bioremediation were also reviewed. Finally, the research team shared their perspectives on the future challenges impacting metabolic engineering than must be overcome in order to achieve advancements in sustainability and health.
Distinguished Professor Sang Yup Lee said, “Replacing fossil resource-based chemical processes with bio-based sustainable processes for the production of chemicals, fuels, and materials using metabolic engineering has become our essential task for the future. By looking back on the 30+ years of metabolic engineering, we aimed to highlight the contributions of metabolic engineering to achieve sustainability and good health.” He added, “Metabolic engineering will play an increasingly important role as a key solution to the climate crisis, environmental pollution, food and energy shortages, and health problems in aging societies.”
< Figure: Metabolic Engineering Timeline >
2023.01.25 View 9518 -
 Scientists re-writes FDA-recommended equation to improve estimation of drug-drug interaction
Drugs absorbed into the body are metabolized and thus removed by enzymes from several organs like the liver. How fast a drug is cleared out of the system can be affected by other drugs that are taken together because added substance can increase the amount of enzyme secretion in the body. This dramatically decreases the concentration of a drug, reducing its efficacy, often leading to the failure of having any effect at all. Therefore, accurately predicting the clearance rate in the presence of drug-drug interaction* is critical in the process of drug prescription and development of a new drug in order to ensure its efficacy and/or to avoid unwanted side-effects.
*Drug-drug interaction: In terms of metabolism, drug-drug interaction is a phenomenon in which one drug changes the metabolism of another drug to promote or inhibit its excretion from the body when two or more drugs are taken together. As a result, it increases the toxicity of medicines or causes loss of efficacy.
Since it is practically impossible to evaluate all interactions between new drug candidates and all marketed drugs during the development process, the FDA recommends indirect evaluation of drug interactions using a formula suggested in their guidance, first published in 1997, revised in January of 2020, in order to evaluate drug interactions and minimize side effects of having to use more than one type of drugs at once.
The formula relies on the 110-year-old Michaelis-Menten (MM) model, which has a fundamental limit of making a very broad and groundless assumption on the part of the presence of the enzymes that metabolizes the drug. While MM equation has been one of the most widely known equations in biochemistry used in more than 220,000 published papers, the MM equation is accurate only when the concentration of the enzyme that metabolizes the drug is almost non-existent, causing the accuracy of the equation highly unsatisfactory – only 38 percent of the predictions had less than two-fold errors.
“To make up for the gap, researcher resorted to plugging in scientifically unjustified constants into the equation,” Professor Jung-woo Chae of Chungnam National University College of Pharmacy said. “This is comparable to having to have the epicyclic orbits introduced to explain the motion of the planets back in the days in order to explain the now-defunct Ptolemaic theory, because it was 'THE' theory back then.”
< (From left) Ph.D. student Yun Min Song (KAIST, co-first authors), Professor Sang Kyum Kim (Chungnam National University, co-corresponding author), Jae Kyoung Kim, CI (KAIST, co-corresponding author), Professor Jung-woo Chae (Chungnam National University, co-corresponding author), Ph.D. students Quyen Thi Tran and Ngoc-Anh Thi Vu (Chungnam National University, co-first authors) >
A joint research team composed of mathematicians from the Biomedical Mathematics Group within the Institute for Basic Science (IBS) and the Korea Advanced Institute of Science and Technology (KAIST) and pharmacological scientists from the Chungnam National University reported that they identified the major causes of the FDA-recommended equation’s inaccuracies and presented a solution.
When estimating the gut bioavailability (Fg), which is the key parameter of the equation, the fraction absorbed from the gut lumen (Fa) is usually assumed to be 1. However, many experiments have shown that Fa is less than 1, obviously since it can’t be expected that all of the orally taken drugs to be completely absorbed by the intestines. To solve this problem, the research team used an “estimated Fa” value based on factors such as the drug’s transit time, intestine radius, and permeability values and used it to re-calculate Fg.
Also, taking a different approach from the MM equation, the team used an alternative model they derived in a previous study back in 2020, which can more accurately predict the drug metabolism rate regardless of the enzyme concentration. Combining these changes, the modified equation with re-calculated Fg had a dramatically increased accuracy of the resulting estimate. The existing FDA formula predicted drug interactions within a 2-fold margin of error at the rate of 38%, whereas the accuracy rate of the revised formula reached 80%.
“Such drastic improvement in drug-drug interaction prediction accuracy is expected to make great contribution to increasing the success rate of new drug development and drug efficacy in clinical trials. As the results of this study were published in one of the top clinical pharmacology journal, it is expected that the FDA guidance will be revised according to the results of this study.” said Professor Sang Kyum Kim from Chungnam National University College of Pharmacy.
Furthermore, this study highlights the importance of collaborative research between research groups in vastly different disciplines, in a field that is as dynamic as drug interactions.
“Thanks to the collaborative research between mathematics and pharmacy, we were able to recify the formula that we have accepted to be the right answer for so long to finally grasp on the leads toward healthier life for mankind.,” said Professor Jae Kyung Kim. He continued, “I hope seeing a ‘K-formula’ entered into the US FDA guidance one day.”
The results of this study were published in the online edition of Clinical Pharmacology and Therapeutics (IF 7.051), an authoritative journal in the field of clinical pharmacology, on December 15, 2022 (Korean time).
Thesis Title: Beyond the Michaelis-Menten: Accurate Prediction of Drug Interactions through Cytochrome P450 3A4 Induction (doi: 10.1002/cpt.2824)
< Figure 1. The formula proposed by the FDA guidance for predicting drug-drug interactions (top) and the formula newly derived by the researchers (bottom). AUCR (the ratio of substrate area under the plasma concentration-time curve) represents the rate of change in drug concentration due to drug interactions. The research team more than doubled the accuracy of drug interaction prediction compared to the existing formula. >
< Figure 2. Existing FDA formulas tend to underestimate the extent of drug-drug interactions (gray dots) than the actual measured values. On the other hand, the newly derived equation (red dot) has a prediction rate that is within the error range of 2 times (0.5 to 2 times) of the measured value, and is more than twice as high as the existing equation. The solid line in the figure represents the predicted value that matches the measured value. The dotted line represents the predicted value with an error of 0.5 to 2 times. >
For further information or to request media assistance, please contact Jae Kyoung Kim at Biomedical Mathematics Group, Institute for Basic Science (IBS) (jaekkim@ibs.re.kr) or William I. Suh at the IBS Communications Team (willisuh@ibs.re.kr).
- About the Institute for Basic Science (IBS)
IBS was founded in 2011 by the government of the Republic of Korea with the sole purpose of driving forward the development of basic science in South Korea. IBS has 4 research institutes and 33 research centers as of January 2023. There are eleven physics, three mathematics, five chemistry, nine life science, two earth science, and three interdisciplinary research centers.
2023.01.18 View 11971
Scientists re-writes FDA-recommended equation to improve estimation of drug-drug interaction
Drugs absorbed into the body are metabolized and thus removed by enzymes from several organs like the liver. How fast a drug is cleared out of the system can be affected by other drugs that are taken together because added substance can increase the amount of enzyme secretion in the body. This dramatically decreases the concentration of a drug, reducing its efficacy, often leading to the failure of having any effect at all. Therefore, accurately predicting the clearance rate in the presence of drug-drug interaction* is critical in the process of drug prescription and development of a new drug in order to ensure its efficacy and/or to avoid unwanted side-effects.
*Drug-drug interaction: In terms of metabolism, drug-drug interaction is a phenomenon in which one drug changes the metabolism of another drug to promote or inhibit its excretion from the body when two or more drugs are taken together. As a result, it increases the toxicity of medicines or causes loss of efficacy.
Since it is practically impossible to evaluate all interactions between new drug candidates and all marketed drugs during the development process, the FDA recommends indirect evaluation of drug interactions using a formula suggested in their guidance, first published in 1997, revised in January of 2020, in order to evaluate drug interactions and minimize side effects of having to use more than one type of drugs at once.
The formula relies on the 110-year-old Michaelis-Menten (MM) model, which has a fundamental limit of making a very broad and groundless assumption on the part of the presence of the enzymes that metabolizes the drug. While MM equation has been one of the most widely known equations in biochemistry used in more than 220,000 published papers, the MM equation is accurate only when the concentration of the enzyme that metabolizes the drug is almost non-existent, causing the accuracy of the equation highly unsatisfactory – only 38 percent of the predictions had less than two-fold errors.
“To make up for the gap, researcher resorted to plugging in scientifically unjustified constants into the equation,” Professor Jung-woo Chae of Chungnam National University College of Pharmacy said. “This is comparable to having to have the epicyclic orbits introduced to explain the motion of the planets back in the days in order to explain the now-defunct Ptolemaic theory, because it was 'THE' theory back then.”
< (From left) Ph.D. student Yun Min Song (KAIST, co-first authors), Professor Sang Kyum Kim (Chungnam National University, co-corresponding author), Jae Kyoung Kim, CI (KAIST, co-corresponding author), Professor Jung-woo Chae (Chungnam National University, co-corresponding author), Ph.D. students Quyen Thi Tran and Ngoc-Anh Thi Vu (Chungnam National University, co-first authors) >
A joint research team composed of mathematicians from the Biomedical Mathematics Group within the Institute for Basic Science (IBS) and the Korea Advanced Institute of Science and Technology (KAIST) and pharmacological scientists from the Chungnam National University reported that they identified the major causes of the FDA-recommended equation’s inaccuracies and presented a solution.
When estimating the gut bioavailability (Fg), which is the key parameter of the equation, the fraction absorbed from the gut lumen (Fa) is usually assumed to be 1. However, many experiments have shown that Fa is less than 1, obviously since it can’t be expected that all of the orally taken drugs to be completely absorbed by the intestines. To solve this problem, the research team used an “estimated Fa” value based on factors such as the drug’s transit time, intestine radius, and permeability values and used it to re-calculate Fg.
Also, taking a different approach from the MM equation, the team used an alternative model they derived in a previous study back in 2020, which can more accurately predict the drug metabolism rate regardless of the enzyme concentration. Combining these changes, the modified equation with re-calculated Fg had a dramatically increased accuracy of the resulting estimate. The existing FDA formula predicted drug interactions within a 2-fold margin of error at the rate of 38%, whereas the accuracy rate of the revised formula reached 80%.
“Such drastic improvement in drug-drug interaction prediction accuracy is expected to make great contribution to increasing the success rate of new drug development and drug efficacy in clinical trials. As the results of this study were published in one of the top clinical pharmacology journal, it is expected that the FDA guidance will be revised according to the results of this study.” said Professor Sang Kyum Kim from Chungnam National University College of Pharmacy.
Furthermore, this study highlights the importance of collaborative research between research groups in vastly different disciplines, in a field that is as dynamic as drug interactions.
“Thanks to the collaborative research between mathematics and pharmacy, we were able to recify the formula that we have accepted to be the right answer for so long to finally grasp on the leads toward healthier life for mankind.,” said Professor Jae Kyung Kim. He continued, “I hope seeing a ‘K-formula’ entered into the US FDA guidance one day.”
The results of this study were published in the online edition of Clinical Pharmacology and Therapeutics (IF 7.051), an authoritative journal in the field of clinical pharmacology, on December 15, 2022 (Korean time).
Thesis Title: Beyond the Michaelis-Menten: Accurate Prediction of Drug Interactions through Cytochrome P450 3A4 Induction (doi: 10.1002/cpt.2824)
< Figure 1. The formula proposed by the FDA guidance for predicting drug-drug interactions (top) and the formula newly derived by the researchers (bottom). AUCR (the ratio of substrate area under the plasma concentration-time curve) represents the rate of change in drug concentration due to drug interactions. The research team more than doubled the accuracy of drug interaction prediction compared to the existing formula. >
< Figure 2. Existing FDA formulas tend to underestimate the extent of drug-drug interactions (gray dots) than the actual measured values. On the other hand, the newly derived equation (red dot) has a prediction rate that is within the error range of 2 times (0.5 to 2 times) of the measured value, and is more than twice as high as the existing equation. The solid line in the figure represents the predicted value that matches the measured value. The dotted line represents the predicted value with an error of 0.5 to 2 times. >
For further information or to request media assistance, please contact Jae Kyoung Kim at Biomedical Mathematics Group, Institute for Basic Science (IBS) (jaekkim@ibs.re.kr) or William I. Suh at the IBS Communications Team (willisuh@ibs.re.kr).
- About the Institute for Basic Science (IBS)
IBS was founded in 2011 by the government of the Republic of Korea with the sole purpose of driving forward the development of basic science in South Korea. IBS has 4 research institutes and 33 research centers as of January 2023. There are eleven physics, three mathematics, five chemistry, nine life science, two earth science, and three interdisciplinary research centers.
2023.01.18 View 11971 -
 “3D sketch” Your Ideas and Bring Them to Life, Instantly!
Professor Seok-Hyung Bae’s research team at the Department of Industrial Design developed a novel 3D sketching system that rapidly creates animated 3D concepts through simple user interactions like sketching on a piece of paper or playing a toy.
Foldable drones, transforming vehicles, and multi-legged robots from sci-fi movies are now becoming commonplace thanks to technological progress. However, designing them remains a difficult challenge even for skilled experts, because complex design decisions must be made regarding not only their form, but also the structure, poses, and motions, which are interdependent on one another.
Creating a 3D concept comprising of multiple moving parts connected by different types of joints using a traditional 3D CAD tool, which is more suited for processing precise and elaborate modeling, is a painstaking and time-consuming process. This presents a major bottleneck for the workflow during the early stage of design, in which it is preferred that as many ideas are tried and discarded out as quickly as possible in order to explore a wide range of possibilities in the shortest amount of time.
A research team led by Professor Bae has focused on designers’ freehand sketches drew up with a pen on a paper that serve as the starting point for virtually all design projects. This led them to develop their 3D sketching technology to generate desired 3D curves from the rough but expressive 2D strokes drawn with a digital stylus on a digital tablet.
Their latest research helps designers bring their 3D sketches to life almost instantly. Using the intuitive set of multi-touch gestures the team successfully designed and implemented, designers can handle the 3D sketches they are working on with their fingers as if they are playing with toys and put them into animation in no time.
< Figure 1. A novel 3D sketching system for rapidly designing articulated 3D concepts with a small set of coherent pen and multi-touch gestures. (a) Sketching: A 3D sketch curve is created by marking a pen stroke that is projected onto a sketch plane widget. (b) Segmenting: Entire or partial sketch curves are added to separate parts that serve as links in the kinematic chain. (c) Rigging: Repeatedly demonstrating the desired motion of a part leaves behind a trail, from which the system infers a joint. (d) Posing: Desired poses can be achieved through actuating joints via forward or inverse kinematics. (e) Filming: A sequence of keyframes specifying desired poses and viewpoints is connected as a smooth motion. >
< Figure 2. (a) Concept drawing of an autonomous excavator. It features (b, c) four caterpillars that swivel for high maneuverability, (d) an extendable boom and a bucket connected by multiple links, and (e) a rotating platform. The concept’s designer, who had 8 years of work experience, estimated that it would take 1-2 weeks to express and communicate such a complex articulated object with existing tools. With the proposed system, it took only 2 hours and 52 minutes. >
The major findings of their work were published under the title “Rapid Design of Articulated Objects” in ACM Transactions on Graphics (impact factor: 7.403), the top international journal in the field of computer graphics, and presented at ACM SIGGRAPH 2022 (h5-index: 103), the world’s largest international academic conference in the field, which was held back in August in Vancouver, Canada with Joon Hyub Lee, a Ph.D. student of the Department of Industrial Design as the first author.
The ACM SIGGRAPH 2022 conference was reportedly attended by over 10,000 participants including researchers, artists, and developers from world-renowned universities; film, animation, and game studies, such as Marvel, Pixar, and Blizzard; high-tech manufacturers, such as Lockheed Martin and Boston Dynamics; and metaverse platform companies, such as Meta and Roblox.
< Figure 3. The findings of Professor Bae’s research team were published in ACM Transactions on Graphics, the top international academic journal in the field of computer graphics, and presented at ACM SIGGRAPH 2022, the largest international academic conference held in conjunction early August in Vancouver, Canada. The team’s live demo at the Emerging Technologies program was highly praised by numerous academics and industry officials and received an Honorable Mention. >
The team was also invited to present their technical paper as a demo and a special talk at the Emerging Technologies program at ACM SIGGRAPH 2022 as one of the top-three impactful technologies. The live performance, in which Hanbit Kim, a Ph.D. student of the Department of Industrial Design at KAIST and a co-author, sketched and animated a sophisticated animal-shaped robot from scratch in a matter of a few minutes, wowed the audience and won the Honorable Mention Award from the jury.
Edwin Catmull, the co-founder of Pixar and a keynote speaker at the SIGGRAPH conference, praised the team’s research on 3D sketching as “really excellent work” and “a kind of tool that would be useful to Pixar's creative model designers.”
This technology, which became virally popular in Japan after featuring in an online IT media outlet and attracting more than 600K views, received a special award from the Digital Content Association of Japan (DCAJ) and was invited and exhibited for three days at Tokyo in November, as a part of Inter BEE 2022, the largest broadcasting and media expo in Japan.
“The more we come to understand how designers think and work, the more effective design tools can be built around that understanding,” said Professor Bae, explaining that “the key is to integrate different algorithms into a harmonious system as intuitive interactions.” He added that “this work wouldn’t have been possible if it weren’t for the convergent research environment cultivated by the Department of Industrial Design at KAIST, in which all students see themselves not only as aspiring creative designers, but also as practical engineers.”
By enabling designers to produce highly expressive animated 3D concepts far more quickly and easily in comparison to using existing methods, this new tool is expected to revolutionize design practices and processes in the content creation, manufacturing, and metaverse-related industries.
This research was funded by the Ministry of Science and ICT, and the National Research Foundation of Korea.
More info: https://sketch.kaist.ac.kr/publications/2022_siggraph_rapid_design
Video: https://www.youtube.com/watch?v=rsBl0QvSDqI
< Figure 4. From left to right: Ph.D. students Hanbit Kim, and Joon Hyub Lee and Professor Bae of the Department of Industrial Design, KAIST >
2022.11.23 View 9100
“3D sketch” Your Ideas and Bring Them to Life, Instantly!
Professor Seok-Hyung Bae’s research team at the Department of Industrial Design developed a novel 3D sketching system that rapidly creates animated 3D concepts through simple user interactions like sketching on a piece of paper or playing a toy.
Foldable drones, transforming vehicles, and multi-legged robots from sci-fi movies are now becoming commonplace thanks to technological progress. However, designing them remains a difficult challenge even for skilled experts, because complex design decisions must be made regarding not only their form, but also the structure, poses, and motions, which are interdependent on one another.
Creating a 3D concept comprising of multiple moving parts connected by different types of joints using a traditional 3D CAD tool, which is more suited for processing precise and elaborate modeling, is a painstaking and time-consuming process. This presents a major bottleneck for the workflow during the early stage of design, in which it is preferred that as many ideas are tried and discarded out as quickly as possible in order to explore a wide range of possibilities in the shortest amount of time.
A research team led by Professor Bae has focused on designers’ freehand sketches drew up with a pen on a paper that serve as the starting point for virtually all design projects. This led them to develop their 3D sketching technology to generate desired 3D curves from the rough but expressive 2D strokes drawn with a digital stylus on a digital tablet.
Their latest research helps designers bring their 3D sketches to life almost instantly. Using the intuitive set of multi-touch gestures the team successfully designed and implemented, designers can handle the 3D sketches they are working on with their fingers as if they are playing with toys and put them into animation in no time.
< Figure 1. A novel 3D sketching system for rapidly designing articulated 3D concepts with a small set of coherent pen and multi-touch gestures. (a) Sketching: A 3D sketch curve is created by marking a pen stroke that is projected onto a sketch plane widget. (b) Segmenting: Entire or partial sketch curves are added to separate parts that serve as links in the kinematic chain. (c) Rigging: Repeatedly demonstrating the desired motion of a part leaves behind a trail, from which the system infers a joint. (d) Posing: Desired poses can be achieved through actuating joints via forward or inverse kinematics. (e) Filming: A sequence of keyframes specifying desired poses and viewpoints is connected as a smooth motion. >
< Figure 2. (a) Concept drawing of an autonomous excavator. It features (b, c) four caterpillars that swivel for high maneuverability, (d) an extendable boom and a bucket connected by multiple links, and (e) a rotating platform. The concept’s designer, who had 8 years of work experience, estimated that it would take 1-2 weeks to express and communicate such a complex articulated object with existing tools. With the proposed system, it took only 2 hours and 52 minutes. >
The major findings of their work were published under the title “Rapid Design of Articulated Objects” in ACM Transactions on Graphics (impact factor: 7.403), the top international journal in the field of computer graphics, and presented at ACM SIGGRAPH 2022 (h5-index: 103), the world’s largest international academic conference in the field, which was held back in August in Vancouver, Canada with Joon Hyub Lee, a Ph.D. student of the Department of Industrial Design as the first author.
The ACM SIGGRAPH 2022 conference was reportedly attended by over 10,000 participants including researchers, artists, and developers from world-renowned universities; film, animation, and game studies, such as Marvel, Pixar, and Blizzard; high-tech manufacturers, such as Lockheed Martin and Boston Dynamics; and metaverse platform companies, such as Meta and Roblox.
< Figure 3. The findings of Professor Bae’s research team were published in ACM Transactions on Graphics, the top international academic journal in the field of computer graphics, and presented at ACM SIGGRAPH 2022, the largest international academic conference held in conjunction early August in Vancouver, Canada. The team’s live demo at the Emerging Technologies program was highly praised by numerous academics and industry officials and received an Honorable Mention. >
The team was also invited to present their technical paper as a demo and a special talk at the Emerging Technologies program at ACM SIGGRAPH 2022 as one of the top-three impactful technologies. The live performance, in which Hanbit Kim, a Ph.D. student of the Department of Industrial Design at KAIST and a co-author, sketched and animated a sophisticated animal-shaped robot from scratch in a matter of a few minutes, wowed the audience and won the Honorable Mention Award from the jury.
Edwin Catmull, the co-founder of Pixar and a keynote speaker at the SIGGRAPH conference, praised the team’s research on 3D sketching as “really excellent work” and “a kind of tool that would be useful to Pixar's creative model designers.”
This technology, which became virally popular in Japan after featuring in an online IT media outlet and attracting more than 600K views, received a special award from the Digital Content Association of Japan (DCAJ) and was invited and exhibited for three days at Tokyo in November, as a part of Inter BEE 2022, the largest broadcasting and media expo in Japan.
“The more we come to understand how designers think and work, the more effective design tools can be built around that understanding,” said Professor Bae, explaining that “the key is to integrate different algorithms into a harmonious system as intuitive interactions.” He added that “this work wouldn’t have been possible if it weren’t for the convergent research environment cultivated by the Department of Industrial Design at KAIST, in which all students see themselves not only as aspiring creative designers, but also as practical engineers.”
By enabling designers to produce highly expressive animated 3D concepts far more quickly and easily in comparison to using existing methods, this new tool is expected to revolutionize design practices and processes in the content creation, manufacturing, and metaverse-related industries.
This research was funded by the Ministry of Science and ICT, and the National Research Foundation of Korea.
More info: https://sketch.kaist.ac.kr/publications/2022_siggraph_rapid_design
Video: https://www.youtube.com/watch?v=rsBl0QvSDqI
< Figure 4. From left to right: Ph.D. students Hanbit Kim, and Joon Hyub Lee and Professor Bae of the Department of Industrial Design, KAIST >
2022.11.23 View 9100 -
 KAIST develops biocompatible adhesive applicable to hair transplants
Aside from being used as a new medical adhesive, the new material can be applied to developing a new method of hair transplants, which cannot be repeated multiple times using current method of implanting the wholly intact follicles into the skin.
Medical adhesives are materials that can be applied to various uses such as wound healing, hemostasis, vascular anastomosis, and tissue engineering, and is expected to contribute greatly to the development of minimally invasive surgery and organ transplants. However, adhesives with high adhesion, low toxicity, and capable of decomposing in the body are rare. Adhesives based on natural proteins, such as fibrin and collagen, have high biocompatibility but insufficient adhesive strength. Synthetic polymer adhesives based on urethane or acrylic have greater adhesion but do not decompose well and may cause an inflammatory reaction in the body.
A joint research team led by Professor Myungeun Seo and Professor Haeshin Lee from the KAIST Department of Chemistry developed a bio-friendly adhesive from biocompatible polymers using tannic acid, the source of astringency in wine.
The research team focused on tannic acid, a natural polyphenolic product. Tannic acid is a polyphenol present in large amounts in fruit peels, nuts, and cacao. It has a high affinity and coating ability on other substances, and we sense the astringent taste in wine when tannic acid sticks to the surface of our tongue. When tannic acid is mixed with hydrophilic polymers, they form coacervates, or small droplets of jelly-like fluids that sink. If the polymers used are biocompatible, the mixture can be applied as a medical adhesive with low toxicity. However, coacervates are fundamentally fluid-like and cannot withstand large forces, which limits their adhesive capabilities. Thus, while research to utilize it as an adhesive has been actively discussed, a biodegradable material exhibiting strong adhesion due to its high shear strength has not yet been developed.
The research team figured out a way to enhance adhesion by mixing two biocompatible FDA-approved polymers, polyethylene glycol (PEG) and polylactic acid (PLA). While PEG, which is used widely in eyedrops and cream, is hydrophilic, PLA, a well-known bioplastic derived from lactic acid, is insoluble in water. The team combined the two into a block copolymer, which forms hydrophilic PLA aggregates in water with PEG blocks surrounding them. A coacervate created by mixing the micelles and tannic acid would behave like a solid due to the hard PLA components, and show an elastic modulus improved by a thousand times compared to PEG, enabling it to withstand much greater force as an adhesive.
Figure 1. (Above) Principle of biodegradable adhesive made by mixing poly(ethylene glycol)-poly(lactic acid) diblock copolymer and tannic acid in water. Yellow coacervate is precipitated through hydrogen bonding between the block copolymer micelles and tannic acid, and exhibits adhesion. After heat treatment, hydrogen bonds are rearranged to further improve adhesion. (Bottom) Adhesion comparison. Compared to using poly(ethylene glycol) polymer (d), it can support 10 times more weight when using block copolymer (e) and 60 times more weight after heat treatment (f). The indicated G' values represent the elastic modulus of the material.
Furthermore, the research team observed that the material’s mechanical properties can be improved by over a hundred times through a heating and cooling process that is used to heat-treat metals. They also discovered that this is due to the enforced interactions between micelle and tannic acid arrays.
The research team used the fact that the material shows minimal irritation to the skin and decomposes well in the body to demonstrate its possible application as an adhesive for hair transplantation through an animal experiment. Professor Haeshin Lee, who has pioneered various application fields including medical adhesives, hemostatic agents, and browning shampoo, focused on the adhesive capacities and low toxicity of polyphenols like tannic acid, and now looks forward to it improving the limitations of current hair transplant methods, which still involve follicle transfer and are difficult to be repeated multiple times.
Figure 2. (a) Overview of a hair transplantation method using a biodegradable adhesive (right) compared to a conventional hair transplantation method (left) that transplants hair containing hair follicles. After applying an adhesive to the tip of the hair, it is fixed to the skin by implanting it through a subcutaneous injection, and repeated treatment is possible. (b) Initial animal test results. One day after 15 hair transplantation, 12 strands of hair remain. If you pull the 3 strands of hair, you can see that the whole body is pulled up, indicating that it is firmly implanted into the skin. All strands of hair applied without the new adhesive material fell off, and in the case of adhesive without heat treatment, the efficiency was 1/7.
This research was conducted by first co-authors Dr. Jongmin Park (currently a senior researcher at the Korea Research Institute of Chemical Technology) from Professor Myeongeun Seo’s team and Dr. Eunsook Park from Professor Haeshin Lee’s team in the KAIST Department of Chemistry, and through joint research with the teams led by Professor Hyungjun Kim from the KAIST Department of Chemistry and Professor Siyoung Choi from the Department of Chemical and Biomolecular Engineering. The research was published online on August 22 in the international journal Au (JACS Au) under the title Biodegradable Block Copolymer-Tannic Acid Glue.
This study was funded by the Support Research Under Protection Project of the National Research Foundation (NRF), Leading Research Center Support Project (Research Center for Multiscale Chiral Structure), Biodegradable Plastics Commercialization and Demonstration Project by the Ministry of Trade and Industry, and institutional funding from the Korea Research Institute of Chemical Technology.
2022.10.07 View 10057
KAIST develops biocompatible adhesive applicable to hair transplants
Aside from being used as a new medical adhesive, the new material can be applied to developing a new method of hair transplants, which cannot be repeated multiple times using current method of implanting the wholly intact follicles into the skin.
Medical adhesives are materials that can be applied to various uses such as wound healing, hemostasis, vascular anastomosis, and tissue engineering, and is expected to contribute greatly to the development of minimally invasive surgery and organ transplants. However, adhesives with high adhesion, low toxicity, and capable of decomposing in the body are rare. Adhesives based on natural proteins, such as fibrin and collagen, have high biocompatibility but insufficient adhesive strength. Synthetic polymer adhesives based on urethane or acrylic have greater adhesion but do not decompose well and may cause an inflammatory reaction in the body.
A joint research team led by Professor Myungeun Seo and Professor Haeshin Lee from the KAIST Department of Chemistry developed a bio-friendly adhesive from biocompatible polymers using tannic acid, the source of astringency in wine.
The research team focused on tannic acid, a natural polyphenolic product. Tannic acid is a polyphenol present in large amounts in fruit peels, nuts, and cacao. It has a high affinity and coating ability on other substances, and we sense the astringent taste in wine when tannic acid sticks to the surface of our tongue. When tannic acid is mixed with hydrophilic polymers, they form coacervates, or small droplets of jelly-like fluids that sink. If the polymers used are biocompatible, the mixture can be applied as a medical adhesive with low toxicity. However, coacervates are fundamentally fluid-like and cannot withstand large forces, which limits their adhesive capabilities. Thus, while research to utilize it as an adhesive has been actively discussed, a biodegradable material exhibiting strong adhesion due to its high shear strength has not yet been developed.
The research team figured out a way to enhance adhesion by mixing two biocompatible FDA-approved polymers, polyethylene glycol (PEG) and polylactic acid (PLA). While PEG, which is used widely in eyedrops and cream, is hydrophilic, PLA, a well-known bioplastic derived from lactic acid, is insoluble in water. The team combined the two into a block copolymer, which forms hydrophilic PLA aggregates in water with PEG blocks surrounding them. A coacervate created by mixing the micelles and tannic acid would behave like a solid due to the hard PLA components, and show an elastic modulus improved by a thousand times compared to PEG, enabling it to withstand much greater force as an adhesive.
Figure 1. (Above) Principle of biodegradable adhesive made by mixing poly(ethylene glycol)-poly(lactic acid) diblock copolymer and tannic acid in water. Yellow coacervate is precipitated through hydrogen bonding between the block copolymer micelles and tannic acid, and exhibits adhesion. After heat treatment, hydrogen bonds are rearranged to further improve adhesion. (Bottom) Adhesion comparison. Compared to using poly(ethylene glycol) polymer (d), it can support 10 times more weight when using block copolymer (e) and 60 times more weight after heat treatment (f). The indicated G' values represent the elastic modulus of the material.
Furthermore, the research team observed that the material’s mechanical properties can be improved by over a hundred times through a heating and cooling process that is used to heat-treat metals. They also discovered that this is due to the enforced interactions between micelle and tannic acid arrays.
The research team used the fact that the material shows minimal irritation to the skin and decomposes well in the body to demonstrate its possible application as an adhesive for hair transplantation through an animal experiment. Professor Haeshin Lee, who has pioneered various application fields including medical adhesives, hemostatic agents, and browning shampoo, focused on the adhesive capacities and low toxicity of polyphenols like tannic acid, and now looks forward to it improving the limitations of current hair transplant methods, which still involve follicle transfer and are difficult to be repeated multiple times.
Figure 2. (a) Overview of a hair transplantation method using a biodegradable adhesive (right) compared to a conventional hair transplantation method (left) that transplants hair containing hair follicles. After applying an adhesive to the tip of the hair, it is fixed to the skin by implanting it through a subcutaneous injection, and repeated treatment is possible. (b) Initial animal test results. One day after 15 hair transplantation, 12 strands of hair remain. If you pull the 3 strands of hair, you can see that the whole body is pulled up, indicating that it is firmly implanted into the skin. All strands of hair applied without the new adhesive material fell off, and in the case of adhesive without heat treatment, the efficiency was 1/7.
This research was conducted by first co-authors Dr. Jongmin Park (currently a senior researcher at the Korea Research Institute of Chemical Technology) from Professor Myeongeun Seo’s team and Dr. Eunsook Park from Professor Haeshin Lee’s team in the KAIST Department of Chemistry, and through joint research with the teams led by Professor Hyungjun Kim from the KAIST Department of Chemistry and Professor Siyoung Choi from the Department of Chemical and Biomolecular Engineering. The research was published online on August 22 in the international journal Au (JACS Au) under the title Biodegradable Block Copolymer-Tannic Acid Glue.
This study was funded by the Support Research Under Protection Project of the National Research Foundation (NRF), Leading Research Center Support Project (Research Center for Multiscale Chiral Structure), Biodegradable Plastics Commercialization and Demonstration Project by the Ministry of Trade and Industry, and institutional funding from the Korea Research Institute of Chemical Technology.
2022.10.07 View 10057