Neurobiology
-
 A Single, Master Switch for Sugar Levels?
When a fly eats sugar, a single brain cell sends simultaneous messages to stimulate one hormone and inhibit another to control glucose levels in the body. Further research into this control system with remarkable precision could shed light on the neural mechanisms of diabetes and obesity in humans .
A single neuron appears to monitor and control sugar levels in the fly body, according to research published this week in Nature. This new insight into the mechanisms in the fly brain that maintain a balance of two key hormones controlling glucose levels, insulin and glucagon, can provide a framework for understanding diabetes and obesity in humans.
Neurons that sense and respond to glucose were identified more than 50 years ago, but what they do in our body has remained unclear. Researchers at the Korea Advanced Institute of Science and Technology (KAIST) and New York University School of Medicine have now found a single “glucose-sensing neuron” that appears to be the master controller in Drosophila, the vinegar fly, for maintaining an ideal glucose balance, called homeostasis.
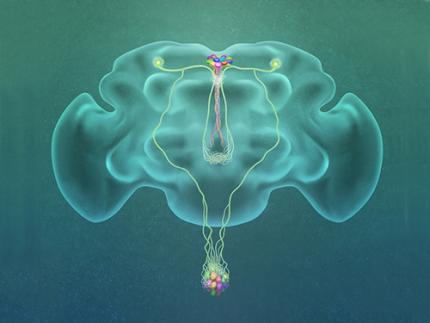
Professor Greg Seong-Bae Suh, Dr. Yangkyun Oh and colleagues identified a key neuron that is excited by glucose, which they called CN neuron. This CN neuron has a unique shape – it has an axon (which is used to transmit information to downstream cells) that is bifurcated. One branch projects to insulin-producing cells, and sends a signal triggering the secretion of the insulin equivalent in flies. The other branch projects to glucagon-producing cells and sends a signal inhibiting the secretion of the glucagon equivalent.
When flies consume food, the levels of glucose in their body increase; this excites the CN neuron, which fires the simultaneous signals to stimulate insulin and inhibit glucagon secretion, thereby maintaining the appropriate balance between the hormones and sugar in the blood. The researchers were able to see this happening in the brain in real time by using a combination of cutting-edge fluorescent calcium imaging technology, as well as measuring hormone and sugar levels and applying highly sophisticated molecular genetic techniques.
When flies were not fed, however, the researchers observed a reduction in the activity of CN neuron, a reduction in insulin secretion and an increase in glucagon secretion. These findings indicate that these key hormones are under the direct control of the glucose-sensing neuron. Furthermore, when they silenced the CN neuron rendering dysfunctional CN neuron in flies, these animals experienced an imbalance, resulting in hyperglycemia – high levels of sugars in the blood, similar to what is observed in diabetes in humans. This further suggests that the CN neuron is critical to maintaining glucose homeostasis in animals.
While further research is required to investigate this process in humans, Suh notes this is a significant step forward in the fields of both neurobiology and endocrinology.
“This work lays the foundation for translational research to better understand how this delicate regulatory process is affected by diabetes, obesity, excessive nutrition and diets high in sugar,” Suh said.
Profile: Greg Seong-Bae Suh
seongbaesuh@kaist.ac.kr
Professor Department of Biological Sciences
KAIST
(Figure: A single glucose-excited CN neuron extends bifurcated axonal branches,
one of which innervates insulin producing cells and stimulates their activity an the other axonal branch projects to glucagon producing cells and inhibits their activity.)
2019.10.24 View 18874
A Single, Master Switch for Sugar Levels?
When a fly eats sugar, a single brain cell sends simultaneous messages to stimulate one hormone and inhibit another to control glucose levels in the body. Further research into this control system with remarkable precision could shed light on the neural mechanisms of diabetes and obesity in humans .
A single neuron appears to monitor and control sugar levels in the fly body, according to research published this week in Nature. This new insight into the mechanisms in the fly brain that maintain a balance of two key hormones controlling glucose levels, insulin and glucagon, can provide a framework for understanding diabetes and obesity in humans.
Neurons that sense and respond to glucose were identified more than 50 years ago, but what they do in our body has remained unclear. Researchers at the Korea Advanced Institute of Science and Technology (KAIST) and New York University School of Medicine have now found a single “glucose-sensing neuron” that appears to be the master controller in Drosophila, the vinegar fly, for maintaining an ideal glucose balance, called homeostasis.
Professor Greg Seong-Bae Suh, Dr. Yangkyun Oh and colleagues identified a key neuron that is excited by glucose, which they called CN neuron. This CN neuron has a unique shape – it has an axon (which is used to transmit information to downstream cells) that is bifurcated. One branch projects to insulin-producing cells, and sends a signal triggering the secretion of the insulin equivalent in flies. The other branch projects to glucagon-producing cells and sends a signal inhibiting the secretion of the glucagon equivalent.
When flies consume food, the levels of glucose in their body increase; this excites the CN neuron, which fires the simultaneous signals to stimulate insulin and inhibit glucagon secretion, thereby maintaining the appropriate balance between the hormones and sugar in the blood. The researchers were able to see this happening in the brain in real time by using a combination of cutting-edge fluorescent calcium imaging technology, as well as measuring hormone and sugar levels and applying highly sophisticated molecular genetic techniques.
When flies were not fed, however, the researchers observed a reduction in the activity of CN neuron, a reduction in insulin secretion and an increase in glucagon secretion. These findings indicate that these key hormones are under the direct control of the glucose-sensing neuron. Furthermore, when they silenced the CN neuron rendering dysfunctional CN neuron in flies, these animals experienced an imbalance, resulting in hyperglycemia – high levels of sugars in the blood, similar to what is observed in diabetes in humans. This further suggests that the CN neuron is critical to maintaining glucose homeostasis in animals.
While further research is required to investigate this process in humans, Suh notes this is a significant step forward in the fields of both neurobiology and endocrinology.
“This work lays the foundation for translational research to better understand how this delicate regulatory process is affected by diabetes, obesity, excessive nutrition and diets high in sugar,” Suh said.
Profile: Greg Seong-Bae Suh
seongbaesuh@kaist.ac.kr
Professor Department of Biological Sciences
KAIST
(Figure: A single glucose-excited CN neuron extends bifurcated axonal branches,
one of which innervates insulin producing cells and stimulates their activity an the other axonal branch projects to glucagon producing cells and inhibits their activity.)
2019.10.24 View 18874 -
 Accurate Detection of Low-Level Somatic Mutation in Intractable Epilepsy
KAIST medical scientists have developed an advanced method for perfectly detecting low-level somatic mutation in patients with intractable epilepsy. Their study showed that deep sequencing replicates of major focal epilepsy genes accurately and efficiently identified low-level somatic mutations in intractable epilepsy.
According to the study, their diagnostic method could increase the accuracy up to 100%, unlike the conventional sequencing analysis, which stands at about 30% accuracy. This work was published in Acta Neuropathologica.
Epilepsy is a neurological disorder common in children. Approximately one third of child patients are diagnosed with intractable epilepsy despite adequate anti-epileptic medication treatment.
Somatic mutations in mTOR pathway genes, SLC35A2, and BRAF are the major genetic causes of intractable epilepsies. A clinical trial to target Focal Cortical Dysplasia type II (FCDII), the mTOR inhibitor is underway at Severance Hospital, their collaborator in Seoul, Korea. However, it is difficult to detect such somatic mutations causing intractable epilepsy because their mutational burden is less than 5%, which is similar to the level of sequencing artifacts. In the clinical field, this has remained a standing challenge for the genetic diagnosis of somatic mutations in intractable epilepsy.
Professor Jeong Ho Lee’s team at the Graduate School of Medical Science and Engineering analyzed paired brain and peripheral tissues from 232 intractable epilepsy patients with various brain pathologies at Severance Hospital using deep sequencing and extracted the major focal epilepsy genes.
They narrowed down target genes to eight major focal epilepsy genes, eliminating almost all of the false positive calls using deep targeted sequencing. As a result, the advanced method robustly increased the accuracy and enabled them to detect low-level somatic mutations in unmatched Formalin Fixed Paraffin Embedded (FFPE) brain samples, the most clinically relevant samples.
Professor Lee conducted this study in collaboration with Professor Dong Suk Kim and Hoon-Chul Kang at Severance Hospital of Yonsei University. He said, “This advanced method of genetic analysis will improve overall patient care by providing more comprehensive genetic counseling and informing decisions on alternative treatments.”
Professor Lee has investigated low-level somatic mutations arising in the brain for a decade. He is developing innovative diagnostics and therapeutics for untreatable brain disorders including intractable epilepsy and glioblastoma at a tech-startup called SoVarGen. “All of the technologies we used during the research were transferred to the company. This research gave us very good momentum to reach the next phase of our startup,” he remarked.
The work was supported by grants from the Suh Kyungbae Foundation, a National Research Foundation of Korea grant funded by the Ministry of Science and ICT, the Korean Health Technology R&D Project from the Ministry of Health & Welfare, and the Netherlands Organization for Health Research and Development.
(Figure: Landscape of somatic and germline mutations identified in intractable epilepsy patients. a Signaling pathways for all of the mutated genes identified in this study. Bold: somatic mutation, Regular: germline mutation. b The distribution of variant allelic frequencies (VAFs) of identified somatic mutations. c The detecting rate and types of identified mutations according to histopathology. Yellow: somatic mutations, green: two-hit mutations, grey: germline mutations.)
2019.08.14 View 30438
Accurate Detection of Low-Level Somatic Mutation in Intractable Epilepsy
KAIST medical scientists have developed an advanced method for perfectly detecting low-level somatic mutation in patients with intractable epilepsy. Their study showed that deep sequencing replicates of major focal epilepsy genes accurately and efficiently identified low-level somatic mutations in intractable epilepsy.
According to the study, their diagnostic method could increase the accuracy up to 100%, unlike the conventional sequencing analysis, which stands at about 30% accuracy. This work was published in Acta Neuropathologica.
Epilepsy is a neurological disorder common in children. Approximately one third of child patients are diagnosed with intractable epilepsy despite adequate anti-epileptic medication treatment.
Somatic mutations in mTOR pathway genes, SLC35A2, and BRAF are the major genetic causes of intractable epilepsies. A clinical trial to target Focal Cortical Dysplasia type II (FCDII), the mTOR inhibitor is underway at Severance Hospital, their collaborator in Seoul, Korea. However, it is difficult to detect such somatic mutations causing intractable epilepsy because their mutational burden is less than 5%, which is similar to the level of sequencing artifacts. In the clinical field, this has remained a standing challenge for the genetic diagnosis of somatic mutations in intractable epilepsy.
Professor Jeong Ho Lee’s team at the Graduate School of Medical Science and Engineering analyzed paired brain and peripheral tissues from 232 intractable epilepsy patients with various brain pathologies at Severance Hospital using deep sequencing and extracted the major focal epilepsy genes.
They narrowed down target genes to eight major focal epilepsy genes, eliminating almost all of the false positive calls using deep targeted sequencing. As a result, the advanced method robustly increased the accuracy and enabled them to detect low-level somatic mutations in unmatched Formalin Fixed Paraffin Embedded (FFPE) brain samples, the most clinically relevant samples.
Professor Lee conducted this study in collaboration with Professor Dong Suk Kim and Hoon-Chul Kang at Severance Hospital of Yonsei University. He said, “This advanced method of genetic analysis will improve overall patient care by providing more comprehensive genetic counseling and informing decisions on alternative treatments.”
Professor Lee has investigated low-level somatic mutations arising in the brain for a decade. He is developing innovative diagnostics and therapeutics for untreatable brain disorders including intractable epilepsy and glioblastoma at a tech-startup called SoVarGen. “All of the technologies we used during the research were transferred to the company. This research gave us very good momentum to reach the next phase of our startup,” he remarked.
The work was supported by grants from the Suh Kyungbae Foundation, a National Research Foundation of Korea grant funded by the Ministry of Science and ICT, the Korean Health Technology R&D Project from the Ministry of Health & Welfare, and the Netherlands Organization for Health Research and Development.
(Figure: Landscape of somatic and germline mutations identified in intractable epilepsy patients. a Signaling pathways for all of the mutated genes identified in this study. Bold: somatic mutation, Regular: germline mutation. b The distribution of variant allelic frequencies (VAFs) of identified somatic mutations. c The detecting rate and types of identified mutations according to histopathology. Yellow: somatic mutations, green: two-hit mutations, grey: germline mutations.)
2019.08.14 View 30438 -
 Deciphering Brain Somatic Mutations Associated with Alzheimer's Disease
Researchers have found a potential link between non-inherited somatic mutations in the brain and the progression of Alzheimer’s disease
Researchers have identified somatic mutations in the brain that could contribute to the development of Alzheimer’s disease (AD). Their findings were published in the journal Nature Communications last week.
Decades worth of research has identified inherited mutations that lead to early-onset familial AD. Inherited mutations, however, are behind at most half the cases of late onset sporadic AD, in which there is no family history of the disease. But the genetic factors causing the other half of these sporadic cases have been unclear.
Professor Jeong Ho Lee at the Graduate School of Medical Science and Engineering and colleagues analysed the DNA present in post-mortem hippocampal formations and in blood samples from people aged 70 to 96 with AD and age-matched controls. They specifically looked for non-inherited somatic mutations in their brains using high-depth whole exome sequencing.
The team developed a bioinformatics pipeline that enabled them to detect low-level brain somatic single nucleotide variations (SNVs) – mutations that involve the substitution of a single nucleotide with another nucleotide. Brain somatic SNVs have been reported on and accumulate throughout our lives and can sometimes be associated with a range of neurological diseases.
The number of somatic SNVs did not differ between individuals with AD and non-demented controls. Interestingly, somatic SNVs in AD brains arise about 4.8 times more slowly than in blood. When the team performed gene-set enrichment tests, 26.9 percent of the AD brain samples had pathogenic brain somatic SNVs known to be linked to hyperphosphorylation of tau proteins, which is one of major hallmarks of AD.
Then, they pinpointed a pathogenic SNV in the PIN1 gene, a cis/trans isomerase that balances phosphorylation in tau proteins, found in one AD patient’s brain. They found the mutation was 4.9 time more abundant in AT8-positive – a marker for hyper-phosphorylated tau proteins– neurons in the entorhinal cortex than the bulk hippocampal tissue. Furthermore, in a series of functional assays, they observed the mutation causing a loss of function in PIN1 and such haploinsufficiency increased the phosphorylation and aggregation of tau proteins.
“Our study provides new insights into the molecular genetic factors behind Alzheimer’s disease and other neurodegenerative diseases potentially linked to somatic mutations in the brain,” said Professor Lee.
The team is planning to expand their study to a larger cohort in order to establish stronger links between these brain somatic mutations and the pathogenesis of Alzheimer’s disease.
(Figure 1. Bioinformatic pipeline for detecting low-level brain somatic mutations in AD and non-AD.)
(Figure 2. Pathogenic brain somatic mutations associated with tau phosphorylation are significantly enriched in AD brains.)
(Figure 3. A pathogenic brain somatic mutation in PIN1 (c. 477 C>T) is a loss-of-function and related functional assays show its haploinsufficiency increases phosphorylation and aggregation of tau.)
2019.07.19 View 37464
Deciphering Brain Somatic Mutations Associated with Alzheimer's Disease
Researchers have found a potential link between non-inherited somatic mutations in the brain and the progression of Alzheimer’s disease
Researchers have identified somatic mutations in the brain that could contribute to the development of Alzheimer’s disease (AD). Their findings were published in the journal Nature Communications last week.
Decades worth of research has identified inherited mutations that lead to early-onset familial AD. Inherited mutations, however, are behind at most half the cases of late onset sporadic AD, in which there is no family history of the disease. But the genetic factors causing the other half of these sporadic cases have been unclear.
Professor Jeong Ho Lee at the Graduate School of Medical Science and Engineering and colleagues analysed the DNA present in post-mortem hippocampal formations and in blood samples from people aged 70 to 96 with AD and age-matched controls. They specifically looked for non-inherited somatic mutations in their brains using high-depth whole exome sequencing.
The team developed a bioinformatics pipeline that enabled them to detect low-level brain somatic single nucleotide variations (SNVs) – mutations that involve the substitution of a single nucleotide with another nucleotide. Brain somatic SNVs have been reported on and accumulate throughout our lives and can sometimes be associated with a range of neurological diseases.
The number of somatic SNVs did not differ between individuals with AD and non-demented controls. Interestingly, somatic SNVs in AD brains arise about 4.8 times more slowly than in blood. When the team performed gene-set enrichment tests, 26.9 percent of the AD brain samples had pathogenic brain somatic SNVs known to be linked to hyperphosphorylation of tau proteins, which is one of major hallmarks of AD.
Then, they pinpointed a pathogenic SNV in the PIN1 gene, a cis/trans isomerase that balances phosphorylation in tau proteins, found in one AD patient’s brain. They found the mutation was 4.9 time more abundant in AT8-positive – a marker for hyper-phosphorylated tau proteins– neurons in the entorhinal cortex than the bulk hippocampal tissue. Furthermore, in a series of functional assays, they observed the mutation causing a loss of function in PIN1 and such haploinsufficiency increased the phosphorylation and aggregation of tau proteins.
“Our study provides new insights into the molecular genetic factors behind Alzheimer’s disease and other neurodegenerative diseases potentially linked to somatic mutations in the brain,” said Professor Lee.
The team is planning to expand their study to a larger cohort in order to establish stronger links between these brain somatic mutations and the pathogenesis of Alzheimer’s disease.
(Figure 1. Bioinformatic pipeline for detecting low-level brain somatic mutations in AD and non-AD.)
(Figure 2. Pathogenic brain somatic mutations associated with tau phosphorylation are significantly enriched in AD brains.)
(Figure 3. A pathogenic brain somatic mutation in PIN1 (c. 477 C>T) is a loss-of-function and related functional assays show its haploinsufficiency increases phosphorylation and aggregation of tau.)
2019.07.19 View 37464 -
 New Bio-Clock gene and its function found
The Ministry of Education, Science and Technology announced that a Korean research team has found a new gene responsible for maintaining the bio-clock (twenty-four) and its mechanism.
Twnety-four was led by Professor Choi Joon Ho and Dr. Lee Jong Bin of KAIST (department of Biology) and was a joint operation with Professor Ravi Allada and Dr.Lim Jeong Hoon of Northwestern University (department of neurobiology) and the result was published in ‘Nature’ magazine.
The research team experimented with transformed small fruit flies for 4 years and found that there was an undiscovered gene that deals with the bio rhythm in the brain which they named ‘twenty-four’.
The understanding with genes prior to twenty-four was that these genes regulate biorhythm in the transcription phase (DNA to mRNA). Twenty-four operates in the step after transcription when the ribosome creates proteins. Especially twenty-four has a great effect on the ‘period protein’ which acts as a sub-atomic clock that regulates the rhythm and life of each cell.
The experiment was innovational in that it was able to scientifically prove the function of the protein produced by the gene.
The result is expected to help solve the problems associated with sleep disorders, jetlags, eating rhythms, bio rhythms, etc.
The name twenty-four was the fact that a day, a cycle, is 24 hours long and the gene’s serial numbers CG4857 adds up to twenty four.
2011.02.23 View 14252
New Bio-Clock gene and its function found
The Ministry of Education, Science and Technology announced that a Korean research team has found a new gene responsible for maintaining the bio-clock (twenty-four) and its mechanism.
Twnety-four was led by Professor Choi Joon Ho and Dr. Lee Jong Bin of KAIST (department of Biology) and was a joint operation with Professor Ravi Allada and Dr.Lim Jeong Hoon of Northwestern University (department of neurobiology) and the result was published in ‘Nature’ magazine.
The research team experimented with transformed small fruit flies for 4 years and found that there was an undiscovered gene that deals with the bio rhythm in the brain which they named ‘twenty-four’.
The understanding with genes prior to twenty-four was that these genes regulate biorhythm in the transcription phase (DNA to mRNA). Twenty-four operates in the step after transcription when the ribosome creates proteins. Especially twenty-four has a great effect on the ‘period protein’ which acts as a sub-atomic clock that regulates the rhythm and life of each cell.
The experiment was innovational in that it was able to scientifically prove the function of the protein produced by the gene.
The result is expected to help solve the problems associated with sleep disorders, jetlags, eating rhythms, bio rhythms, etc.
The name twenty-four was the fact that a day, a cycle, is 24 hours long and the gene’s serial numbers CG4857 adds up to twenty four.
2011.02.23 View 14252