microbial
-
 KAIST Accelerates Synthetic Microbe Design by Discovering Novel Enzymes Using AI
< (From left) Professor Sang Yup Lee of the Department of Chemical and Biomolecular Engineering (top), Hongkeun Ji, PhD candidate of the Department of Chemical and Biomolecular Engineering (top), Ha Rim Kim, PhD candidate of the Department of Chemical and Biomolecular Engineering, and Dr. Gi Bae Kim of the BioProcess Engineering Research Center >
Enzymes are proteins that catalyze biochemical reactions within cells and play a pivotal role in metabolic processes. Accordingly, identifying the functions of novel enzymes is a critical task in the construction of microbial cell factories.
A KAIST research team has leveraged artificial intelligence (AI) to design novel enzymes that do not exist in nature, significantly accelerating microbial cell factory development and boosting the potential for next-generation biotechnological applications such as drug development and biofuel production.
KAIST (represented by President Kwang-Hyung Lee) announced on the 21st of April that Distinguished Professor Sang Yup Lee and his team from the Department of Chemical and Biomolecular Engineering have published a review titled “Enzyme Functional Classification Using Artificial Intelligence,” which outlines the advancement of AI-based enzyme function prediction technologies and analyzes how AI has contributed to the discovery and design of new enzymes.
Professor Lee’s team systematically reviewed the development of enzyme function prediction technologies utilizing machine learning and deep learning, offering a comprehensive analysis.
From sequence similarity-based prediction methods to the integration of convolutional neural networks (CNNs), recurrent neural networks (RNNs), graph neural networks (GNNs), and transformer-based large language models, the paper covers a broad range of AI applications. It analyzes how these technologies extract meaningful information from protein sequences and enhance prediction accuracy.
In particular, enzyme function prediction using deep learning goes beyond simple sequence similarity analysis. By automatically extracting structural and evolutionary features embedded in amino acid sequences, deep learning enables more precise predictions of catalytic functions.
This highlights the unique advantages of AI models compared to traditional bioinformatics approaches.
Moreover, the review suggests that the advancement of generative AI will move future research beyond predicting existing functions to generating entirely new enzymes with functions not found in nature. This shift is expected to profoundly impact the trajectory of biotechnology and synthetic biology.
< Figure 1. Extraction of enzyme characteristics and function prediction using various deep learning structures >
Ha Rim Kim, a Ph.D. candidate and co-first author from the Department of Chemical and Biomolecular Engineering, stated, “AI-based enzyme function prediction and enzyme design are highly important across various fields including metabolic engineering, synthetic biology, and healthcare.”
Distinguished Professor Sang Yup Lee added, “AI-powered enzyme function prediction shows the potential to solve diverse biological problems and will significantly contribute to accelerating research across the entire field.”
The review was published on March 28 in Trends in Biotechnology, a leading biotechnology journal issued by Cell Press.
※ Title: Enzyme Functional Classification Using Artificial Intelligence
※DOI: https://doi.org/10.1016/j.tibtech.2025.03.003
※ Author Information: Ha Rim Kim (KAIST, Co-first author), Hongkeun Ji (KAIST, Co-first author), Gi Bae Kim (KAIST, Third author), Sang Yup Lee (KAIST, Corresponding author)
This research was supported by the Ministry of Science and ICT under the project Development of Core Technologies for Advanced Synthetic Biology to Lead the Bio-Manufacturing Industry (aimed at replacing petroleum-based chemicals), and also by joint support from the Ministry of Science and ICT and the Ministry of Health and Welfare for the project Development of Novel Antibiotic Structures Using Deep Learning-Based Synthetic Biology.
2025.04.07 View 3283
KAIST Accelerates Synthetic Microbe Design by Discovering Novel Enzymes Using AI
< (From left) Professor Sang Yup Lee of the Department of Chemical and Biomolecular Engineering (top), Hongkeun Ji, PhD candidate of the Department of Chemical and Biomolecular Engineering (top), Ha Rim Kim, PhD candidate of the Department of Chemical and Biomolecular Engineering, and Dr. Gi Bae Kim of the BioProcess Engineering Research Center >
Enzymes are proteins that catalyze biochemical reactions within cells and play a pivotal role in metabolic processes. Accordingly, identifying the functions of novel enzymes is a critical task in the construction of microbial cell factories.
A KAIST research team has leveraged artificial intelligence (AI) to design novel enzymes that do not exist in nature, significantly accelerating microbial cell factory development and boosting the potential for next-generation biotechnological applications such as drug development and biofuel production.
KAIST (represented by President Kwang-Hyung Lee) announced on the 21st of April that Distinguished Professor Sang Yup Lee and his team from the Department of Chemical and Biomolecular Engineering have published a review titled “Enzyme Functional Classification Using Artificial Intelligence,” which outlines the advancement of AI-based enzyme function prediction technologies and analyzes how AI has contributed to the discovery and design of new enzymes.
Professor Lee’s team systematically reviewed the development of enzyme function prediction technologies utilizing machine learning and deep learning, offering a comprehensive analysis.
From sequence similarity-based prediction methods to the integration of convolutional neural networks (CNNs), recurrent neural networks (RNNs), graph neural networks (GNNs), and transformer-based large language models, the paper covers a broad range of AI applications. It analyzes how these technologies extract meaningful information from protein sequences and enhance prediction accuracy.
In particular, enzyme function prediction using deep learning goes beyond simple sequence similarity analysis. By automatically extracting structural and evolutionary features embedded in amino acid sequences, deep learning enables more precise predictions of catalytic functions.
This highlights the unique advantages of AI models compared to traditional bioinformatics approaches.
Moreover, the review suggests that the advancement of generative AI will move future research beyond predicting existing functions to generating entirely new enzymes with functions not found in nature. This shift is expected to profoundly impact the trajectory of biotechnology and synthetic biology.
< Figure 1. Extraction of enzyme characteristics and function prediction using various deep learning structures >
Ha Rim Kim, a Ph.D. candidate and co-first author from the Department of Chemical and Biomolecular Engineering, stated, “AI-based enzyme function prediction and enzyme design are highly important across various fields including metabolic engineering, synthetic biology, and healthcare.”
Distinguished Professor Sang Yup Lee added, “AI-powered enzyme function prediction shows the potential to solve diverse biological problems and will significantly contribute to accelerating research across the entire field.”
The review was published on March 28 in Trends in Biotechnology, a leading biotechnology journal issued by Cell Press.
※ Title: Enzyme Functional Classification Using Artificial Intelligence
※DOI: https://doi.org/10.1016/j.tibtech.2025.03.003
※ Author Information: Ha Rim Kim (KAIST, Co-first author), Hongkeun Ji (KAIST, Co-first author), Gi Bae Kim (KAIST, Third author), Sang Yup Lee (KAIST, Corresponding author)
This research was supported by the Ministry of Science and ICT under the project Development of Core Technologies for Advanced Synthetic Biology to Lead the Bio-Manufacturing Industry (aimed at replacing petroleum-based chemicals), and also by joint support from the Ministry of Science and ICT and the Ministry of Health and Welfare for the project Development of Novel Antibiotic Structures Using Deep Learning-Based Synthetic Biology.
2025.04.07 View 3283 -
 KAIST Develops Eco-Friendly, Nylon-Like Plastic Using Microorganisms
Poly(ester amide) amide is a next-generation material that combines the advantages of PET (polyester) and nylon (polyamide), two widely used plastics. However, it could only be produced from fossil fuels, which posed environmental concerns. Using microorganisms, KAIST researchers have successfully developed a new bio-based plastic to replace conventional plastic.
KAIST (represented by President Kwang Hyung Lee) announced on the 20th of March that a research team led by Distinguished Professor Sang Yup Lee from the Department of Chemical and Biomolecular Engineering has developed microbial strains through systems metabolic engineering to produce various eco-friendly, bio-based poly(ester amide)s. The team collaborated with researchers from the Korea Research Institute of Chemical Technology (KRICT, President Young-Kook Lee) to analyze and confirm the properties of the resulting plastic.
Professor Sang Yup Lee’s research team designed new metabolic pathways that do not naturally exist in microorganisms, and developed a platform microbial strain capable of producing nine different types of poly(ester amide)s, including poly(3-hydroxybutyrate-ran-3-aminopropionate) and poly(3-hydroxybutyrate-ran-4-aminobutyrate).
Using glucose derived from abundant biomass sources such as waste wood and weeds, the team successfully produced poly(ester amide)s in an eco-friendly manner. The researchers also confirmed the potential for industrial-scale production by demonstrating high production efficiency (54.57 g/L) using fed-batch fermentation of the engineered strain.
In collaboration with researchers Haemin Jeong and Jihoon Shin from KRICT, the KAIST team analyzed the properties of the bio-based plastic and found that it exhibited characteristics similar to high-density polyethylene (HDPE). This means the new plastic is not only eco-friendly but also strong and durable enough to replace conventional plastics.
The engineered strains and strategies developed in this study are expected to be useful not only for producing various poly(ester amide)s but also for constructing metabolic pathways for the biosynthesis of other types of polymers.
Professor Sang Yup Lee stated, “This study is the first to demonstrate the possibility of producing poly(ester amide)s (plastics) through a renewable bio-based chemical process rather than relying on the petroleum-based chemical industry. We plan to further enhance the production yield and efficiency through continued research.”
The study was published online on March 17 in the international journal Nature Chemical Biology.
·Title: Biosynthesis of poly(ester amide)s in engineered Escherichia coli
·DOI: 10.1038/s41589-025-01842-2
·Authors: A total of seven authors including Tong Un Chae (KAIST, first author), So Young Choi (KAIST, second author), Da-Hee Ahn (KAIST, third author), Woo Dae Jang (KAIST, fourth author), Haemin Jeong (KRICT, fifth author), Jihoon Shin (KRICT, sixth author), and Sang Yup Lee (KAIST, corresponding author).
This research was supported by the Ministry of Science and ICT (MSIT) under the Eco-Friendly Chemical Technology Development Project as part of the "Next-Generation Biorefinery Technology Development to Lead the Bio-Chemical Industry" initiative (project led by Distinguished Professor Sang Yup Lee at KAIST).
2025.03.24 View 4939
KAIST Develops Eco-Friendly, Nylon-Like Plastic Using Microorganisms
Poly(ester amide) amide is a next-generation material that combines the advantages of PET (polyester) and nylon (polyamide), two widely used plastics. However, it could only be produced from fossil fuels, which posed environmental concerns. Using microorganisms, KAIST researchers have successfully developed a new bio-based plastic to replace conventional plastic.
KAIST (represented by President Kwang Hyung Lee) announced on the 20th of March that a research team led by Distinguished Professor Sang Yup Lee from the Department of Chemical and Biomolecular Engineering has developed microbial strains through systems metabolic engineering to produce various eco-friendly, bio-based poly(ester amide)s. The team collaborated with researchers from the Korea Research Institute of Chemical Technology (KRICT, President Young-Kook Lee) to analyze and confirm the properties of the resulting plastic.
Professor Sang Yup Lee’s research team designed new metabolic pathways that do not naturally exist in microorganisms, and developed a platform microbial strain capable of producing nine different types of poly(ester amide)s, including poly(3-hydroxybutyrate-ran-3-aminopropionate) and poly(3-hydroxybutyrate-ran-4-aminobutyrate).
Using glucose derived from abundant biomass sources such as waste wood and weeds, the team successfully produced poly(ester amide)s in an eco-friendly manner. The researchers also confirmed the potential for industrial-scale production by demonstrating high production efficiency (54.57 g/L) using fed-batch fermentation of the engineered strain.
In collaboration with researchers Haemin Jeong and Jihoon Shin from KRICT, the KAIST team analyzed the properties of the bio-based plastic and found that it exhibited characteristics similar to high-density polyethylene (HDPE). This means the new plastic is not only eco-friendly but also strong and durable enough to replace conventional plastics.
The engineered strains and strategies developed in this study are expected to be useful not only for producing various poly(ester amide)s but also for constructing metabolic pathways for the biosynthesis of other types of polymers.
Professor Sang Yup Lee stated, “This study is the first to demonstrate the possibility of producing poly(ester amide)s (plastics) through a renewable bio-based chemical process rather than relying on the petroleum-based chemical industry. We plan to further enhance the production yield and efficiency through continued research.”
The study was published online on March 17 in the international journal Nature Chemical Biology.
·Title: Biosynthesis of poly(ester amide)s in engineered Escherichia coli
·DOI: 10.1038/s41589-025-01842-2
·Authors: A total of seven authors including Tong Un Chae (KAIST, first author), So Young Choi (KAIST, second author), Da-Hee Ahn (KAIST, third author), Woo Dae Jang (KAIST, fourth author), Haemin Jeong (KRICT, fifth author), Jihoon Shin (KRICT, sixth author), and Sang Yup Lee (KAIST, corresponding author).
This research was supported by the Ministry of Science and ICT (MSIT) under the Eco-Friendly Chemical Technology Development Project as part of the "Next-Generation Biorefinery Technology Development to Lead the Bio-Chemical Industry" initiative (project led by Distinguished Professor Sang Yup Lee at KAIST).
2025.03.24 View 4939 -
 KAIST finds ways for Bacteria to produce PET-like materials
Among various eco-friendly polymers, polyhydroxyalkanoates (PHA) stand out for their excellent biodegradability and biocompatibility. They decompose naturally in soil and marine environments and are used in applications such as food packaging and medical products. However, natural PHA produced to date has faced challenges meeting various physical property requirements, such as durability and thermal stability, and has been limited in its commercial application due to low production concentrations. In light of this, KAIST researchers have recently developed a technology that could play a crucial role in solving the environmental pollution problem caused by plastics.
KAIST (represented by President Kwang-Hyung Lee) announced on August 26th that a research team led by Distinguished Professor Sang Yup Lee from the Department of Chemical and Biomolecular Engineering, including Dr. Youngjoon Lee and master's student Minju Kang, has successfully developed a microbial strain that efficiently produces aromatic polyester* using systems metabolic engineering.
※ Aromatic polyester: A polymer containing aromatic compounds (specific carbon ring structures like benzene) and ester bonds.
In this study, the research team used metabolic engineering to enhance the metabolic flux of the biosynthetic pathway for the aromatic monomer phenyllactate (PhLA) in E. coli. They manipulated the metabolic pathway to increase the polymer fraction accumulated within the cells and employed computer simulations to predict the structure of PHA synthase and improve the enzyme based on the structure-function relationship.
Through subsequent fermentation optimization, the team achieved the world’s highest concentration (12.3±0.1 g/L) for the efficient production of poly (PhLA) and successfully produced polyester through a 30L scale fed-batch fermentation, demonstrating the possibility of industrial-level production. The produced aromatic polyesters showed enhanced thermal properties, improved mechanical properties, and potential for use as drug delivery carriers.
< Figure 1. Development schematics of aromatic polyester producing microorganisms >
The research team also demonstrated that an exogenous phasin protein* plays a crucial role in increasing the intracellular polymer accumulation fraction, which is directly related to the economic feasibility and efficiency of non-natural PHA production. They improved PHA synthase using a rational enzyme design approach, predicting the three-dimensional structure of the enzyme through homology modeling (a method of predicting the three-dimensional structure of a new protein based on the structure of similar proteins) followed by molecular docking simulations (simulations that predict how well a monomer can bind to an enzyme) and molecular dynamics simulations (simulations that predict how molecules move and interact over time) to upgrade the enzyme into a mutant enzyme with enhanced monomer polymerization efficiency.
※ Exogenous phasin protein: Phasin is a protein related to PHA production, interacting with the cytoplasmic environment on the surface of granules of PHA, and playing a role in polymer accumulation and controlling the number and size of granules. In this study, genes encoding phasin proteins derived from various natural PHA-producing microorganisms were selected and introduced.
Dr. Youngjoon Lee, co-first author of the paper, explained, "The significance of this study lies in the fact that we have achieved the world's highest concentration of microbial-based aromatic polyester production using eco-friendly materials and methods. This technology is expected to play a crucial role in addressing the environmental pollution caused by plastics." Distinguished Professor Sang Yup Lee added, "This study, which presents various strategies for the high-efficiency production of useful polymers via systems metabolic engineering, is expected to make a significant contribution to solving climate change issues, particularly the recent plastic problem."
< Figure 2. Detailed development strategy for aromatic polyester producing microorganisms >
The research findings were published on August 21st in Trends in Biotechnology, published by Cell, an international academic journal.
※ Paper Title: “Microbial production of an aromatic homopolyester”
※ Author Information: Youngjoon Lee (KAIST, co-first author), Minju Kang (KAIST, co-first author), Woo Dae Jang (KAIST, second author), So Young Choi (KAIST, third author), Jung Eun Yang (KAIST, fourth author), Sang Yup Lee (KAIST, corresponding author), totaling six authors.
This research was supported by the "Development of Next-Generation Biorefinery Platform Technologies for Leading the Bio-based Chemicals Industry" project led by Distinguished Professor Sang Yup Lee at KAIST, under the eco-friendly chemical technology development project aimed at substituting petroleum, funded by the Ministry of Science and ICT. It was also supported by the "Development of Platform Technology for the Production of Novel Aromatic Bioplastic Using Microbial Cell Factories" project (Project Leader: Si Jae Park, Ewha Woman’s University).
2024.08.28 View 6796
KAIST finds ways for Bacteria to produce PET-like materials
Among various eco-friendly polymers, polyhydroxyalkanoates (PHA) stand out for their excellent biodegradability and biocompatibility. They decompose naturally in soil and marine environments and are used in applications such as food packaging and medical products. However, natural PHA produced to date has faced challenges meeting various physical property requirements, such as durability and thermal stability, and has been limited in its commercial application due to low production concentrations. In light of this, KAIST researchers have recently developed a technology that could play a crucial role in solving the environmental pollution problem caused by plastics.
KAIST (represented by President Kwang-Hyung Lee) announced on August 26th that a research team led by Distinguished Professor Sang Yup Lee from the Department of Chemical and Biomolecular Engineering, including Dr. Youngjoon Lee and master's student Minju Kang, has successfully developed a microbial strain that efficiently produces aromatic polyester* using systems metabolic engineering.
※ Aromatic polyester: A polymer containing aromatic compounds (specific carbon ring structures like benzene) and ester bonds.
In this study, the research team used metabolic engineering to enhance the metabolic flux of the biosynthetic pathway for the aromatic monomer phenyllactate (PhLA) in E. coli. They manipulated the metabolic pathway to increase the polymer fraction accumulated within the cells and employed computer simulations to predict the structure of PHA synthase and improve the enzyme based on the structure-function relationship.
Through subsequent fermentation optimization, the team achieved the world’s highest concentration (12.3±0.1 g/L) for the efficient production of poly (PhLA) and successfully produced polyester through a 30L scale fed-batch fermentation, demonstrating the possibility of industrial-level production. The produced aromatic polyesters showed enhanced thermal properties, improved mechanical properties, and potential for use as drug delivery carriers.
< Figure 1. Development schematics of aromatic polyester producing microorganisms >
The research team also demonstrated that an exogenous phasin protein* plays a crucial role in increasing the intracellular polymer accumulation fraction, which is directly related to the economic feasibility and efficiency of non-natural PHA production. They improved PHA synthase using a rational enzyme design approach, predicting the three-dimensional structure of the enzyme through homology modeling (a method of predicting the three-dimensional structure of a new protein based on the structure of similar proteins) followed by molecular docking simulations (simulations that predict how well a monomer can bind to an enzyme) and molecular dynamics simulations (simulations that predict how molecules move and interact over time) to upgrade the enzyme into a mutant enzyme with enhanced monomer polymerization efficiency.
※ Exogenous phasin protein: Phasin is a protein related to PHA production, interacting with the cytoplasmic environment on the surface of granules of PHA, and playing a role in polymer accumulation and controlling the number and size of granules. In this study, genes encoding phasin proteins derived from various natural PHA-producing microorganisms were selected and introduced.
Dr. Youngjoon Lee, co-first author of the paper, explained, "The significance of this study lies in the fact that we have achieved the world's highest concentration of microbial-based aromatic polyester production using eco-friendly materials and methods. This technology is expected to play a crucial role in addressing the environmental pollution caused by plastics." Distinguished Professor Sang Yup Lee added, "This study, which presents various strategies for the high-efficiency production of useful polymers via systems metabolic engineering, is expected to make a significant contribution to solving climate change issues, particularly the recent plastic problem."
< Figure 2. Detailed development strategy for aromatic polyester producing microorganisms >
The research findings were published on August 21st in Trends in Biotechnology, published by Cell, an international academic journal.
※ Paper Title: “Microbial production of an aromatic homopolyester”
※ Author Information: Youngjoon Lee (KAIST, co-first author), Minju Kang (KAIST, co-first author), Woo Dae Jang (KAIST, second author), So Young Choi (KAIST, third author), Jung Eun Yang (KAIST, fourth author), Sang Yup Lee (KAIST, corresponding author), totaling six authors.
This research was supported by the "Development of Next-Generation Biorefinery Platform Technologies for Leading the Bio-based Chemicals Industry" project led by Distinguished Professor Sang Yup Lee at KAIST, under the eco-friendly chemical technology development project aimed at substituting petroleum, funded by the Ministry of Science and ICT. It was also supported by the "Development of Platform Technology for the Production of Novel Aromatic Bioplastic Using Microbial Cell Factories" project (Project Leader: Si Jae Park, Ewha Woman’s University).
2024.08.28 View 6796 -
 KAIST Develops Microbial Liquid Egg Substitute
A team of researchers published a paper on developing a substitute for eggs using microorganisms, grabbing international attention. It is expected that the development of egg substitutes using non-animal raw materials will solve the problems of factory farming, which causes problems like increased emission of greenhouse gas and waste, and contribute to building a sustainable food system that allows easy protein intake.
KAIST (President Kwang-Hyung Lee) announced that Research Professor Kyeong Rok Choi from the Biological Process Research Center and Distinguished Professor Sang Yup Lee from the Department of Chemical and Biomolecular Engineering have published a paper on the development of an "Eco-Friendly Liquid Egg Substitute Derived from Microorganisms."
Eggs play a crucial role in various culinary applications due to their unique physicochemical properties such as gelling, foaming, and emulsifying, while also providing essential nutrients. However, traditional egg production is not only unethical and resource-intensive but also has significant environmental impacts such as greenhouse gas emissions and waste issues. Additionally, factors such as wars and trade regulations have led to significant increases in egg prices, highlighting food security concerns. In response to these issues, there has been growing interest in egg substitutes made from non-animal sources to establish a sustainable food system.
Although there has been progress in developing non-animal protein-based egg substitutes, no substitute has been able to fully replicate the essential functional properties of liquid eggs, such as gelling and foaming, while also providing complete nutrition. In this context, the research team aimed to develop a liquid egg substitute using microbial biomass, which has a protein content comparable to that of meat per unit dry mass. Various microorganisms, such as yeast, Bacillus, lactic acid bacteria, and other probiotics, have been proven safe through long-term human consumption. Microbial biomass requires fewer resources like water and land during production, and possesses high-quality nutrients, making it a promising sustainable food resource.
< Figure 1. Comparison of heat treatment results of microbial pellets and microbial lysates >
However, the semi-solid microbial biomass recovered through microbial cultivation was observed to turn liquid upon heating, unlike liquid egg. To address this, the research team devised a microbial lysate by breaking down the cell walls and cell membranes of microorganisms, which correspond to the eggshell. They found that the microbial lysate's proteins coagulated when heated and formed a gel similar to that of liquid egg. The gel formed from the heated microbial lysate was found to have microscopic structures and physical properties similar to those of boiled eggs. The addition of microbial-derived edible enzymes or plant-based materials allowed for the adjustment of its properties, enabling the creation of various textures.
Furthermore, the researchers demonstrated that the microbial lysate could form stable foams widely used in baking, such as meringues (made from egg whites). They successfully baked meringue cookies using this lysate, showing its potential as a functional liquid egg substitute.
Distinguished Professor Sang Yup Lee stated, "This substitute has excellent nutritional components, making it suitable for regular food consumption. It is especially promising as emergency food for long-term space travel, wartime situations, and other emergencies. More importantly, it contributes to securing a sustainable food system."
< Figure 2. Example of foaming ability of microbial lysate and meringue cookie production >
< Figure 3. Example of foaming ability of microbial lysate and meringue cookie production >
The paper was published online in the journal npj Science of Food, issued by Nature.
- Paper Title: Microbial lysates repurposed as liquid egg substitutes
- Authors: Kyeong Rok Choi (first author), Da-Hee Ahn, Seok Yeong Jung, YuHyun Lee, and Sang Yup Lee (corresponding author)
This research was supported by the Ministry of Science and ICT's project for developing eco-friendly chemical technologies to replace petroleum (Project Leader: Distinguished Professor Sang Yup Lee, KAIST) and the Rural Development Administration's Agricultural Microorganisms Project Group (Director: Professor Pan-sik Jang, Seoul National University) for developing protein production technology from inorganic substances through microbial metabolic system control (Project Leader: Research Professor Kyeong Rok Choi, KAIST).
2024.07.05 View 7007
KAIST Develops Microbial Liquid Egg Substitute
A team of researchers published a paper on developing a substitute for eggs using microorganisms, grabbing international attention. It is expected that the development of egg substitutes using non-animal raw materials will solve the problems of factory farming, which causes problems like increased emission of greenhouse gas and waste, and contribute to building a sustainable food system that allows easy protein intake.
KAIST (President Kwang-Hyung Lee) announced that Research Professor Kyeong Rok Choi from the Biological Process Research Center and Distinguished Professor Sang Yup Lee from the Department of Chemical and Biomolecular Engineering have published a paper on the development of an "Eco-Friendly Liquid Egg Substitute Derived from Microorganisms."
Eggs play a crucial role in various culinary applications due to their unique physicochemical properties such as gelling, foaming, and emulsifying, while also providing essential nutrients. However, traditional egg production is not only unethical and resource-intensive but also has significant environmental impacts such as greenhouse gas emissions and waste issues. Additionally, factors such as wars and trade regulations have led to significant increases in egg prices, highlighting food security concerns. In response to these issues, there has been growing interest in egg substitutes made from non-animal sources to establish a sustainable food system.
Although there has been progress in developing non-animal protein-based egg substitutes, no substitute has been able to fully replicate the essential functional properties of liquid eggs, such as gelling and foaming, while also providing complete nutrition. In this context, the research team aimed to develop a liquid egg substitute using microbial biomass, which has a protein content comparable to that of meat per unit dry mass. Various microorganisms, such as yeast, Bacillus, lactic acid bacteria, and other probiotics, have been proven safe through long-term human consumption. Microbial biomass requires fewer resources like water and land during production, and possesses high-quality nutrients, making it a promising sustainable food resource.
< Figure 1. Comparison of heat treatment results of microbial pellets and microbial lysates >
However, the semi-solid microbial biomass recovered through microbial cultivation was observed to turn liquid upon heating, unlike liquid egg. To address this, the research team devised a microbial lysate by breaking down the cell walls and cell membranes of microorganisms, which correspond to the eggshell. They found that the microbial lysate's proteins coagulated when heated and formed a gel similar to that of liquid egg. The gel formed from the heated microbial lysate was found to have microscopic structures and physical properties similar to those of boiled eggs. The addition of microbial-derived edible enzymes or plant-based materials allowed for the adjustment of its properties, enabling the creation of various textures.
Furthermore, the researchers demonstrated that the microbial lysate could form stable foams widely used in baking, such as meringues (made from egg whites). They successfully baked meringue cookies using this lysate, showing its potential as a functional liquid egg substitute.
Distinguished Professor Sang Yup Lee stated, "This substitute has excellent nutritional components, making it suitable for regular food consumption. It is especially promising as emergency food for long-term space travel, wartime situations, and other emergencies. More importantly, it contributes to securing a sustainable food system."
< Figure 2. Example of foaming ability of microbial lysate and meringue cookie production >
< Figure 3. Example of foaming ability of microbial lysate and meringue cookie production >
The paper was published online in the journal npj Science of Food, issued by Nature.
- Paper Title: Microbial lysates repurposed as liquid egg substitutes
- Authors: Kyeong Rok Choi (first author), Da-Hee Ahn, Seok Yeong Jung, YuHyun Lee, and Sang Yup Lee (corresponding author)
This research was supported by the Ministry of Science and ICT's project for developing eco-friendly chemical technologies to replace petroleum (Project Leader: Distinguished Professor Sang Yup Lee, KAIST) and the Rural Development Administration's Agricultural Microorganisms Project Group (Director: Professor Pan-sik Jang, Seoul National University) for developing protein production technology from inorganic substances through microbial metabolic system control (Project Leader: Research Professor Kyeong Rok Choi, KAIST).
2024.07.05 View 7007 -
 KAIST introduces microbial food as a strategy food production of the future
The global food crisis is increasing due to rapid population growth and declining food productivity to climate change. Moreover, today's food production and supply system emit a huge amount of carbon dioxide, reaching 30% of the total amount emitted by humanity, aggravating climate change. Sustainable and nutritious microbial food is attracting attention as a key to overcoming this impasse.
KAIST (President Kwang Hyung Lee) announced on April 12th that Research Professor Kyeong Rok Choi of the BioProcess Engineering Research Center and Distinguished Professor Sang Yup Lee from the Department of Chemical and Biomolecular Engineering published a paper that proposes a direction of research on ‘microbial food production from sustainable raw materials.’
Microbial food refers to various foods and food ingredients produced using microorganisms. Microbial biomass contains a large amount of protein per unit in dry mass, comparable to that of meat, and emits the smallest amount of carbon dioxide and is required to produce a unit mass compared to various livestock, fish, shellfish, and crops. Since the amount of water and space requirement is small, it can be an eco-friendly, sustainable and highly nutritious food resource.
Fermented foods are the most readily available microbial foods around us. Although the proportion of microbial biomass in fermented foods is small, compounds with relatively low nutritional value, such as carbohydrates, are consumed during the fermentation process, and as microorganisms proliferate, the content of nutrients with higher nutritional value, such as proteins and vitamins, increases.
Various food compounds isolated and purified from biomass or culture media obtained through microbial culture are also a branch of microbial food. Examples that can be found around us include various amino acids, including monosodium glutamate, food proteins, enzymes, flavoring compounds, food colorings, and bioactive substances.
< Figure 1. Schematic diagram portraying various microbial biomass production strategies utlizing sustainable feedstocks >
Lastly, the most ultimate and fundamental form of microbial food can be said to be microbial biomass or extracts produced through microbial culture and foods cooked using them. A representative example is single-cell protein, which collectively refers to microbial biomass or microbial proteins extracted from it.
In this paper, the researchers comprehensively covered various non-edible raw materials and strategies for using them that can be used to produce microbial food in a more sustainable way. Furthermore, it covers various microbial foods that are actually produced in the industry using the relevant raw materials and their characteristics, as well as prospects for the production and generalization of sustainable microbial foods.
Research Professor Kyeong Rok Choi, the first author of this paper, said, “Microbial foods produced from various sustainable raw materials will soon be commonly encountered at our tables.” Second author Seok Yeong Jung, a doctoral student, also said, “Microbial foods of the future will not be limited foods consumed only out of a sense of obligation to the environment, but will be complete foods that are consumed by choice because of their nutritional value and taste.” In addition, Distinguished Professor Sang Yup Lee said, “It is time for the industry and academia, as well as the public and private sectors, to cooperate more closely so that more diverse microbial foods can be developed and supplied in order to create a sustainable society for ourselves and our descendants.”
< Figure 2. Compositions and environmental footprints of animal, plant and microbial biomass. >
This paper was published online on April 9 in ‘Nature Microbiology’ published by Nature.
※ Paper title: From sustainable feedstocks to microbial foods
※ Author information: Kyeong Rok Choi (first author), Seok Yeong Jung (second author) and Sang Yup Lee (corresponding author)
This research was conducted under the development of platform technologies of microbial cell factories for the next-generation biorefineries project (project leader KAIST Distinguished Professor Sang Yup Lee) supported by the Ministry of Science and ICT and the Cooperative Research Program for Agriculture Science and Technology Development (Project leader KAIST Research Professor Kyeong Rok Choi) of the Agricultural Microbiology Project Group (Director, Professor Pahn-Shick Chang) supported by the Rural Development Administration.
2024.04.12 View 8029
KAIST introduces microbial food as a strategy food production of the future
The global food crisis is increasing due to rapid population growth and declining food productivity to climate change. Moreover, today's food production and supply system emit a huge amount of carbon dioxide, reaching 30% of the total amount emitted by humanity, aggravating climate change. Sustainable and nutritious microbial food is attracting attention as a key to overcoming this impasse.
KAIST (President Kwang Hyung Lee) announced on April 12th that Research Professor Kyeong Rok Choi of the BioProcess Engineering Research Center and Distinguished Professor Sang Yup Lee from the Department of Chemical and Biomolecular Engineering published a paper that proposes a direction of research on ‘microbial food production from sustainable raw materials.’
Microbial food refers to various foods and food ingredients produced using microorganisms. Microbial biomass contains a large amount of protein per unit in dry mass, comparable to that of meat, and emits the smallest amount of carbon dioxide and is required to produce a unit mass compared to various livestock, fish, shellfish, and crops. Since the amount of water and space requirement is small, it can be an eco-friendly, sustainable and highly nutritious food resource.
Fermented foods are the most readily available microbial foods around us. Although the proportion of microbial biomass in fermented foods is small, compounds with relatively low nutritional value, such as carbohydrates, are consumed during the fermentation process, and as microorganisms proliferate, the content of nutrients with higher nutritional value, such as proteins and vitamins, increases.
Various food compounds isolated and purified from biomass or culture media obtained through microbial culture are also a branch of microbial food. Examples that can be found around us include various amino acids, including monosodium glutamate, food proteins, enzymes, flavoring compounds, food colorings, and bioactive substances.
< Figure 1. Schematic diagram portraying various microbial biomass production strategies utlizing sustainable feedstocks >
Lastly, the most ultimate and fundamental form of microbial food can be said to be microbial biomass or extracts produced through microbial culture and foods cooked using them. A representative example is single-cell protein, which collectively refers to microbial biomass or microbial proteins extracted from it.
In this paper, the researchers comprehensively covered various non-edible raw materials and strategies for using them that can be used to produce microbial food in a more sustainable way. Furthermore, it covers various microbial foods that are actually produced in the industry using the relevant raw materials and their characteristics, as well as prospects for the production and generalization of sustainable microbial foods.
Research Professor Kyeong Rok Choi, the first author of this paper, said, “Microbial foods produced from various sustainable raw materials will soon be commonly encountered at our tables.” Second author Seok Yeong Jung, a doctoral student, also said, “Microbial foods of the future will not be limited foods consumed only out of a sense of obligation to the environment, but will be complete foods that are consumed by choice because of their nutritional value and taste.” In addition, Distinguished Professor Sang Yup Lee said, “It is time for the industry and academia, as well as the public and private sectors, to cooperate more closely so that more diverse microbial foods can be developed and supplied in order to create a sustainable society for ourselves and our descendants.”
< Figure 2. Compositions and environmental footprints of animal, plant and microbial biomass. >
This paper was published online on April 9 in ‘Nature Microbiology’ published by Nature.
※ Paper title: From sustainable feedstocks to microbial foods
※ Author information: Kyeong Rok Choi (first author), Seok Yeong Jung (second author) and Sang Yup Lee (corresponding author)
This research was conducted under the development of platform technologies of microbial cell factories for the next-generation biorefineries project (project leader KAIST Distinguished Professor Sang Yup Lee) supported by the Ministry of Science and ICT and the Cooperative Research Program for Agriculture Science and Technology Development (Project leader KAIST Research Professor Kyeong Rok Choi) of the Agricultural Microbiology Project Group (Director, Professor Pahn-Shick Chang) supported by the Rural Development Administration.
2024.04.12 View 8029 -
 KAIST presents strategies for environmentally friendly and sustainable polyamides production
- Provides current research trends in bio-based polyamide production
- Research on bio-based polyamides production gains importance for achieving a carbon-neutral society
Global industries focused on carbon neutrality, under the slogan "Net-Zero," are gaining increasing attention. In particular, research on microbial production of polymers, replacing traditional chemical methods with biological approaches, is actively progressing.
Polyamides, represented by nylon, are linear polymers widely used in various industries such as automotive, electronics, textiles, and medical fields. They possess beneficial properties such as high tensile strength, electrical insulation, heat resistance, wear resistance, and biocompatibility. Since the commercialization of nylon in 1938, approximately 7 million tons of polyamides are produced worldwide annually. Considering their broad applications and significance, producing polyamides through bio-based methods holds considerable environmental and industrial importance.
KAIST (President Kwang-Hyung Lee) announced that a research team led by Distinguished Professor Sang Yup Lee, including Dr. Jong An Lee and doctoral candidate Ji Yeon Kim from the Department of Chemical and Biomolecular Engineering, published a paper titled "Current Advancements in Bio-Based Production of Polyamides”. The paper was featured on the cover of the monthly issue of "Trends in Chemistry” by Cell Press.
As part of climate change response technologies, bio-refineries involve using biotechnological and chemical methods to produce industrially important chemicals and biofuels from renewable biomass without relying on fossil resources. Notably, systems metabolic engineering, pioneered by KAIST's Distinguished Professor Sang Yup Lee, is a research field that effectively manipulates microbial metabolic pathways to produce valuable chemicals, forming the core technology for bio-refineries. The research team has successfully developed high-performance strains producing a variety of compounds, including succinic acid, biodegradable plastics, biofuels, and natural products, using systems metabolic engineering tools and strategies.
The research team predicted that if bio-based polyamide production technology, which is widely used in the production of clothing and textiles, becomes widespread, it will attract attention as a future technology that can respond to the climate crisis due to its environment-friendly production technology. In this study, the research team comprehensively reviewed the bio-based polyamide production strategies. They provided insights into the advancements in polyamide monomer production using metabolically engineered microorganisms and highlighted the recent trends in bio-based polyamide advancements utilizing these monomers. Additionally, they reviewed the strategies for synthesizing bio-based polyamides through chemical conversion of natural oils and discussed the biodegradability and recycling of the polyamides. Furthermore, the paper presented the future direction in which metabolic engineering can be applied for the bio-based polyamide production, contributing to environmentally friendly and sustainable society.
Ji Yeon Kim, the co-first author of this paper from KAIST, stated "The importance of utilizing systems metabolic engineering tools and strategies for bio-based polyamides production is becoming increasingly prominent in achieving carbon neutrality."
Professor Sang Yup Lee emphasized, "Amid growing concerns about climate change, the significance of environmentally friendly and sustainable industrial development is greater than ever. Systems metabolic engineering is expected to have a significant impact not only on the chemical industry but also in various fields."
< [Figure 1] A schematic overview of the overall process for polyamides production >
This paper by Dr. Jong An Lee, PhD student Ji Yeon Kim, Dr. Jung Ho Ahn, and Master Yeah-Ji Ahn from the Department of Chemical and Biomolecular Engineering at KAIST was published in the December issue of 'Trends in Chemistry', an authoritative review journal in the field of chemistry published by Cell. It was published on December 7 as the cover paper and featured review.
※ Paper title: Current advancements in the bio-based production of polyamides
※ Author information: Jong An Lee, Ji Yeon Kim, Jung Ho Ahn, Yeah-Ji Ahn, and Sang Yup Lee
This research was conducted with the support from the development of platform technologies of microbial cell factories for the next-generation biorefineries project and C1 gas refinery program by Korean Ministry of Science and ICT.
< [Figure 2] Cover paper of the December issue of Trends in Chemistry >
2023.12.21 View 7458
KAIST presents strategies for environmentally friendly and sustainable polyamides production
- Provides current research trends in bio-based polyamide production
- Research on bio-based polyamides production gains importance for achieving a carbon-neutral society
Global industries focused on carbon neutrality, under the slogan "Net-Zero," are gaining increasing attention. In particular, research on microbial production of polymers, replacing traditional chemical methods with biological approaches, is actively progressing.
Polyamides, represented by nylon, are linear polymers widely used in various industries such as automotive, electronics, textiles, and medical fields. They possess beneficial properties such as high tensile strength, electrical insulation, heat resistance, wear resistance, and biocompatibility. Since the commercialization of nylon in 1938, approximately 7 million tons of polyamides are produced worldwide annually. Considering their broad applications and significance, producing polyamides through bio-based methods holds considerable environmental and industrial importance.
KAIST (President Kwang-Hyung Lee) announced that a research team led by Distinguished Professor Sang Yup Lee, including Dr. Jong An Lee and doctoral candidate Ji Yeon Kim from the Department of Chemical and Biomolecular Engineering, published a paper titled "Current Advancements in Bio-Based Production of Polyamides”. The paper was featured on the cover of the monthly issue of "Trends in Chemistry” by Cell Press.
As part of climate change response technologies, bio-refineries involve using biotechnological and chemical methods to produce industrially important chemicals and biofuels from renewable biomass without relying on fossil resources. Notably, systems metabolic engineering, pioneered by KAIST's Distinguished Professor Sang Yup Lee, is a research field that effectively manipulates microbial metabolic pathways to produce valuable chemicals, forming the core technology for bio-refineries. The research team has successfully developed high-performance strains producing a variety of compounds, including succinic acid, biodegradable plastics, biofuels, and natural products, using systems metabolic engineering tools and strategies.
The research team predicted that if bio-based polyamide production technology, which is widely used in the production of clothing and textiles, becomes widespread, it will attract attention as a future technology that can respond to the climate crisis due to its environment-friendly production technology. In this study, the research team comprehensively reviewed the bio-based polyamide production strategies. They provided insights into the advancements in polyamide monomer production using metabolically engineered microorganisms and highlighted the recent trends in bio-based polyamide advancements utilizing these monomers. Additionally, they reviewed the strategies for synthesizing bio-based polyamides through chemical conversion of natural oils and discussed the biodegradability and recycling of the polyamides. Furthermore, the paper presented the future direction in which metabolic engineering can be applied for the bio-based polyamide production, contributing to environmentally friendly and sustainable society.
Ji Yeon Kim, the co-first author of this paper from KAIST, stated "The importance of utilizing systems metabolic engineering tools and strategies for bio-based polyamides production is becoming increasingly prominent in achieving carbon neutrality."
Professor Sang Yup Lee emphasized, "Amid growing concerns about climate change, the significance of environmentally friendly and sustainable industrial development is greater than ever. Systems metabolic engineering is expected to have a significant impact not only on the chemical industry but also in various fields."
< [Figure 1] A schematic overview of the overall process for polyamides production >
This paper by Dr. Jong An Lee, PhD student Ji Yeon Kim, Dr. Jung Ho Ahn, and Master Yeah-Ji Ahn from the Department of Chemical and Biomolecular Engineering at KAIST was published in the December issue of 'Trends in Chemistry', an authoritative review journal in the field of chemistry published by Cell. It was published on December 7 as the cover paper and featured review.
※ Paper title: Current advancements in the bio-based production of polyamides
※ Author information: Jong An Lee, Ji Yeon Kim, Jung Ho Ahn, Yeah-Ji Ahn, and Sang Yup Lee
This research was conducted with the support from the development of platform technologies of microbial cell factories for the next-generation biorefineries project and C1 gas refinery program by Korean Ministry of Science and ICT.
< [Figure 2] Cover paper of the December issue of Trends in Chemistry >
2023.12.21 View 7458 -
 KAIST proposes alternatives to chemical factories through “iBridge”
- A computer simulation program “iBridge” was developed at KAIST that can put together microbial cell factories quickly and efficiently to produce cosmetics and food additives, and raw materials for nylons
- Eco-friendly and sustainable fermentation process to establish an alternative to chemical plants
As climate change and environmental concerns intensify, sustainable microbial cell factories garner significant attention as candidates to replace chemical plants. To develop microorganisms to be used in the microbial cell factories, it is crucial to modify their metabolic processes to induce efficient target chemical production by modulating its gene expressions. Yet, the challenge persists in determining which gene expressions to amplify and suppress, and the experimental verification of these modification targets is a time- and resource-intensive process even for experts. The challenges were addressed by a team of researchers at KAIST (President Kwang-Hyung Lee) led by Distinguished Professor Sang Yup Lee.
It was announced on the 9th by the school that a method for building a microbial factory at low cost, quickly and efficiently, was presented by a novel computer simulation program developed by the team under Professor Lee’s guidance, which is named “iBridge”. This innovative system is designed to predict gene targets to either overexpress or downregulate in the goal of producing a desired compound to enable the cost-effective and efficient construction of microbial cell factories specifically tailored for producing the chemical compound in demand from renewable biomass.
Systems metabolic engineering is a field of research and engineering pioneered by KAIST’s Distinguished Professor Sang Yup Lee that seeks to produce valuable compounds in industrial demands using microorganisms that are re-configured by a combination of methods including, but not limited to, metabolic engineering, synthetic biology, systems biology, and fermentation engineering.
In order to improve microorganisms’ capability to produce useful compounds, it is essential to delete, suppress, or overexpress microbial genes. However, it is difficult even for the experts to identify the gene targets to modify without experimental confirmations for each of them, which can take up immeasurable amount of time and resources.
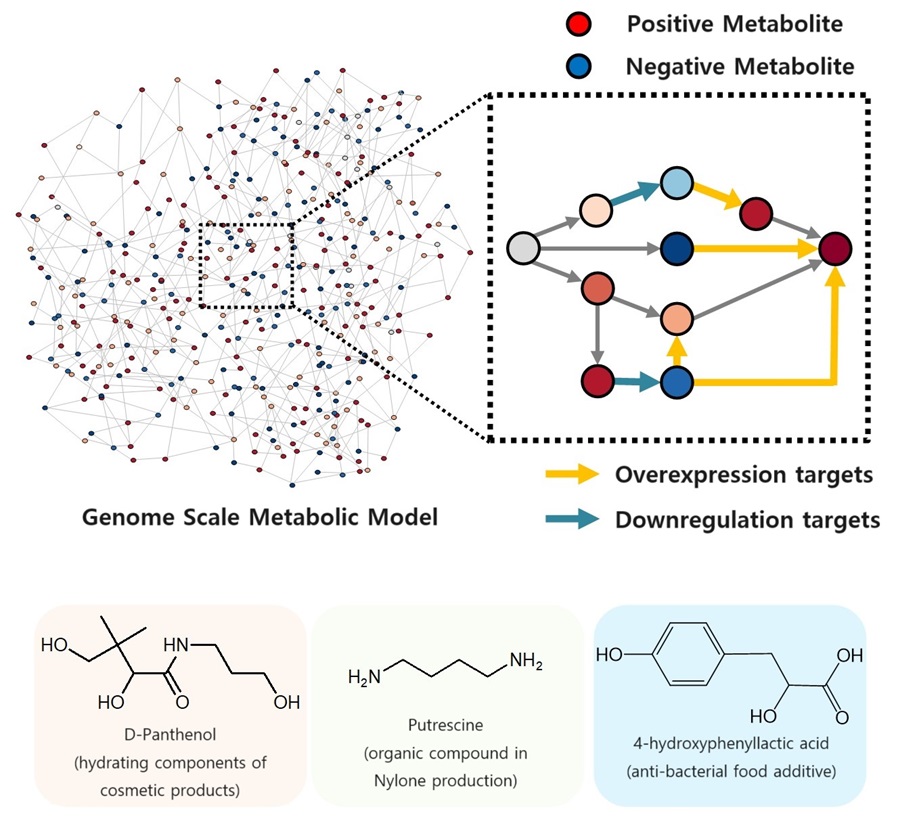
The newly developed iBridge identifies positive and negative metabolites within cells, which exert positive and/or negative impact on formation of the products, by calculating the sum of covariances of their outgoing (consuming) reaction fluxes for a target chemical. Subsequently, it pinpoints "bridge" reactions responsible for converting negative metabolites into positive ones as candidates for overexpression, while identifying the opposites as targets for downregulation.
The research team successfully utilized the iBridge simulation to establish E. coli microbial cell factories each capable of producing three of the compounds that are in high demands at a production capacity that has not been reported around the world. They developed E. coli strains that can each produce panthenol, a moisturizing agent found in many cosmetics, putrescine, which is one of the key components in nylon production, and 4-hydroxyphenyllactic acid, an anti-bacterial food additive. In addition to these three compounds, the study presents predictions for overexpression and suppression genes to construct microbial factories for 298 other industrially valuable compounds.
Dr. Youngjoon Lee, the co-first author of this paper from KAIST, emphasized the accelerated construction of various microbial factories the newly developed simulation enabled. He stated, "With the use of this simulation, multiple microbial cell factories have been established significantly faster than it would have been using the conventional methods. Microbial cell factories producing a wider range of valuable compounds can now be constructed quickly using this technology."
Professor Sang Yup Lee said, "Systems metabolic engineering is a crucial technology for addressing the current climate change issues." He added, "This simulation could significantly expedite the transition from resorting to conventional chemical factories to utilizing environmentally friendly microbial factories."
< Figure. Conceptual diagram of the flow of iBridge simulation >
The team’s work on iBridge is described in a paper titled "Genome-Wide Identification of Overexpression and Downregulation Gene Targets Based on the Sum of Covariances of the Outgoing Reaction Fluxes" written by Dr. Won Jun Kim, and Dr. Youngjoon Lee of the Bioprocess Research Center and Professors Hyun Uk Kim and Sang Yup Lee of the Department of Chemical and Biomolecular Engineering of KAIST. The paper was published via peer-review on the 6th of November on “Cell Systems” by Cell Press.
This research was conducted with the support from the Development of Platform Technologies of Microbial Cell Factories for the Next-generation Biorefineries Project (Project Leader: Distinguished Professor Sang Yup Lee, KAIST) and Development of Platform Technology for the Production of Novel Aromatic Bioplastic using Microbial Cell Factories Project (Project Leader: Research Professor So Young Choi, KAIST) of the Korean Ministry of Science and ICT.
2023.11.09 View 8773
KAIST proposes alternatives to chemical factories through “iBridge”
- A computer simulation program “iBridge” was developed at KAIST that can put together microbial cell factories quickly and efficiently to produce cosmetics and food additives, and raw materials for nylons
- Eco-friendly and sustainable fermentation process to establish an alternative to chemical plants
As climate change and environmental concerns intensify, sustainable microbial cell factories garner significant attention as candidates to replace chemical plants. To develop microorganisms to be used in the microbial cell factories, it is crucial to modify their metabolic processes to induce efficient target chemical production by modulating its gene expressions. Yet, the challenge persists in determining which gene expressions to amplify and suppress, and the experimental verification of these modification targets is a time- and resource-intensive process even for experts. The challenges were addressed by a team of researchers at KAIST (President Kwang-Hyung Lee) led by Distinguished Professor Sang Yup Lee.
It was announced on the 9th by the school that a method for building a microbial factory at low cost, quickly and efficiently, was presented by a novel computer simulation program developed by the team under Professor Lee’s guidance, which is named “iBridge”. This innovative system is designed to predict gene targets to either overexpress or downregulate in the goal of producing a desired compound to enable the cost-effective and efficient construction of microbial cell factories specifically tailored for producing the chemical compound in demand from renewable biomass.
Systems metabolic engineering is a field of research and engineering pioneered by KAIST’s Distinguished Professor Sang Yup Lee that seeks to produce valuable compounds in industrial demands using microorganisms that are re-configured by a combination of methods including, but not limited to, metabolic engineering, synthetic biology, systems biology, and fermentation engineering.
In order to improve microorganisms’ capability to produce useful compounds, it is essential to delete, suppress, or overexpress microbial genes. However, it is difficult even for the experts to identify the gene targets to modify without experimental confirmations for each of them, which can take up immeasurable amount of time and resources.
The newly developed iBridge identifies positive and negative metabolites within cells, which exert positive and/or negative impact on formation of the products, by calculating the sum of covariances of their outgoing (consuming) reaction fluxes for a target chemical. Subsequently, it pinpoints "bridge" reactions responsible for converting negative metabolites into positive ones as candidates for overexpression, while identifying the opposites as targets for downregulation.
The research team successfully utilized the iBridge simulation to establish E. coli microbial cell factories each capable of producing three of the compounds that are in high demands at a production capacity that has not been reported around the world. They developed E. coli strains that can each produce panthenol, a moisturizing agent found in many cosmetics, putrescine, which is one of the key components in nylon production, and 4-hydroxyphenyllactic acid, an anti-bacterial food additive. In addition to these three compounds, the study presents predictions for overexpression and suppression genes to construct microbial factories for 298 other industrially valuable compounds.
Dr. Youngjoon Lee, the co-first author of this paper from KAIST, emphasized the accelerated construction of various microbial factories the newly developed simulation enabled. He stated, "With the use of this simulation, multiple microbial cell factories have been established significantly faster than it would have been using the conventional methods. Microbial cell factories producing a wider range of valuable compounds can now be constructed quickly using this technology."
Professor Sang Yup Lee said, "Systems metabolic engineering is a crucial technology for addressing the current climate change issues." He added, "This simulation could significantly expedite the transition from resorting to conventional chemical factories to utilizing environmentally friendly microbial factories."
< Figure. Conceptual diagram of the flow of iBridge simulation >
The team’s work on iBridge is described in a paper titled "Genome-Wide Identification of Overexpression and Downregulation Gene Targets Based on the Sum of Covariances of the Outgoing Reaction Fluxes" written by Dr. Won Jun Kim, and Dr. Youngjoon Lee of the Bioprocess Research Center and Professors Hyun Uk Kim and Sang Yup Lee of the Department of Chemical and Biomolecular Engineering of KAIST. The paper was published via peer-review on the 6th of November on “Cell Systems” by Cell Press.
This research was conducted with the support from the Development of Platform Technologies of Microbial Cell Factories for the Next-generation Biorefineries Project (Project Leader: Distinguished Professor Sang Yup Lee, KAIST) and Development of Platform Technology for the Production of Novel Aromatic Bioplastic using Microbial Cell Factories Project (Project Leader: Research Professor So Young Choi, KAIST) of the Korean Ministry of Science and ICT.
2023.11.09 View 8773 -
 Synthetic sRNAs to knockdown genes in medical and industrial bacteria
Bacteria are intimately involved in our daily lives. These microorganisms have been used in human history for food such as cheese, yogurt, and wine, In more recent years, through metabolic engineering, microorganisms been used extensively as microbial cell factories to manufacture plastics, feed for livestock, dietary supplements, and drugs. However, in addition to these bacteria that are beneficial to human lives, pathogens such as Pneumonia, Salmonella, and Staphylococcus that cause various infectious diseases are also ubiquitously present. It is important to be able to metabolically control these beneficial industrial bacteria for high value-added chemicals production and to manipulate harmful pathogens to suppress its pathogenic traits.
KAIST (President Kwang Hyung Lee) announced on the 10th that a research team led by Distinguished Professor Sang Yup Lee of the Department of Biochemical Engineering has developed a new sRNA tool that can effectively inhibit target genes in various bacteria, including both Gram-negative and Gram-positive bacteria. The research results were published online on April 24 in Nature Communications.
※ Thesis title: Targeted and high-throughput gene knockdown in diverse bacteria using synthetic sRNAs
※ Author information : Jae Sung Cho (co-1st), Dongsoo Yang (co-1st), Cindy Pricilia Surya Prabowo (co-author), Mohammad Rifqi Ghiffary (co-author), Taehee Han (co-author), Kyeong Rok Choi (co-author), Cheon Woo Moon (co-author), Hengrui Zhou (co-author), Jae Yong Ryu (co-author), Hyun Uk Kim (co-author) and Sang Yup Lee (corresponding author).
sRNA is an effective tool for synthesizing and regulating target genes in E. coli, but it has been difficult to apply to industrially useful Gram-positive bacteria such as Bacillus subtilis and Corynebacterium in addition to Gram-negative bacteria such as E. coli.
To address this issue, a research team led by Distinguished Professor Lee Sang Yup Lee of the Department of Chemical and Biomolecular Engineering at KAIST developed a new sRNA platform that can effectively suppress target genes in various bacteria, including both Gram-negative and positive bacteria. The research team surveyed thousands of microbial-derived sRNA systems in the microbial database, and eventually designated the sRNA system derived from 'Bacillus subtilis' that showed the highest gene knockdown efficiency, and designated it as “Broad-Host-Range sRNA”, or BHR-sRNA.
A similar well-known system is the CRISPR interference (CRISPRi) system, which is a modified CRISPR system that knocks down gene expression by suppressing the gene transcription process. However, the Cas9 protein in the CRISPRi system has a very high molecular weight, and there have been reports growth inhibition in bacteria. The BHR-sRNA system developed in this study did not affect bacterial growth while showing similar gene knockdown efficiencies to CRISPRi.
< Figure 1. a) Schematic illustration demonstrating the mechanism of syntetic sRNA b) Phylogenetic tree of the 16 Gram-negative and Gram-positive bacterial species tested for gene knockdown by the BHR-sRNA system. >
To validate the versatility of the BHR-sRNA system, 16 different gram-negative and gram-positive bacteria were selected and tested, where the BHR-sRNA system worked successfully in 15 of them. In addition, it was demonstrated that the gene knockdown capability was more effective than that of the existing E. coli-based sRNA system in 10 bacteria. The BHR-sRNA system proved to be a universal tool capable of effectively inhibiting gene expression in various bacteria.
In order to address the problem of antibiotic-resistant pathogens that have recently become more serious, the BHR-sRNA was demonstrated to suppress the pathogenicity by suppressing the gene producing the virulence factor. By using BHR-sRNA, biofilm formation, one of the factors resulting in antibiotic resistance, was inhibited by 73% in Staphylococcus epidermidis a pathogen that can cause hospital-acquired infections. Antibiotic resistance was also weakened by 58% in the pneumonia causing bacteria Klebsiella pneumoniae. In addition, BHR-sRNA was applied to industrial bacteria to develop microbial cell factories to produce high value-added chemicals with better production performance. Notably, superior industrial strains were constructed with the aid of BHR-sRNA to produce the following chemicals: valerolactam, a raw material for polyamide polymers, methyl-anthranilate, a grape-flavor food additive, and indigoidine, a blue-toned natural dye.
The BHR-sRNA developed through this study will help expedite the commercialization of bioprocesses to produce high value-added compounds and materials such as artificial meat, jet fuel, health supplements, pharmaceuticals, and plastics. It is also anticipated that to help eradicating antibiotic-resistant pathogens in preparation for another upcoming pandemic. “In the past, we could only develop new tools for gene knockdown for each bacterium, but now we have developed a tool that works for a variety of bacteria” said Distinguished Professor Sang Yup Lee.
This work was supported by the Development of Next-generation Biorefinery Platform Technologies for Leading Bio-based Chemicals Industry Project and the Development of Platform Technologies of Microbial Cell Factories for the Next-generation Biorefineries Project from NRF supported by the Korean MSIT.
2023.05.10 View 8575
Synthetic sRNAs to knockdown genes in medical and industrial bacteria
Bacteria are intimately involved in our daily lives. These microorganisms have been used in human history for food such as cheese, yogurt, and wine, In more recent years, through metabolic engineering, microorganisms been used extensively as microbial cell factories to manufacture plastics, feed for livestock, dietary supplements, and drugs. However, in addition to these bacteria that are beneficial to human lives, pathogens such as Pneumonia, Salmonella, and Staphylococcus that cause various infectious diseases are also ubiquitously present. It is important to be able to metabolically control these beneficial industrial bacteria for high value-added chemicals production and to manipulate harmful pathogens to suppress its pathogenic traits.
KAIST (President Kwang Hyung Lee) announced on the 10th that a research team led by Distinguished Professor Sang Yup Lee of the Department of Biochemical Engineering has developed a new sRNA tool that can effectively inhibit target genes in various bacteria, including both Gram-negative and Gram-positive bacteria. The research results were published online on April 24 in Nature Communications.
※ Thesis title: Targeted and high-throughput gene knockdown in diverse bacteria using synthetic sRNAs
※ Author information : Jae Sung Cho (co-1st), Dongsoo Yang (co-1st), Cindy Pricilia Surya Prabowo (co-author), Mohammad Rifqi Ghiffary (co-author), Taehee Han (co-author), Kyeong Rok Choi (co-author), Cheon Woo Moon (co-author), Hengrui Zhou (co-author), Jae Yong Ryu (co-author), Hyun Uk Kim (co-author) and Sang Yup Lee (corresponding author).
sRNA is an effective tool for synthesizing and regulating target genes in E. coli, but it has been difficult to apply to industrially useful Gram-positive bacteria such as Bacillus subtilis and Corynebacterium in addition to Gram-negative bacteria such as E. coli.
To address this issue, a research team led by Distinguished Professor Lee Sang Yup Lee of the Department of Chemical and Biomolecular Engineering at KAIST developed a new sRNA platform that can effectively suppress target genes in various bacteria, including both Gram-negative and positive bacteria. The research team surveyed thousands of microbial-derived sRNA systems in the microbial database, and eventually designated the sRNA system derived from 'Bacillus subtilis' that showed the highest gene knockdown efficiency, and designated it as “Broad-Host-Range sRNA”, or BHR-sRNA.
A similar well-known system is the CRISPR interference (CRISPRi) system, which is a modified CRISPR system that knocks down gene expression by suppressing the gene transcription process. However, the Cas9 protein in the CRISPRi system has a very high molecular weight, and there have been reports growth inhibition in bacteria. The BHR-sRNA system developed in this study did not affect bacterial growth while showing similar gene knockdown efficiencies to CRISPRi.
< Figure 1. a) Schematic illustration demonstrating the mechanism of syntetic sRNA b) Phylogenetic tree of the 16 Gram-negative and Gram-positive bacterial species tested for gene knockdown by the BHR-sRNA system. >
To validate the versatility of the BHR-sRNA system, 16 different gram-negative and gram-positive bacteria were selected and tested, where the BHR-sRNA system worked successfully in 15 of them. In addition, it was demonstrated that the gene knockdown capability was more effective than that of the existing E. coli-based sRNA system in 10 bacteria. The BHR-sRNA system proved to be a universal tool capable of effectively inhibiting gene expression in various bacteria.
In order to address the problem of antibiotic-resistant pathogens that have recently become more serious, the BHR-sRNA was demonstrated to suppress the pathogenicity by suppressing the gene producing the virulence factor. By using BHR-sRNA, biofilm formation, one of the factors resulting in antibiotic resistance, was inhibited by 73% in Staphylococcus epidermidis a pathogen that can cause hospital-acquired infections. Antibiotic resistance was also weakened by 58% in the pneumonia causing bacteria Klebsiella pneumoniae. In addition, BHR-sRNA was applied to industrial bacteria to develop microbial cell factories to produce high value-added chemicals with better production performance. Notably, superior industrial strains were constructed with the aid of BHR-sRNA to produce the following chemicals: valerolactam, a raw material for polyamide polymers, methyl-anthranilate, a grape-flavor food additive, and indigoidine, a blue-toned natural dye.
The BHR-sRNA developed through this study will help expedite the commercialization of bioprocesses to produce high value-added compounds and materials such as artificial meat, jet fuel, health supplements, pharmaceuticals, and plastics. It is also anticipated that to help eradicating antibiotic-resistant pathogens in preparation for another upcoming pandemic. “In the past, we could only develop new tools for gene knockdown for each bacterium, but now we have developed a tool that works for a variety of bacteria” said Distinguished Professor Sang Yup Lee.
This work was supported by the Development of Next-generation Biorefinery Platform Technologies for Leading Bio-based Chemicals Industry Project and the Development of Platform Technologies of Microbial Cell Factories for the Next-generation Biorefineries Project from NRF supported by the Korean MSIT.
2023.05.10 View 8575 -
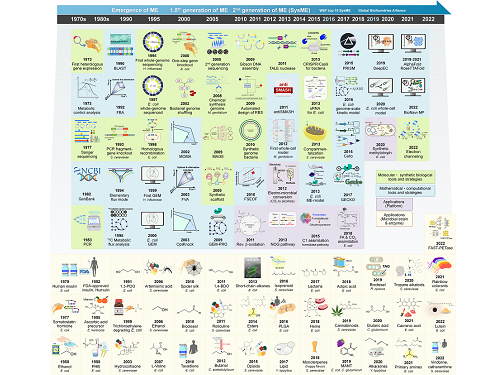 Overview of the 30-year history of metabolic engineering
< Distinguished Professor Sang Yup Lee from the Department of Chemical and Biomolecular Engineering at KAIST >
A research team comprised of Gi Bae Kim, Dr. So Young Choi, Dr. In Jin Cho, Da-Hee Ahn, and Distinguished Professor Sang Yup Lee from the Department of Chemical and Biomolecular Engineering at KAIST reported the 30-year history of metabolic engineering, highlighting examples of recent progress in the field and contributions to sustainability and health. Their paper “Metabolic engineering for sustainability and health” was published online in the 40th anniversary special issue of Trends in Biotechnology on January 10, 2023.
Metabolic engineering, a discipline of engineering that modifies cell phenotypes through molecular and genetic-level manipulations to improve cellular activities, has been studied since the early 1990s, and has progressed significantly over the past 30 years. In particular, metabolic engineering has enabled the engineering of microorganisms for the development of microbial cell factories capable of efficiently producing chemicals and materials as well as degrading recalcitrant contaminants.
This review article revisited how metabolic engineering has advanced over the past 30 years, from the advent of genetic engineering techniques such as recombinant DNA technologies to recent breakthroughs in systems metabolic engineering and data science aided by artificial intelligence. The research team highlighted momentous events and achievements in metabolic engineering, providing both trends and future directions in the field. Metabolic engineering’s contributions to bio-based sustainable chemicals and clean energy, health, and bioremediation were also reviewed. Finally, the research team shared their perspectives on the future challenges impacting metabolic engineering than must be overcome in order to achieve advancements in sustainability and health.
Distinguished Professor Sang Yup Lee said, “Replacing fossil resource-based chemical processes with bio-based sustainable processes for the production of chemicals, fuels, and materials using metabolic engineering has become our essential task for the future. By looking back on the 30+ years of metabolic engineering, we aimed to highlight the contributions of metabolic engineering to achieve sustainability and good health.” He added, “Metabolic engineering will play an increasingly important role as a key solution to the climate crisis, environmental pollution, food and energy shortages, and health problems in aging societies.”
< Figure: Metabolic Engineering Timeline >
2023.01.25 View 11638
Overview of the 30-year history of metabolic engineering
< Distinguished Professor Sang Yup Lee from the Department of Chemical and Biomolecular Engineering at KAIST >
A research team comprised of Gi Bae Kim, Dr. So Young Choi, Dr. In Jin Cho, Da-Hee Ahn, and Distinguished Professor Sang Yup Lee from the Department of Chemical and Biomolecular Engineering at KAIST reported the 30-year history of metabolic engineering, highlighting examples of recent progress in the field and contributions to sustainability and health. Their paper “Metabolic engineering for sustainability and health” was published online in the 40th anniversary special issue of Trends in Biotechnology on January 10, 2023.
Metabolic engineering, a discipline of engineering that modifies cell phenotypes through molecular and genetic-level manipulations to improve cellular activities, has been studied since the early 1990s, and has progressed significantly over the past 30 years. In particular, metabolic engineering has enabled the engineering of microorganisms for the development of microbial cell factories capable of efficiently producing chemicals and materials as well as degrading recalcitrant contaminants.
This review article revisited how metabolic engineering has advanced over the past 30 years, from the advent of genetic engineering techniques such as recombinant DNA technologies to recent breakthroughs in systems metabolic engineering and data science aided by artificial intelligence. The research team highlighted momentous events and achievements in metabolic engineering, providing both trends and future directions in the field. Metabolic engineering’s contributions to bio-based sustainable chemicals and clean energy, health, and bioremediation were also reviewed. Finally, the research team shared their perspectives on the future challenges impacting metabolic engineering than must be overcome in order to achieve advancements in sustainability and health.
Distinguished Professor Sang Yup Lee said, “Replacing fossil resource-based chemical processes with bio-based sustainable processes for the production of chemicals, fuels, and materials using metabolic engineering has become our essential task for the future. By looking back on the 30+ years of metabolic engineering, we aimed to highlight the contributions of metabolic engineering to achieve sustainability and good health.” He added, “Metabolic engineering will play an increasingly important role as a key solution to the climate crisis, environmental pollution, food and energy shortages, and health problems in aging societies.”
< Figure: Metabolic Engineering Timeline >
2023.01.25 View 11638 -
 Interactive Map of Metabolical Synthesis of Chemicals
An interactive map that compiled the chemicals produced by biological, chemical and combined reactions has been distributed on the web
- A team led by Distinguished Professor Sang Yup Lee of the Department of Chemical and Biomolecular Engineering, organized and distributed an all-inclusive listing of chemical substances that can be synthesized using microorganisms
- It is expected to be used by researchers around the world as it enables easy assessment of the synthetic pathway through the web.
A research team comprised of Woo Dae Jang, Gi Bae Kim, and Distinguished Professor Sang Yup Lee of the Department of Chemical and Biomolecular Engineering at KAIST reported an interactive metabolic map of bio-based chemicals. Their research paper “An interactive metabolic map of bio-based chemicals” was published online in Trends in Biotechnology on August 10, 2022.
As a response to rapid climate change and environmental pollution, research on the production of petrochemical products using microorganisms is receiving attention as a sustainable alternative to existing methods of productions. In order to synthesize various chemical substances, materials, and fuel using microorganisms, it is necessary to first construct the biosynthetic pathway toward desired product by exploration and discovery and introduce them into microorganisms. In addition, in order to efficiently synthesize various chemical substances, it is sometimes necessary to employ chemical methods along with bioengineering methods using microorganisms at the same time. For the production of non-native chemicals, novel pathways are designed by recruiting enzymes from heterologous sources or employing enzymes designed though rational engineering, directed evolution, or ab initio design.
The research team had completed a map of chemicals which compiled all available pathways of biological and/or chemical reactions that lead to the production of various bio-based chemicals back in 2019 and published the map in Nature Catalysis. The map was distributed in the form of a poster to industries and academia so that the synthesis paths of bio-based chemicals could be checked at a glance.
The research team has expanded the bio-based chemicals map this time in the form of an interactive map on the web so that anyone with internet access can quickly explore efficient paths to synthesize desired products. The web-based map provides interactive visual tools to allow interactive visualization, exploration, and analysis of complex networks of biological and/or chemical reactions toward the desired products. In addition, the reported paper also discusses the production of natural compounds that are used for diverse purposes such as food and medicine, which will help designing novel pathways through similar approaches or by exploiting the promiscuity of enzymes described in the map. The published bio-based chemicals map is also available at http://systemsbiotech.co.kr.
The co-first authors, Dr. Woo Dae Jang and Ph.D. student Gi Bae Kim, said, “We conducted this study to address the demand for updating the previously distributed chemicals map and enhancing its versatility.” “The map is expected to be utilized in a variety of research and in efforts to set strategies and prospects for chemical production incorporating bio and chemical methods that are detailed in the map.”
Distinguished Professor Sang Yup Lee said, “The interactive bio-based chemicals map is expected to help design and optimization of the metabolic pathways for the biosynthesis of target chemicals together with the strategies of chemical conversions, serving as a blueprint for developing further ideas on the production of desired chemicals through biological and/or chemical reactions.”
The interactive metabolic map of bio-based chemicals.
2022.08.11 View 16600
Interactive Map of Metabolical Synthesis of Chemicals
An interactive map that compiled the chemicals produced by biological, chemical and combined reactions has been distributed on the web
- A team led by Distinguished Professor Sang Yup Lee of the Department of Chemical and Biomolecular Engineering, organized and distributed an all-inclusive listing of chemical substances that can be synthesized using microorganisms
- It is expected to be used by researchers around the world as it enables easy assessment of the synthetic pathway through the web.
A research team comprised of Woo Dae Jang, Gi Bae Kim, and Distinguished Professor Sang Yup Lee of the Department of Chemical and Biomolecular Engineering at KAIST reported an interactive metabolic map of bio-based chemicals. Their research paper “An interactive metabolic map of bio-based chemicals” was published online in Trends in Biotechnology on August 10, 2022.
As a response to rapid climate change and environmental pollution, research on the production of petrochemical products using microorganisms is receiving attention as a sustainable alternative to existing methods of productions. In order to synthesize various chemical substances, materials, and fuel using microorganisms, it is necessary to first construct the biosynthetic pathway toward desired product by exploration and discovery and introduce them into microorganisms. In addition, in order to efficiently synthesize various chemical substances, it is sometimes necessary to employ chemical methods along with bioengineering methods using microorganisms at the same time. For the production of non-native chemicals, novel pathways are designed by recruiting enzymes from heterologous sources or employing enzymes designed though rational engineering, directed evolution, or ab initio design.
The research team had completed a map of chemicals which compiled all available pathways of biological and/or chemical reactions that lead to the production of various bio-based chemicals back in 2019 and published the map in Nature Catalysis. The map was distributed in the form of a poster to industries and academia so that the synthesis paths of bio-based chemicals could be checked at a glance.
The research team has expanded the bio-based chemicals map this time in the form of an interactive map on the web so that anyone with internet access can quickly explore efficient paths to synthesize desired products. The web-based map provides interactive visual tools to allow interactive visualization, exploration, and analysis of complex networks of biological and/or chemical reactions toward the desired products. In addition, the reported paper also discusses the production of natural compounds that are used for diverse purposes such as food and medicine, which will help designing novel pathways through similar approaches or by exploiting the promiscuity of enzymes described in the map. The published bio-based chemicals map is also available at http://systemsbiotech.co.kr.
The co-first authors, Dr. Woo Dae Jang and Ph.D. student Gi Bae Kim, said, “We conducted this study to address the demand for updating the previously distributed chemicals map and enhancing its versatility.” “The map is expected to be utilized in a variety of research and in efforts to set strategies and prospects for chemical production incorporating bio and chemical methods that are detailed in the map.”
Distinguished Professor Sang Yup Lee said, “The interactive bio-based chemicals map is expected to help design and optimization of the metabolic pathways for the biosynthesis of target chemicals together with the strategies of chemical conversions, serving as a blueprint for developing further ideas on the production of desired chemicals through biological and/or chemical reactions.”
The interactive metabolic map of bio-based chemicals.
2022.08.11 View 16600 -
 Microbial Production of a Natural Red Colorant Carminic Acid
Metabolic engineering and computer-simulated enzyme engineering led to the production of carminic acid, a natural red colorant, from bacteria for the first time
A research group at KAIST has engineered a bacterium capable of producing a natural red colorant, carminic acid, which is widely used for food and cosmetics. The research team reported the complete biosynthesis of carminic acid from glucose in engineered Escherichia coli. The strategies will be useful for the design and construction of biosynthetic pathways involving unknown enzymes and consequently the production of diverse industrially important natural products for the food, pharmaceutical, and cosmetic industries.
Carminic acid is a natural red colorant widely being used for products such as strawberry milk and lipstick. However, carminic acid has been produced by farming cochineals, a scale insect which only grows in the region around Peru and Canary Islands, followed by complicated multi-step purification processes. Moreover, carminic acid often contains protein contaminants that cause allergies so many people are unwilling to consume products made of insect-driven colorants. On that account, manufacturers around the world are using alternative red colorants despite the fact that carminic acid is one of the most stable natural red colorants.
These challenges inspired the metabolic engineering research group at KAIST to address this issue. Its members include postdoctoral researchers Dongsoo Yang and Woo Dae Jang, and Distinguished Professor Sang Yup Lee of the Department of Chemical and Biomolecular Engineering. This study entitled “Production of carminic acid by metabolically engineered Escherichia coli” was published online in the Journal of the American Chemical Society (JACS) on April 2.
This research reports for the first time the development of a bacterial strain capable of producing carminic acid from glucose via metabolic engineering and computer simulation-assisted enzyme engineering. The research group optimized the type II polyketide synthase machinery to efficiently produce the precursor of carminic acid, flavokermesic acid.
Since the enzymes responsible for the remaining two reactions were neither discovered nor functional, biochemical reaction analysis was performed to identify enzymes that can convert flavokermesic acid into carminic acid. Then, homology modeling and docking simulations were performed to enhance the activities of the two identified enzymes. The team could confirm that the final engineered strain could produce carminic acid directly from glucose. The C-glucosyltransferase developed in this study was found to be generally applicable for other natural products as showcased by the successful production of an additional product, aloesin, which is found in aloe leaves.
“The most important part of this research is that unknown enzymes for the production of target natural products were identified and improved by biochemical reaction analyses and computer simulation-assisted enzyme engineering,” says Dr. Dongsoo Yang. He explained the development of a generally applicable C-glucosyltransferase is also useful since C-glucosylation is a relatively unexplored reaction in bacteria including Escherichia coli. Using the C-glucosyltransferase developed in this study, both carminic acid and aloesin were successfully produced from glucose.
“A sustainable and insect-free method of producing carminic acid was achieved for the first time in this study. Unknown or inefficient enzymes have always been a major problem in natural product biosynthesis, and here we suggest one effective solution for solving this problem. As maintaining good health in the aging society is becoming increasingly important, we expect that the technology and strategies developed here will play pivotal roles in producing other valuable natural products of medical or nutritional importance,” said Distinguished Professor Sang Yup Lee.
This work was supported by the Technology Development Program to Solve Climate Changes on Systems Metabolic Engineering for Biorefineries of the Ministry of Science and ICT (MSIT) through the National Research Foundation (NRF) of Korea and the KAIST Cross-Generation Collaborative Lab project; Sang Yup Lee and Dongsoo Yang were also supported by Novo Nordisk Foundation in Denmark.
Publication:
Dongsoo Yang, Woo Dae Jang, and Sang Yup Lee. Production of carminic acid by metabolically engineered Escherichia coli. at the Journal of the American Chemical Society. https://doi.org.10.1021/jacs.0c12406
Profile:
Sang Yup Lee, PhD
Distinguished Professor
leesy@kaist.ac.kr
http://mbel.kaist.ac.kr
Metabolic &Biomolecular Engineering National Research Laboratory
Department of Chemical and Biomolecular Engineering
KAIST
2021.04.06 View 13769
Microbial Production of a Natural Red Colorant Carminic Acid
Metabolic engineering and computer-simulated enzyme engineering led to the production of carminic acid, a natural red colorant, from bacteria for the first time
A research group at KAIST has engineered a bacterium capable of producing a natural red colorant, carminic acid, which is widely used for food and cosmetics. The research team reported the complete biosynthesis of carminic acid from glucose in engineered Escherichia coli. The strategies will be useful for the design and construction of biosynthetic pathways involving unknown enzymes and consequently the production of diverse industrially important natural products for the food, pharmaceutical, and cosmetic industries.
Carminic acid is a natural red colorant widely being used for products such as strawberry milk and lipstick. However, carminic acid has been produced by farming cochineals, a scale insect which only grows in the region around Peru and Canary Islands, followed by complicated multi-step purification processes. Moreover, carminic acid often contains protein contaminants that cause allergies so many people are unwilling to consume products made of insect-driven colorants. On that account, manufacturers around the world are using alternative red colorants despite the fact that carminic acid is one of the most stable natural red colorants.
These challenges inspired the metabolic engineering research group at KAIST to address this issue. Its members include postdoctoral researchers Dongsoo Yang and Woo Dae Jang, and Distinguished Professor Sang Yup Lee of the Department of Chemical and Biomolecular Engineering. This study entitled “Production of carminic acid by metabolically engineered Escherichia coli” was published online in the Journal of the American Chemical Society (JACS) on April 2.
This research reports for the first time the development of a bacterial strain capable of producing carminic acid from glucose via metabolic engineering and computer simulation-assisted enzyme engineering. The research group optimized the type II polyketide synthase machinery to efficiently produce the precursor of carminic acid, flavokermesic acid.
Since the enzymes responsible for the remaining two reactions were neither discovered nor functional, biochemical reaction analysis was performed to identify enzymes that can convert flavokermesic acid into carminic acid. Then, homology modeling and docking simulations were performed to enhance the activities of the two identified enzymes. The team could confirm that the final engineered strain could produce carminic acid directly from glucose. The C-glucosyltransferase developed in this study was found to be generally applicable for other natural products as showcased by the successful production of an additional product, aloesin, which is found in aloe leaves.
“The most important part of this research is that unknown enzymes for the production of target natural products were identified and improved by biochemical reaction analyses and computer simulation-assisted enzyme engineering,” says Dr. Dongsoo Yang. He explained the development of a generally applicable C-glucosyltransferase is also useful since C-glucosylation is a relatively unexplored reaction in bacteria including Escherichia coli. Using the C-glucosyltransferase developed in this study, both carminic acid and aloesin were successfully produced from glucose.
“A sustainable and insect-free method of producing carminic acid was achieved for the first time in this study. Unknown or inefficient enzymes have always been a major problem in natural product biosynthesis, and here we suggest one effective solution for solving this problem. As maintaining good health in the aging society is becoming increasingly important, we expect that the technology and strategies developed here will play pivotal roles in producing other valuable natural products of medical or nutritional importance,” said Distinguished Professor Sang Yup Lee.
This work was supported by the Technology Development Program to Solve Climate Changes on Systems Metabolic Engineering for Biorefineries of the Ministry of Science and ICT (MSIT) through the National Research Foundation (NRF) of Korea and the KAIST Cross-Generation Collaborative Lab project; Sang Yup Lee and Dongsoo Yang were also supported by Novo Nordisk Foundation in Denmark.
Publication:
Dongsoo Yang, Woo Dae Jang, and Sang Yup Lee. Production of carminic acid by metabolically engineered Escherichia coli. at the Journal of the American Chemical Society. https://doi.org.10.1021/jacs.0c12406
Profile:
Sang Yup Lee, PhD
Distinguished Professor
leesy@kaist.ac.kr
http://mbel.kaist.ac.kr
Metabolic &Biomolecular Engineering National Research Laboratory
Department of Chemical and Biomolecular Engineering
KAIST
2021.04.06 View 13769 -
 Electrosprayed Micro Droplets Help Kill Bacteria and Viruses
With COVID-19 raging around the globe, researchers are doubling down on methods for developing diverse antimicrobial technologies that could be effective in killing a virus, but harmless to humans and the environment.
A recent study by a KAIST research team will be one of the responses to such efforts. Professor Seung Seob Lee and Dr. Ji-hun Jeong from the Department of Mechanical Engineering developed a harmless air sterilization prototype featuring electrosprayed water from a polymer micro-nozzle array. This study is one of the projects being supported by the KAIST New Deal R&D Initiative in response to COVID-19. Their study was reported in Polymer.
The electrosprayed microdroplets encapsulate reactive oxygen species such as hydroxyl radicals, superoxides that are known to have an antimicrobial function. The encapsulation prolongs the life of reactive oxygen species, which enable the droplets to perform their antimicrobial function effectively. Prior research has already proven the antimicrobial and encapsulation effects of electrosprayed droplets.
Despite its potential for antimicrobial applications, electrosprayed water generally operates under an electrical discharge condition, which can generate ozone. The inhalation of ozone is known to cause damage to the respiratory system of humans. Another technical barrier for electrospraying is the low flow rate problem. Since electrospraying exhibits the dependence of droplet size on the flow rate, there is a limit for the amount of water microdroplets a single nozzle can produce.
With this in mind, the research team developed a dielectric polymer micro-nozzle array to perform the multiplexed electrospraying of water without electrical discharge. The polymer micro-nozzle array was fabricated using the MEMS (Micro Electro-Mechanical System) process. According to the research team, the nozzle can carry five to 19 micro-nozzles depending on the required application.
The high aspect ratio of the micro-nozzle and an in-plane extractor were proposed to concentrate the electric field at the tip of the micro-nozzle, which prevents the electrical discharge caused by the high surface tension of water. A micro-pillar array with a hydrophobic coating around the micro-nozzle was also proposed to prevent the wetting of the micro-nozzle array.
The polymer micro-nozzle array performed in steady cone jet mode without electrical discharge as confirmed by high-speed imaging and nanosecond pulsed imaging. The water microdroplets were measured to be in the range of six to 10 μm and displayed an antimicrobial effect on Escherichia coli and Staphylococcus aureus.
Professor Lee said, “We believe that this research can be applied to air conditioning products in areas that require antimicrobial and humidifying functions.”
Publication:
Jeong, J. H., et al. (2020) Polymer micro-atomizer for water electrospray in the cone jet mode. Polymer. Vol. No. 194, 122405. Available online at https://doi.org/10.1016/j.polymer.2020.122405
Profile: Seung Seob Lee, Ph.D.
sslee97@kaist.ac.kr
http://mmst.kaist.ac.kr/
Professor
Department of Mechanical Engineering (ME)
Korea Advanced Institute of Science and Technology (KAIST)
https://www.kaist.ac.kr
Daejeon 34141, Korea
Profile: Ji-hun Jeong, Ph.D.
jiuni6022@kaist.ac.kr
Postdoctoral researcher
Department of Mechanical Engineering (ME)
Korea Advanced Institute of Science and Technology (KAIST)
https://www.kaist.ac.kr
Daejeon 34141, Korea
(END)
2020.12.21 View 15539
Electrosprayed Micro Droplets Help Kill Bacteria and Viruses
With COVID-19 raging around the globe, researchers are doubling down on methods for developing diverse antimicrobial technologies that could be effective in killing a virus, but harmless to humans and the environment.
A recent study by a KAIST research team will be one of the responses to such efforts. Professor Seung Seob Lee and Dr. Ji-hun Jeong from the Department of Mechanical Engineering developed a harmless air sterilization prototype featuring electrosprayed water from a polymer micro-nozzle array. This study is one of the projects being supported by the KAIST New Deal R&D Initiative in response to COVID-19. Their study was reported in Polymer.
The electrosprayed microdroplets encapsulate reactive oxygen species such as hydroxyl radicals, superoxides that are known to have an antimicrobial function. The encapsulation prolongs the life of reactive oxygen species, which enable the droplets to perform their antimicrobial function effectively. Prior research has already proven the antimicrobial and encapsulation effects of electrosprayed droplets.
Despite its potential for antimicrobial applications, electrosprayed water generally operates under an electrical discharge condition, which can generate ozone. The inhalation of ozone is known to cause damage to the respiratory system of humans. Another technical barrier for electrospraying is the low flow rate problem. Since electrospraying exhibits the dependence of droplet size on the flow rate, there is a limit for the amount of water microdroplets a single nozzle can produce.
With this in mind, the research team developed a dielectric polymer micro-nozzle array to perform the multiplexed electrospraying of water without electrical discharge. The polymer micro-nozzle array was fabricated using the MEMS (Micro Electro-Mechanical System) process. According to the research team, the nozzle can carry five to 19 micro-nozzles depending on the required application.
The high aspect ratio of the micro-nozzle and an in-plane extractor were proposed to concentrate the electric field at the tip of the micro-nozzle, which prevents the electrical discharge caused by the high surface tension of water. A micro-pillar array with a hydrophobic coating around the micro-nozzle was also proposed to prevent the wetting of the micro-nozzle array.
The polymer micro-nozzle array performed in steady cone jet mode without electrical discharge as confirmed by high-speed imaging and nanosecond pulsed imaging. The water microdroplets were measured to be in the range of six to 10 μm and displayed an antimicrobial effect on Escherichia coli and Staphylococcus aureus.
Professor Lee said, “We believe that this research can be applied to air conditioning products in areas that require antimicrobial and humidifying functions.”
Publication:
Jeong, J. H., et al. (2020) Polymer micro-atomizer for water electrospray in the cone jet mode. Polymer. Vol. No. 194, 122405. Available online at https://doi.org/10.1016/j.polymer.2020.122405
Profile: Seung Seob Lee, Ph.D.
sslee97@kaist.ac.kr
http://mmst.kaist.ac.kr/
Professor
Department of Mechanical Engineering (ME)
Korea Advanced Institute of Science and Technology (KAIST)
https://www.kaist.ac.kr
Daejeon 34141, Korea
Profile: Ji-hun Jeong, Ph.D.
jiuni6022@kaist.ac.kr
Postdoctoral researcher
Department of Mechanical Engineering (ME)
Korea Advanced Institute of Science and Technology (KAIST)
https://www.kaist.ac.kr
Daejeon 34141, Korea
(END)
2020.12.21 View 15539