biosynthesis
-
 Microbial Production of a Natural Red Colorant Carminic Acid
Metabolic engineering and computer-simulated enzyme engineering led to the production of carminic acid, a natural red colorant, from bacteria for the first time
A research group at KAIST has engineered a bacterium capable of producing a natural red colorant, carminic acid, which is widely used for food and cosmetics. The research team reported the complete biosynthesis of carminic acid from glucose in engineered Escherichia coli. The strategies will be useful for the design and construction of biosynthetic pathways involving unknown enzymes and consequently the production of diverse industrially important natural products for the food, pharmaceutical, and cosmetic industries.
Carminic acid is a natural red colorant widely being used for products such as strawberry milk and lipstick. However, carminic acid has been produced by farming cochineals, a scale insect which only grows in the region around Peru and Canary Islands, followed by complicated multi-step purification processes. Moreover, carminic acid often contains protein contaminants that cause allergies so many people are unwilling to consume products made of insect-driven colorants. On that account, manufacturers around the world are using alternative red colorants despite the fact that carminic acid is one of the most stable natural red colorants.
These challenges inspired the metabolic engineering research group at KAIST to address this issue. Its members include postdoctoral researchers Dongsoo Yang and Woo Dae Jang, and Distinguished Professor Sang Yup Lee of the Department of Chemical and Biomolecular Engineering. This study entitled “Production of carminic acid by metabolically engineered Escherichia coli” was published online in the Journal of the American Chemical Society (JACS) on April 2.
This research reports for the first time the development of a bacterial strain capable of producing carminic acid from glucose via metabolic engineering and computer simulation-assisted enzyme engineering. The research group optimized the type II polyketide synthase machinery to efficiently produce the precursor of carminic acid, flavokermesic acid.
Since the enzymes responsible for the remaining two reactions were neither discovered nor functional, biochemical reaction analysis was performed to identify enzymes that can convert flavokermesic acid into carminic acid. Then, homology modeling and docking simulations were performed to enhance the activities of the two identified enzymes. The team could confirm that the final engineered strain could produce carminic acid directly from glucose. The C-glucosyltransferase developed in this study was found to be generally applicable for other natural products as showcased by the successful production of an additional product, aloesin, which is found in aloe leaves.
“The most important part of this research is that unknown enzymes for the production of target natural products were identified and improved by biochemical reaction analyses and computer simulation-assisted enzyme engineering,” says Dr. Dongsoo Yang. He explained the development of a generally applicable C-glucosyltransferase is also useful since C-glucosylation is a relatively unexplored reaction in bacteria including Escherichia coli. Using the C-glucosyltransferase developed in this study, both carminic acid and aloesin were successfully produced from glucose.
“A sustainable and insect-free method of producing carminic acid was achieved for the first time in this study. Unknown or inefficient enzymes have always been a major problem in natural product biosynthesis, and here we suggest one effective solution for solving this problem. As maintaining good health in the aging society is becoming increasingly important, we expect that the technology and strategies developed here will play pivotal roles in producing other valuable natural products of medical or nutritional importance,” said Distinguished Professor Sang Yup Lee.
This work was supported by the Technology Development Program to Solve Climate Changes on Systems Metabolic Engineering for Biorefineries of the Ministry of Science and ICT (MSIT) through the National Research Foundation (NRF) of Korea and the KAIST Cross-Generation Collaborative Lab project; Sang Yup Lee and Dongsoo Yang were also supported by Novo Nordisk Foundation in Denmark.
Publication:
Dongsoo Yang, Woo Dae Jang, and Sang Yup Lee. Production of carminic acid by metabolically engineered Escherichia coli. at the Journal of the American Chemical Society. https://doi.org.10.1021/jacs.0c12406
Profile:
Sang Yup Lee, PhD
Distinguished Professor
leesy@kaist.ac.kr
http://mbel.kaist.ac.kr
Metabolic &Biomolecular Engineering National Research Laboratory
Department of Chemical and Biomolecular Engineering
KAIST
2021.04.06 View 12041
Microbial Production of a Natural Red Colorant Carminic Acid
Metabolic engineering and computer-simulated enzyme engineering led to the production of carminic acid, a natural red colorant, from bacteria for the first time
A research group at KAIST has engineered a bacterium capable of producing a natural red colorant, carminic acid, which is widely used for food and cosmetics. The research team reported the complete biosynthesis of carminic acid from glucose in engineered Escherichia coli. The strategies will be useful for the design and construction of biosynthetic pathways involving unknown enzymes and consequently the production of diverse industrially important natural products for the food, pharmaceutical, and cosmetic industries.
Carminic acid is a natural red colorant widely being used for products such as strawberry milk and lipstick. However, carminic acid has been produced by farming cochineals, a scale insect which only grows in the region around Peru and Canary Islands, followed by complicated multi-step purification processes. Moreover, carminic acid often contains protein contaminants that cause allergies so many people are unwilling to consume products made of insect-driven colorants. On that account, manufacturers around the world are using alternative red colorants despite the fact that carminic acid is one of the most stable natural red colorants.
These challenges inspired the metabolic engineering research group at KAIST to address this issue. Its members include postdoctoral researchers Dongsoo Yang and Woo Dae Jang, and Distinguished Professor Sang Yup Lee of the Department of Chemical and Biomolecular Engineering. This study entitled “Production of carminic acid by metabolically engineered Escherichia coli” was published online in the Journal of the American Chemical Society (JACS) on April 2.
This research reports for the first time the development of a bacterial strain capable of producing carminic acid from glucose via metabolic engineering and computer simulation-assisted enzyme engineering. The research group optimized the type II polyketide synthase machinery to efficiently produce the precursor of carminic acid, flavokermesic acid.
Since the enzymes responsible for the remaining two reactions were neither discovered nor functional, biochemical reaction analysis was performed to identify enzymes that can convert flavokermesic acid into carminic acid. Then, homology modeling and docking simulations were performed to enhance the activities of the two identified enzymes. The team could confirm that the final engineered strain could produce carminic acid directly from glucose. The C-glucosyltransferase developed in this study was found to be generally applicable for other natural products as showcased by the successful production of an additional product, aloesin, which is found in aloe leaves.
“The most important part of this research is that unknown enzymes for the production of target natural products were identified and improved by biochemical reaction analyses and computer simulation-assisted enzyme engineering,” says Dr. Dongsoo Yang. He explained the development of a generally applicable C-glucosyltransferase is also useful since C-glucosylation is a relatively unexplored reaction in bacteria including Escherichia coli. Using the C-glucosyltransferase developed in this study, both carminic acid and aloesin were successfully produced from glucose.
“A sustainable and insect-free method of producing carminic acid was achieved for the first time in this study. Unknown or inefficient enzymes have always been a major problem in natural product biosynthesis, and here we suggest one effective solution for solving this problem. As maintaining good health in the aging society is becoming increasingly important, we expect that the technology and strategies developed here will play pivotal roles in producing other valuable natural products of medical or nutritional importance,” said Distinguished Professor Sang Yup Lee.
This work was supported by the Technology Development Program to Solve Climate Changes on Systems Metabolic Engineering for Biorefineries of the Ministry of Science and ICT (MSIT) through the National Research Foundation (NRF) of Korea and the KAIST Cross-Generation Collaborative Lab project; Sang Yup Lee and Dongsoo Yang were also supported by Novo Nordisk Foundation in Denmark.
Publication:
Dongsoo Yang, Woo Dae Jang, and Sang Yup Lee. Production of carminic acid by metabolically engineered Escherichia coli. at the Journal of the American Chemical Society. https://doi.org.10.1021/jacs.0c12406
Profile:
Sang Yup Lee, PhD
Distinguished Professor
leesy@kaist.ac.kr
http://mbel.kaist.ac.kr
Metabolic &Biomolecular Engineering National Research Laboratory
Department of Chemical and Biomolecular Engineering
KAIST
2021.04.06 View 12041 -
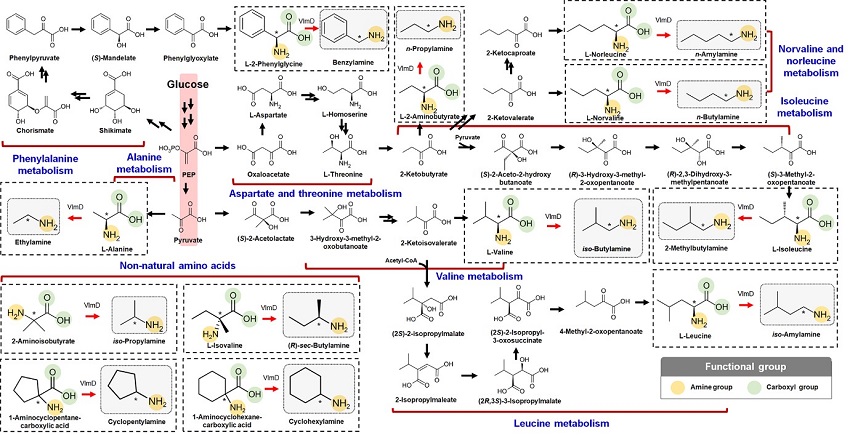 Expanding the Biosynthetic Pathway via Retrobiosynthesis
- Researchers reports a new strategy for the microbial production of multiple short-chain primary amines via retrobiosynthesis. -
KAIST metabolic engineers presented the bio-based production of multiple short-chain primary amines that have a wide range of applications in chemical industries for the first time. The research team led by Distinguished Professor Sang Yup Lee from the Department of Chemical and Biomolecular Engineering designed the novel biosynthetic pathways for short-chain primary amines by combining retrobiosynthesis and a precursor selection step.
The research team verified the newly designed pathways by confirming the in vivo production of 10 short-chain primary amines by supplying the precursors. Furthermore, the platform Escherichia coli strains were metabolically engineered to produce three proof-of-concept short-chain primary amines from glucose, demonstrating the possibility of the bio-based production of diverse short-chain primary amines from renewable resources. The research team said this study expands the strategy of systematically designing biosynthetic pathways for the production of a group of related chemicals as demonstrated by multiple short-chain primary amines as examples.
Currently, most of the industrial chemicals used in our daily lives are produced with petroleum-based products. However, there are several serious issues with the petroleum industry such as the depletion of fossil fuel reserves and environmental problems including global warming. To solve these problems, the sustainable production of industrial chemicals and materials is being explored with microorganisms as cell factories and renewable non-food biomass as raw materials for alternative to petroleum-based products. The engineering of these microorganisms has increasingly become more efficient and effective with the help of systems metabolic engineering – a practice of engineering the metabolism of a living organism toward the production of a desired metabolite. In this regard, the number of chemicals produced using biomass as a raw material has substantially increased.
Although the scope of chemicals that are producible using microorganisms continues to expand through advances in systems metabolic engineering, the biological production of short-chain primary amines has not yet been reported despite their industrial importance. Short-chain primary amines are the chemicals that have an alkyl or aryl group in the place of a hydrogen atom in ammonia with carbon chain lengths ranging from C1 to C7. Short-chain primary amines have a wide range of applications in chemical industries, for example, as a precursor for pharmaceuticals (e.g., antidiabetic and antihypertensive drugs), agrochemicals (e.g., herbicides, fungicides and insecticides), solvents, and vulcanization accelerators for rubber and plasticizers. The market size of short-chain primary amines was estimated to be more than 4 billion US dollars in 2014.
The main reason why the bio-based production of short-chain primary amines was not yet possible was due to their unknown biosynthetic pathways. Therefore, the team designed synthetic biosynthetic pathways for short-chain primary amines by combining retrobiosynthesis and a precursor selection step. The retrobiosynthesis allowed the systematic design of a biosynthetic pathway for short-chain primary amines by using a set of biochemical reaction rules that describe chemical transformation patterns between a substrate and product molecules at an atomic level.
These multiple precursors predicted for the possible biosynthesis of each short-chain primary amine were sequentially narrowed down by using the precursor selection step for efficient metabolic engineering experiments.
“Our research demonstrates the possibility of the renewable production of short-chain primary amines for the first time. We are planning to increase production efficiencies of short-chain primary amines. We believe that our study will play an important role in the development of sustainable and eco-friendly bio-based industries and the reorganization of the chemical industry, which is mandatory for solving the environmental problems threating the survival of mankind,” said Professor Lee.
This paper titled “Microbial production of multiple short-chain primary amines via retrobiosynthesis” was published in Nature Communications. This work was supported by the Technology Development Program to Solve Climate Changes on Systems Metabolic Engineering for Biorefineries from the Ministry of Science and ICT through the National Research Foundation (NRF) of Korea.
-Publication
Dong In Kim, Tong Un Chae, Hyun Uk Kim, Woo Dae Jang, and Sang Yup Lee. Microbial production of multiple short-chain primary amines via retrobiosynthesis. Nature Communications ( https://www.nature.com/articles/s41467-020-20423-6)
-Profile
Distinguished Professor Sang Yup Lee
leesy@kaist.ac.kr
Metabolic &Biomolecular Engineering National Research Laboratory
http://mbel.kaist.ac.kr
Department of Chemical and Biomolecular Engineering
KAIST
2021.01.14 View 11955
Expanding the Biosynthetic Pathway via Retrobiosynthesis
- Researchers reports a new strategy for the microbial production of multiple short-chain primary amines via retrobiosynthesis. -
KAIST metabolic engineers presented the bio-based production of multiple short-chain primary amines that have a wide range of applications in chemical industries for the first time. The research team led by Distinguished Professor Sang Yup Lee from the Department of Chemical and Biomolecular Engineering designed the novel biosynthetic pathways for short-chain primary amines by combining retrobiosynthesis and a precursor selection step.
The research team verified the newly designed pathways by confirming the in vivo production of 10 short-chain primary amines by supplying the precursors. Furthermore, the platform Escherichia coli strains were metabolically engineered to produce three proof-of-concept short-chain primary amines from glucose, demonstrating the possibility of the bio-based production of diverse short-chain primary amines from renewable resources. The research team said this study expands the strategy of systematically designing biosynthetic pathways for the production of a group of related chemicals as demonstrated by multiple short-chain primary amines as examples.
Currently, most of the industrial chemicals used in our daily lives are produced with petroleum-based products. However, there are several serious issues with the petroleum industry such as the depletion of fossil fuel reserves and environmental problems including global warming. To solve these problems, the sustainable production of industrial chemicals and materials is being explored with microorganisms as cell factories and renewable non-food biomass as raw materials for alternative to petroleum-based products. The engineering of these microorganisms has increasingly become more efficient and effective with the help of systems metabolic engineering – a practice of engineering the metabolism of a living organism toward the production of a desired metabolite. In this regard, the number of chemicals produced using biomass as a raw material has substantially increased.
Although the scope of chemicals that are producible using microorganisms continues to expand through advances in systems metabolic engineering, the biological production of short-chain primary amines has not yet been reported despite their industrial importance. Short-chain primary amines are the chemicals that have an alkyl or aryl group in the place of a hydrogen atom in ammonia with carbon chain lengths ranging from C1 to C7. Short-chain primary amines have a wide range of applications in chemical industries, for example, as a precursor for pharmaceuticals (e.g., antidiabetic and antihypertensive drugs), agrochemicals (e.g., herbicides, fungicides and insecticides), solvents, and vulcanization accelerators for rubber and plasticizers. The market size of short-chain primary amines was estimated to be more than 4 billion US dollars in 2014.
The main reason why the bio-based production of short-chain primary amines was not yet possible was due to their unknown biosynthetic pathways. Therefore, the team designed synthetic biosynthetic pathways for short-chain primary amines by combining retrobiosynthesis and a precursor selection step. The retrobiosynthesis allowed the systematic design of a biosynthetic pathway for short-chain primary amines by using a set of biochemical reaction rules that describe chemical transformation patterns between a substrate and product molecules at an atomic level.
These multiple precursors predicted for the possible biosynthesis of each short-chain primary amine were sequentially narrowed down by using the precursor selection step for efficient metabolic engineering experiments.
“Our research demonstrates the possibility of the renewable production of short-chain primary amines for the first time. We are planning to increase production efficiencies of short-chain primary amines. We believe that our study will play an important role in the development of sustainable and eco-friendly bio-based industries and the reorganization of the chemical industry, which is mandatory for solving the environmental problems threating the survival of mankind,” said Professor Lee.
This paper titled “Microbial production of multiple short-chain primary amines via retrobiosynthesis” was published in Nature Communications. This work was supported by the Technology Development Program to Solve Climate Changes on Systems Metabolic Engineering for Biorefineries from the Ministry of Science and ICT through the National Research Foundation (NRF) of Korea.
-Publication
Dong In Kim, Tong Un Chae, Hyun Uk Kim, Woo Dae Jang, and Sang Yup Lee. Microbial production of multiple short-chain primary amines via retrobiosynthesis. Nature Communications ( https://www.nature.com/articles/s41467-020-20423-6)
-Profile
Distinguished Professor Sang Yup Lee
leesy@kaist.ac.kr
Metabolic &Biomolecular Engineering National Research Laboratory
http://mbel.kaist.ac.kr
Department of Chemical and Biomolecular Engineering
KAIST
2021.01.14 View 11955 -
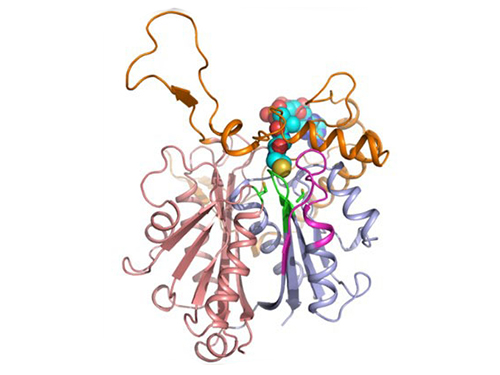 Discovery of Redox-Switch of KEenzyme Involved in N-Butanol Biosynthesis
Research teams at KAIST and Kyungpook National University (KNU) have succeeded in uncovering the redox-switch of thiolase, a key enzyme for n-butanol production in Clostridium acetobutylicum, one of the best known butanol-producing bacteria.
Biological n-butanol production was first reported by Louis Pasteur in 1861, and the bioprocess was industrialized usingClostridium acetobutylicum. The fermentation process by Clostridium strains has been known to be the most efficient one for n-butanol production. Due to growing world-wide issues such as energy security and climate change, the biological production of n-butanol has been receiving much renewed interest. This is because n-butanol possesses much better fuel characteristics compared to ethanol, such as higher energy content (29.2 MJ/L vs 19.6 MJ/L), less corrosiveness, less hygroscopy, and the ease with which it can be blended with gasoline and diesel.
In the paper published in Nature Communications, a broad-scope, online-only, and open access journal issued by the Nature Publishing Group (NPG), on September 22, 2015, Professor Kyung-Jin Kim at the School of Life Sciences, KNU, and Distinguished Professor Sang Yup Lee at the Department of Chemical and Biomolecular Engineering, KAIST, have proved that the redox-switch of thiolase plays a role in a regulation of metabolic flux in C. acetobutylicum by using in silico modeling and simulation tools.
The research team has redesigned thiolase with enhanced activity on the basis of the 3D structure of the wild-type enzyme. To reinforce a metabolic flux toward butanol production, the metabolic network of C. acetobutylicum strain was engineered with the redesigned enzyme. The combination of the discovery of 3D enzyme structure and systems metabolic engineering approaches resulted in increased n-butanol production in C. acetobutylicum, which allows the production of this important industrial chemical to be cost competitive.
Professors Kim and Lee said, "We have reported the 3D structure of C. acetobutylicum thiolase-a key enzyme involved in n-butanol biosynthesis, for the first time. Further study will be done to produce butanol more economically on the basis of the 3D structure of C. acetobutylicum thiolase."
This work was published online in Nature Communications on September 22, 2015.
Reference: Kim et al. "Redox-switch regulatory mechanism of thiolase from Clostridium acetobutylicum," Nature Communications
This research was supported by the Technology Development Program to Solve Climate Changes from the Ministry of Education, Science and Technology (MEST), Korea, the National Research Foundation of Korea, and the Advanced Biomass Center through the Global Frontier Research Program of the MEST, Korea.
For further information, contact Dr. Sang Yup Lee, Distinguished Professor, KAIST, Daejeon, Korea (leesy@kaist.ac.kr, +82-42-350-3930); and Dr. Kyung-Jin Kim, Professor, KNU, Daegu, Korea (kkim@knu.ac.kr, +82-53-950-6088).
Figure 1: A redox-switch of thiolase involves in butanol biosynthesis in Clostridium acetobutylicum. Thiolase condenses two acetyl-CoA molecules for initiating four carbon flux towards butanol.
Figure 2: Thiolase catalyzes the condensation reaction of acetyl-CoA to acetoacetyl-CoA. Two catalytic cysteine residues at 88th and 378th are oxidized and formed an intermolecular disulfide bond in an oxidized status, which results in inactivation of the enzyme for n-butanol biosynthesis. The intermolecular disulfide bond is broken enabling the n-butanol biosynthesis, when the environment status is reduced.
2015.09.23 View 10781
Discovery of Redox-Switch of KEenzyme Involved in N-Butanol Biosynthesis
Research teams at KAIST and Kyungpook National University (KNU) have succeeded in uncovering the redox-switch of thiolase, a key enzyme for n-butanol production in Clostridium acetobutylicum, one of the best known butanol-producing bacteria.
Biological n-butanol production was first reported by Louis Pasteur in 1861, and the bioprocess was industrialized usingClostridium acetobutylicum. The fermentation process by Clostridium strains has been known to be the most efficient one for n-butanol production. Due to growing world-wide issues such as energy security and climate change, the biological production of n-butanol has been receiving much renewed interest. This is because n-butanol possesses much better fuel characteristics compared to ethanol, such as higher energy content (29.2 MJ/L vs 19.6 MJ/L), less corrosiveness, less hygroscopy, and the ease with which it can be blended with gasoline and diesel.
In the paper published in Nature Communications, a broad-scope, online-only, and open access journal issued by the Nature Publishing Group (NPG), on September 22, 2015, Professor Kyung-Jin Kim at the School of Life Sciences, KNU, and Distinguished Professor Sang Yup Lee at the Department of Chemical and Biomolecular Engineering, KAIST, have proved that the redox-switch of thiolase plays a role in a regulation of metabolic flux in C. acetobutylicum by using in silico modeling and simulation tools.
The research team has redesigned thiolase with enhanced activity on the basis of the 3D structure of the wild-type enzyme. To reinforce a metabolic flux toward butanol production, the metabolic network of C. acetobutylicum strain was engineered with the redesigned enzyme. The combination of the discovery of 3D enzyme structure and systems metabolic engineering approaches resulted in increased n-butanol production in C. acetobutylicum, which allows the production of this important industrial chemical to be cost competitive.
Professors Kim and Lee said, "We have reported the 3D structure of C. acetobutylicum thiolase-a key enzyme involved in n-butanol biosynthesis, for the first time. Further study will be done to produce butanol more economically on the basis of the 3D structure of C. acetobutylicum thiolase."
This work was published online in Nature Communications on September 22, 2015.
Reference: Kim et al. "Redox-switch regulatory mechanism of thiolase from Clostridium acetobutylicum," Nature Communications
This research was supported by the Technology Development Program to Solve Climate Changes from the Ministry of Education, Science and Technology (MEST), Korea, the National Research Foundation of Korea, and the Advanced Biomass Center through the Global Frontier Research Program of the MEST, Korea.
For further information, contact Dr. Sang Yup Lee, Distinguished Professor, KAIST, Daejeon, Korea (leesy@kaist.ac.kr, +82-42-350-3930); and Dr. Kyung-Jin Kim, Professor, KNU, Daegu, Korea (kkim@knu.ac.kr, +82-53-950-6088).
Figure 1: A redox-switch of thiolase involves in butanol biosynthesis in Clostridium acetobutylicum. Thiolase condenses two acetyl-CoA molecules for initiating four carbon flux towards butanol.
Figure 2: Thiolase catalyzes the condensation reaction of acetyl-CoA to acetoacetyl-CoA. Two catalytic cysteine residues at 88th and 378th are oxidized and formed an intermolecular disulfide bond in an oxidized status, which results in inactivation of the enzyme for n-butanol biosynthesis. The intermolecular disulfide bond is broken enabling the n-butanol biosynthesis, when the environment status is reduced.
2015.09.23 View 10781