%62%69%6f%2d%69%6d%61%67%69%6e%67
-
 Microscopy Approach Poised to Offer New Insights into Liver Diseases
Researchers have developed a new way to visualize the progression of nonalcoholic fatty liver disease (NAFLD) in mouse models of the disease. The new microscopy method provides a high-resolution 3D view that could lead to important new insights into NAFLD, a condition in which too much fat is stored in the liver.
“It is estimated that a quarter of the adult global population has NAFLD, yet an effective treatment strategy has not been found,” said professor Pilhan Kim from the Graduate School of Medical Science and Engineering at KAIST. “NAFLD is associated with obesity and type 2 diabetes and can sometimes progress to liver failure in serious case.”
In the Optical Society (OSA) journal Biomedical Optics Express, Professor Kim and colleagues reported their new imaging technique and showed that it can be used to observe how tiny droplets of fat, or lipids, accumulate in the liver cells of living mice over time.
“It has been challenging to find a treatment strategy for NAFLD because most studies examine excised liver tissue that represents just one timepoint in disease progression,” said Professor Kim. “Our technique can capture details of lipid accumulation over time, providing a highly useful research tool for identifying the multiple parameters that likely contribute to the disease and could be targeted with treatment.”
Capturing the dynamics of NAFLD in living mouse models of the disease requires the ability to observe quickly changing interactions of biological components in intact tissue in real-time. To accomplish this, the researchers developed a custom intravital confocal and two-photon microscopy system that acquires images of multiple fluorescent labels at video-rate with cellular resolution.
“With video-rate imaging capability, the continuous movement of liver tissue in live mice due to breathing and heart beating could be tracked in real time and precisely compensated,” said Professor Kim. “This provided motion-artifact free high-resolution images of cellular and sub-cellular sized individual lipid droplets.”
The key to fast imaging was a polygonal mirror that rotated at more than 240 miles per hour to provide extremely fast laser scanning. The researchers also incorporated four different lasers and four high-sensitivity optical detectors into the setup so that they could acquire multi-color images to capture different color fluorescent probes used to label the lipid droplets and microvasculature in the livers of live mice.
“Our approach can capture real-time changes in cell behavior and morphology, vascular structure and function, and the spatiotemporal localization of biological components while directly visualizing of lipid droplet development in NAFLD progression,” said Professor Kim. “It also allows the analysis of the highly complex behaviors of various immune cells as NAFLD progresses.”
The researchers demonstrated their approach by using it to observe the development and spatial distribution of lipid droplets in individual mice with NAFLD induced by a methionine and choline-deficient diet. Next, they plan to use it to study how the liver microenvironment changes during NAFLD progression by imaging the same mouse over time. They also want to use their microscope technique to visualize various immune cells and lipid droplets to better understand the complex liver microenvironment in NAFLD progression.
2020.08.21 View 14628
Microscopy Approach Poised to Offer New Insights into Liver Diseases
Researchers have developed a new way to visualize the progression of nonalcoholic fatty liver disease (NAFLD) in mouse models of the disease. The new microscopy method provides a high-resolution 3D view that could lead to important new insights into NAFLD, a condition in which too much fat is stored in the liver.
“It is estimated that a quarter of the adult global population has NAFLD, yet an effective treatment strategy has not been found,” said professor Pilhan Kim from the Graduate School of Medical Science and Engineering at KAIST. “NAFLD is associated with obesity and type 2 diabetes and can sometimes progress to liver failure in serious case.”
In the Optical Society (OSA) journal Biomedical Optics Express, Professor Kim and colleagues reported their new imaging technique and showed that it can be used to observe how tiny droplets of fat, or lipids, accumulate in the liver cells of living mice over time.
“It has been challenging to find a treatment strategy for NAFLD because most studies examine excised liver tissue that represents just one timepoint in disease progression,” said Professor Kim. “Our technique can capture details of lipid accumulation over time, providing a highly useful research tool for identifying the multiple parameters that likely contribute to the disease and could be targeted with treatment.”
Capturing the dynamics of NAFLD in living mouse models of the disease requires the ability to observe quickly changing interactions of biological components in intact tissue in real-time. To accomplish this, the researchers developed a custom intravital confocal and two-photon microscopy system that acquires images of multiple fluorescent labels at video-rate with cellular resolution.
“With video-rate imaging capability, the continuous movement of liver tissue in live mice due to breathing and heart beating could be tracked in real time and precisely compensated,” said Professor Kim. “This provided motion-artifact free high-resolution images of cellular and sub-cellular sized individual lipid droplets.”
The key to fast imaging was a polygonal mirror that rotated at more than 240 miles per hour to provide extremely fast laser scanning. The researchers also incorporated four different lasers and four high-sensitivity optical detectors into the setup so that they could acquire multi-color images to capture different color fluorescent probes used to label the lipid droplets and microvasculature in the livers of live mice.
“Our approach can capture real-time changes in cell behavior and morphology, vascular structure and function, and the spatiotemporal localization of biological components while directly visualizing of lipid droplet development in NAFLD progression,” said Professor Kim. “It also allows the analysis of the highly complex behaviors of various immune cells as NAFLD progresses.”
The researchers demonstrated their approach by using it to observe the development and spatial distribution of lipid droplets in individual mice with NAFLD induced by a methionine and choline-deficient diet. Next, they plan to use it to study how the liver microenvironment changes during NAFLD progression by imaging the same mouse over time. They also want to use their microscope technique to visualize various immune cells and lipid droplets to better understand the complex liver microenvironment in NAFLD progression.
2020.08.21 View 14628 -
 The new era of personalized cancer diagnosis and treatment
Professor Tae-Young Yoon
- Succeeded in observing carcinogenic protein at the molecular level
- “Paved the way to customized cancer treatment through accurate analysis of carcinogenic protein”
The joint KAIST research team of Professor Tae Young Yoon of the Department of Physics and Professor Won Do Huh of the Department of Biological Sciences have developed the technology to monitor characteristics of carcinogenic protein in cancer tissue – for the first time in the world.
The technology makes it possible to analyse the mechanism of cancer development through a small amount of carcinogenic protein from a cancer patient. Therefore, a personalised approach to diagnosis and treatment using the knowledge of the specific mechanism of cancer development in the patient may be possible in the future.
Until recently, modern medicine could only speculate on the cause of cancer through statistics. Although developed countries, such as the United States, are known to use a large sequencing technology that analyses the patient’s DNA, identification of the interactions between proteins responsible for causing cancer remained an unanswered question for a long time in medicine.
Firstly, Professor Yoon’s research team has developed a fluorescent microscope that can observe even a single molecule. Then, the “Immunoprecipitation method”, a technology to extract a specific protein exploiting the high affinity between antigens and antibodies was developed. Using this technology and the microscope, “Real-Time Single Molecule co-Immunoprecipitation Method” was created. In this way, the team succeeded in observing the interactions between carcinogenic and other proteins at a molecular level, in real time.
To validate the developed technology, the team investigated Ras, a carcinogenic protein; its mutation statistically is known to cause around 30% of cancers.
The experimental results confirmed that 30-50% of Ras protein was expressed in mouse tumour and human cancer cells. In normal cells, less than 5% of Ras protein was expressed. Thus, the experiment showed that unusual increase in activation of Ras protein induces cancer.
The increase in the ratio of active Ras protein can be inferred from existing research data but the measurement of specific numerical data has never been done before.
The team suggested a new molecular level diagnosis technique of identifying the progress of cancer in patients through measuring the percentage of activated carcinogenic protein in cancer tissue.
Professor Yoon Tae-young said, “This newly developed technology does not require a separate procedure of protein expression or refining, hence the existing proteins in real biological tissues or cancer cells can be observed directly.” He also said, “Since carcinogenic protein can be analyzed accurately, it has opened up the path to customized cancer treatment in the future.”
“Since the observation is possible on a molecular level, the technology confers the advantage that researchers can carry out various examinations on a small sample of the cancer patient.” He added, “The clinical trial will start in December 2012 and in a few years customized cancer diagnosis and treatment will be possible.”
Meanwhile, the research has been published in Nature Communications (February 19). Many researchers from various fields have participated, regardless of the differences in their speciality, and successfully produced interdisciplinary research. Professor Tae Young Yoon of the Department of Physics and Professors Dae Sik Lim and Won Do Huh of Biological Sciences at KAIST, and Professor Chang Bong Hyun of Computational Science of KIAS contributed to developing the technique.
Figure 1: Schematic diagram of observed interactions at the molecular level in real time using fluorescent microscope. The carcinogenic protein from a mouse tumour is fixed on the microchip, and its molecular characteristics are observed live.
Figure 2: Molecular interaction data using a molecular level fluorescent microscope. A signal in the form of spike is shown when two proteins combine. This is monitored live using an Electron Multiplying Charge Coupled Device (EMCCD). It shows signal results in bright dots.
An organism has an immune system as a defence mechanism to foreign intruders. The immune system is activated when unwanted pathogens or foreign protein are in the body. Antibodies form in recognition of the specific antigen to protect itself. Organisms evolved to form antibodies with high specificity to a certain antigen. Antibodies only react to its complementary antigens. The field of molecular biology uses the affinity between antigens and antibodies to extract specific proteins; a technology called immunoprecipitation. Even in a mixture of many proteins, the protein sought can be extracted using antibodies. Thus immunoprecipitation is widely used to detect pathogens or to extract specific proteins.
Technology co-IP is a well-known example that uses immunoprecipitation. The research on interactions between proteins uses co-IP in general. The basis of fixing the antigen on the antibody to extract antigen protein is the same as immunoprecipitation. Then, researchers inject and observe its reaction with the partner protein to observe the interactions and precipitate the antibodies. If the reaction occurs, the partner protein will be found with the antibodies in the precipitations. If not, then the partner protein will not be found. This shows that the two proteins interact.
However, the traditional co-IP can be used to infer the interactions between the two proteins although the information of the dynamics on how the reaction occurs is lost. To overcome these shortcomings, the Real-Time Single Molecule co-IP Method enables observation on individual protein level in real time. Therefore, the significance of the new technique is in making observation of interactions more direct and quantitative.
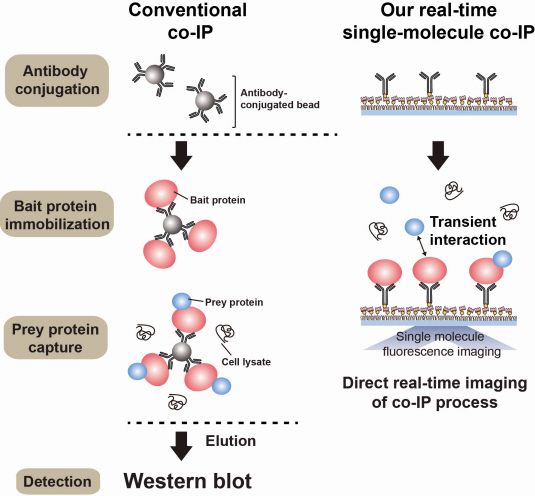
Additional Figure 1: Comparison between Conventional co-IP and Real-Time Single Molecule co-IP
2013.04.01 View 26474
The new era of personalized cancer diagnosis and treatment
Professor Tae-Young Yoon
- Succeeded in observing carcinogenic protein at the molecular level
- “Paved the way to customized cancer treatment through accurate analysis of carcinogenic protein”
The joint KAIST research team of Professor Tae Young Yoon of the Department of Physics and Professor Won Do Huh of the Department of Biological Sciences have developed the technology to monitor characteristics of carcinogenic protein in cancer tissue – for the first time in the world.
The technology makes it possible to analyse the mechanism of cancer development through a small amount of carcinogenic protein from a cancer patient. Therefore, a personalised approach to diagnosis and treatment using the knowledge of the specific mechanism of cancer development in the patient may be possible in the future.
Until recently, modern medicine could only speculate on the cause of cancer through statistics. Although developed countries, such as the United States, are known to use a large sequencing technology that analyses the patient’s DNA, identification of the interactions between proteins responsible for causing cancer remained an unanswered question for a long time in medicine.
Firstly, Professor Yoon’s research team has developed a fluorescent microscope that can observe even a single molecule. Then, the “Immunoprecipitation method”, a technology to extract a specific protein exploiting the high affinity between antigens and antibodies was developed. Using this technology and the microscope, “Real-Time Single Molecule co-Immunoprecipitation Method” was created. In this way, the team succeeded in observing the interactions between carcinogenic and other proteins at a molecular level, in real time.
To validate the developed technology, the team investigated Ras, a carcinogenic protein; its mutation statistically is known to cause around 30% of cancers.
The experimental results confirmed that 30-50% of Ras protein was expressed in mouse tumour and human cancer cells. In normal cells, less than 5% of Ras protein was expressed. Thus, the experiment showed that unusual increase in activation of Ras protein induces cancer.
The increase in the ratio of active Ras protein can be inferred from existing research data but the measurement of specific numerical data has never been done before.
The team suggested a new molecular level diagnosis technique of identifying the progress of cancer in patients through measuring the percentage of activated carcinogenic protein in cancer tissue.
Professor Yoon Tae-young said, “This newly developed technology does not require a separate procedure of protein expression or refining, hence the existing proteins in real biological tissues or cancer cells can be observed directly.” He also said, “Since carcinogenic protein can be analyzed accurately, it has opened up the path to customized cancer treatment in the future.”
“Since the observation is possible on a molecular level, the technology confers the advantage that researchers can carry out various examinations on a small sample of the cancer patient.” He added, “The clinical trial will start in December 2012 and in a few years customized cancer diagnosis and treatment will be possible.”
Meanwhile, the research has been published in Nature Communications (February 19). Many researchers from various fields have participated, regardless of the differences in their speciality, and successfully produced interdisciplinary research. Professor Tae Young Yoon of the Department of Physics and Professors Dae Sik Lim and Won Do Huh of Biological Sciences at KAIST, and Professor Chang Bong Hyun of Computational Science of KIAS contributed to developing the technique.
Figure 1: Schematic diagram of observed interactions at the molecular level in real time using fluorescent microscope. The carcinogenic protein from a mouse tumour is fixed on the microchip, and its molecular characteristics are observed live.
Figure 2: Molecular interaction data using a molecular level fluorescent microscope. A signal in the form of spike is shown when two proteins combine. This is monitored live using an Electron Multiplying Charge Coupled Device (EMCCD). It shows signal results in bright dots.
An organism has an immune system as a defence mechanism to foreign intruders. The immune system is activated when unwanted pathogens or foreign protein are in the body. Antibodies form in recognition of the specific antigen to protect itself. Organisms evolved to form antibodies with high specificity to a certain antigen. Antibodies only react to its complementary antigens. The field of molecular biology uses the affinity between antigens and antibodies to extract specific proteins; a technology called immunoprecipitation. Even in a mixture of many proteins, the protein sought can be extracted using antibodies. Thus immunoprecipitation is widely used to detect pathogens or to extract specific proteins.
Technology co-IP is a well-known example that uses immunoprecipitation. The research on interactions between proteins uses co-IP in general. The basis of fixing the antigen on the antibody to extract antigen protein is the same as immunoprecipitation. Then, researchers inject and observe its reaction with the partner protein to observe the interactions and precipitate the antibodies. If the reaction occurs, the partner protein will be found with the antibodies in the precipitations. If not, then the partner protein will not be found. This shows that the two proteins interact.
However, the traditional co-IP can be used to infer the interactions between the two proteins although the information of the dynamics on how the reaction occurs is lost. To overcome these shortcomings, the Real-Time Single Molecule co-IP Method enables observation on individual protein level in real time. Therefore, the significance of the new technique is in making observation of interactions more direct and quantitative.
Additional Figure 1: Comparison between Conventional co-IP and Real-Time Single Molecule co-IP
2013.04.01 View 26474