%4e%61%74%75%72%65%20%43%6f%6d%6d%75%6e%69%63%61%74%69%6f%6e%73
-
 “One Experiment Is All It Takes”: KAIST Team Revolutionizes Drug Interaction Testing, Replacing 60,000 Studies
A groundbreaking new method developed by researchers at KAIST and Chungnam National University could drastically streamline drug interaction testing — replacing dozens of traditional experiments with just one.
The research, led by Professor Jae Kyoung Kim of KAIST Department of Mathematical Sciences & IBS Biomedical Mathematics Group and Professor Sang Kyum Kim of Chungnam National University's College of Pharmacy, introduces a novel analysis technique called 50-BOA, published in Nature Communications on June 5, 2025.
< Photo 1. (From left) Professor Sang Kyum Kim (Chungnam National University College of Pharmacy, co-corresponding author), Dr. Yun Min Song (IBS Biomedical Mathematics Group, formerly KAIST Department of Mathematical Sciences, co-first author), undergraduate student Hyeong Jun Jang (KAIST, co-first author), Professor Jae Kyoung Kim (KAIST and IBS Biomedical Mathematics Group, co-corresponding author) (Top left in the bubble) Professor Hwi-yeol Yun (Chungnam National University College of Pharmacy, co-author) >
For decades, scientists have had to repeat drug inhibition experiments across a wide range of concentrations to estimate inhibition constants — a process seen in over 60,000 scientific publications. But the KAIST-led team discovered that a single, well-chosen inhibitor concentration can yield even more accurate results.
< Figure 1. Graphical summary of 50-BOA. 50-BOA improves the accuracy and efficiency of inhibition constant estimation by using only a single inhibitor concentration instead of the traditionally used method of employing multiple inhibitor concentrations. >
“This approach challenges long-standing assumptions in experimental pharmacology,” says Prof. Kim. “It shows how mathematics can fundamentally redesign life science experiments.”
By mathematically analyzing the sources of error in conventional methods, the team found that over half the data typically collected adds no value or even skews results. Their new method not only cuts experimental effort by over 75%, but also enhances reproducibility and accuracy.
To help researchers adopt the method quickly, the team developed a user-friendly tool that takes simple Excel files as input, now freely available on GitHub:
☞ https://github.com/Mathbiomed/50-BOA
< Figure 2. The MATLAB and R package of 50-BOA at GitHub >
The work holds promise for faster and more reliable drug development, especially in assessing potential interactions in combination therapies. The U.S. FDA already emphasizes the importance of accurate enzyme inhibition assessment during early-stage drug evaluation — and this method could soon become a new gold standard.
2025.06.16 View 8801
“One Experiment Is All It Takes”: KAIST Team Revolutionizes Drug Interaction Testing, Replacing 60,000 Studies
A groundbreaking new method developed by researchers at KAIST and Chungnam National University could drastically streamline drug interaction testing — replacing dozens of traditional experiments with just one.
The research, led by Professor Jae Kyoung Kim of KAIST Department of Mathematical Sciences & IBS Biomedical Mathematics Group and Professor Sang Kyum Kim of Chungnam National University's College of Pharmacy, introduces a novel analysis technique called 50-BOA, published in Nature Communications on June 5, 2025.
< Photo 1. (From left) Professor Sang Kyum Kim (Chungnam National University College of Pharmacy, co-corresponding author), Dr. Yun Min Song (IBS Biomedical Mathematics Group, formerly KAIST Department of Mathematical Sciences, co-first author), undergraduate student Hyeong Jun Jang (KAIST, co-first author), Professor Jae Kyoung Kim (KAIST and IBS Biomedical Mathematics Group, co-corresponding author) (Top left in the bubble) Professor Hwi-yeol Yun (Chungnam National University College of Pharmacy, co-author) >
For decades, scientists have had to repeat drug inhibition experiments across a wide range of concentrations to estimate inhibition constants — a process seen in over 60,000 scientific publications. But the KAIST-led team discovered that a single, well-chosen inhibitor concentration can yield even more accurate results.
< Figure 1. Graphical summary of 50-BOA. 50-BOA improves the accuracy and efficiency of inhibition constant estimation by using only a single inhibitor concentration instead of the traditionally used method of employing multiple inhibitor concentrations. >
“This approach challenges long-standing assumptions in experimental pharmacology,” says Prof. Kim. “It shows how mathematics can fundamentally redesign life science experiments.”
By mathematically analyzing the sources of error in conventional methods, the team found that over half the data typically collected adds no value or even skews results. Their new method not only cuts experimental effort by over 75%, but also enhances reproducibility and accuracy.
To help researchers adopt the method quickly, the team developed a user-friendly tool that takes simple Excel files as input, now freely available on GitHub:
☞ https://github.com/Mathbiomed/50-BOA
< Figure 2. The MATLAB and R package of 50-BOA at GitHub >
The work holds promise for faster and more reliable drug development, especially in assessing potential interactions in combination therapies. The U.S. FDA already emphasizes the importance of accurate enzyme inhibition assessment during early-stage drug evaluation — and this method could soon become a new gold standard.
2025.06.16 View 8801 -
 KAIST Develops Virtual Staining Technology for 3D Histopathology
Moving beyond traditional methods of observing thinly sliced and stained cancer tissues, a collaborative international research team led by KAIST has successfully developed a groundbreaking technology. This innovation uses advanced optical techniques combined with an artificial intelligence-based deep learning algorithm to create realistic, virtually stained 3D images of cancer tissue without the need for serial sectioning nor staining. This breakthrough is anticipated to pave the way for next-generation non-invasive pathological diagnosis.
< Photo 1. (From left) Juyeon Park (Ph.D. Candidate, Department of Physics), Professor YongKeun Park (Department of Physics) (Top left) Professor Su-Jin Shin (Gangnam Severance Hospital), Professor Tae Hyun Hwang (Vanderbilt University School of Medicine) >
KAIST (President Kwang Hyung Lee) announced on the 26th that a research team led by Professor YongKeun Park of the Department of Physics, in collaboration with Professor Su-Jin Shin's team at Yonsei University Gangnam Severance Hospital, Professor Tae Hyun Hwang's team at Mayo Clinic, and Tomocube's AI research team, has developed an innovative technology capable of vividly displaying the 3D structure of cancer tissues without separate staining.
For over 200 years, conventional pathology has relied on observing cancer tissues under a microscope, a method that only shows specific cross-sections of the 3D cancer tissue. This has limited the ability to understand the three-dimensional connections and spatial arrangements between cells.
To overcome this, the research team utilized holotomography (HT), an advanced optical technology, to measure the 3D refractive index information of tissues. They then integrated an AI-based deep learning algorithm to successfully generate virtual H&E* images.* H&E (Hematoxylin & Eosin): The most widely used staining method for observing pathological tissues. Hematoxylin stains cell nuclei blue, and eosin stains cytoplasm pink.
The research team quantitatively demonstrated that the images generated by this technology are highly similar to actual stained tissue images. Furthermore, the technology exhibited consistent performance across various organs and tissues, proving its versatility and reliability as a next-generation pathological analysis tool.
< Figure 1. Comparison of conventional 3D tissue pathology procedure and the 3D virtual H&E staining technology proposed in this study. The traditional method requires preparing and staining dozens of tissue slides, while the proposed technology can reduce the number of slides by up to 10 times and quickly generate H&E images without the staining process. >
Moreover, by validating the feasibility of this technology through joint research with hospitals and research institutions in Korea and the United States, utilizing Tomocube's holotomography equipment, the team demonstrated its potential for full-scale adoption in real-world pathological research settings.
Professor YongKeun Park stated, "This research marks a major advancement by transitioning pathological analysis from conventional 2D methods to comprehensive 3D imaging. It will greatly enhance biomedical research and clinical diagnostics, particularly in understanding cancer tumor boundaries and the intricate spatial arrangements of cells within tumor microenvironments."
< Figure 2. Results of AI-based 3D virtual H&E staining and quantitative analysis of pathological tissue. The virtually stained images enabled 3D reconstruction of key pathological features such as cell nuclei and glandular lumens. Based on this, various quantitative indicators, including cell nuclear distribution, volume, and surface area, could be extracted. >
This research, with Juyeon Park, a student of the Integrated Master’s and Ph.D. Program at KAIST, as the first author, was published online in the prestigious journal Nature Communications on May 22.
(Paper title: Revealing 3D microanatomical structures of unlabeled thick cancer tissues using holotomography and virtual H&E staining.
[https://doi.org/10.1038/s41467-025-59820-0]
This study was supported by the Leader Researcher Program of the National Research Foundation of Korea, the Global Industry Technology Cooperation Center Project of the Korea Institute for Advancement of Technology, and the Korea Health Industry Development Institute.
2025.05.26 View 11303
KAIST Develops Virtual Staining Technology for 3D Histopathology
Moving beyond traditional methods of observing thinly sliced and stained cancer tissues, a collaborative international research team led by KAIST has successfully developed a groundbreaking technology. This innovation uses advanced optical techniques combined with an artificial intelligence-based deep learning algorithm to create realistic, virtually stained 3D images of cancer tissue without the need for serial sectioning nor staining. This breakthrough is anticipated to pave the way for next-generation non-invasive pathological diagnosis.
< Photo 1. (From left) Juyeon Park (Ph.D. Candidate, Department of Physics), Professor YongKeun Park (Department of Physics) (Top left) Professor Su-Jin Shin (Gangnam Severance Hospital), Professor Tae Hyun Hwang (Vanderbilt University School of Medicine) >
KAIST (President Kwang Hyung Lee) announced on the 26th that a research team led by Professor YongKeun Park of the Department of Physics, in collaboration with Professor Su-Jin Shin's team at Yonsei University Gangnam Severance Hospital, Professor Tae Hyun Hwang's team at Mayo Clinic, and Tomocube's AI research team, has developed an innovative technology capable of vividly displaying the 3D structure of cancer tissues without separate staining.
For over 200 years, conventional pathology has relied on observing cancer tissues under a microscope, a method that only shows specific cross-sections of the 3D cancer tissue. This has limited the ability to understand the three-dimensional connections and spatial arrangements between cells.
To overcome this, the research team utilized holotomography (HT), an advanced optical technology, to measure the 3D refractive index information of tissues. They then integrated an AI-based deep learning algorithm to successfully generate virtual H&E* images.* H&E (Hematoxylin & Eosin): The most widely used staining method for observing pathological tissues. Hematoxylin stains cell nuclei blue, and eosin stains cytoplasm pink.
The research team quantitatively demonstrated that the images generated by this technology are highly similar to actual stained tissue images. Furthermore, the technology exhibited consistent performance across various organs and tissues, proving its versatility and reliability as a next-generation pathological analysis tool.
< Figure 1. Comparison of conventional 3D tissue pathology procedure and the 3D virtual H&E staining technology proposed in this study. The traditional method requires preparing and staining dozens of tissue slides, while the proposed technology can reduce the number of slides by up to 10 times and quickly generate H&E images without the staining process. >
Moreover, by validating the feasibility of this technology through joint research with hospitals and research institutions in Korea and the United States, utilizing Tomocube's holotomography equipment, the team demonstrated its potential for full-scale adoption in real-world pathological research settings.
Professor YongKeun Park stated, "This research marks a major advancement by transitioning pathological analysis from conventional 2D methods to comprehensive 3D imaging. It will greatly enhance biomedical research and clinical diagnostics, particularly in understanding cancer tumor boundaries and the intricate spatial arrangements of cells within tumor microenvironments."
< Figure 2. Results of AI-based 3D virtual H&E staining and quantitative analysis of pathological tissue. The virtually stained images enabled 3D reconstruction of key pathological features such as cell nuclei and glandular lumens. Based on this, various quantitative indicators, including cell nuclear distribution, volume, and surface area, could be extracted. >
This research, with Juyeon Park, a student of the Integrated Master’s and Ph.D. Program at KAIST, as the first author, was published online in the prestigious journal Nature Communications on May 22.
(Paper title: Revealing 3D microanatomical structures of unlabeled thick cancer tissues using holotomography and virtual H&E staining.
[https://doi.org/10.1038/s41467-025-59820-0]
This study was supported by the Leader Researcher Program of the National Research Foundation of Korea, the Global Industry Technology Cooperation Center Project of the Korea Institute for Advancement of Technology, and the Korea Health Industry Development Institute.
2025.05.26 View 11303 -
 No More Touch Issues on Rainy Days! KAIST Develops Human-Like Tactile Sensor
Recent advancements in robotics have enabled machines to handle delicate objects like eggs with precision, thanks to highly integrated pressure sensors that provide detailed tactile feedback. However, even the most advanced robots struggle to accurately detect pressure in complex environments involving water, bending, or electromagnetic interference. A research team at KAIST has successfully developed a pressure sensor that operates stably without external interference, even on wet surfaces like a smartphone screen covered in water, achieving human-level tactile sensitivity.
KAIST (represented by President Kwang Hyung Lee) announced on the 10th of March that a research team led by Professor Jun-Bo Yoon from the School of Electrical Engineering has developed a high-resolution pressure sensor that remains unaffected by external interference such as "ghost touches" caused by moisture on touchscreens.
Capacitive pressure sensors, widely used in touch systems due to their simple structure and durability, are essential components of human-machine interface (HMI) technologies in smartphones, wearable devices, and robots. However, they are prone to malfunctions caused by water droplets, electromagnetic interference, and curves.
To address these issues, the research team investigated the root causes of interference in capacitive pressure sensors. They identified that the "fringe field" generated at the sensor’s edges is particularly susceptible to external disturbances.
The researchers concluded that, to fundamentally resolve this issue, suppressing the fringe field was necessary. Through theoretical analysis, they determined that reducing the electrode spacing to the nanometer scale could effectively minimize the fringe field to below a few percent.
Utilizing proprietary micro/nanofabrication techniques, the team developed a nanogap pressure sensor with an electrode spacing of 900 nanometers (nm). This newly developed sensor reliably detected pressure regardless of the material exerting force and remained unaffected by bending or electromagnetic interference.
Furthermore, the team successfully implemented an artificial tactile system utilizing the developed sensor’s characteristics. Human skin contains specialized pressure receptors called Merkel’s disks. To artificially mimic them, the exclusive detection of pressure was necessary, but hadn’t been achieved by conventional sensors.
Professor Yoon’s research team overcame these challenges, developing a sensor achieving a density comparable to Merkel’s discs and enabling wireless, high-precision pressure sensing.
To explore potential applications, the researcher also developed a force touch pad system, demonstrating its ability to capture pressure magnitude and distribution with high resolution and without interference.
Professor Yoon stated, “Our nanogap pressure sensor operates reliably even in rainy conditions or sweaty environments, eliminating common touch malfunctions. We believe this innovation will significantly enhance everyday user experiences.”
He added, “This technology has the potential to revolutionize various fields, including precision tactile sensors for robotics, medical wearable devices, and next-generation augmented reality (AR) and virtual reality (VR) interfaces.”
The study was led by Jae-Soon Yang (Ph.D.), Myung-Kun Chung (Ph.D. candidate), and Jae-Young Yoo (Assistant Professor at Sungkyunkwan University, a KAIST Ph.D. graduate). The research findings were published in Nature Communications on February 27, 2025. (Paper title: “Interference-Free Nanogap Pressure Sensor Array with High Spatial Resolution for Wireless Human-Machine Interface Applications”, DOI: 10.1038/s41467-025-57232-8)
This study was supported by the National Research Foundation of Korea’s Mid-Career Researcher Program and Leading Research Center Support Program.
2025.03.14 View 8635
No More Touch Issues on Rainy Days! KAIST Develops Human-Like Tactile Sensor
Recent advancements in robotics have enabled machines to handle delicate objects like eggs with precision, thanks to highly integrated pressure sensors that provide detailed tactile feedback. However, even the most advanced robots struggle to accurately detect pressure in complex environments involving water, bending, or electromagnetic interference. A research team at KAIST has successfully developed a pressure sensor that operates stably without external interference, even on wet surfaces like a smartphone screen covered in water, achieving human-level tactile sensitivity.
KAIST (represented by President Kwang Hyung Lee) announced on the 10th of March that a research team led by Professor Jun-Bo Yoon from the School of Electrical Engineering has developed a high-resolution pressure sensor that remains unaffected by external interference such as "ghost touches" caused by moisture on touchscreens.
Capacitive pressure sensors, widely used in touch systems due to their simple structure and durability, are essential components of human-machine interface (HMI) technologies in smartphones, wearable devices, and robots. However, they are prone to malfunctions caused by water droplets, electromagnetic interference, and curves.
To address these issues, the research team investigated the root causes of interference in capacitive pressure sensors. They identified that the "fringe field" generated at the sensor’s edges is particularly susceptible to external disturbances.
The researchers concluded that, to fundamentally resolve this issue, suppressing the fringe field was necessary. Through theoretical analysis, they determined that reducing the electrode spacing to the nanometer scale could effectively minimize the fringe field to below a few percent.
Utilizing proprietary micro/nanofabrication techniques, the team developed a nanogap pressure sensor with an electrode spacing of 900 nanometers (nm). This newly developed sensor reliably detected pressure regardless of the material exerting force and remained unaffected by bending or electromagnetic interference.
Furthermore, the team successfully implemented an artificial tactile system utilizing the developed sensor’s characteristics. Human skin contains specialized pressure receptors called Merkel’s disks. To artificially mimic them, the exclusive detection of pressure was necessary, but hadn’t been achieved by conventional sensors.
Professor Yoon’s research team overcame these challenges, developing a sensor achieving a density comparable to Merkel’s discs and enabling wireless, high-precision pressure sensing.
To explore potential applications, the researcher also developed a force touch pad system, demonstrating its ability to capture pressure magnitude and distribution with high resolution and without interference.
Professor Yoon stated, “Our nanogap pressure sensor operates reliably even in rainy conditions or sweaty environments, eliminating common touch malfunctions. We believe this innovation will significantly enhance everyday user experiences.”
He added, “This technology has the potential to revolutionize various fields, including precision tactile sensors for robotics, medical wearable devices, and next-generation augmented reality (AR) and virtual reality (VR) interfaces.”
The study was led by Jae-Soon Yang (Ph.D.), Myung-Kun Chung (Ph.D. candidate), and Jae-Young Yoo (Assistant Professor at Sungkyunkwan University, a KAIST Ph.D. graduate). The research findings were published in Nature Communications on February 27, 2025. (Paper title: “Interference-Free Nanogap Pressure Sensor Array with High Spatial Resolution for Wireless Human-Machine Interface Applications”, DOI: 10.1038/s41467-025-57232-8)
This study was supported by the National Research Foundation of Korea’s Mid-Career Researcher Program and Leading Research Center Support Program.
2025.03.14 View 8635 -
 KAIST Research Team Develops an AI Framework Capable of Overcoming the Strength-Ductility Dilemma in Additive-manufactured Titanium Alloys
<(From Left) Ph.D. Student Jaejung Park and Professor Seungchul Lee of KAIST Department of Mechanical Engineering and , Professor Hyoung Seop Kim of POSTECH, and M.S.–Ph.D. Integrated Program Student Jeong Ah Lee of POSTECH. >
The KAIST research team led by Professor Seungchul Lee from Department of Mechanical Engineering, in collaboration with Professor Hyoung Seop Kim’s team at POSTECH, successfully overcame the strength–ductility dilemma of Ti 6Al 4V alloy using artificial intelligence, enabling the production of high strength, high ductility metal products. The AI developed by the team accurately predicts mechanical properties based on various 3D printing process parameters while also providing uncertainty information, and it uses both to recommend process parameters that hold high promise for 3D printing.
Among various 3D printing technologies, laser powder bed fusion is an innovative method for manufacturing Ti-6Al-4V alloy, renowned for its high strength and bio-compatibility. However, this alloy made via 3D printing has traditionally faced challenges in simultaneously achieving high strength and high ductility. Although there have been attempts to address this issue by adjusting both the printing process parameters and heat treatment conditions, the vast number of possible combinations made it difficult to explore them all through experiments and simulations alone.
The active learning framework developed by the team quickly explores a wide range of 3D printing process parameters and heat treatment conditions to recommend those expected to improve both strength and ductility of the alloy. These recommendations are based on the AI model’s predictions of ultimate tensile strength and total elongation along with associated uncertainty information for each set of process parameters and heat treatment conditions. The recommended conditions are then validated by performing 3D printing and tensile tests to obtain the true mechanical property values. These new data are incorporated into further AI model training, and through iterative exploration, the optimal process parameters and heat treatment conditions for producing high-performance alloys were determined in only five iterations. With these optimized conditions, the 3D printed Ti-6Al-4V alloy achieved an ultimate tensile strength of 1190 MPa and a total elongation of 16.5%, successfully overcoming the strength–ductility dilemma.
Professor Seungchul Lee commented, “In this study, by optimizing the 3D printing process parameters and heat treatment conditions, we were able to develop a high-strength, high-ductility Ti-6Al-4V alloy with minimal experimentation trials. Compared to previous studies, we produced an alloy with a similar ultimate tensile strength but higher total elongation, as well as that with a similar elongation but greater ultimate tensile strength.” He added, “Furthermore, if our approach is applied not only to mechanical properties but also to other properties such as thermal conductivity and thermal expansion, we anticipate that it will enable efficient exploration of 3D printing process parameters and heat treatment conditions.”
This study was published in Nature Communications on January 22 (https://doi.org/10.1038/s41467-025-56267-1), and the research was supported by the National Research Foundation of Korea’s Nano & Material Technology Development Program and the Leading Research Center Program.
2025.02.21 View 12971
KAIST Research Team Develops an AI Framework Capable of Overcoming the Strength-Ductility Dilemma in Additive-manufactured Titanium Alloys
<(From Left) Ph.D. Student Jaejung Park and Professor Seungchul Lee of KAIST Department of Mechanical Engineering and , Professor Hyoung Seop Kim of POSTECH, and M.S.–Ph.D. Integrated Program Student Jeong Ah Lee of POSTECH. >
The KAIST research team led by Professor Seungchul Lee from Department of Mechanical Engineering, in collaboration with Professor Hyoung Seop Kim’s team at POSTECH, successfully overcame the strength–ductility dilemma of Ti 6Al 4V alloy using artificial intelligence, enabling the production of high strength, high ductility metal products. The AI developed by the team accurately predicts mechanical properties based on various 3D printing process parameters while also providing uncertainty information, and it uses both to recommend process parameters that hold high promise for 3D printing.
Among various 3D printing technologies, laser powder bed fusion is an innovative method for manufacturing Ti-6Al-4V alloy, renowned for its high strength and bio-compatibility. However, this alloy made via 3D printing has traditionally faced challenges in simultaneously achieving high strength and high ductility. Although there have been attempts to address this issue by adjusting both the printing process parameters and heat treatment conditions, the vast number of possible combinations made it difficult to explore them all through experiments and simulations alone.
The active learning framework developed by the team quickly explores a wide range of 3D printing process parameters and heat treatment conditions to recommend those expected to improve both strength and ductility of the alloy. These recommendations are based on the AI model’s predictions of ultimate tensile strength and total elongation along with associated uncertainty information for each set of process parameters and heat treatment conditions. The recommended conditions are then validated by performing 3D printing and tensile tests to obtain the true mechanical property values. These new data are incorporated into further AI model training, and through iterative exploration, the optimal process parameters and heat treatment conditions for producing high-performance alloys were determined in only five iterations. With these optimized conditions, the 3D printed Ti-6Al-4V alloy achieved an ultimate tensile strength of 1190 MPa and a total elongation of 16.5%, successfully overcoming the strength–ductility dilemma.
Professor Seungchul Lee commented, “In this study, by optimizing the 3D printing process parameters and heat treatment conditions, we were able to develop a high-strength, high-ductility Ti-6Al-4V alloy with minimal experimentation trials. Compared to previous studies, we produced an alloy with a similar ultimate tensile strength but higher total elongation, as well as that with a similar elongation but greater ultimate tensile strength.” He added, “Furthermore, if our approach is applied not only to mechanical properties but also to other properties such as thermal conductivity and thermal expansion, we anticipate that it will enable efficient exploration of 3D printing process parameters and heat treatment conditions.”
This study was published in Nature Communications on January 22 (https://doi.org/10.1038/s41467-025-56267-1), and the research was supported by the National Research Foundation of Korea’s Nano & Material Technology Development Program and the Leading Research Center Program.
2025.02.21 View 12971 -
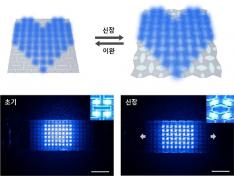 KAIST Develops Stretchable Displays Featuring 25% Expansion Without Image Distortion
Stretchable displays, praised for their spatial efficiency, design flexibility, and human-like flexibility, are seen as the next generation of display technology. A team of Korean researchers has developed a stretchable display that can expand by 25% while maintaining clear image quality without distortion. It can also stretch and contract up to 5,000 times at 15% expansion without any performance degradation, making it the first deformation-free stretchable display with a negative Poisson's ratio* developed in Korea.
*Poisson’s ratio of -1: A ratio where both width and length stretch equally, expressed as a negative value. A positive Poisson's ratio represents the ratio where horizontal stretching leads to vertical contraction, which is the case for most materials.
KAIST (represented by President Kwang-Hyung Lee) announced on the 20th of August that a research team led by Professor Byeong-Soo Bae of the Department of Materials Science and Engineering (Director of the Wearable Platform Materials Technology Center) , in collaboration with the Korea Institute of Machinery & Materials (President Seoghyeon Ryu), successfully developed a stretchable display substrate that suppresses image distortion through omnidirectional stretchability.
Currently, most stretchable displays are made with highly elastic elastomer* materials, but these materials possess a positive Poisson's ratio, causing unavoidable image distortion when the display is stretched.
*Elastomer: A polymer with elasticity similar to rubber.
To address this, the introduction of auxetic* meta-structures has been gaining attention. Unlike conventional materials, auxetic structures have a unique 'negative Poisson's ratio,' expanding in all directions when stretched in just one direction. However, traditional auxetic structures contain many empty spaces, limiting their stability and usability in display substrates.
*Auxetic structure: A special geometric structure that exhibits a negative Poisson's ratio.
To tackle the issue of image distortion, Professor Bae's research team developed a method to create a seamless surface for the auxetic meta-structure, achieving the ideal negative Poisson's ratio of -1 and overcoming the biggest challenge in auxetic meta-structures.
To overcome the second issue of elastic modulus*, the team inserted a textile made of glass fiber bundles with a diameter of just 25 micrometers (a quarter of the thickness of human hair) into the elastomer material. They then filled the empty spaces with the same elastomer, creating a flat and stable integrated film without gaps.
*Elastic Modulus: The ratio that indicates the extent of deformation when force is applied to a material. A higher elastic modulus means that the material is less likely to deform under force.
The research team theoretically identified that the difference in elasticity between the auxetic structure and the elastomer material directly influences the negative Poisson's ratio and successfully achieved an elasticity difference of over 230,000 times, producing a film with a Poisson's ratio of -1, the theoretical limit.
< Figure 2. Deformation of S-AUX film. a) Configurations and visualized principal strain distribution of the optimized S-AUX film at various strain rates. b) Biaxial stretching image. While pristine elastomer shrinks in the directions that were not stretched, S-AUX film developed in this study expands in all directions simultaneously while maintaining its original shape. >
Professor Byeong-Soo Bae, who led the study, explained, "Preventing image distortion using auxetic structures in stretchable displays is a core technology, but it has faced challenges due to the many empty spaces in the surface, making it difficult to use as a substrate. This research outcome is expected to significantly accelerate commercialization through high-resolution, distortion-free stretchable display applications that utilize the entire surface."
This study, co-authored by Dr. Yung Lee from KAIST’s Department of Materials Science and Engineering and Dr. Bongkyun Jang from the Korea Institute of Machinery & Materials, was published on August 20th in the international journal Nature Communications under the title "A seamless auxetic substrate with a negative Poisson's ratio of –1".
The research was supported by the Wearable Platform Materials Technology Center at KAIST, the Korea Institute of Machinery & Materials, and LG Display.
< Figure 3. Structural configuration of the distortion-free display components on the S-AUX film and a contour image of a micro-LED chip transferred onto the S-AUX film. >
< Figure 4. Schematic illustrations and photographic images of the S-AUX film-based image: distortion-free display in its stretched state and released state. >
2024.09.20 View 11304
KAIST Develops Stretchable Displays Featuring 25% Expansion Without Image Distortion
Stretchable displays, praised for their spatial efficiency, design flexibility, and human-like flexibility, are seen as the next generation of display technology. A team of Korean researchers has developed a stretchable display that can expand by 25% while maintaining clear image quality without distortion. It can also stretch and contract up to 5,000 times at 15% expansion without any performance degradation, making it the first deformation-free stretchable display with a negative Poisson's ratio* developed in Korea.
*Poisson’s ratio of -1: A ratio where both width and length stretch equally, expressed as a negative value. A positive Poisson's ratio represents the ratio where horizontal stretching leads to vertical contraction, which is the case for most materials.
KAIST (represented by President Kwang-Hyung Lee) announced on the 20th of August that a research team led by Professor Byeong-Soo Bae of the Department of Materials Science and Engineering (Director of the Wearable Platform Materials Technology Center) , in collaboration with the Korea Institute of Machinery & Materials (President Seoghyeon Ryu), successfully developed a stretchable display substrate that suppresses image distortion through omnidirectional stretchability.
Currently, most stretchable displays are made with highly elastic elastomer* materials, but these materials possess a positive Poisson's ratio, causing unavoidable image distortion when the display is stretched.
*Elastomer: A polymer with elasticity similar to rubber.
To address this, the introduction of auxetic* meta-structures has been gaining attention. Unlike conventional materials, auxetic structures have a unique 'negative Poisson's ratio,' expanding in all directions when stretched in just one direction. However, traditional auxetic structures contain many empty spaces, limiting their stability and usability in display substrates.
*Auxetic structure: A special geometric structure that exhibits a negative Poisson's ratio.
To tackle the issue of image distortion, Professor Bae's research team developed a method to create a seamless surface for the auxetic meta-structure, achieving the ideal negative Poisson's ratio of -1 and overcoming the biggest challenge in auxetic meta-structures.
To overcome the second issue of elastic modulus*, the team inserted a textile made of glass fiber bundles with a diameter of just 25 micrometers (a quarter of the thickness of human hair) into the elastomer material. They then filled the empty spaces with the same elastomer, creating a flat and stable integrated film without gaps.
*Elastic Modulus: The ratio that indicates the extent of deformation when force is applied to a material. A higher elastic modulus means that the material is less likely to deform under force.
The research team theoretically identified that the difference in elasticity between the auxetic structure and the elastomer material directly influences the negative Poisson's ratio and successfully achieved an elasticity difference of over 230,000 times, producing a film with a Poisson's ratio of -1, the theoretical limit.
< Figure 2. Deformation of S-AUX film. a) Configurations and visualized principal strain distribution of the optimized S-AUX film at various strain rates. b) Biaxial stretching image. While pristine elastomer shrinks in the directions that were not stretched, S-AUX film developed in this study expands in all directions simultaneously while maintaining its original shape. >
Professor Byeong-Soo Bae, who led the study, explained, "Preventing image distortion using auxetic structures in stretchable displays is a core technology, but it has faced challenges due to the many empty spaces in the surface, making it difficult to use as a substrate. This research outcome is expected to significantly accelerate commercialization through high-resolution, distortion-free stretchable display applications that utilize the entire surface."
This study, co-authored by Dr. Yung Lee from KAIST’s Department of Materials Science and Engineering and Dr. Bongkyun Jang from the Korea Institute of Machinery & Materials, was published on August 20th in the international journal Nature Communications under the title "A seamless auxetic substrate with a negative Poisson's ratio of –1".
The research was supported by the Wearable Platform Materials Technology Center at KAIST, the Korea Institute of Machinery & Materials, and LG Display.
< Figure 3. Structural configuration of the distortion-free display components on the S-AUX film and a contour image of a micro-LED chip transferred onto the S-AUX film. >
< Figure 4. Schematic illustrations and photographic images of the S-AUX film-based image: distortion-free display in its stretched state and released state. >
2024.09.20 View 11304 -
 KAIST Research Team Breaks Down Musical Instincts with AI
Music, often referred to as the universal language, is known to be a common component in all cultures. Then, could ‘musical instinct’ be something that is shared to some degree despite the extensive environmental differences amongst cultures?
On January 16, a KAIST research team led by Professor Hawoong Jung from the Department of Physics announced to have identified the principle by which musical instincts emerge from the human brain without special learning using an artificial neural network model.
Previously, many researchers have attempted to identify the similarities and differences between the music that exist in various different cultures, and tried to understand the origin of the universality. A paper published in Science in 2019 had revealed that music is produced in all ethnographically distinct cultures, and that similar forms of beats and tunes are used. Neuroscientist have also previously found out that a specific part of the human brain, namely the auditory cortex, is responsible for processing musical information.
Professor Jung’s team used an artificial neural network model to show that cognitive functions for music forms spontaneously as a result of processing auditory information received from nature, without being taught music. The research team utilized AudioSet, a large-scale collection of sound data provided by Google, and taught the artificial neural network to learn the various sounds. Interestingly, the research team discovered that certain neurons within the network model would respond selectively to music. In other words, they observed the spontaneous generation of neurons that reacted minimally to various other sounds like those of animals, nature, or machines, but showed high levels of response to various forms of music including both instrumental and vocal.
The neurons in the artificial neural network model showed similar reactive behaviours to those in the auditory cortex of a real brain. For example, artificial neurons responded less to the sound of music that was cropped into short intervals and were rearranged. This indicates that the spontaneously-generated music-selective neurons encode the temporal structure of music. This property was not limited to a specific genre of music, but emerged across 25 different genres including classic, pop, rock, jazz, and electronic.
< Figure 1. Illustration of the musicality of the brain and artificial neural network (created with DALL·E3 AI based on the paper content) >
Furthermore, suppressing the activity of the music-selective neurons was found to greatly impede the cognitive accuracy for other natural sounds. That is to say, the neural function that processes musical information helps process other sounds, and that ‘musical ability’ may be an instinct formed as a result of an evolutionary adaptation acquired to better process sounds from nature.
Professor Hawoong Jung, who advised the research, said, “The results of our study imply that evolutionary pressure has contributed to forming the universal basis for processing musical information in various cultures.” As for the significance of the research, he explained, “We look forward for this artificially built model with human-like musicality to become an original model for various applications including AI music generation, musical therapy, and for research in musical cognition.” He also commented on its limitations, adding, “This research however does not take into consideration the developmental process that follows the learning of music, and it must be noted that this is a study on the foundation of processing musical information in early development.”
< Figure 2. The artificial neural network that learned to recognize non-musical natural sounds in the cyber space distinguishes between music and non-music. >
This research, conducted by first author Dr. Gwangsu Kim of the KAIST Department of Physics (current affiliation: MIT Department of Brain and Cognitive Sciences) and Dr. Dong-Kyum Kim (current affiliation: IBS) was published in Nature Communications under the title, “Spontaneous emergence of rudimentary music detectors in deep neural networks”.
This research was supported by the National Research Foundation of Korea.
2024.01.23 View 13337
KAIST Research Team Breaks Down Musical Instincts with AI
Music, often referred to as the universal language, is known to be a common component in all cultures. Then, could ‘musical instinct’ be something that is shared to some degree despite the extensive environmental differences amongst cultures?
On January 16, a KAIST research team led by Professor Hawoong Jung from the Department of Physics announced to have identified the principle by which musical instincts emerge from the human brain without special learning using an artificial neural network model.
Previously, many researchers have attempted to identify the similarities and differences between the music that exist in various different cultures, and tried to understand the origin of the universality. A paper published in Science in 2019 had revealed that music is produced in all ethnographically distinct cultures, and that similar forms of beats and tunes are used. Neuroscientist have also previously found out that a specific part of the human brain, namely the auditory cortex, is responsible for processing musical information.
Professor Jung’s team used an artificial neural network model to show that cognitive functions for music forms spontaneously as a result of processing auditory information received from nature, without being taught music. The research team utilized AudioSet, a large-scale collection of sound data provided by Google, and taught the artificial neural network to learn the various sounds. Interestingly, the research team discovered that certain neurons within the network model would respond selectively to music. In other words, they observed the spontaneous generation of neurons that reacted minimally to various other sounds like those of animals, nature, or machines, but showed high levels of response to various forms of music including both instrumental and vocal.
The neurons in the artificial neural network model showed similar reactive behaviours to those in the auditory cortex of a real brain. For example, artificial neurons responded less to the sound of music that was cropped into short intervals and were rearranged. This indicates that the spontaneously-generated music-selective neurons encode the temporal structure of music. This property was not limited to a specific genre of music, but emerged across 25 different genres including classic, pop, rock, jazz, and electronic.
< Figure 1. Illustration of the musicality of the brain and artificial neural network (created with DALL·E3 AI based on the paper content) >
Furthermore, suppressing the activity of the music-selective neurons was found to greatly impede the cognitive accuracy for other natural sounds. That is to say, the neural function that processes musical information helps process other sounds, and that ‘musical ability’ may be an instinct formed as a result of an evolutionary adaptation acquired to better process sounds from nature.
Professor Hawoong Jung, who advised the research, said, “The results of our study imply that evolutionary pressure has contributed to forming the universal basis for processing musical information in various cultures.” As for the significance of the research, he explained, “We look forward for this artificially built model with human-like musicality to become an original model for various applications including AI music generation, musical therapy, and for research in musical cognition.” He also commented on its limitations, adding, “This research however does not take into consideration the developmental process that follows the learning of music, and it must be noted that this is a study on the foundation of processing musical information in early development.”
< Figure 2. The artificial neural network that learned to recognize non-musical natural sounds in the cyber space distinguishes between music and non-music. >
This research, conducted by first author Dr. Gwangsu Kim of the KAIST Department of Physics (current affiliation: MIT Department of Brain and Cognitive Sciences) and Dr. Dong-Kyum Kim (current affiliation: IBS) was published in Nature Communications under the title, “Spontaneous emergence of rudimentary music detectors in deep neural networks”.
This research was supported by the National Research Foundation of Korea.
2024.01.23 View 13337 -
 North Korea and Beyond: AI-Powered Satellite Analysis Reveals the Unseen Economic Landscape of Underdeveloped Nations
- A joint research team in computer science, economics, and geography has developed an artificial intelligence (AI) technology to measure grid-level economic development within six-square-kilometer regions.
- This AI technology is applicable in regions with limited statistical data (e.g., North Korea), supporting international efforts to propose policies for economic growth and poverty reduction in underdeveloped countries.
- The research team plans to make this technology freely available for use to contribute to the United Nations' Sustainable Development Goals (SDGs).
The United Nations reports that more than 700 million people are in extreme poverty, earning less than two dollars a day. However, an accurate assessment of poverty remains a global challenge. For example, 53 countries have not conducted agricultural surveys in the past 15 years, and 17 countries have not published a population census. To fill this data gap, new technologies are being explored to estimate poverty using alternative sources such as street views, aerial photos, and satellite images.
The paper published in Nature Communications demonstrates how artificial intelligence (AI) can help analyze economic conditions from daytime satellite imagery. This new technology can even apply to the least developed countries - such as North Korea - that do not have reliable statistical data for typical machine learning training.
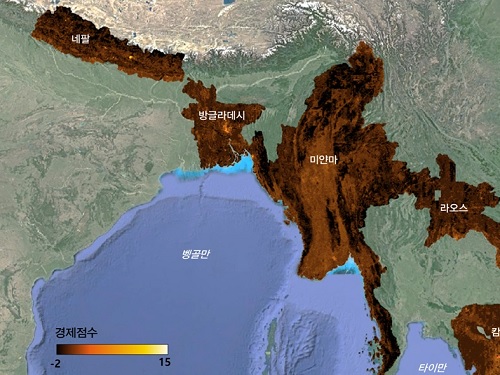
The researchers used Sentinel-2 satellite images from the European Space Agency (ESA) that are publicly available. They split these images into small six-square-kilometer grids. At this zoom level, visual information such as buildings, roads, and greenery can be used to quantify economic indicators. As a result, the team obtained the first ever fine-grained economic map of regions like North Korea. The same algorithm was applied to other underdeveloped countries in Asia: North Korea, Nepal, Laos, Myanmar, Bangladesh, and Cambodia (see Image 1).
The key feature of their research model is the "human-machine collaborative approach," which lets researchers combine human input with AI predictions for areas with scarce data. In this research, ten human experts compared satellite images and judged the economic conditions in the area, with the AI learning from this human data and giving economic scores to each image. The results showed that the Human-AI collaborative approach outperformed machine-only learning algorithms.
< Image 1. Nightlight satellite images of North Korea (Top-left: Background photo provided by NASA's Earth Observatory). South Korea appears brightly lit compared to North Korea, which is mostly dark except for Pyongyang. In contrast, the model developed by the research team uses daytime satellite imagery to predict more detailed economic predictions for North Korea (top-right) and five Asian countries (Bottom: Background photo from Google Earth). >
The research was led by an interdisciplinary team of computer scientists, economists, and a geographer from KAIST & IBS (Donghyun Ahn, Meeyoung Cha, Jihee Kim), Sogang University (Hyunjoo Yang), HKUST (Sangyoon Park), and NUS (Jeasurk Yang). Dr Charles Axelsson, Associate Editor at Nature Communications, handled this paper during the peer review process at the journal.
The research team found that the scores showed a strong correlation with traditional socio-economic metrics such as population density, employment, and number of businesses. This demonstrates the wide applicability and scalability of the approach, particularly in data-scarce countries. Furthermore, the model's strength lies in its ability to detect annual changes in economic conditions at a more detailed geospatial level without using any survey data (see Image 2).
< Image 2. Differences in satellite imagery and economic scores in North Korea between 2016 and 2019. Significant development was found in the Wonsan Kalma area (top), one of the tourist development zones, but no changes were observed in the Wiwon Industrial Development Zone (bottom). (Background photo: Sentinel-2 satellite imagery provided by the European Space Agency (ESA)). >
This model would be especially valuable for rapidly monitoring the progress of Sustainable Development Goals such as reducing poverty and promoting more equitable and sustainable growth on an international scale. The model can also be adapted to measure various social and environmental indicators. For example, it can be trained to identify regions with high vulnerability to climate change and disasters to provide timely guidance on disaster relief efforts.
As an example, the researchers explored how North Korea changed before and after the United Nations sanctions against the country. By applying the model to satellite images of North Korea both in 2016 and in 2019, the researchers discovered three key trends in the country's economic development between 2016 and 2019. First, economic growth in North Korea became more concentrated in Pyongyang and major cities, exacerbating the urban-rural divide. Second, satellite imagery revealed significant changes in areas designated for tourism and economic development, such as new building construction and other meaningful alterations. Third, traditional industrial and export development zones showed relatively minor changes.
Meeyoung Cha, a data scientist in the team explained, "This is an important interdisciplinary effort to address global challenges like poverty. We plan to apply our AI algorithm to other international issues, such as monitoring carbon emissions, disaster damage detection, and the impact of climate change."
An economist on the research team, Jihee Kim, commented that this approach would enable detailed examinations of economic conditions in the developing world at a low cost, reducing data disparities between developed and developing nations. She further emphasized that this is most essential because many public policies require economic measurements to achieve their goals, whether they are for growth, equality, or sustainability.
The research team has made the source code publicly available via GitHub and plans to continue improving the technology, applying it to new satellite images updated annually. The results of this study, with Ph.D. candidate Donghyun Ahn at KAIST and Ph.D. candidate Jeasurk Yang at NUS as joint first authors, were published in Nature Communications under the title "A human-machine collaborative approach measures economic development using satellite imagery."
< Photos of the main authors. 1. Donghyun Ahn, PhD candidate at KAIST School of Computing 2. Jeasurk Yang, PhD candidate at the Department of Geography of National University of Singapore 3. Meeyoung Cha, Professor of KAIST School of Computing and CI at IBS 4. Jihee Kim, Professor of KAIST School of Business and Technology Management 5. Sangyoon Park, Professor of the Division of Social Science at Hong Kong University of Science and Technology 6. Hyunjoo Yang, Professor of the Department of Economics at Sogang University >
2023.12.07 View 15074
North Korea and Beyond: AI-Powered Satellite Analysis Reveals the Unseen Economic Landscape of Underdeveloped Nations
- A joint research team in computer science, economics, and geography has developed an artificial intelligence (AI) technology to measure grid-level economic development within six-square-kilometer regions.
- This AI technology is applicable in regions with limited statistical data (e.g., North Korea), supporting international efforts to propose policies for economic growth and poverty reduction in underdeveloped countries.
- The research team plans to make this technology freely available for use to contribute to the United Nations' Sustainable Development Goals (SDGs).
The United Nations reports that more than 700 million people are in extreme poverty, earning less than two dollars a day. However, an accurate assessment of poverty remains a global challenge. For example, 53 countries have not conducted agricultural surveys in the past 15 years, and 17 countries have not published a population census. To fill this data gap, new technologies are being explored to estimate poverty using alternative sources such as street views, aerial photos, and satellite images.
The paper published in Nature Communications demonstrates how artificial intelligence (AI) can help analyze economic conditions from daytime satellite imagery. This new technology can even apply to the least developed countries - such as North Korea - that do not have reliable statistical data for typical machine learning training.
The researchers used Sentinel-2 satellite images from the European Space Agency (ESA) that are publicly available. They split these images into small six-square-kilometer grids. At this zoom level, visual information such as buildings, roads, and greenery can be used to quantify economic indicators. As a result, the team obtained the first ever fine-grained economic map of regions like North Korea. The same algorithm was applied to other underdeveloped countries in Asia: North Korea, Nepal, Laos, Myanmar, Bangladesh, and Cambodia (see Image 1).
The key feature of their research model is the "human-machine collaborative approach," which lets researchers combine human input with AI predictions for areas with scarce data. In this research, ten human experts compared satellite images and judged the economic conditions in the area, with the AI learning from this human data and giving economic scores to each image. The results showed that the Human-AI collaborative approach outperformed machine-only learning algorithms.
< Image 1. Nightlight satellite images of North Korea (Top-left: Background photo provided by NASA's Earth Observatory). South Korea appears brightly lit compared to North Korea, which is mostly dark except for Pyongyang. In contrast, the model developed by the research team uses daytime satellite imagery to predict more detailed economic predictions for North Korea (top-right) and five Asian countries (Bottom: Background photo from Google Earth). >
The research was led by an interdisciplinary team of computer scientists, economists, and a geographer from KAIST & IBS (Donghyun Ahn, Meeyoung Cha, Jihee Kim), Sogang University (Hyunjoo Yang), HKUST (Sangyoon Park), and NUS (Jeasurk Yang). Dr Charles Axelsson, Associate Editor at Nature Communications, handled this paper during the peer review process at the journal.
The research team found that the scores showed a strong correlation with traditional socio-economic metrics such as population density, employment, and number of businesses. This demonstrates the wide applicability and scalability of the approach, particularly in data-scarce countries. Furthermore, the model's strength lies in its ability to detect annual changes in economic conditions at a more detailed geospatial level without using any survey data (see Image 2).
< Image 2. Differences in satellite imagery and economic scores in North Korea between 2016 and 2019. Significant development was found in the Wonsan Kalma area (top), one of the tourist development zones, but no changes were observed in the Wiwon Industrial Development Zone (bottom). (Background photo: Sentinel-2 satellite imagery provided by the European Space Agency (ESA)). >
This model would be especially valuable for rapidly monitoring the progress of Sustainable Development Goals such as reducing poverty and promoting more equitable and sustainable growth on an international scale. The model can also be adapted to measure various social and environmental indicators. For example, it can be trained to identify regions with high vulnerability to climate change and disasters to provide timely guidance on disaster relief efforts.
As an example, the researchers explored how North Korea changed before and after the United Nations sanctions against the country. By applying the model to satellite images of North Korea both in 2016 and in 2019, the researchers discovered three key trends in the country's economic development between 2016 and 2019. First, economic growth in North Korea became more concentrated in Pyongyang and major cities, exacerbating the urban-rural divide. Second, satellite imagery revealed significant changes in areas designated for tourism and economic development, such as new building construction and other meaningful alterations. Third, traditional industrial and export development zones showed relatively minor changes.
Meeyoung Cha, a data scientist in the team explained, "This is an important interdisciplinary effort to address global challenges like poverty. We plan to apply our AI algorithm to other international issues, such as monitoring carbon emissions, disaster damage detection, and the impact of climate change."
An economist on the research team, Jihee Kim, commented that this approach would enable detailed examinations of economic conditions in the developing world at a low cost, reducing data disparities between developed and developing nations. She further emphasized that this is most essential because many public policies require economic measurements to achieve their goals, whether they are for growth, equality, or sustainability.
The research team has made the source code publicly available via GitHub and plans to continue improving the technology, applying it to new satellite images updated annually. The results of this study, with Ph.D. candidate Donghyun Ahn at KAIST and Ph.D. candidate Jeasurk Yang at NUS as joint first authors, were published in Nature Communications under the title "A human-machine collaborative approach measures economic development using satellite imagery."
< Photos of the main authors. 1. Donghyun Ahn, PhD candidate at KAIST School of Computing 2. Jeasurk Yang, PhD candidate at the Department of Geography of National University of Singapore 3. Meeyoung Cha, Professor of KAIST School of Computing and CI at IBS 4. Jihee Kim, Professor of KAIST School of Business and Technology Management 5. Sangyoon Park, Professor of the Division of Social Science at Hong Kong University of Science and Technology 6. Hyunjoo Yang, Professor of the Department of Economics at Sogang University >
2023.12.07 View 15074 -
 KAIST builds a high-resolution 3D holographic sensor using a single mask
Holographic cameras can provide more realistic images than ordinary cameras thanks to their ability to acquire 3D information about objects. However, existing holographic cameras use interferometers that measure the wavelength and refraction of light through the interference of light waves, which makes them complex and sensitive to their surrounding environment.
On August 23, a KAIST research team led by Professor YongKeun Park from the Department of Physics announced a new leap forward in 3D holographic imaging sensor technology.
The team proposed an innovative holographic camera technology that does not use complex interferometry. Instead, it uses a mask to precisely measure the phase information of light and reconstruct the 3D information of an object with higher accuracy.
< Figure 1. Structure and principle of the proposed holographic camera. The amplitude and phase information of light scattered from a holographic camera can be measured. >
The team used a mask that fulfills certain mathematical conditions and incorporated it into an ordinary camera, and the light scattered from a laser is measured through the mask and analyzed using a computer. This does not require a complex interferometer and allows the phase information of light to be collected through a simplified optical system. With this technique, the mask that is placed between the two lenses and behind an object plays an important role. The mask selectively filters specific parts of light,, and the intensity of the light passing through the lens can be measured using an ordinary commercial camera. This technique combines the image data received from the camera with the unique pattern received from the mask and reconstructs an object’s precise 3D information using an algorithm.
This method allows a high-resolution 3D image of an object to be captured in any position. In practical situations, one can construct a laser-based holographic 3D image sensor by adding a mask with a simple design to a general image sensor. This makes the design and construction of the optical system much easier. In particular, this novel technology can capture high-resolution holographic images of objects moving at high speeds, which widens its potential field of application.
< Figure 2. A moving doll captured by a conventional camera and the proposed holographic camera. When taking a picture without focusing on the object, only a blurred image of the doll can be obtained from a general camera, but the proposed holographic camera can restore the blurred image of the doll into a clear image. >
The results of this study, conducted by Dr. Jeonghun Oh from the KAIST Department of Physics as the first author, were published in Nature Communications on August 12 under the title, "Non-interferometric stand-alone single-shot holographic camera using reciprocal diffractive imaging".
Dr. Oh said, “The holographic camera module we are suggesting can be built by adding a filter to an ordinary camera, which would allow even non-experts to handle it easily in everyday life if it were to be commercialized.” He added, “In particular, it is a promising candidate with the potential to replace existing remote sensing technologies.”
This research was supported by the National Research Foundation’s Leader Research Project, the Korean Ministry of Science and ICT’s Core Hologram Technology Support Project, and the Nano and Material Technology Development Project.
2023.09.05 View 15256
KAIST builds a high-resolution 3D holographic sensor using a single mask
Holographic cameras can provide more realistic images than ordinary cameras thanks to their ability to acquire 3D information about objects. However, existing holographic cameras use interferometers that measure the wavelength and refraction of light through the interference of light waves, which makes them complex and sensitive to their surrounding environment.
On August 23, a KAIST research team led by Professor YongKeun Park from the Department of Physics announced a new leap forward in 3D holographic imaging sensor technology.
The team proposed an innovative holographic camera technology that does not use complex interferometry. Instead, it uses a mask to precisely measure the phase information of light and reconstruct the 3D information of an object with higher accuracy.
< Figure 1. Structure and principle of the proposed holographic camera. The amplitude and phase information of light scattered from a holographic camera can be measured. >
The team used a mask that fulfills certain mathematical conditions and incorporated it into an ordinary camera, and the light scattered from a laser is measured through the mask and analyzed using a computer. This does not require a complex interferometer and allows the phase information of light to be collected through a simplified optical system. With this technique, the mask that is placed between the two lenses and behind an object plays an important role. The mask selectively filters specific parts of light,, and the intensity of the light passing through the lens can be measured using an ordinary commercial camera. This technique combines the image data received from the camera with the unique pattern received from the mask and reconstructs an object’s precise 3D information using an algorithm.
This method allows a high-resolution 3D image of an object to be captured in any position. In practical situations, one can construct a laser-based holographic 3D image sensor by adding a mask with a simple design to a general image sensor. This makes the design and construction of the optical system much easier. In particular, this novel technology can capture high-resolution holographic images of objects moving at high speeds, which widens its potential field of application.
< Figure 2. A moving doll captured by a conventional camera and the proposed holographic camera. When taking a picture without focusing on the object, only a blurred image of the doll can be obtained from a general camera, but the proposed holographic camera can restore the blurred image of the doll into a clear image. >
The results of this study, conducted by Dr. Jeonghun Oh from the KAIST Department of Physics as the first author, were published in Nature Communications on August 12 under the title, "Non-interferometric stand-alone single-shot holographic camera using reciprocal diffractive imaging".
Dr. Oh said, “The holographic camera module we are suggesting can be built by adding a filter to an ordinary camera, which would allow even non-experts to handle it easily in everyday life if it were to be commercialized.” He added, “In particular, it is a promising candidate with the potential to replace existing remote sensing technologies.”
This research was supported by the National Research Foundation’s Leader Research Project, the Korean Ministry of Science and ICT’s Core Hologram Technology Support Project, and the Nano and Material Technology Development Project.
2023.09.05 View 15256 -
 Professor Shinhyun Choi’s team, selected for Nature Communications Editors’ highlight
[ From left, Ph.D. candidates See-On Park and Hakcheon Jeong, along with Master's student Jong-Yong Park and Professor Shinhyun Choi ]
See-On Park, Hakcheon Jeong, Jong-Yong Park - a team of researchers under the leadership of Professor Shinhyun Choi of the School of Electrical Engineering, developed a highly reliable variable resistor (memristor) array that simulates the behavior of neurons using a metal oxide layer with an oxygen concentration gradient, and published their work in Nature Communications. The study was selected as the Nature Communications' Editor's highlight, and as the featured article posted on the main page of the journal's website.
Link : https://www.nature.com/ncomms/
[ Figure 1. The featured image on the main page of the Nature Communications' website introducing the research by Professor Choi's team on the memristor for artificial neurons ]
Thesis title: Experimental demonstration of highly reliable dynamic memristor for artificial neuron and neuromorphic computing.
( https://doi.org/10.1038/s41467-022-30539-6 )
At KAIST, their research was introduced on the 2022 Fall issue of Breakthroughs, the biannual newsletter published by KAIST College of Engineering.
This research was conducted with the support from the Samsung Research Funding & Incubation Center of Samsung Electronics.
2022.11.01 View 15949
Professor Shinhyun Choi’s team, selected for Nature Communications Editors’ highlight
[ From left, Ph.D. candidates See-On Park and Hakcheon Jeong, along with Master's student Jong-Yong Park and Professor Shinhyun Choi ]
See-On Park, Hakcheon Jeong, Jong-Yong Park - a team of researchers under the leadership of Professor Shinhyun Choi of the School of Electrical Engineering, developed a highly reliable variable resistor (memristor) array that simulates the behavior of neurons using a metal oxide layer with an oxygen concentration gradient, and published their work in Nature Communications. The study was selected as the Nature Communications' Editor's highlight, and as the featured article posted on the main page of the journal's website.
Link : https://www.nature.com/ncomms/
[ Figure 1. The featured image on the main page of the Nature Communications' website introducing the research by Professor Choi's team on the memristor for artificial neurons ]
Thesis title: Experimental demonstration of highly reliable dynamic memristor for artificial neuron and neuromorphic computing.
( https://doi.org/10.1038/s41467-022-30539-6 )
At KAIST, their research was introduced on the 2022 Fall issue of Breakthroughs, the biannual newsletter published by KAIST College of Engineering.
This research was conducted with the support from the Samsung Research Funding & Incubation Center of Samsung Electronics.
2022.11.01 View 15949 -
 Acoustic Graphene Plasmons Study Paves Way for Optoelectronic Applications
- The first images of mid-infrared optical waves compressed 1,000 times captured using a highly sensitive scattering-type scanning near-field optical microscope. -
KAIST researchers and their collaborators at home and abroad have successfully demonstrated a new methodology for direct near-field optical imaging of acoustic graphene plasmon fields. This strategy will provide a breakthrough for the practical applications of acoustic graphene plasmon platforms in next-generation, high-performance, graphene-based optoelectronic devices with enhanced light-matter interactions and lower propagation loss.
It was recently demonstrated that ‘graphene plasmons’ – collective oscillations of free electrons in graphene coupled to electromagnetic waves of light – can be used to trap and compress optical waves inside a very thin dielectric layer separating graphene from a metallic sheet. In such a configuration, graphene’s conduction electrons are “reflected” in the metal, so when the light waves “push” the electrons in graphene, their image charges in metal also start to oscillate. This new type of collective electronic oscillation mode is called ‘acoustic graphene plasmon (AGP)’.
The existence of AGP could previously be observed only via indirect methods such as far-field infrared spectroscopy and photocurrent mapping. This indirect observation was the price that researchers had to pay for the strong compression of optical waves inside nanometer-thin structures. It was believed that the intensity of electromagnetic fields outside the device was insufficient for direct near-field optical imaging of AGP.
Challenged by these limitations, three research groups combined their efforts to bring together a unique experimental technique using advanced nanofabrication methods. Their findings were published in Nature Communications on February 19.
A KAIST research team led by Professor Min Seok Jang from the School of Electrical Engineering used a highly sensitive scattering-type scanning near-field optical microscope (s-SNOM) to directly measure the optical fields of the AGP waves propagating in a nanometer-thin waveguide, visualizing thousand-fold compression of mid-infrared light for the first time.
Professor Jang and a post-doc researcher in his group, Sergey G. Menabde, successfully obtained direct images of AGP waves by taking advantage of their rapidly decaying yet always present electric field above graphene. They showed that AGPs are detectable even when most of their energy is flowing inside the dielectric below the graphene.
This became possible due to the ultra-smooth surfaces inside the nano-waveguides where plasmonic waves can propagate at longer distances. The AGP mode probed by the researchers was up to 2.3 times more confined and exhibited a 1.4 times higher figure of merit in terms of the normalized propagation length compared to the graphene surface plasmon under similar conditions.
These ultra-smooth nanostructures of the waveguides used in the experiment were created using a template-stripping method by Professor Sang-Hyun Oh and a post-doc researcher, In-Ho Lee, from the Department of Electrical and Computer Engineering at the University of Minnesota.
Professor Young Hee Lee and his researchers at the Center for Integrated Nanostructure Physics (CINAP) of the Institute of Basic Science (IBS) at Sungkyunkwan University synthesized the graphene with a monocrystalline structure, and this high-quality, large-area graphene enabled low-loss plasmonic propagation.
The chemical and physical properties of many important organic molecules can be detected and evaluated by their absorption signatures in the mid-infrared spectrum. However, conventional detection methods require a large number of molecules for successful detection, whereas the ultra-compressed AGP fields can provide strong light-matter interactions at the microscopic level, thus significantly improving the detection sensitivity down to a single molecule.
Furthermore, the study conducted by Professor Jang and the team demonstrated that the mid-infrared AGPs are inherently less sensitive to losses in graphene due to their fields being mostly confined within the dielectric. The research team’s reported results suggest that AGPs could become a promising platform for electrically tunable graphene-based optoelectronic devices that typically suffer from higher absorption rates in graphene such as metasurfaces, optical switches, photovoltaics, and other optoelectronic applications operating at infrared frequencies.
Professor Jang said, “Our research revealed that the ultra-compressed electromagnetic fields of acoustic graphene plasmons can be directly accessed through near-field optical microscopy methods. I hope this realization will motivate other researchers to apply AGPs to various problems where strong light-matter interactions and lower propagation loss are needed.”
This research was primarily funded by the Samsung Research Funding & Incubation Center of Samsung Electronics. The National Research Foundation of Korea (NRF), the U.S. National Science Foundation (NSF), Samsung Global Research Outreach (GRO) Program, and Institute for Basic Science of Korea (IBS) also supported the work.
Publication:
Menabde, S. G., et al. (2021) Real-space imaging of acoustic plasmons in large-area graphene grown by chemical vapor deposition. Nature Communications 12, Article No. 938. Available online at https://doi.org/10.1038/s41467-021-21193-5
Profile:
Min Seok Jang, MS, PhD
Associate Professorjang.minseok@kaist.ac.krhttp://jlab.kaist.ac.kr/
Min Seok Jang Research GroupSchool of Electrical Engineering
http://kaist.ac.kr/en/Korea Advanced Institute of Science and Technology (KAIST)Daejeon, Republic of Korea
(END)
2021.03.16 View 20743
Acoustic Graphene Plasmons Study Paves Way for Optoelectronic Applications
- The first images of mid-infrared optical waves compressed 1,000 times captured using a highly sensitive scattering-type scanning near-field optical microscope. -
KAIST researchers and their collaborators at home and abroad have successfully demonstrated a new methodology for direct near-field optical imaging of acoustic graphene plasmon fields. This strategy will provide a breakthrough for the practical applications of acoustic graphene plasmon platforms in next-generation, high-performance, graphene-based optoelectronic devices with enhanced light-matter interactions and lower propagation loss.
It was recently demonstrated that ‘graphene plasmons’ – collective oscillations of free electrons in graphene coupled to electromagnetic waves of light – can be used to trap and compress optical waves inside a very thin dielectric layer separating graphene from a metallic sheet. In such a configuration, graphene’s conduction electrons are “reflected” in the metal, so when the light waves “push” the electrons in graphene, their image charges in metal also start to oscillate. This new type of collective electronic oscillation mode is called ‘acoustic graphene plasmon (AGP)’.
The existence of AGP could previously be observed only via indirect methods such as far-field infrared spectroscopy and photocurrent mapping. This indirect observation was the price that researchers had to pay for the strong compression of optical waves inside nanometer-thin structures. It was believed that the intensity of electromagnetic fields outside the device was insufficient for direct near-field optical imaging of AGP.
Challenged by these limitations, three research groups combined their efforts to bring together a unique experimental technique using advanced nanofabrication methods. Their findings were published in Nature Communications on February 19.
A KAIST research team led by Professor Min Seok Jang from the School of Electrical Engineering used a highly sensitive scattering-type scanning near-field optical microscope (s-SNOM) to directly measure the optical fields of the AGP waves propagating in a nanometer-thin waveguide, visualizing thousand-fold compression of mid-infrared light for the first time.
Professor Jang and a post-doc researcher in his group, Sergey G. Menabde, successfully obtained direct images of AGP waves by taking advantage of their rapidly decaying yet always present electric field above graphene. They showed that AGPs are detectable even when most of their energy is flowing inside the dielectric below the graphene.
This became possible due to the ultra-smooth surfaces inside the nano-waveguides where plasmonic waves can propagate at longer distances. The AGP mode probed by the researchers was up to 2.3 times more confined and exhibited a 1.4 times higher figure of merit in terms of the normalized propagation length compared to the graphene surface plasmon under similar conditions.
These ultra-smooth nanostructures of the waveguides used in the experiment were created using a template-stripping method by Professor Sang-Hyun Oh and a post-doc researcher, In-Ho Lee, from the Department of Electrical and Computer Engineering at the University of Minnesota.
Professor Young Hee Lee and his researchers at the Center for Integrated Nanostructure Physics (CINAP) of the Institute of Basic Science (IBS) at Sungkyunkwan University synthesized the graphene with a monocrystalline structure, and this high-quality, large-area graphene enabled low-loss plasmonic propagation.
The chemical and physical properties of many important organic molecules can be detected and evaluated by their absorption signatures in the mid-infrared spectrum. However, conventional detection methods require a large number of molecules for successful detection, whereas the ultra-compressed AGP fields can provide strong light-matter interactions at the microscopic level, thus significantly improving the detection sensitivity down to a single molecule.
Furthermore, the study conducted by Professor Jang and the team demonstrated that the mid-infrared AGPs are inherently less sensitive to losses in graphene due to their fields being mostly confined within the dielectric. The research team’s reported results suggest that AGPs could become a promising platform for electrically tunable graphene-based optoelectronic devices that typically suffer from higher absorption rates in graphene such as metasurfaces, optical switches, photovoltaics, and other optoelectronic applications operating at infrared frequencies.
Professor Jang said, “Our research revealed that the ultra-compressed electromagnetic fields of acoustic graphene plasmons can be directly accessed through near-field optical microscopy methods. I hope this realization will motivate other researchers to apply AGPs to various problems where strong light-matter interactions and lower propagation loss are needed.”
This research was primarily funded by the Samsung Research Funding & Incubation Center of Samsung Electronics. The National Research Foundation of Korea (NRF), the U.S. National Science Foundation (NSF), Samsung Global Research Outreach (GRO) Program, and Institute for Basic Science of Korea (IBS) also supported the work.
Publication:
Menabde, S. G., et al. (2021) Real-space imaging of acoustic plasmons in large-area graphene grown by chemical vapor deposition. Nature Communications 12, Article No. 938. Available online at https://doi.org/10.1038/s41467-021-21193-5
Profile:
Min Seok Jang, MS, PhD
Associate Professorjang.minseok@kaist.ac.krhttp://jlab.kaist.ac.kr/
Min Seok Jang Research GroupSchool of Electrical Engineering
http://kaist.ac.kr/en/Korea Advanced Institute of Science and Technology (KAIST)Daejeon, Republic of Korea
(END)
2021.03.16 View 20743 -
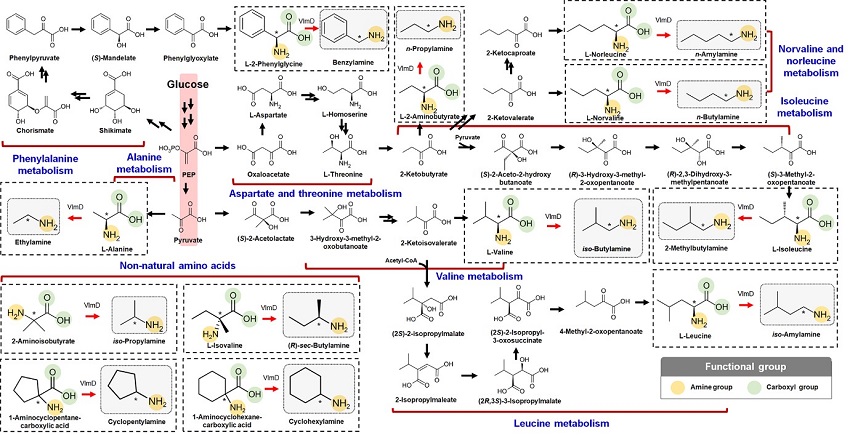 Expanding the Biosynthetic Pathway via Retrobiosynthesis
- Researchers reports a new strategy for the microbial production of multiple short-chain primary amines via retrobiosynthesis. -
KAIST metabolic engineers presented the bio-based production of multiple short-chain primary amines that have a wide range of applications in chemical industries for the first time. The research team led by Distinguished Professor Sang Yup Lee from the Department of Chemical and Biomolecular Engineering designed the novel biosynthetic pathways for short-chain primary amines by combining retrobiosynthesis and a precursor selection step.
The research team verified the newly designed pathways by confirming the in vivo production of 10 short-chain primary amines by supplying the precursors. Furthermore, the platform Escherichia coli strains were metabolically engineered to produce three proof-of-concept short-chain primary amines from glucose, demonstrating the possibility of the bio-based production of diverse short-chain primary amines from renewable resources. The research team said this study expands the strategy of systematically designing biosynthetic pathways for the production of a group of related chemicals as demonstrated by multiple short-chain primary amines as examples.
Currently, most of the industrial chemicals used in our daily lives are produced with petroleum-based products. However, there are several serious issues with the petroleum industry such as the depletion of fossil fuel reserves and environmental problems including global warming. To solve these problems, the sustainable production of industrial chemicals and materials is being explored with microorganisms as cell factories and renewable non-food biomass as raw materials for alternative to petroleum-based products. The engineering of these microorganisms has increasingly become more efficient and effective with the help of systems metabolic engineering – a practice of engineering the metabolism of a living organism toward the production of a desired metabolite. In this regard, the number of chemicals produced using biomass as a raw material has substantially increased.
Although the scope of chemicals that are producible using microorganisms continues to expand through advances in systems metabolic engineering, the biological production of short-chain primary amines has not yet been reported despite their industrial importance. Short-chain primary amines are the chemicals that have an alkyl or aryl group in the place of a hydrogen atom in ammonia with carbon chain lengths ranging from C1 to C7. Short-chain primary amines have a wide range of applications in chemical industries, for example, as a precursor for pharmaceuticals (e.g., antidiabetic and antihypertensive drugs), agrochemicals (e.g., herbicides, fungicides and insecticides), solvents, and vulcanization accelerators for rubber and plasticizers. The market size of short-chain primary amines was estimated to be more than 4 billion US dollars in 2014.
The main reason why the bio-based production of short-chain primary amines was not yet possible was due to their unknown biosynthetic pathways. Therefore, the team designed synthetic biosynthetic pathways for short-chain primary amines by combining retrobiosynthesis and a precursor selection step. The retrobiosynthesis allowed the systematic design of a biosynthetic pathway for short-chain primary amines by using a set of biochemical reaction rules that describe chemical transformation patterns between a substrate and product molecules at an atomic level.
These multiple precursors predicted for the possible biosynthesis of each short-chain primary amine were sequentially narrowed down by using the precursor selection step for efficient metabolic engineering experiments.
“Our research demonstrates the possibility of the renewable production of short-chain primary amines for the first time. We are planning to increase production efficiencies of short-chain primary amines. We believe that our study will play an important role in the development of sustainable and eco-friendly bio-based industries and the reorganization of the chemical industry, which is mandatory for solving the environmental problems threating the survival of mankind,” said Professor Lee.
This paper titled “Microbial production of multiple short-chain primary amines via retrobiosynthesis” was published in Nature Communications. This work was supported by the Technology Development Program to Solve Climate Changes on Systems Metabolic Engineering for Biorefineries from the Ministry of Science and ICT through the National Research Foundation (NRF) of Korea.
-Publication
Dong In Kim, Tong Un Chae, Hyun Uk Kim, Woo Dae Jang, and Sang Yup Lee. Microbial production of multiple short-chain primary amines via retrobiosynthesis. Nature Communications ( https://www.nature.com/articles/s41467-020-20423-6)
-Profile
Distinguished Professor Sang Yup Lee
leesy@kaist.ac.kr
Metabolic &Biomolecular Engineering National Research Laboratory
http://mbel.kaist.ac.kr
Department of Chemical and Biomolecular Engineering
KAIST
2021.01.14 View 17946
Expanding the Biosynthetic Pathway via Retrobiosynthesis
- Researchers reports a new strategy for the microbial production of multiple short-chain primary amines via retrobiosynthesis. -
KAIST metabolic engineers presented the bio-based production of multiple short-chain primary amines that have a wide range of applications in chemical industries for the first time. The research team led by Distinguished Professor Sang Yup Lee from the Department of Chemical and Biomolecular Engineering designed the novel biosynthetic pathways for short-chain primary amines by combining retrobiosynthesis and a precursor selection step.
The research team verified the newly designed pathways by confirming the in vivo production of 10 short-chain primary amines by supplying the precursors. Furthermore, the platform Escherichia coli strains were metabolically engineered to produce three proof-of-concept short-chain primary amines from glucose, demonstrating the possibility of the bio-based production of diverse short-chain primary amines from renewable resources. The research team said this study expands the strategy of systematically designing biosynthetic pathways for the production of a group of related chemicals as demonstrated by multiple short-chain primary amines as examples.
Currently, most of the industrial chemicals used in our daily lives are produced with petroleum-based products. However, there are several serious issues with the petroleum industry such as the depletion of fossil fuel reserves and environmental problems including global warming. To solve these problems, the sustainable production of industrial chemicals and materials is being explored with microorganisms as cell factories and renewable non-food biomass as raw materials for alternative to petroleum-based products. The engineering of these microorganisms has increasingly become more efficient and effective with the help of systems metabolic engineering – a practice of engineering the metabolism of a living organism toward the production of a desired metabolite. In this regard, the number of chemicals produced using biomass as a raw material has substantially increased.
Although the scope of chemicals that are producible using microorganisms continues to expand through advances in systems metabolic engineering, the biological production of short-chain primary amines has not yet been reported despite their industrial importance. Short-chain primary amines are the chemicals that have an alkyl or aryl group in the place of a hydrogen atom in ammonia with carbon chain lengths ranging from C1 to C7. Short-chain primary amines have a wide range of applications in chemical industries, for example, as a precursor for pharmaceuticals (e.g., antidiabetic and antihypertensive drugs), agrochemicals (e.g., herbicides, fungicides and insecticides), solvents, and vulcanization accelerators for rubber and plasticizers. The market size of short-chain primary amines was estimated to be more than 4 billion US dollars in 2014.
The main reason why the bio-based production of short-chain primary amines was not yet possible was due to their unknown biosynthetic pathways. Therefore, the team designed synthetic biosynthetic pathways for short-chain primary amines by combining retrobiosynthesis and a precursor selection step. The retrobiosynthesis allowed the systematic design of a biosynthetic pathway for short-chain primary amines by using a set of biochemical reaction rules that describe chemical transformation patterns between a substrate and product molecules at an atomic level.
These multiple precursors predicted for the possible biosynthesis of each short-chain primary amine were sequentially narrowed down by using the precursor selection step for efficient metabolic engineering experiments.
“Our research demonstrates the possibility of the renewable production of short-chain primary amines for the first time. We are planning to increase production efficiencies of short-chain primary amines. We believe that our study will play an important role in the development of sustainable and eco-friendly bio-based industries and the reorganization of the chemical industry, which is mandatory for solving the environmental problems threating the survival of mankind,” said Professor Lee.
This paper titled “Microbial production of multiple short-chain primary amines via retrobiosynthesis” was published in Nature Communications. This work was supported by the Technology Development Program to Solve Climate Changes on Systems Metabolic Engineering for Biorefineries from the Ministry of Science and ICT through the National Research Foundation (NRF) of Korea.
-Publication
Dong In Kim, Tong Un Chae, Hyun Uk Kim, Woo Dae Jang, and Sang Yup Lee. Microbial production of multiple short-chain primary amines via retrobiosynthesis. Nature Communications ( https://www.nature.com/articles/s41467-020-20423-6)
-Profile
Distinguished Professor Sang Yup Lee
leesy@kaist.ac.kr
Metabolic &Biomolecular Engineering National Research Laboratory
http://mbel.kaist.ac.kr
Department of Chemical and Biomolecular Engineering
KAIST
2021.01.14 View 17946 -
 A Deep-Learned E-Skin Decodes Complex Human Motion
A deep-learning powered single-strained electronic skin sensor can capture human motion from a distance. The single strain sensor placed on the wrist decodes complex five-finger motions in real time with a virtual 3D hand that mirrors the original motions. The deep neural network boosted by rapid situation learning (RSL) ensures stable operation regardless of its position on the surface of the skin.
Conventional approaches require many sensor networks that cover the entire curvilinear surfaces of the target area. Unlike conventional wafer-based fabrication, this laser fabrication provides a new sensing paradigm for motion tracking.
The research team, led by Professor Sungho Jo from the School of Computing, collaborated with Professor Seunghwan Ko from Seoul National University to design this new measuring system that extracts signals corresponding to multiple finger motions by generating cracks in metal nanoparticle films using laser technology. The sensor patch was then attached to a user’s wrist to detect the movement of the fingers.
The concept of this research started from the idea that pinpointing a single area would be more efficient for identifying movements than affixing sensors to every joint and muscle. To make this targeting strategy work, it needs to accurately capture the signals from different areas at the point where they all converge, and then decoupling the information entangled in the converged signals. To maximize users’ usability and mobility, the research team used a single-channeled sensor to generate the signals corresponding to complex hand motions.
The rapid situation learning (RSL) system collects data from arbitrary parts on the wrist and automatically trains the model in a real-time demonstration with a virtual 3D hand that mirrors the original motions. To enhance the sensitivity of the sensor, researchers used laser-induced nanoscale cracking.
This sensory system can track the motion of the entire body with a small sensory network and facilitate the indirect remote measurement of human motions, which is applicable for wearable VR/AR systems.
The research team said they focused on two tasks while developing the sensor. First, they analyzed the sensor signal patterns into a latent space encapsulating temporal sensor behavior and then they mapped the latent vectors to finger motion metric spaces.
Professor Jo said, “Our system is expandable to other body parts. We already confirmed that the sensor is also capable of extracting gait motions from a pelvis. This technology is expected to provide a turning point in health-monitoring, motion tracking, and soft robotics.”
This study was featured in Nature Communications.
Publication:
Kim, K. K., et al. (2020) A deep-learned skin sensor decoding the epicentral human motions. Nature Communications. 11. 2149. https://doi.org/10.1038/s41467-020-16040-y29
Link to download the full-text paper:
https://www.nature.com/articles/s41467-020-16040-y.pdf
Profile: Professor Sungho Jo
shjo@kaist.ac.kr
http://nmail.kaist.ac.kr
Neuro-Machine Augmented Intelligence Lab
School of Computing
College of Engineering
KAIST
2020.06.10 View 18209
A Deep-Learned E-Skin Decodes Complex Human Motion
A deep-learning powered single-strained electronic skin sensor can capture human motion from a distance. The single strain sensor placed on the wrist decodes complex five-finger motions in real time with a virtual 3D hand that mirrors the original motions. The deep neural network boosted by rapid situation learning (RSL) ensures stable operation regardless of its position on the surface of the skin.
Conventional approaches require many sensor networks that cover the entire curvilinear surfaces of the target area. Unlike conventional wafer-based fabrication, this laser fabrication provides a new sensing paradigm for motion tracking.
The research team, led by Professor Sungho Jo from the School of Computing, collaborated with Professor Seunghwan Ko from Seoul National University to design this new measuring system that extracts signals corresponding to multiple finger motions by generating cracks in metal nanoparticle films using laser technology. The sensor patch was then attached to a user’s wrist to detect the movement of the fingers.
The concept of this research started from the idea that pinpointing a single area would be more efficient for identifying movements than affixing sensors to every joint and muscle. To make this targeting strategy work, it needs to accurately capture the signals from different areas at the point where they all converge, and then decoupling the information entangled in the converged signals. To maximize users’ usability and mobility, the research team used a single-channeled sensor to generate the signals corresponding to complex hand motions.
The rapid situation learning (RSL) system collects data from arbitrary parts on the wrist and automatically trains the model in a real-time demonstration with a virtual 3D hand that mirrors the original motions. To enhance the sensitivity of the sensor, researchers used laser-induced nanoscale cracking.
This sensory system can track the motion of the entire body with a small sensory network and facilitate the indirect remote measurement of human motions, which is applicable for wearable VR/AR systems.
The research team said they focused on two tasks while developing the sensor. First, they analyzed the sensor signal patterns into a latent space encapsulating temporal sensor behavior and then they mapped the latent vectors to finger motion metric spaces.
Professor Jo said, “Our system is expandable to other body parts. We already confirmed that the sensor is also capable of extracting gait motions from a pelvis. This technology is expected to provide a turning point in health-monitoring, motion tracking, and soft robotics.”
This study was featured in Nature Communications.
Publication:
Kim, K. K., et al. (2020) A deep-learned skin sensor decoding the epicentral human motions. Nature Communications. 11. 2149. https://doi.org/10.1038/s41467-020-16040-y29
Link to download the full-text paper:
https://www.nature.com/articles/s41467-020-16040-y.pdf
Profile: Professor Sungho Jo
shjo@kaist.ac.kr
http://nmail.kaist.ac.kr
Neuro-Machine Augmented Intelligence Lab
School of Computing
College of Engineering
KAIST
2020.06.10 View 18209