%45%4d%53
-
 Semiconductor Leadership Spotlighted in Nature Sister Journal
<(From Left) Prof. Shinhyun Choi, Prof. Young Gyu Yoon, Prof.Seunghyub Yoo from the School of Electrical Engineering, Prof. Kyung Min Kim from Materials Science and Engineering>
KAIST (President Kwang Hyung Lee) announced on the 5th of September that its semiconductor research and education achievements were highlighted on August 18 in Nature Reviews Electrical Engineering, a sister journal of the world-renowned scientific journal Nature.
Title: Semiconductor-related research and education at KAIST DOI: 10.1038/s44287-025-00204-3
This special "Focus" article provides a detailed look at KAIST's leadership in next-generation semiconductor research, talent development, and global industry-academia collaboration, presenting a future blueprint for Korea's semiconductor industry. Editor Silvia Conti personally conducted the interviews, with KAIST professors including Kyung Min Kim from the Department of Materials Science and Engineering, and Young Gyu Yoon, Shinhyun Choi, Sung-Yool Choi, and Seunghyub Yoo from the School of Electrical Engineering, participating.
KAIST operates educational programs such as the School of Electrical Engineering, the Department of Semiconductor Systems Engineering, and the Graduate School of Semiconductor Engineering. It is leading next-generation semiconductor research in areas like neuromorphic computing, in-memory computing, and 2D new material-based devices. Building on this foundation, researchers are developing new architectures and devices that transcend the limitations of existing silicon, driving innovation in various application fields such as artificial intelligence, robotics, and medicine.
Notably, research on implementing biological functions like synapses and neurons into hardware platforms using new types of memory such as RRAM and PRAM is gaining international attention. This work opens up possibilities for applications in robots, edge computing, and on-sensor AI systems.
Furthermore, KAIST has operated EPSS (Samsung Advanced Human Resources Training Program) and KEPSI (SK Hynix Semiconductor Advanced Human Resources Training Program) based on long-standing partnerships with Samsung Electronics and SK Hynix. Graduate students in these programs receive full scholarships and are guaranteed employment after graduation. The Department of Semiconductor Systems Engineering, newly established in 2022, selects 100 undergraduate students each year to provide systematic education. Additionally, the KAIST–Samsung Electronics Industry-Academia Cooperation Center, which involves more than 70 labs annually, serves as a long-term hub for joint industry-academia research, contributing to solving critical issues within the industry.
The article emphasizes KAIST's growth beyond a simple research institution into an international research hub. KAIST is enhancing diversity and inclusivity by expanding the hiring of female faculty and establishing a Global Talent Visa Center to support foreign professors and students, attracting outstanding talent from around the world. As a core university within the Daedeok Research Complex (Daedeok Innopolis), it serves as the heart of "Korea's Silicon Valley."
KAIST researchers predict that the future of semiconductor technology is not in simple device miniaturization but in a convergent approach involving neuromorphic technology, 3D packaging technology, and AI applications. This article shows that KAIST's strategic research direction and leadership are gaining attention from both the global academic and industrial communities.
Professor Kyung Min Kim stated, "I am very pleased that KAIST's next-generation semiconductor research and talent development strategy has been widely publicized to domestic and international academia and industry through this article, and we will continue to contribute to the development of future semiconductor technology with innovative convergence research."
KAIST President Kwang Hyung Lee remarked, "Being highlighted for our semiconductor research and education achievements in a world-renowned science journal is a testament to the dedication and pioneering spirit of our university members. I am delighted that KAIST's growth as a global research hub is gaining recognition, and we will continue to expand industry-academia collaboration to lead next-generation semiconductor innovation and play a key role in helping Korea become a future semiconductor powerhouse."
2025.09.05 View 5176
Semiconductor Leadership Spotlighted in Nature Sister Journal
<(From Left) Prof. Shinhyun Choi, Prof. Young Gyu Yoon, Prof.Seunghyub Yoo from the School of Electrical Engineering, Prof. Kyung Min Kim from Materials Science and Engineering>
KAIST (President Kwang Hyung Lee) announced on the 5th of September that its semiconductor research and education achievements were highlighted on August 18 in Nature Reviews Electrical Engineering, a sister journal of the world-renowned scientific journal Nature.
Title: Semiconductor-related research and education at KAIST DOI: 10.1038/s44287-025-00204-3
This special "Focus" article provides a detailed look at KAIST's leadership in next-generation semiconductor research, talent development, and global industry-academia collaboration, presenting a future blueprint for Korea's semiconductor industry. Editor Silvia Conti personally conducted the interviews, with KAIST professors including Kyung Min Kim from the Department of Materials Science and Engineering, and Young Gyu Yoon, Shinhyun Choi, Sung-Yool Choi, and Seunghyub Yoo from the School of Electrical Engineering, participating.
KAIST operates educational programs such as the School of Electrical Engineering, the Department of Semiconductor Systems Engineering, and the Graduate School of Semiconductor Engineering. It is leading next-generation semiconductor research in areas like neuromorphic computing, in-memory computing, and 2D new material-based devices. Building on this foundation, researchers are developing new architectures and devices that transcend the limitations of existing silicon, driving innovation in various application fields such as artificial intelligence, robotics, and medicine.
Notably, research on implementing biological functions like synapses and neurons into hardware platforms using new types of memory such as RRAM and PRAM is gaining international attention. This work opens up possibilities for applications in robots, edge computing, and on-sensor AI systems.
Furthermore, KAIST has operated EPSS (Samsung Advanced Human Resources Training Program) and KEPSI (SK Hynix Semiconductor Advanced Human Resources Training Program) based on long-standing partnerships with Samsung Electronics and SK Hynix. Graduate students in these programs receive full scholarships and are guaranteed employment after graduation. The Department of Semiconductor Systems Engineering, newly established in 2022, selects 100 undergraduate students each year to provide systematic education. Additionally, the KAIST–Samsung Electronics Industry-Academia Cooperation Center, which involves more than 70 labs annually, serves as a long-term hub for joint industry-academia research, contributing to solving critical issues within the industry.
The article emphasizes KAIST's growth beyond a simple research institution into an international research hub. KAIST is enhancing diversity and inclusivity by expanding the hiring of female faculty and establishing a Global Talent Visa Center to support foreign professors and students, attracting outstanding talent from around the world. As a core university within the Daedeok Research Complex (Daedeok Innopolis), it serves as the heart of "Korea's Silicon Valley."
KAIST researchers predict that the future of semiconductor technology is not in simple device miniaturization but in a convergent approach involving neuromorphic technology, 3D packaging technology, and AI applications. This article shows that KAIST's strategic research direction and leadership are gaining attention from both the global academic and industrial communities.
Professor Kyung Min Kim stated, "I am very pleased that KAIST's next-generation semiconductor research and talent development strategy has been widely publicized to domestic and international academia and industry through this article, and we will continue to contribute to the development of future semiconductor technology with innovative convergence research."
KAIST President Kwang Hyung Lee remarked, "Being highlighted for our semiconductor research and education achievements in a world-renowned science journal is a testament to the dedication and pioneering spirit of our university members. I am delighted that KAIST's growth as a global research hub is gaining recognition, and we will continue to expand industry-academia collaboration to lead next-generation semiconductor innovation and play a key role in helping Korea become a future semiconductor powerhouse."
2025.09.05 View 5176 -
 KAIST succeeds in controlling complex altered gene networks to restore them to normal
Previously, research on controlling gene networks has been carried out based on a single stimulus-response of cells. More recently, studies have been proposed to precisely analyze complex gene networks to identify control targets. A KAIST research team has succeeded in developing a universal technology that identifies gene control targets in altered cellular gene networks and restores them. This achievement is expected to be widely applied to new anticancer therapies such as cancer reversibility, drug development, precision medicine, and reprogramming for cell therapy.
KAIST (President Kwang Hyung Lee) announced on the 28th of August that Professor Kwang-Hyun Cho’s research team from the Department of Bio and Brain Engineering has developed a technology to systematically identify gene control targets that can restore the altered stimulus-response patterns of cells to normal by using an algebraic approach. The algebraic approach expresses gene networks as mathematical equations and identifies control targets through algebraic computations.
The research team represented the complex interactions among genes within a cell as a "logic circuit diagram" (Boolean network). Based on this, they visualized how a cell responds to external stimuli as a "landscape map" (phenotype landscape).
By applying a mathematical method called the "semi-tensor product,*" they developed a way to quickly and accurately calculate how the overall cellular response would change if a specific gene were controlled.
*Semi-tensor product: a method that calculates all possible gene combinations and control effects in a single algebraic formula
However, because the key genes that determine actual cellular responses number in the thousands, the calculations are extremely complex. To address this, the research team applied a numerical approximation method (Taylor approximation) to simplify the calculations. In simple terms, they transformed a complex problem into a simpler formula while still yielding nearly identical results.
Through this, the team was able to calculate which stable state (attractor) a cell would reach and predict how the cell’s state would change when a particular gene was controlled. As a result, they were able to identify core gene control targets that could restore abnormal cellular responses to states most similar to normal.
Professor Cho’s team applied the developed control technology to various gene networks and verified that it can accurately predict gene control targets that restore altered stimulus-response patterns of cells back to normal.
In particular, by applying it to bladder cancer cell networks, they identified gene control targets capable of restoring altered responses to normal. They also discovered gene control targets in large-scale distorted gene networks during immune cell differentiation that are capable of restoring normal stimulus-response patterns. This enabled them to solve problems that previously required only approximate searches through lengthy computer simulations in a fast and systematic way.
Professor Cho said, “This study is evaluated as a core original technology for the development of the Digital Cell Twin model*, which analyzes and controls the phenotype landscape of gene networks that determine cell fate. In the future, it is expected to be widely applicable across the life sciences and medicine, including new anticancer therapies through cancer reversibility, drug development, precision medicine, and reprogramming for cell therapy.”
*Digital Cell Twin model: a technology that digitally models the complex reactions occurring within cells, enabling virtual simulations of cellular responses instead of actual experiments
KAIST master’s student Insoo Jung, PhD student Corbin Hopper, PhD student Seong-Hoon Jang, and PhD student Hyunsoo Yeo participated in this study. The results were published online on August 22 in Science Advances, an international journal published by the American Association for the Advancement of Science (AAAS).
※ Paper title: “Reverse Control of Biological Networks to Restore Phenotype Landscapes”
※ DOI: https://www.science.org/doi/10.1126/sciadv.adw3995
This research was supported by the Mid-Career Researcher Program and the Basic Research Laboratory Program of the National Research Foundation of Korea, funded by the Ministry of Science and ICT.
2025.08.29 View 4476
KAIST succeeds in controlling complex altered gene networks to restore them to normal
Previously, research on controlling gene networks has been carried out based on a single stimulus-response of cells. More recently, studies have been proposed to precisely analyze complex gene networks to identify control targets. A KAIST research team has succeeded in developing a universal technology that identifies gene control targets in altered cellular gene networks and restores them. This achievement is expected to be widely applied to new anticancer therapies such as cancer reversibility, drug development, precision medicine, and reprogramming for cell therapy.
KAIST (President Kwang Hyung Lee) announced on the 28th of August that Professor Kwang-Hyun Cho’s research team from the Department of Bio and Brain Engineering has developed a technology to systematically identify gene control targets that can restore the altered stimulus-response patterns of cells to normal by using an algebraic approach. The algebraic approach expresses gene networks as mathematical equations and identifies control targets through algebraic computations.
The research team represented the complex interactions among genes within a cell as a "logic circuit diagram" (Boolean network). Based on this, they visualized how a cell responds to external stimuli as a "landscape map" (phenotype landscape).
By applying a mathematical method called the "semi-tensor product,*" they developed a way to quickly and accurately calculate how the overall cellular response would change if a specific gene were controlled.
*Semi-tensor product: a method that calculates all possible gene combinations and control effects in a single algebraic formula
However, because the key genes that determine actual cellular responses number in the thousands, the calculations are extremely complex. To address this, the research team applied a numerical approximation method (Taylor approximation) to simplify the calculations. In simple terms, they transformed a complex problem into a simpler formula while still yielding nearly identical results.
Through this, the team was able to calculate which stable state (attractor) a cell would reach and predict how the cell’s state would change when a particular gene was controlled. As a result, they were able to identify core gene control targets that could restore abnormal cellular responses to states most similar to normal.
Professor Cho’s team applied the developed control technology to various gene networks and verified that it can accurately predict gene control targets that restore altered stimulus-response patterns of cells back to normal.
In particular, by applying it to bladder cancer cell networks, they identified gene control targets capable of restoring altered responses to normal. They also discovered gene control targets in large-scale distorted gene networks during immune cell differentiation that are capable of restoring normal stimulus-response patterns. This enabled them to solve problems that previously required only approximate searches through lengthy computer simulations in a fast and systematic way.
Professor Cho said, “This study is evaluated as a core original technology for the development of the Digital Cell Twin model*, which analyzes and controls the phenotype landscape of gene networks that determine cell fate. In the future, it is expected to be widely applicable across the life sciences and medicine, including new anticancer therapies through cancer reversibility, drug development, precision medicine, and reprogramming for cell therapy.”
*Digital Cell Twin model: a technology that digitally models the complex reactions occurring within cells, enabling virtual simulations of cellular responses instead of actual experiments
KAIST master’s student Insoo Jung, PhD student Corbin Hopper, PhD student Seong-Hoon Jang, and PhD student Hyunsoo Yeo participated in this study. The results were published online on August 22 in Science Advances, an international journal published by the American Association for the Advancement of Science (AAAS).
※ Paper title: “Reverse Control of Biological Networks to Restore Phenotype Landscapes”
※ DOI: https://www.science.org/doi/10.1126/sciadv.adw3995
This research was supported by the Mid-Career Researcher Program and the Basic Research Laboratory Program of the National Research Foundation of Korea, funded by the Ministry of Science and ICT.
2025.08.29 View 4476 -
 High-Resolution Spectrometer that Fits into Smartphones Developed by KAIST Researchers
- Professor Mooseok Jang's research team at the Department of Bio and Brain Engineering develops an ultra-compact, high-resolution spectrometer using 'double-layer disordered metasurfaces' that generate unique random patterns depending on light's color.
- Unlike conventional dispersion-based spectrometers that were difficult to apply to portable devices, this new concept spectrometer technology achieves 1nm-level high resolution in a device smaller than 1cm, comparable in size to a fingernail.
- It can be utilized as a built-in spectrometer in smartphones and wearable devices in the future, and can be expanded to advanced optical technologies such as hyperspectral imaging and ultrafast imaging.
< Photo 1. (From left) Professor Mooseok Jang, Dong-gu Lee (Ph.D. candidate), Gookho Song (Ph.D. candidate) >
Color, as the way light's wavelength is perceived by the human eye, goes beyond a simple aesthetic element, containing important scientific information like a substance's composition or state. Spectrometers are optical devices that analyze material properties by decomposing light into its constituent wavelengths, and they are widely used in various scientific and industrial fields, including material analysis, chemical component detection, and life science research. Existing high-resolution spectrometers were large and complex, making them difficult for widespread daily use. However, thanks to the ultra-compact, high-resolution spectrometer developed by KAIST researchers, it is now expected that light's color information can be utilized even within smartphones or wearable devices.
KAIST (President Kwang Hyung Lee) announced on the 13th that Professor Mooseok Jang's research team at the Department of Bio and Brain Engineering has successfully developed a reconstruction-based spectrometer technology using double-layer disordered metasurfaces*.
*Double-layer disordered metasurface: An innovative optical device that complexly scatters light through two layers of disordered nanostructures, creating unique and predictable speckle patterns for each wavelength.
Existing high-resolution spectrometers have a large form factor, on the order of tens of centimeters, and require complex calibration processes to maintain accuracy. This fundamentally stems from the operating principle of traditional dispersive elements, such as gratings and prisms, which separate light wavelengths along the propagation direction, much like a rainbow separates colors. Consequently, despite the potential for light's color information to be widely useful in daily life, spectroscopic technology has been limited to laboratory or industrial manufacturing environments.
< Figure 1. Through a simple structure consisting of a double layer of disordered metasurfaces and an image sensor, it was shown that speckles of predictable spectral channels with high spectral resolution can be generated in a compact form factor. The high similarity between the measured and calculated speckles was used to solve the inverse problem and verify the ability to reconstruct the spectrum. >
The research team devised a method that departs from the conventional spectroscopic paradigm of using diffraction gratings or prisms, which establish a one-to-one correspondence between light's color information and its propagation direction, by utilizing designed disordered structures as optical components. In this process, they employed metasurfaces, which can freely control the light propagation process using structures tens to hundreds of nanometers in size, to accurately implement 'complex random patterns (speckle*)'.
*Speckle: An irregular pattern of light intensity created by the interference of multiple wavefronts of light.
Specifically, they developed a method that involves implementing a double-layer disordered metasurface to generate wavelength-specific speckle patterns and then reconstructing precise color information (wavelength) of the light from the random patterns measured by a camera.
As a result, they successfully developed a new concept spectrometer technology that can accurately measure light across a broad range of visible to infrared (440-1,300nm) with a high resolution of 1 nanometer (nm) in a device smaller than a fingernail (less than 1cm) using only a single image capture.
< Figure 2. A disordered metasurface is a metasurface with irregularly arranged structures ranging from tens to hundreds of nanometers in size. In a double-layer structure, a propagation space is placed between the two metasurfaces to control the output speckle with high degrees of freedom, thereby achieving a spectral resolution of 1 nm even in a form factor smaller than 1 cm. >
Dong-gu Lee, a lead author of this study, stated, "This technology is implemented in a way that is directly integrated with commercial image sensors, and we expect that it will enable easy acquisition and utilization of light's wavelength information in daily life when built into mobile devices in the future."
Professor Mooseok Jang said, "This technology overcomes the limitations of existing RGB three-color based machine vision fields, which only distinguish and recognize three color components (red, green, blue), and has diverse applications. We anticipate various applied research for this technology, which expands the horizon of laboratory-level technology to daily-level machine vision technology for applications such as food component analysis, crop health diagnosis, skin health measurement, environmental pollution detection, and bio/medical diagnostics." He added, "Furthermore, it can be extended to various advanced optical technologies such as hyperspectral imaging, which records wavelength and spatial information simultaneously with high resolution, 3D optical trapping technology, which precisely controls light of multiple wavelengths into desired forms, and ultrafast imaging technology, which captures phenomena occurring in very short periods."
This research was collaboratively led by Dong-gu Lee (Ph.D. candidate) and Gookho Song (Ph.D. candidate) from the KAIST Department of Bio and Brain Engineering as co-first authors, with Professor Mooseok Jang as the corresponding author. The findings were published online in the international journal Science Advances on May 28, 2025.* Paper Title: Reconstructive spectrometer using double-layer disordered metasurfaces* DOI: 10.1126/sciadv.adv2376
This research was supported by the Samsung Research Funding and Incubation Center of Samsung Electronics grant, the National Research Foundation of Korea (NRF) grant funded by the Korea government (MSIT), and the Bio & Medical Technology Development Program of the National Research Foundation (NRF) funded by the Korean government (MSIT).
2025.06.13 View 8021
High-Resolution Spectrometer that Fits into Smartphones Developed by KAIST Researchers
- Professor Mooseok Jang's research team at the Department of Bio and Brain Engineering develops an ultra-compact, high-resolution spectrometer using 'double-layer disordered metasurfaces' that generate unique random patterns depending on light's color.
- Unlike conventional dispersion-based spectrometers that were difficult to apply to portable devices, this new concept spectrometer technology achieves 1nm-level high resolution in a device smaller than 1cm, comparable in size to a fingernail.
- It can be utilized as a built-in spectrometer in smartphones and wearable devices in the future, and can be expanded to advanced optical technologies such as hyperspectral imaging and ultrafast imaging.
< Photo 1. (From left) Professor Mooseok Jang, Dong-gu Lee (Ph.D. candidate), Gookho Song (Ph.D. candidate) >
Color, as the way light's wavelength is perceived by the human eye, goes beyond a simple aesthetic element, containing important scientific information like a substance's composition or state. Spectrometers are optical devices that analyze material properties by decomposing light into its constituent wavelengths, and they are widely used in various scientific and industrial fields, including material analysis, chemical component detection, and life science research. Existing high-resolution spectrometers were large and complex, making them difficult for widespread daily use. However, thanks to the ultra-compact, high-resolution spectrometer developed by KAIST researchers, it is now expected that light's color information can be utilized even within smartphones or wearable devices.
KAIST (President Kwang Hyung Lee) announced on the 13th that Professor Mooseok Jang's research team at the Department of Bio and Brain Engineering has successfully developed a reconstruction-based spectrometer technology using double-layer disordered metasurfaces*.
*Double-layer disordered metasurface: An innovative optical device that complexly scatters light through two layers of disordered nanostructures, creating unique and predictable speckle patterns for each wavelength.
Existing high-resolution spectrometers have a large form factor, on the order of tens of centimeters, and require complex calibration processes to maintain accuracy. This fundamentally stems from the operating principle of traditional dispersive elements, such as gratings and prisms, which separate light wavelengths along the propagation direction, much like a rainbow separates colors. Consequently, despite the potential for light's color information to be widely useful in daily life, spectroscopic technology has been limited to laboratory or industrial manufacturing environments.
< Figure 1. Through a simple structure consisting of a double layer of disordered metasurfaces and an image sensor, it was shown that speckles of predictable spectral channels with high spectral resolution can be generated in a compact form factor. The high similarity between the measured and calculated speckles was used to solve the inverse problem and verify the ability to reconstruct the spectrum. >
The research team devised a method that departs from the conventional spectroscopic paradigm of using diffraction gratings or prisms, which establish a one-to-one correspondence between light's color information and its propagation direction, by utilizing designed disordered structures as optical components. In this process, they employed metasurfaces, which can freely control the light propagation process using structures tens to hundreds of nanometers in size, to accurately implement 'complex random patterns (speckle*)'.
*Speckle: An irregular pattern of light intensity created by the interference of multiple wavefronts of light.
Specifically, they developed a method that involves implementing a double-layer disordered metasurface to generate wavelength-specific speckle patterns and then reconstructing precise color information (wavelength) of the light from the random patterns measured by a camera.
As a result, they successfully developed a new concept spectrometer technology that can accurately measure light across a broad range of visible to infrared (440-1,300nm) with a high resolution of 1 nanometer (nm) in a device smaller than a fingernail (less than 1cm) using only a single image capture.
< Figure 2. A disordered metasurface is a metasurface with irregularly arranged structures ranging from tens to hundreds of nanometers in size. In a double-layer structure, a propagation space is placed between the two metasurfaces to control the output speckle with high degrees of freedom, thereby achieving a spectral resolution of 1 nm even in a form factor smaller than 1 cm. >
Dong-gu Lee, a lead author of this study, stated, "This technology is implemented in a way that is directly integrated with commercial image sensors, and we expect that it will enable easy acquisition and utilization of light's wavelength information in daily life when built into mobile devices in the future."
Professor Mooseok Jang said, "This technology overcomes the limitations of existing RGB three-color based machine vision fields, which only distinguish and recognize three color components (red, green, blue), and has diverse applications. We anticipate various applied research for this technology, which expands the horizon of laboratory-level technology to daily-level machine vision technology for applications such as food component analysis, crop health diagnosis, skin health measurement, environmental pollution detection, and bio/medical diagnostics." He added, "Furthermore, it can be extended to various advanced optical technologies such as hyperspectral imaging, which records wavelength and spatial information simultaneously with high resolution, 3D optical trapping technology, which precisely controls light of multiple wavelengths into desired forms, and ultrafast imaging technology, which captures phenomena occurring in very short periods."
This research was collaboratively led by Dong-gu Lee (Ph.D. candidate) and Gookho Song (Ph.D. candidate) from the KAIST Department of Bio and Brain Engineering as co-first authors, with Professor Mooseok Jang as the corresponding author. The findings were published online in the international journal Science Advances on May 28, 2025.* Paper Title: Reconstructive spectrometer using double-layer disordered metasurfaces* DOI: 10.1126/sciadv.adv2376
This research was supported by the Samsung Research Funding and Incubation Center of Samsung Electronics grant, the National Research Foundation of Korea (NRF) grant funded by the Korea government (MSIT), and the Bio & Medical Technology Development Program of the National Research Foundation (NRF) funded by the Korean government (MSIT).
2025.06.13 View 8021 -
 KAIST to Develop a Korean-style ChatGPT Platform Specifically Geared Toward Medical Diagnosis and Drug Discovery
On May 23rd, KAIST (President Kwang-Hyung Lee) announced that its Digital Bio-Health AI Research Center (Director: Professor JongChul Ye of KAIST Kim Jaechul Graduate School of AI) has been selected for the Ministry of Science and ICT's 'AI Top-Tier Young Researcher Support Program (AI Star Fellowship Project).' With a total investment of ₩11.5 billion from May 2025 to December 2030, the center will embark on the full-scale development of AI technology and a platform capable of independently inferring and determining the kinds of diseases, and discovering new drugs.
< Photo. On May 20th, a kick-off meeting for the AI Star Fellowship Project was held at KAIST Kim Jaechul Graduate School of AI’s Yangjae Research Center with the KAIST research team and participating organizations of Samsung Medical Center, NAVER Cloud, and HITS. [From left to right in the front row] Professor Jaegul Joo (KAIST), Professor Yoonjae Choi (KAIST), Professor Woo Youn Kim (KAIST/HITS), Professor JongChul Ye (KAIST), Professor Sungsoo Ahn (KAIST), Dr. Haanju Yoo (NAVER Cloud), Yoonho Lee (KAIST), HyeYoon Moon (Samsung Medical Center), Dr. Su Min Kim (Samsung Medical Center) >
This project aims to foster an innovative AI research ecosystem centered on young researchers and develop an inferential AI agent that can utilize and automatically expand specialized knowledge systems in the bio and medical fields.
Professor JongChul Ye of the Kim Jaechul Graduate School of AI will serve as the lead researcher, with young researchers from KAIST including Professors Yoonjae Choi, Kimin Lee, Sungsoo Ahn, and Chanyoung Park, along with mid-career researchers like Professors Jaegul Joo and Woo Youn Kim, jointly undertaking the project. They will collaborate with various laboratories within KAIST to conduct comprehensive research covering the entire cycle from the theoretical foundations of AI inference to its practical application.
Specifically, the main goals include: - Building high-performance inference models that integrate diverse medical knowledge systems to enhance the precision and reliability of diagnosis and treatment. - Developing a convergence inference platform that efficiently combines symbol-based inference with neural network models. - Securing AI technology for new drug development and biomarker discovery based on 'cell ontology.'
Furthermore, through close collaboration with industry and medical institutions such as Samsung Medical Center, NAVER Cloud, and HITS Co., Ltd., the project aims to achieve: - Clinical diagnostic AI utilizing medical knowledge systems. - AI-based molecular target exploration for new drug development. - Commercialization of an extendible AI inference platform.
Professor JongChul Ye, Director of KAIST's Digital Bio-Health AI Research Center, stated, "At a time when competition in AI inference model development is intensifying, it is a great honor for KAIST to lead the development of AI technology specialized in the bio and medical fields with world-class young researchers." He added, "We will do our best to ensure that the participating young researchers reach a world-leading level in terms of research achievements after the completion of this seven-year project starting in 2025."
The AI Star Fellowship is a newly established program where post-doctoral researchers and faculty members within seven years of appointment participate as project leaders (PLs) to independently lead research. Multiple laboratories within a university and demand-side companies form a consortium to operate the program.
Through this initiative, KAIST plans to nurture bio-medical convergence AI talent and simultaneously promote the commercialization of core technologies in collaboration with Samsung Medical Center, NAVER Cloud, and HITS.
2025.05.26 View 14866
KAIST to Develop a Korean-style ChatGPT Platform Specifically Geared Toward Medical Diagnosis and Drug Discovery
On May 23rd, KAIST (President Kwang-Hyung Lee) announced that its Digital Bio-Health AI Research Center (Director: Professor JongChul Ye of KAIST Kim Jaechul Graduate School of AI) has been selected for the Ministry of Science and ICT's 'AI Top-Tier Young Researcher Support Program (AI Star Fellowship Project).' With a total investment of ₩11.5 billion from May 2025 to December 2030, the center will embark on the full-scale development of AI technology and a platform capable of independently inferring and determining the kinds of diseases, and discovering new drugs.
< Photo. On May 20th, a kick-off meeting for the AI Star Fellowship Project was held at KAIST Kim Jaechul Graduate School of AI’s Yangjae Research Center with the KAIST research team and participating organizations of Samsung Medical Center, NAVER Cloud, and HITS. [From left to right in the front row] Professor Jaegul Joo (KAIST), Professor Yoonjae Choi (KAIST), Professor Woo Youn Kim (KAIST/HITS), Professor JongChul Ye (KAIST), Professor Sungsoo Ahn (KAIST), Dr. Haanju Yoo (NAVER Cloud), Yoonho Lee (KAIST), HyeYoon Moon (Samsung Medical Center), Dr. Su Min Kim (Samsung Medical Center) >
This project aims to foster an innovative AI research ecosystem centered on young researchers and develop an inferential AI agent that can utilize and automatically expand specialized knowledge systems in the bio and medical fields.
Professor JongChul Ye of the Kim Jaechul Graduate School of AI will serve as the lead researcher, with young researchers from KAIST including Professors Yoonjae Choi, Kimin Lee, Sungsoo Ahn, and Chanyoung Park, along with mid-career researchers like Professors Jaegul Joo and Woo Youn Kim, jointly undertaking the project. They will collaborate with various laboratories within KAIST to conduct comprehensive research covering the entire cycle from the theoretical foundations of AI inference to its practical application.
Specifically, the main goals include: - Building high-performance inference models that integrate diverse medical knowledge systems to enhance the precision and reliability of diagnosis and treatment. - Developing a convergence inference platform that efficiently combines symbol-based inference with neural network models. - Securing AI technology for new drug development and biomarker discovery based on 'cell ontology.'
Furthermore, through close collaboration with industry and medical institutions such as Samsung Medical Center, NAVER Cloud, and HITS Co., Ltd., the project aims to achieve: - Clinical diagnostic AI utilizing medical knowledge systems. - AI-based molecular target exploration for new drug development. - Commercialization of an extendible AI inference platform.
Professor JongChul Ye, Director of KAIST's Digital Bio-Health AI Research Center, stated, "At a time when competition in AI inference model development is intensifying, it is a great honor for KAIST to lead the development of AI technology specialized in the bio and medical fields with world-class young researchers." He added, "We will do our best to ensure that the participating young researchers reach a world-leading level in terms of research achievements after the completion of this seven-year project starting in 2025."
The AI Star Fellowship is a newly established program where post-doctoral researchers and faculty members within seven years of appointment participate as project leaders (PLs) to independently lead research. Multiple laboratories within a university and demand-side companies form a consortium to operate the program.
Through this initiative, KAIST plans to nurture bio-medical convergence AI talent and simultaneously promote the commercialization of core technologies in collaboration with Samsung Medical Center, NAVER Cloud, and HITS.
2025.05.26 View 14866 -
 KAIST's Pioneering VR Precision Technology & Choreography Tool Receive Spotlights at CHI 2025
Accurate pointing in virtual spaces is essential for seamless interaction. If pointing is not precise, selecting the desired object becomes challenging, breaking user immersion and reducing overall experience quality. KAIST researchers have developed a technology that offers a vivid, lifelike experience in virtual space, alongside a new tool that assists choreographers throughout the creative process.
KAIST (President Kwang-Hyung Lee) announced on May 13th that a research team led by Professor Sang Ho Yoon of the Graduate School of Culture Technology, in collaboration with Professor Yang Zhang of the University of California, Los Angeles (UCLA), has developed the ‘T2IRay’ technology and the ‘ChoreoCraft’ platform, which enables choreographers to work more freely and creatively in virtual reality. These technologies received two Honorable Mention awards, recognizing the top 5% of papers, at CHI 2025*, the best international conference in the field of human-computer interaction, hosted by the Association for Computing Machinery (ACM) from April 25 to May 1.
< (From left) PhD candidates Jina Kim and Kyungeun Jung along with Master's candidate, Hyunyoung Han and Professor Sang Ho Yoon of KAIST Graduate School of Culture Technology and Professor Yang Zhang (top) of UCLA >
T2IRay: Enabling Virtual Input with Precision
T2IRay introduces a novel input method that allows for precise object pointing in virtual environments by expanding traditional thumb-to-index gestures. This approach overcomes previous limitations, such as interruptions or reduced accuracy due to changes in hand position or orientation.
The technology uses a local coordinate system based on finger relationships, ensuring continuous input even as hand positions shift. It accurately captures subtle thumb movements within this coordinate system, integrating natural head movements to allow fluid, intuitive control across a wide range.
< Figure 1. T2IRay framework utilizing the delicate movements of the thumb and index fingers for AR/VR pointing >
Professor Sang Ho Yoon explained, “T2IRay can significantly enhance the user experience in AR/VR by enabling smooth, stable control even when the user’s hands are in motion.”
This study, led by first author Jina Kim, was supported by the Excellent New Researcher Support Project of the National Research Foundation of Korea under the Ministry of Science and ICT, as well as the University ICT Research Center (ITRC) Support Project of the Institute of Information and Communications Technology Planning and Evaluation (IITP).
▴ Paper title: T2IRay: Design of Thumb-to-Index Based Indirect Pointing for Continuous and Robust AR/VR Input▴ Paper link: https://doi.org/10.1145/3706598.3713442
▴ T2IRay demo video: https://youtu.be/ElJlcJbkJPY
ChoreoCraft: Creativity Support through VR for Choreographers
In addition, Professor Yoon’s team developed ‘ChoreoCraft,’ a virtual reality tool designed to support choreographers by addressing the unique challenges they face, such as memorizing complex movements, overcoming creative blocks, and managing subjective feedback.
ChoreoCraft reduces reliance on memory by allowing choreographers to save and refine movements directly within a VR space, using a motion-capture avatar for real-time interaction. It also enhances creativity by suggesting movements that naturally fit with prior choreography and musical elements. Furthermore, the system provides quantitative feedback by analyzing kinematic factors like motion stability and engagement, helping choreographers make data-driven creative decisions.
< Figure 2. ChoreoCraft's approaches to encourage creative process >
Professor Yoon noted, “ChoreoCraft is a tool designed to address the core challenges faced by choreographers, enhancing both creativity and efficiency. In user tests with professional choreographers, it received high marks for its ability to spark creative ideas and provide valuable quantitative feedback.”
This research was conducted in collaboration with doctoral candidate Kyungeun Jung and master’s candidate Hyunyoung Han, alongside the Electronics and Telecommunications Research Institute (ETRI) and One Million Co., Ltd. (CEO Hye-rang Kim), with support from the Cultural and Arts Immersive Service Development Project by the Ministry of Culture, Sports and Tourism.
▴ Paper title: ChoreoCraft: In-situ Crafting of Choreography in Virtual Reality through Creativity Support Tools▴ Paper link: https://doi.org/10.1145/3706598.3714220
▴ ChoreoCraft demo video: https://youtu.be/Ms1fwiSBjjw
*CHI (Conference on Human Factors in Computing Systems): The premier international conference on human-computer interaction, organized by the ACM, was held this year from April 25 to May 1, 2025.
2025.05.13 View 11750
KAIST's Pioneering VR Precision Technology & Choreography Tool Receive Spotlights at CHI 2025
Accurate pointing in virtual spaces is essential for seamless interaction. If pointing is not precise, selecting the desired object becomes challenging, breaking user immersion and reducing overall experience quality. KAIST researchers have developed a technology that offers a vivid, lifelike experience in virtual space, alongside a new tool that assists choreographers throughout the creative process.
KAIST (President Kwang-Hyung Lee) announced on May 13th that a research team led by Professor Sang Ho Yoon of the Graduate School of Culture Technology, in collaboration with Professor Yang Zhang of the University of California, Los Angeles (UCLA), has developed the ‘T2IRay’ technology and the ‘ChoreoCraft’ platform, which enables choreographers to work more freely and creatively in virtual reality. These technologies received two Honorable Mention awards, recognizing the top 5% of papers, at CHI 2025*, the best international conference in the field of human-computer interaction, hosted by the Association for Computing Machinery (ACM) from April 25 to May 1.
< (From left) PhD candidates Jina Kim and Kyungeun Jung along with Master's candidate, Hyunyoung Han and Professor Sang Ho Yoon of KAIST Graduate School of Culture Technology and Professor Yang Zhang (top) of UCLA >
T2IRay: Enabling Virtual Input with Precision
T2IRay introduces a novel input method that allows for precise object pointing in virtual environments by expanding traditional thumb-to-index gestures. This approach overcomes previous limitations, such as interruptions or reduced accuracy due to changes in hand position or orientation.
The technology uses a local coordinate system based on finger relationships, ensuring continuous input even as hand positions shift. It accurately captures subtle thumb movements within this coordinate system, integrating natural head movements to allow fluid, intuitive control across a wide range.
< Figure 1. T2IRay framework utilizing the delicate movements of the thumb and index fingers for AR/VR pointing >
Professor Sang Ho Yoon explained, “T2IRay can significantly enhance the user experience in AR/VR by enabling smooth, stable control even when the user’s hands are in motion.”
This study, led by first author Jina Kim, was supported by the Excellent New Researcher Support Project of the National Research Foundation of Korea under the Ministry of Science and ICT, as well as the University ICT Research Center (ITRC) Support Project of the Institute of Information and Communications Technology Planning and Evaluation (IITP).
▴ Paper title: T2IRay: Design of Thumb-to-Index Based Indirect Pointing for Continuous and Robust AR/VR Input▴ Paper link: https://doi.org/10.1145/3706598.3713442
▴ T2IRay demo video: https://youtu.be/ElJlcJbkJPY
ChoreoCraft: Creativity Support through VR for Choreographers
In addition, Professor Yoon’s team developed ‘ChoreoCraft,’ a virtual reality tool designed to support choreographers by addressing the unique challenges they face, such as memorizing complex movements, overcoming creative blocks, and managing subjective feedback.
ChoreoCraft reduces reliance on memory by allowing choreographers to save and refine movements directly within a VR space, using a motion-capture avatar for real-time interaction. It also enhances creativity by suggesting movements that naturally fit with prior choreography and musical elements. Furthermore, the system provides quantitative feedback by analyzing kinematic factors like motion stability and engagement, helping choreographers make data-driven creative decisions.
< Figure 2. ChoreoCraft's approaches to encourage creative process >
Professor Yoon noted, “ChoreoCraft is a tool designed to address the core challenges faced by choreographers, enhancing both creativity and efficiency. In user tests with professional choreographers, it received high marks for its ability to spark creative ideas and provide valuable quantitative feedback.”
This research was conducted in collaboration with doctoral candidate Kyungeun Jung and master’s candidate Hyunyoung Han, alongside the Electronics and Telecommunications Research Institute (ETRI) and One Million Co., Ltd. (CEO Hye-rang Kim), with support from the Cultural and Arts Immersive Service Development Project by the Ministry of Culture, Sports and Tourism.
▴ Paper title: ChoreoCraft: In-situ Crafting of Choreography in Virtual Reality through Creativity Support Tools▴ Paper link: https://doi.org/10.1145/3706598.3714220
▴ ChoreoCraft demo video: https://youtu.be/Ms1fwiSBjjw
*CHI (Conference on Human Factors in Computing Systems): The premier international conference on human-computer interaction, organized by the ACM, was held this year from April 25 to May 1, 2025.
2025.05.13 View 11750 -
 KAIST & CMU Unveils Amuse, a Songwriting AI-Collaborator to Help Create Music
Wouldn't it be great if music creators had someone to brainstorm with, help them when they're stuck, and explore different musical directions together? Researchers of KAIST and Carnegie Mellon University (CMU) have developed AI technology similar to a fellow songwriter who helps create music.
KAIST (President Kwang-Hyung Lee) has developed an AI-based music creation support system, Amuse, by a research team led by Professor Sung-Ju Lee of the School of Electrical Engineering in collaboration with CMU. The research was presented at the ACM Conference on Human Factors in Computing Systems (CHI), one of the world’s top conferences in human-computer interaction, held in Yokohama, Japan from April 26 to May 1. It received the Best Paper Award, given to only the top 1% of all submissions.
< (From left) Professor Chris Donahue of Carnegie Mellon University, Ph.D. Student Yewon Kim and Professor Sung-Ju Lee of the School of Electrical Engineering >
The system developed by Professor Sung-Ju Lee’s research team, Amuse, is an AI-based system that converts various forms of inspiration such as text, images, and audio into harmonic structures (chord progressions) to support composition.
For example, if a user inputs a phrase, image, or sound clip such as “memories of a warm summer beach”, Amuse automatically generates and suggests chord progressions that match the inspiration.
Unlike existing generative AI, Amuse is differentiated in that it respects the user's creative flow and naturally induces creative exploration through an interactive method that allows flexible integration and modification of AI suggestions.
The core technology of the Amuse system is a generation method that blends two approaches: a large language model creates music code based on the user's prompt and inspiration, while another AI model, trained on real music data, filters out awkward or unnatural results using rejection sampling.
< Figure 1. Amuse system configuration. After extracting music keywords from user input, a large language model-based code progression is generated and refined through rejection sampling (left). Code extraction from audio input is also possible (right). The bottom is an example visualizing the chord structure of the generated code. >
The research team conducted a user study targeting actual musicians and evaluated that Amuse has high potential as a creative companion, or a Co-Creative AI, a concept in which people and AI collaborate, rather than having a generative AI simply put together a song.
The paper, in which a Ph.D. student Yewon Kim and Professor Sung-Ju Lee of KAIST School of Electrical and Electronic Engineering and Carnegie Mellon University Professor Chris Donahue participated, demonstrated the potential of creative AI system design in both academia and industry. ※ Paper title: Amuse: Human-AI Collaborative Songwriting with Multimodal Inspirations DOI: https://doi.org/10.1145/3706598.3713818
※ Research demo video: https://youtu.be/udilkRSnftI?si=FNXccC9EjxHOCrm1
※ Research homepage: https://nmsl.kaist.ac.kr/projects/amuse/
Professor Sung-Ju Lee said, “Recent generative AI technology has raised concerns in that it directly imitates copyrighted content, thereby violating the copyright of the creator, or generating results one-way regardless of the creator’s intention. Accordingly, the research team was aware of this trend, paid attention to what the creator actually needs, and focused on designing an AI system centered on the creator.”
He continued, “Amuse is an attempt to explore the possibility of collaboration with AI while maintaining the initiative of the creator, and is expected to be a starting point for suggesting a more creator-friendly direction in the development of music creation tools and generative AI systems in the future.”
This research was conducted with the support of the National Research Foundation of Korea with funding from the government (Ministry of Science and ICT). (RS-2024-00337007)
2025.05.07 View 15345
KAIST & CMU Unveils Amuse, a Songwriting AI-Collaborator to Help Create Music
Wouldn't it be great if music creators had someone to brainstorm with, help them when they're stuck, and explore different musical directions together? Researchers of KAIST and Carnegie Mellon University (CMU) have developed AI technology similar to a fellow songwriter who helps create music.
KAIST (President Kwang-Hyung Lee) has developed an AI-based music creation support system, Amuse, by a research team led by Professor Sung-Ju Lee of the School of Electrical Engineering in collaboration with CMU. The research was presented at the ACM Conference on Human Factors in Computing Systems (CHI), one of the world’s top conferences in human-computer interaction, held in Yokohama, Japan from April 26 to May 1. It received the Best Paper Award, given to only the top 1% of all submissions.
< (From left) Professor Chris Donahue of Carnegie Mellon University, Ph.D. Student Yewon Kim and Professor Sung-Ju Lee of the School of Electrical Engineering >
The system developed by Professor Sung-Ju Lee’s research team, Amuse, is an AI-based system that converts various forms of inspiration such as text, images, and audio into harmonic structures (chord progressions) to support composition.
For example, if a user inputs a phrase, image, or sound clip such as “memories of a warm summer beach”, Amuse automatically generates and suggests chord progressions that match the inspiration.
Unlike existing generative AI, Amuse is differentiated in that it respects the user's creative flow and naturally induces creative exploration through an interactive method that allows flexible integration and modification of AI suggestions.
The core technology of the Amuse system is a generation method that blends two approaches: a large language model creates music code based on the user's prompt and inspiration, while another AI model, trained on real music data, filters out awkward or unnatural results using rejection sampling.
< Figure 1. Amuse system configuration. After extracting music keywords from user input, a large language model-based code progression is generated and refined through rejection sampling (left). Code extraction from audio input is also possible (right). The bottom is an example visualizing the chord structure of the generated code. >
The research team conducted a user study targeting actual musicians and evaluated that Amuse has high potential as a creative companion, or a Co-Creative AI, a concept in which people and AI collaborate, rather than having a generative AI simply put together a song.
The paper, in which a Ph.D. student Yewon Kim and Professor Sung-Ju Lee of KAIST School of Electrical and Electronic Engineering and Carnegie Mellon University Professor Chris Donahue participated, demonstrated the potential of creative AI system design in both academia and industry. ※ Paper title: Amuse: Human-AI Collaborative Songwriting with Multimodal Inspirations DOI: https://doi.org/10.1145/3706598.3713818
※ Research demo video: https://youtu.be/udilkRSnftI?si=FNXccC9EjxHOCrm1
※ Research homepage: https://nmsl.kaist.ac.kr/projects/amuse/
Professor Sung-Ju Lee said, “Recent generative AI technology has raised concerns in that it directly imitates copyrighted content, thereby violating the copyright of the creator, or generating results one-way regardless of the creator’s intention. Accordingly, the research team was aware of this trend, paid attention to what the creator actually needs, and focused on designing an AI system centered on the creator.”
He continued, “Amuse is an attempt to explore the possibility of collaboration with AI while maintaining the initiative of the creator, and is expected to be a starting point for suggesting a more creator-friendly direction in the development of music creation tools and generative AI systems in the future.”
This research was conducted with the support of the National Research Foundation of Korea with funding from the government (Ministry of Science and ICT). (RS-2024-00337007)
2025.05.07 View 15345 -
 KAIST Identifies Master Regulator Blocking Immunotherapy, Paving the Way for a New Lung Cancer Treatment
Immune checkpoint inhibitors, a class of immunotherapies that help immune cells attack cancer more effectively, have revolutionized cancer treatment. However, fewer than 20% of patients respond to these treatments, highlighting the urgent need for new strategies tailored to both responders and non-responders.
KAIST researchers have discovered that 'DEAD-box helicases 54 (DDX54)', a type of RNA-binding protein, is the master regulator that hinders the effectiveness of immunotherapy—opening a new path for lung cancer treatment. This breakthrough technology has been transferred to faculty startup BioRevert Inc., where it is currently being developed as a companion therapeutic and is expected to enter clinical trials by 2028.
< Photo 1. (From left) Researcher Jungeun Lee, Professor Kwang-Hyun Cho and Postdoctoral Researcher Jeong-Ryeol Gong of the Department of Bio and Brain Engineering at KAIST >
KAIST (represented by President Kwang-Hyung Lee) announced on April 8 that a research team led by Professor Kwang-Hyun Cho from the Department of Bio and Brain Engineering had identified DDX54 as a critical factor that determines the immune evasion capacity of lung cancer cells. They demonstrated that suppressing DDX54 enhances immune cell infiltration into tumors and significantly improves the efficacy of immunotherapy.
Immunotherapy using anti-PD-1 or anti-PD-L1 antibodies is considered a powerful approach in cancer treatment. However, its low response rate limits the number of patients who actually benefit.
To identify likely responders, tumor mutational burden (TMB) has recently been approved by the FDA as a key biomarker for immunotherapy. Cancers with high mutation rates are thought to be more responsive to immune checkpoint inhibitors. However, even tumors with high TMB can display an “immune-desert” phenotype—where immune cell infiltration is severely limited—resulting in poor treatment responses.
< Figure 1. DDX54 was identified as the master regulator that induces resistance to immunotherapy by orchestrating suppression of immune cell infiltration through cancer tissues as lung cancer cells become immune-evasive >
Professor Kwang-Hyun Cho's research team compared transcriptome and genome data of lung cancer patients with immune evasion capabilities through gene regulatory network analysis (A) and discovered DDX54, a master regulator that induces resistance to immunotherapy (B-F).
This study is especially significant in that it successfully demonstrated that suppressing DDX54 in immune-desert lung tumors can overcome immunotherapy resistance and improve treatment outcomes.
The team used transcriptomic and genomic data from immune-evasive lung cancer patients and employed systems biology techniques to infer gene regulatory networks. Through this analysis, they identified DDX54 as a central regulator in the immune evasion of lung cancer cells.
In a syngeneic mouse model, the suppression of DDX54 led to significant increases in the infiltration of anti-cancer immune cells such as T cells and NK cells, and greatly improved the response to immunotherapy.
Single-cell transcriptomic and spatial transcriptomic analyses further showed that combination therapy targeting DDX54 promoted the differentiation of T cells and memory T cells that suppress tumors, while reducing the infiltration of regulatory T cells and exhausted T cells that support tumor growth.
< Figure 2. In the syngeneic mouse model made of lung cancer cells, it was confirmed that inhibiting DDX54 reversed the immune-evasion ability of cancer cells and enhanced the sensitivity to anti-PD-1 therapy >
In a syngeneic mouse model made of lung cancer cells exhibiting immunotherapy resistance, the treatment applied after DDX54 inhibition resulted in statistically significant inhibition of lung cancer growth (B-D) and a significant increase in immune cell infiltration into the tumor tissue (E, F).
The mechanism is believed to involve DDX54 suppression inactivating signaling pathways such as JAK-STAT, MYC, and NF-κB, thereby downregulating immune-evasive proteins CD38 and CD47. This also reduced the infiltration of circulating monocytes—which promote tumor development—and promoted the differentiation of M1 macrophages that play anti-tumor roles.
Professor Kwang-Hyun Cho stated, “We have, for the first time, identified a master regulatory factor that enables immune evasion in lung cancer cells. By targeting this factor, we developed a new therapeutic strategy that can induce responsiveness to immunotherapy in previously resistant cancers.”
He added, “The discovery of DDX54—hidden within the complex molecular networks of cancer cells—was made possible through the systematic integration of systems biology, combining IT and BT.”
The study, led by Professor Kwang-Hyun Cho, was published in the Proceedings of the National Academy of Sciences of the United States of America (PNAS) on April 2, 2025, with Jeong-Ryeol Gong being the first author, Jungeun Lee, a co-first author, and Younghyun Han, a co-author of the article.
< Figure 3. Single-cell transcriptome and spatial transcriptome analysis confirmed that knockdown of DDX54 increased immune cell infiltration into cancer tissues >
In a syngeneic mouse model made of lung cancer cells that underwent immunotherapy in combination with DDX54 inhibition, single-cell transcriptome (H-L) and spatial transcriptome (A-G) analysis of immune cells infiltrating inside cancer tissues were performed. As a result, it was confirmed that anticancer immune cells such as T cells, B cells, and NK cells actively infiltrated the core of lung cancer tissues when DDX54 inhibition and immunotherapy were concurrently administered.
(Paper title: “DDX54 downregulation enhances anti-PD1 therapy in immune-desert lung tumors with high tumor mutational burden,” DOI: https://doi.org/10.1073/pnas.2412310122)
This work was supported by the Ministry of Science and ICT and the National Research Foundation of Korea through the Mid-Career Research Program and Basic Research Laboratory Program.
< Figure 4. The identified master regulator DDX54 was confirmed to induce CD38 and CD47 expression through Jak-Stat3, MYC, and NF-κB activation. >
DDX54 activates the Jak-Stat3, MYC, and NF-κB pathways in lung cancer cells to increase CD38 and CD47 expression (A-G). This creates a cancer microenvironment that contributes to cancer development (H) and ultimately induces immune anticancer treatment resistance.
< Figure 5. It was confirmed that an immune-inflamed environment can be created by combining DDX54 inhibition and immune checkpoint inhibitor (ICI) therapy. >
When DDX54 inhibition and ICI therapy are simultaneously administered, the cancer cell characteristics change, the immune evasion ability is restored, and the environment is transformed into an ‘immune-activated’ environment in which immune cells easily infiltrate cancer tissues. This strengthens the anticancer immune response, thereby increasing the sensitivity of immunotherapy even in lung cancer tissues that previously had low responsiveness to immunotherapy.
2025.04.08 View 14175
KAIST Identifies Master Regulator Blocking Immunotherapy, Paving the Way for a New Lung Cancer Treatment
Immune checkpoint inhibitors, a class of immunotherapies that help immune cells attack cancer more effectively, have revolutionized cancer treatment. However, fewer than 20% of patients respond to these treatments, highlighting the urgent need for new strategies tailored to both responders and non-responders.
KAIST researchers have discovered that 'DEAD-box helicases 54 (DDX54)', a type of RNA-binding protein, is the master regulator that hinders the effectiveness of immunotherapy—opening a new path for lung cancer treatment. This breakthrough technology has been transferred to faculty startup BioRevert Inc., where it is currently being developed as a companion therapeutic and is expected to enter clinical trials by 2028.
< Photo 1. (From left) Researcher Jungeun Lee, Professor Kwang-Hyun Cho and Postdoctoral Researcher Jeong-Ryeol Gong of the Department of Bio and Brain Engineering at KAIST >
KAIST (represented by President Kwang-Hyung Lee) announced on April 8 that a research team led by Professor Kwang-Hyun Cho from the Department of Bio and Brain Engineering had identified DDX54 as a critical factor that determines the immune evasion capacity of lung cancer cells. They demonstrated that suppressing DDX54 enhances immune cell infiltration into tumors and significantly improves the efficacy of immunotherapy.
Immunotherapy using anti-PD-1 or anti-PD-L1 antibodies is considered a powerful approach in cancer treatment. However, its low response rate limits the number of patients who actually benefit.
To identify likely responders, tumor mutational burden (TMB) has recently been approved by the FDA as a key biomarker for immunotherapy. Cancers with high mutation rates are thought to be more responsive to immune checkpoint inhibitors. However, even tumors with high TMB can display an “immune-desert” phenotype—where immune cell infiltration is severely limited—resulting in poor treatment responses.
< Figure 1. DDX54 was identified as the master regulator that induces resistance to immunotherapy by orchestrating suppression of immune cell infiltration through cancer tissues as lung cancer cells become immune-evasive >
Professor Kwang-Hyun Cho's research team compared transcriptome and genome data of lung cancer patients with immune evasion capabilities through gene regulatory network analysis (A) and discovered DDX54, a master regulator that induces resistance to immunotherapy (B-F).
This study is especially significant in that it successfully demonstrated that suppressing DDX54 in immune-desert lung tumors can overcome immunotherapy resistance and improve treatment outcomes.
The team used transcriptomic and genomic data from immune-evasive lung cancer patients and employed systems biology techniques to infer gene regulatory networks. Through this analysis, they identified DDX54 as a central regulator in the immune evasion of lung cancer cells.
In a syngeneic mouse model, the suppression of DDX54 led to significant increases in the infiltration of anti-cancer immune cells such as T cells and NK cells, and greatly improved the response to immunotherapy.
Single-cell transcriptomic and spatial transcriptomic analyses further showed that combination therapy targeting DDX54 promoted the differentiation of T cells and memory T cells that suppress tumors, while reducing the infiltration of regulatory T cells and exhausted T cells that support tumor growth.
< Figure 2. In the syngeneic mouse model made of lung cancer cells, it was confirmed that inhibiting DDX54 reversed the immune-evasion ability of cancer cells and enhanced the sensitivity to anti-PD-1 therapy >
In a syngeneic mouse model made of lung cancer cells exhibiting immunotherapy resistance, the treatment applied after DDX54 inhibition resulted in statistically significant inhibition of lung cancer growth (B-D) and a significant increase in immune cell infiltration into the tumor tissue (E, F).
The mechanism is believed to involve DDX54 suppression inactivating signaling pathways such as JAK-STAT, MYC, and NF-κB, thereby downregulating immune-evasive proteins CD38 and CD47. This also reduced the infiltration of circulating monocytes—which promote tumor development—and promoted the differentiation of M1 macrophages that play anti-tumor roles.
Professor Kwang-Hyun Cho stated, “We have, for the first time, identified a master regulatory factor that enables immune evasion in lung cancer cells. By targeting this factor, we developed a new therapeutic strategy that can induce responsiveness to immunotherapy in previously resistant cancers.”
He added, “The discovery of DDX54—hidden within the complex molecular networks of cancer cells—was made possible through the systematic integration of systems biology, combining IT and BT.”
The study, led by Professor Kwang-Hyun Cho, was published in the Proceedings of the National Academy of Sciences of the United States of America (PNAS) on April 2, 2025, with Jeong-Ryeol Gong being the first author, Jungeun Lee, a co-first author, and Younghyun Han, a co-author of the article.
< Figure 3. Single-cell transcriptome and spatial transcriptome analysis confirmed that knockdown of DDX54 increased immune cell infiltration into cancer tissues >
In a syngeneic mouse model made of lung cancer cells that underwent immunotherapy in combination with DDX54 inhibition, single-cell transcriptome (H-L) and spatial transcriptome (A-G) analysis of immune cells infiltrating inside cancer tissues were performed. As a result, it was confirmed that anticancer immune cells such as T cells, B cells, and NK cells actively infiltrated the core of lung cancer tissues when DDX54 inhibition and immunotherapy were concurrently administered.
(Paper title: “DDX54 downregulation enhances anti-PD1 therapy in immune-desert lung tumors with high tumor mutational burden,” DOI: https://doi.org/10.1073/pnas.2412310122)
This work was supported by the Ministry of Science and ICT and the National Research Foundation of Korea through the Mid-Career Research Program and Basic Research Laboratory Program.
< Figure 4. The identified master regulator DDX54 was confirmed to induce CD38 and CD47 expression through Jak-Stat3, MYC, and NF-κB activation. >
DDX54 activates the Jak-Stat3, MYC, and NF-κB pathways in lung cancer cells to increase CD38 and CD47 expression (A-G). This creates a cancer microenvironment that contributes to cancer development (H) and ultimately induces immune anticancer treatment resistance.
< Figure 5. It was confirmed that an immune-inflamed environment can be created by combining DDX54 inhibition and immune checkpoint inhibitor (ICI) therapy. >
When DDX54 inhibition and ICI therapy are simultaneously administered, the cancer cell characteristics change, the immune evasion ability is restored, and the environment is transformed into an ‘immune-activated’ environment in which immune cells easily infiltrate cancer tissues. This strengthens the anticancer immune response, thereby increasing the sensitivity of immunotherapy even in lung cancer tissues that previously had low responsiveness to immunotherapy.
2025.04.08 View 14175 -
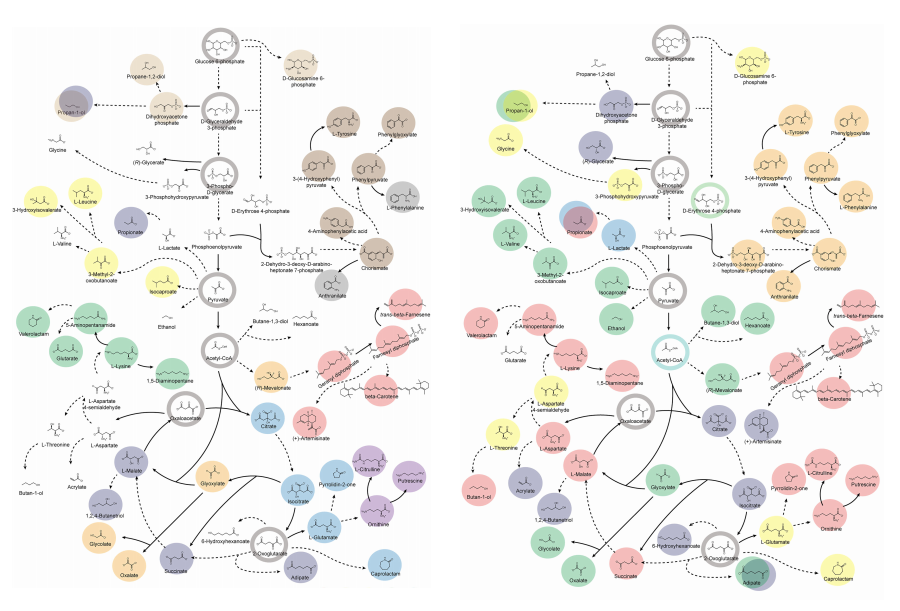 KAIST provides a comprehensive resource on microbial cell factories for sustainable chemical production
In silico analysis of five industrial microorganisms identifies optimal strains and metabolic engineering strategies for producing 235 valuable chemicals
Climate change and the depletion of fossil fuels have raised the global need for sustainable chemical production. In response to these environmental challenges, microbial cell factories are gaining attention as eco-friendly platforms for producing chemicals using renewable resources, while metabolic engineering technologies to enhance these cell factories are becoming crucial tools for maximizing production efficiency. However, difficulties in selecting suitable microbial strains and optimizing complex metabolic pathways continue to pose significant obstacles to practical industrial applications.
KAIST (President Kwang-Hyung Lee) announced on 27th of March that Distinguished Professor Sang Yup Lee’s research team in the Department of Chemical and Biomolecular Engineering comprehensively evaluated the production capabilities of various industrial microbial cell factories using in silico simulations and, based on these findings, identified the most suitable microbial strains for producing specific chemicals as well as optimal metabolic engineering strategies.
Previously, researchers attempted to determine the best strains and efficient metabolic engineering strategies among numerous microbial candidates through extensive biological experiments and meticulous verification processes. However, this approach required substantial time and costs. Recently, the introduction of genome-scale metabolic models (GEMs), which reconstruct the metabolic networks within an organism based on its entire genome information, has enabled systematic analysis of metabolic fluxes via computer simulations. This development offers a new way to overcome limitations of conventional experimental approaches, revolutionizing both strain selection and metabolic pathway design.
Accordingly, Professor Lee’s team at the Department of Chemical and Biomolecular Engineering, KAIST, evaluated the production capabilities of five representative industrial microorganisms—Escherichia coli, Saccharomyces cerevisiae, Bacillus subtilis, Corynebacterium glutamicum, and Pseudomonas putida—for 235 bio-based chemicals. Using GEMs, the researchers calculated both the maximum theoretical yields and the maximum achievable yields under industrial conditions for each chemical, thereby establishing criteria to identify the most suitable strains for each target compound.
< Figure 1. Outline of the strategy for improving microbial cell factories using a genome-scale metabolic model (GEM) >
The team specifically proposed strategies such as introducing heterologous enzyme reactions derived from other organisms and exchanging cofactors used by microbes to expand metabolic pathways. These strategies were shown to increase yields beyond the innate metabolic capacities of the microorganisms, resulting in higher production of industrially important chemicals such as mevalonic acid, propanol, fatty acids, and isoprenoids.
Moreover, by applying a computational approach to analyze metabolic fluxes in silico, the researchers suggested strategies for improving microbial strains to maximize the production of various chemicals. They quantitatively identified the relationships between specific enzyme reactions and target chemical production, as well as the relationships between enzymes and metabolites, determining which enzyme reactions should be up- or down-regulated. Through this, the team presented strategies not only to achieve high theoretical yields but also to maximize actual production capacities.
< Figure 2. Comparison of production routes and maximum yields of useful chemicals using representative industrial microorganisms >
Dr. Gi Bae Kim, the first author of this paper from the KAIST BioProcess Engineering Research Center, explained, “By introducing metabolic pathways derived from other organisms and exchanging cofactors, it is possible to design new microbial cell factories that surpass existing limitations. The strategies presented in this study will play a pivotal role in making microbial-based production processes more economical and efficient.” In addition, Distinguished Professor Sang Yup Lee noted, “This research serves as a key resource in the field of systems metabolic engineering, reducing difficulties in strain selection and pathway design, and enabling more efficient development of microbial cell factories. We expect it to greatly contribute to the future development of technologies for producing various eco-friendly chemicals, such as biofuels, bioplastics, and functional food materials.”
This research was conducted with the support from the Development of platform technologies of microbial cell factories for the next-generation biorefineries project and Development of advanced synthetic biology source technologies for leading the biomanufacturing industry project (Project Leader: Distinguished Professor Sang Yup Lee, KAIST) from National Research Foundation supported by the Korean Ministry of Science and ICT.
2025.03.27 View 9752
KAIST provides a comprehensive resource on microbial cell factories for sustainable chemical production
In silico analysis of five industrial microorganisms identifies optimal strains and metabolic engineering strategies for producing 235 valuable chemicals
Climate change and the depletion of fossil fuels have raised the global need for sustainable chemical production. In response to these environmental challenges, microbial cell factories are gaining attention as eco-friendly platforms for producing chemicals using renewable resources, while metabolic engineering technologies to enhance these cell factories are becoming crucial tools for maximizing production efficiency. However, difficulties in selecting suitable microbial strains and optimizing complex metabolic pathways continue to pose significant obstacles to practical industrial applications.
KAIST (President Kwang-Hyung Lee) announced on 27th of March that Distinguished Professor Sang Yup Lee’s research team in the Department of Chemical and Biomolecular Engineering comprehensively evaluated the production capabilities of various industrial microbial cell factories using in silico simulations and, based on these findings, identified the most suitable microbial strains for producing specific chemicals as well as optimal metabolic engineering strategies.
Previously, researchers attempted to determine the best strains and efficient metabolic engineering strategies among numerous microbial candidates through extensive biological experiments and meticulous verification processes. However, this approach required substantial time and costs. Recently, the introduction of genome-scale metabolic models (GEMs), which reconstruct the metabolic networks within an organism based on its entire genome information, has enabled systematic analysis of metabolic fluxes via computer simulations. This development offers a new way to overcome limitations of conventional experimental approaches, revolutionizing both strain selection and metabolic pathway design.
Accordingly, Professor Lee’s team at the Department of Chemical and Biomolecular Engineering, KAIST, evaluated the production capabilities of five representative industrial microorganisms—Escherichia coli, Saccharomyces cerevisiae, Bacillus subtilis, Corynebacterium glutamicum, and Pseudomonas putida—for 235 bio-based chemicals. Using GEMs, the researchers calculated both the maximum theoretical yields and the maximum achievable yields under industrial conditions for each chemical, thereby establishing criteria to identify the most suitable strains for each target compound.
< Figure 1. Outline of the strategy for improving microbial cell factories using a genome-scale metabolic model (GEM) >
The team specifically proposed strategies such as introducing heterologous enzyme reactions derived from other organisms and exchanging cofactors used by microbes to expand metabolic pathways. These strategies were shown to increase yields beyond the innate metabolic capacities of the microorganisms, resulting in higher production of industrially important chemicals such as mevalonic acid, propanol, fatty acids, and isoprenoids.
Moreover, by applying a computational approach to analyze metabolic fluxes in silico, the researchers suggested strategies for improving microbial strains to maximize the production of various chemicals. They quantitatively identified the relationships between specific enzyme reactions and target chemical production, as well as the relationships between enzymes and metabolites, determining which enzyme reactions should be up- or down-regulated. Through this, the team presented strategies not only to achieve high theoretical yields but also to maximize actual production capacities.
< Figure 2. Comparison of production routes and maximum yields of useful chemicals using representative industrial microorganisms >
Dr. Gi Bae Kim, the first author of this paper from the KAIST BioProcess Engineering Research Center, explained, “By introducing metabolic pathways derived from other organisms and exchanging cofactors, it is possible to design new microbial cell factories that surpass existing limitations. The strategies presented in this study will play a pivotal role in making microbial-based production processes more economical and efficient.” In addition, Distinguished Professor Sang Yup Lee noted, “This research serves as a key resource in the field of systems metabolic engineering, reducing difficulties in strain selection and pathway design, and enabling more efficient development of microbial cell factories. We expect it to greatly contribute to the future development of technologies for producing various eco-friendly chemicals, such as biofuels, bioplastics, and functional food materials.”
This research was conducted with the support from the Development of platform technologies of microbial cell factories for the next-generation biorefineries project and Development of advanced synthetic biology source technologies for leading the biomanufacturing industry project (Project Leader: Distinguished Professor Sang Yup Lee, KAIST) from National Research Foundation supported by the Korean Ministry of Science and ICT.
2025.03.27 View 9752 -
 KAIST Develops Eco-Friendly, Nylon-Like Plastic Using Microorganisms
Poly(ester amide) amide is a next-generation material that combines the advantages of PET (polyester) and nylon (polyamide), two widely used plastics. However, it could only be produced from fossil fuels, which posed environmental concerns. Using microorganisms, KAIST researchers have successfully developed a new bio-based plastic to replace conventional plastic.
KAIST (represented by President Kwang Hyung Lee) announced on the 20th of March that a research team led by Distinguished Professor Sang Yup Lee from the Department of Chemical and Biomolecular Engineering has developed microbial strains through systems metabolic engineering to produce various eco-friendly, bio-based poly(ester amide)s. The team collaborated with researchers from the Korea Research Institute of Chemical Technology (KRICT, President Young-Kook Lee) to analyze and confirm the properties of the resulting plastic.
Professor Sang Yup Lee’s research team designed new metabolic pathways that do not naturally exist in microorganisms, and developed a platform microbial strain capable of producing nine different types of poly(ester amide)s, including poly(3-hydroxybutyrate-ran-3-aminopropionate) and poly(3-hydroxybutyrate-ran-4-aminobutyrate).
Using glucose derived from abundant biomass sources such as waste wood and weeds, the team successfully produced poly(ester amide)s in an eco-friendly manner. The researchers also confirmed the potential for industrial-scale production by demonstrating high production efficiency (54.57 g/L) using fed-batch fermentation of the engineered strain.
In collaboration with researchers Haemin Jeong and Jihoon Shin from KRICT, the KAIST team analyzed the properties of the bio-based plastic and found that it exhibited characteristics similar to high-density polyethylene (HDPE). This means the new plastic is not only eco-friendly but also strong and durable enough to replace conventional plastics.
The engineered strains and strategies developed in this study are expected to be useful not only for producing various poly(ester amide)s but also for constructing metabolic pathways for the biosynthesis of other types of polymers.
Professor Sang Yup Lee stated, “This study is the first to demonstrate the possibility of producing poly(ester amide)s (plastics) through a renewable bio-based chemical process rather than relying on the petroleum-based chemical industry. We plan to further enhance the production yield and efficiency through continued research.”
The study was published online on March 17 in the international journal Nature Chemical Biology.
·Title: Biosynthesis of poly(ester amide)s in engineered Escherichia coli
·DOI: 10.1038/s41589-025-01842-2
·Authors: A total of seven authors including Tong Un Chae (KAIST, first author), So Young Choi (KAIST, second author), Da-Hee Ahn (KAIST, third author), Woo Dae Jang (KAIST, fourth author), Haemin Jeong (KRICT, fifth author), Jihoon Shin (KRICT, sixth author), and Sang Yup Lee (KAIST, corresponding author).
This research was supported by the Ministry of Science and ICT (MSIT) under the Eco-Friendly Chemical Technology Development Project as part of the "Next-Generation Biorefinery Technology Development to Lead the Bio-Chemical Industry" initiative (project led by Distinguished Professor Sang Yup Lee at KAIST).
2025.03.24 View 12584
KAIST Develops Eco-Friendly, Nylon-Like Plastic Using Microorganisms
Poly(ester amide) amide is a next-generation material that combines the advantages of PET (polyester) and nylon (polyamide), two widely used plastics. However, it could only be produced from fossil fuels, which posed environmental concerns. Using microorganisms, KAIST researchers have successfully developed a new bio-based plastic to replace conventional plastic.
KAIST (represented by President Kwang Hyung Lee) announced on the 20th of March that a research team led by Distinguished Professor Sang Yup Lee from the Department of Chemical and Biomolecular Engineering has developed microbial strains through systems metabolic engineering to produce various eco-friendly, bio-based poly(ester amide)s. The team collaborated with researchers from the Korea Research Institute of Chemical Technology (KRICT, President Young-Kook Lee) to analyze and confirm the properties of the resulting plastic.
Professor Sang Yup Lee’s research team designed new metabolic pathways that do not naturally exist in microorganisms, and developed a platform microbial strain capable of producing nine different types of poly(ester amide)s, including poly(3-hydroxybutyrate-ran-3-aminopropionate) and poly(3-hydroxybutyrate-ran-4-aminobutyrate).
Using glucose derived from abundant biomass sources such as waste wood and weeds, the team successfully produced poly(ester amide)s in an eco-friendly manner. The researchers also confirmed the potential for industrial-scale production by demonstrating high production efficiency (54.57 g/L) using fed-batch fermentation of the engineered strain.
In collaboration with researchers Haemin Jeong and Jihoon Shin from KRICT, the KAIST team analyzed the properties of the bio-based plastic and found that it exhibited characteristics similar to high-density polyethylene (HDPE). This means the new plastic is not only eco-friendly but also strong and durable enough to replace conventional plastics.
The engineered strains and strategies developed in this study are expected to be useful not only for producing various poly(ester amide)s but also for constructing metabolic pathways for the biosynthesis of other types of polymers.
Professor Sang Yup Lee stated, “This study is the first to demonstrate the possibility of producing poly(ester amide)s (plastics) through a renewable bio-based chemical process rather than relying on the petroleum-based chemical industry. We plan to further enhance the production yield and efficiency through continued research.”
The study was published online on March 17 in the international journal Nature Chemical Biology.
·Title: Biosynthesis of poly(ester amide)s in engineered Escherichia coli
·DOI: 10.1038/s41589-025-01842-2
·Authors: A total of seven authors including Tong Un Chae (KAIST, first author), So Young Choi (KAIST, second author), Da-Hee Ahn (KAIST, third author), Woo Dae Jang (KAIST, fourth author), Haemin Jeong (KRICT, fifth author), Jihoon Shin (KRICT, sixth author), and Sang Yup Lee (KAIST, corresponding author).
This research was supported by the Ministry of Science and ICT (MSIT) under the Eco-Friendly Chemical Technology Development Project as part of the "Next-Generation Biorefinery Technology Development to Lead the Bio-Chemical Industry" initiative (project led by Distinguished Professor Sang Yup Lee at KAIST).
2025.03.24 View 12584 -
 KAIST Discovers Molecular Switch that Reverses Cancerous Transformation at the Critical Moment of Transition
< (From left) PhD student Seoyoon D. Jeong, (bottom) Professor Kwang-Hyun Cho, (top) Dr. Dongkwan Shin, Dr. Jeong-Ryeol Gong >
Professor Kwang-Hyun Cho’s research team has recently been highlighted for their work on developing an original technology for cancer reversal treatment that does not kill cancer cells but only changes their characteristics to reverse them to a state similar to normal cells. This time, they have succeeded in revealing for the first time that a molecular switch that can induce cancer reversal at the moment when normal cells change into cancer cells is hidden in the genetic network.
KAIST (President Kwang-Hyung Lee) announced on the 5th of February that Professor Kwang-Hyun Cho's research team of the Department of Bio and Brain Engineering has succeeded in developing a fundamental technology to capture the critical transition phenomenon at the moment when normal cells change into cancer cells and analyze it to discover a molecular switch that can revert cancer cells back into normal cells.
A critical transition is a phenomenon in which a sudden change in state occurs at a specific point in time, like water changing into steam at 100℃. This critical transition phenomenon also occurs in the process in which normal cells change into cancer cells at a specific point in time due to the accumulation of genetic and epigenetic changes.
The research team discovered that normal cells can enter an unstable critical transition state where normal cells and cancer cells coexist just before they change into cancer cells during tumorigenesis, the production or development of tumors, and analyzed this critical transition state using a systems biology method to develop a cancer reversal molecular switch identification technology that can reverse the cancerization process. They then applied this to colon cancer cells and confirmed through molecular cell experiments that cancer cells can recover the characteristics of normal cells.
This is an original technology that automatically infers a computer model of the genetic network that controls the critical transition of cancer development from single-cell RNA sequencing data, and systematically finds molecular switches for cancer reversion by simulation analysis. It is expected that this technology will be applied to the development of reversion therapies for other cancers in the future.
Professor Kwang-Hyun Cho said, "We have discovered a molecular switch that can revert the fate of cancer cells back to a normal state by capturing the moment of critical transition right before normal cells are changed into an irreversible cancerous state."
< Figure 1. Overall conceptual framework of the technology that automatically constructs a molecular regulatory network from single-cell RNA sequencing data of colon cancer cells to discover molecular switches for cancer reversion through computer simulation analysis. Professor Kwang-Hyun Cho's research team established a fundamental technology for automatic construction of a computer model of a core gene network by analyzing the entire process of tumorigenesis of colon cells turning into cancer cells, and developed an original technology for discovering the molecular switches that can induce cancer cell reversal through attractor landscape analysis. >
He continued, "In particular, this study has revealed in detail, at the genetic network level, what changes occur within cells behind the process of cancer development, which has been considered a mystery until now." He emphasized, "This is the first study to reveal that an important clue that can revert the fate of tumorigenesis is hidden at this very critical moment of change."
< Figure 2. Identification of tumor transition state using single-cell RNA sequencing data from colorectal cancer. Using single-cell RNA sequencing data from colorectal cancer patient-derived organoids for normal and cancerous tissues, a critical transition was identified in which normal and cancerous cells coexist and instability increases (a-d). The critical transition was confirmed to show intermediate levels of major phenotypic features related to cancer or normal tissues that are indicative of the states between the normal and cancerous cells (e). >
The results of this study, conducted by KAIST Dr. Dongkwan Shin (currently at the National Cancer Center), Dr. Jeong-Ryeol Gong, and doctoral student Seoyoon D. Jeong jointly with a research team at Seoul National University that provided the organoids (in vitro cultured tissues) from colon cancer patient, were published as an online paper in the international journal ‘Advanced Science’ published by Wiley on January 22nd.
(Paper title: Attractor landscape analysis reveals a reversion switch in the transition of colorectal tumorigenesis) (DOI: https://doi.org/10.1002/advs.202412503)
< Figure 3. Reconstruction of a dynamic network model for the transition state of colorectal cancer.
A new technology was established to build a gene network computer model that can simulate the dynamic changes between genes by integrating single-cell RNA sequencing data and existing experimental results on gene-to-gene interactions in the critical transition of cancer. (a). Using this technology, a gene network computer model for the critical transition of colorectal cancer was constructed, and the distribution of attractors representing normal and cancer cell phenotypes was investigated through attractor landscape analysis (b-e). >
This study was conducted with the support of the National Research Foundation of Korea under the Ministry of Science and ICT through the Mid-Career Researcher Program and Basic Research Laboratory Program and the Disease-Centered Translational Research Project of the Korea Health Industry Development Institute (KHIDI) of the Ministry of Health and Welfare.
< Figure 4. Quantification of attractor landscapes and discovery of transcription factors for cancer reversibility through perturbation simulation analysis. A methodology for implementing discontinuous attractor landscapes continuously from a computer model of gene networks and quantifying them as cancer scores was introduced (a), and attractor landscapes for the critical transition of colorectal cancer were secured (b-d). By tracking the change patterns of normal and cancer cell attractors through perturbation simulation analysis for each gene, the optimal combination of transcription factors for cancer reversion was discovered (e-h). This was confirmed in various parameter combinations as well (i). >
< Figure 5. Identification and experimental validation of the optimal target gene for cancer reversion. Among the common target genes of the discovered transcription factor combinations, we identified cancer reversing molecular switches that are predicted to suppress cancer cell proliferation and restore the characteristics of normal colon cells (a-d). When inhibitors for the molecular switches were treated to organoids derived from colon cancer patients, it was confirmed that cancer cell proliferation was suppressed and the expression of key genes related to cancer development was inhibited (e-h), and a group of genes related to normal colon epithelium was activated and transformed into a state similar to normal colon cells (i-j). >
< Figure 6. Schematic diagram of the research results. Professor Kwang-Hyun Cho's research team developed an original technology to systematically discover key molecular switches that can induce reversion of colon cancer cells through a systems biology approach using an attractor landscape analysis of a genetic network model for the critical transition at the moment of transformation from normal cells to cancer cells, and verified the reversing effect of actual colon cancer through cellular experiments. >
2025.02.05 View 43386
KAIST Discovers Molecular Switch that Reverses Cancerous Transformation at the Critical Moment of Transition
< (From left) PhD student Seoyoon D. Jeong, (bottom) Professor Kwang-Hyun Cho, (top) Dr. Dongkwan Shin, Dr. Jeong-Ryeol Gong >
Professor Kwang-Hyun Cho’s research team has recently been highlighted for their work on developing an original technology for cancer reversal treatment that does not kill cancer cells but only changes their characteristics to reverse them to a state similar to normal cells. This time, they have succeeded in revealing for the first time that a molecular switch that can induce cancer reversal at the moment when normal cells change into cancer cells is hidden in the genetic network.
KAIST (President Kwang-Hyung Lee) announced on the 5th of February that Professor Kwang-Hyun Cho's research team of the Department of Bio and Brain Engineering has succeeded in developing a fundamental technology to capture the critical transition phenomenon at the moment when normal cells change into cancer cells and analyze it to discover a molecular switch that can revert cancer cells back into normal cells.
A critical transition is a phenomenon in which a sudden change in state occurs at a specific point in time, like water changing into steam at 100℃. This critical transition phenomenon also occurs in the process in which normal cells change into cancer cells at a specific point in time due to the accumulation of genetic and epigenetic changes.
The research team discovered that normal cells can enter an unstable critical transition state where normal cells and cancer cells coexist just before they change into cancer cells during tumorigenesis, the production or development of tumors, and analyzed this critical transition state using a systems biology method to develop a cancer reversal molecular switch identification technology that can reverse the cancerization process. They then applied this to colon cancer cells and confirmed through molecular cell experiments that cancer cells can recover the characteristics of normal cells.
This is an original technology that automatically infers a computer model of the genetic network that controls the critical transition of cancer development from single-cell RNA sequencing data, and systematically finds molecular switches for cancer reversion by simulation analysis. It is expected that this technology will be applied to the development of reversion therapies for other cancers in the future.
Professor Kwang-Hyun Cho said, "We have discovered a molecular switch that can revert the fate of cancer cells back to a normal state by capturing the moment of critical transition right before normal cells are changed into an irreversible cancerous state."
< Figure 1. Overall conceptual framework of the technology that automatically constructs a molecular regulatory network from single-cell RNA sequencing data of colon cancer cells to discover molecular switches for cancer reversion through computer simulation analysis. Professor Kwang-Hyun Cho's research team established a fundamental technology for automatic construction of a computer model of a core gene network by analyzing the entire process of tumorigenesis of colon cells turning into cancer cells, and developed an original technology for discovering the molecular switches that can induce cancer cell reversal through attractor landscape analysis. >
He continued, "In particular, this study has revealed in detail, at the genetic network level, what changes occur within cells behind the process of cancer development, which has been considered a mystery until now." He emphasized, "This is the first study to reveal that an important clue that can revert the fate of tumorigenesis is hidden at this very critical moment of change."
< Figure 2. Identification of tumor transition state using single-cell RNA sequencing data from colorectal cancer. Using single-cell RNA sequencing data from colorectal cancer patient-derived organoids for normal and cancerous tissues, a critical transition was identified in which normal and cancerous cells coexist and instability increases (a-d). The critical transition was confirmed to show intermediate levels of major phenotypic features related to cancer or normal tissues that are indicative of the states between the normal and cancerous cells (e). >
The results of this study, conducted by KAIST Dr. Dongkwan Shin (currently at the National Cancer Center), Dr. Jeong-Ryeol Gong, and doctoral student Seoyoon D. Jeong jointly with a research team at Seoul National University that provided the organoids (in vitro cultured tissues) from colon cancer patient, were published as an online paper in the international journal ‘Advanced Science’ published by Wiley on January 22nd.
(Paper title: Attractor landscape analysis reveals a reversion switch in the transition of colorectal tumorigenesis) (DOI: https://doi.org/10.1002/advs.202412503)
< Figure 3. Reconstruction of a dynamic network model for the transition state of colorectal cancer.
A new technology was established to build a gene network computer model that can simulate the dynamic changes between genes by integrating single-cell RNA sequencing data and existing experimental results on gene-to-gene interactions in the critical transition of cancer. (a). Using this technology, a gene network computer model for the critical transition of colorectal cancer was constructed, and the distribution of attractors representing normal and cancer cell phenotypes was investigated through attractor landscape analysis (b-e). >
This study was conducted with the support of the National Research Foundation of Korea under the Ministry of Science and ICT through the Mid-Career Researcher Program and Basic Research Laboratory Program and the Disease-Centered Translational Research Project of the Korea Health Industry Development Institute (KHIDI) of the Ministry of Health and Welfare.
< Figure 4. Quantification of attractor landscapes and discovery of transcription factors for cancer reversibility through perturbation simulation analysis. A methodology for implementing discontinuous attractor landscapes continuously from a computer model of gene networks and quantifying them as cancer scores was introduced (a), and attractor landscapes for the critical transition of colorectal cancer were secured (b-d). By tracking the change patterns of normal and cancer cell attractors through perturbation simulation analysis for each gene, the optimal combination of transcription factors for cancer reversion was discovered (e-h). This was confirmed in various parameter combinations as well (i). >
< Figure 5. Identification and experimental validation of the optimal target gene for cancer reversion. Among the common target genes of the discovered transcription factor combinations, we identified cancer reversing molecular switches that are predicted to suppress cancer cell proliferation and restore the characteristics of normal colon cells (a-d). When inhibitors for the molecular switches were treated to organoids derived from colon cancer patients, it was confirmed that cancer cell proliferation was suppressed and the expression of key genes related to cancer development was inhibited (e-h), and a group of genes related to normal colon epithelium was activated and transformed into a state similar to normal colon cells (i-j). >
< Figure 6. Schematic diagram of the research results. Professor Kwang-Hyun Cho's research team developed an original technology to systematically discover key molecular switches that can induce reversion of colon cancer cells through a systems biology approach using an attractor landscape analysis of a genetic network model for the critical transition at the moment of transformation from normal cells to cancer cells, and verified the reversing effect of actual colon cancer through cellular experiments. >
2025.02.05 View 43386 -
 KAIST Uncovers the Principles of Gene Expression Regulation in Cancer and Cellular Functions
< (From left) Professor Seyun Kim, Professor Gwangrog Lee, Dr. Hyoungjoon Ahn, Dr. Jeongmin Yu, Professor Won-Ki Cho, and (below) PhD candidate Kwangmin Ryu of the Department of Biological Sciences>
A research team at KAIST has identified the core gene expression networks regulated by key proteins that fundamentally drive phenomena such as cancer development, metastasis, tissue differentiation from stem cells, and neural activation processes. This discovery lays the foundation for developing innovative therapeutic technologies.
On the 22nd of January, KAIST (represented by President Kwang Hyung Lee) announced that the joint research team led by Professors Seyun Kim, Gwangrog Lee, and Won-Ki Cho from the Department of Biological Sciences had uncovered essential mechanisms controlling gene expression in animal cells.
Inositol phosphate metabolites produced by inositol metabolism enzymes serve as vital secondary messengers in eukaryotic cell signaling systems and are broadly implicated in cancer, obesity, diabetes, and neurological disorders.
The research team demonstrated that the inositol polyphosphate multikinase (IPMK) enzyme, a key player in the inositol metabolism system, acts as a critical transcriptional activator within the core gene expression networks of animal cells. Notably, although IPMK was previously reported to play an important role in the transcription process governed by serum response factor (SRF), a representative transcription factor in animal cells, the precise mechanism of its action was unclear.
SRF is a transcription factor directly controlling the expression of at least 200–300 genes, regulating cell growth, proliferation, apoptosis, and motility, and is indispensable for organ development, such as in the heart.
The team discovered that IPMK binds directly to SRF, altering the three-dimensional structure of the SRF protein. This interaction facilitates the transcriptional activity of various genes through the SRF activated by IPMK, demonstrating that IPMK acts as a critical regulatory switch to enhance SRF's protein activity.
< Figure 1. The serum response factor (SRF) protein, a key transcription factor in animal cells, directly binds to inositol polyphosphate multikinase (IPMK) enzyme and undergoes structural change to acquire DNA binding ability, and precisely regulates growth and differentiation of animal cells through transcriptional activation. >
The team further verified that disruptions in the direct interaction between IPMK and SRF lead to the reduced functionality and activity of SRF, causing severe impairments in gene expression.
By highlighting the significance of the intrinsically disordered region (IDR) in SRF, the researchers underscored the biological importance of intrinsically disordered proteins (IDPs). Unlike most proteins that adopt distinct structures through folding, IDPs, including those with IDRs, do not exhibit specific structures but play crucial biological roles, attracting significant attention in the scientific community.
Professor Seyun Kim commented, "This study provides a vital mechanism proving that IPMK, a key enzyme in the inositol metabolism system, is a major transcriptional activator in the core gene expression network of animal cells. By understanding fundamental processes such as cancer development and metastasis, tissue differentiation from stem cells, and neural activation through SRF, we hope this discovery will lead to the broad application of innovative therapeutic technologies."
The findings were published on January 7th in the international journal Nucleic Acids Research (IF=16.7, top 1.8% in Biochemistry and Molecular Biology), under the title “Single-molecule analysis reveals that IPMK enhances the DNA-binding activity of the transcription factor SRF" (DOI: 10.1093/nar/gkae1281).
This research was supported by the National Research Foundation of Korea's Mid-career Research Program, Leading Research Center Program, and Global Research Laboratory Program, as well as by the Suh Kyungbae Science Foundation and the Samsung Future Technology Development Program.
2025.01.24 View 18519
KAIST Uncovers the Principles of Gene Expression Regulation in Cancer and Cellular Functions
< (From left) Professor Seyun Kim, Professor Gwangrog Lee, Dr. Hyoungjoon Ahn, Dr. Jeongmin Yu, Professor Won-Ki Cho, and (below) PhD candidate Kwangmin Ryu of the Department of Biological Sciences>
A research team at KAIST has identified the core gene expression networks regulated by key proteins that fundamentally drive phenomena such as cancer development, metastasis, tissue differentiation from stem cells, and neural activation processes. This discovery lays the foundation for developing innovative therapeutic technologies.
On the 22nd of January, KAIST (represented by President Kwang Hyung Lee) announced that the joint research team led by Professors Seyun Kim, Gwangrog Lee, and Won-Ki Cho from the Department of Biological Sciences had uncovered essential mechanisms controlling gene expression in animal cells.
Inositol phosphate metabolites produced by inositol metabolism enzymes serve as vital secondary messengers in eukaryotic cell signaling systems and are broadly implicated in cancer, obesity, diabetes, and neurological disorders.
The research team demonstrated that the inositol polyphosphate multikinase (IPMK) enzyme, a key player in the inositol metabolism system, acts as a critical transcriptional activator within the core gene expression networks of animal cells. Notably, although IPMK was previously reported to play an important role in the transcription process governed by serum response factor (SRF), a representative transcription factor in animal cells, the precise mechanism of its action was unclear.
SRF is a transcription factor directly controlling the expression of at least 200–300 genes, regulating cell growth, proliferation, apoptosis, and motility, and is indispensable for organ development, such as in the heart.
The team discovered that IPMK binds directly to SRF, altering the three-dimensional structure of the SRF protein. This interaction facilitates the transcriptional activity of various genes through the SRF activated by IPMK, demonstrating that IPMK acts as a critical regulatory switch to enhance SRF's protein activity.
< Figure 1. The serum response factor (SRF) protein, a key transcription factor in animal cells, directly binds to inositol polyphosphate multikinase (IPMK) enzyme and undergoes structural change to acquire DNA binding ability, and precisely regulates growth and differentiation of animal cells through transcriptional activation. >
The team further verified that disruptions in the direct interaction between IPMK and SRF lead to the reduced functionality and activity of SRF, causing severe impairments in gene expression.
By highlighting the significance of the intrinsically disordered region (IDR) in SRF, the researchers underscored the biological importance of intrinsically disordered proteins (IDPs). Unlike most proteins that adopt distinct structures through folding, IDPs, including those with IDRs, do not exhibit specific structures but play crucial biological roles, attracting significant attention in the scientific community.
Professor Seyun Kim commented, "This study provides a vital mechanism proving that IPMK, a key enzyme in the inositol metabolism system, is a major transcriptional activator in the core gene expression network of animal cells. By understanding fundamental processes such as cancer development and metastasis, tissue differentiation from stem cells, and neural activation through SRF, we hope this discovery will lead to the broad application of innovative therapeutic technologies."
The findings were published on January 7th in the international journal Nucleic Acids Research (IF=16.7, top 1.8% in Biochemistry and Molecular Biology), under the title “Single-molecule analysis reveals that IPMK enhances the DNA-binding activity of the transcription factor SRF" (DOI: 10.1093/nar/gkae1281).
This research was supported by the National Research Foundation of Korea's Mid-career Research Program, Leading Research Center Program, and Global Research Laboratory Program, as well as by the Suh Kyungbae Science Foundation and the Samsung Future Technology Development Program.
2025.01.24 View 18519 -
 KAIST Develops Neuromorphic Semiconductor Chip that Learns and Corrects Itself
< Photo. The research team of the School of Electrical Engineering posed by the newly deveoped processor. (From center to the right) Professor Young-Gyu Yoon, Integrated Master's and Doctoral Program Students Seungjae Han and Hakcheon Jeong and Professor Shinhyun Choi >
- Professor Shinhyun Choi and Professor Young-Gyu Yoon’s Joint Research Team from the School of Electrical Engineering developed a computing chip that can learn, correct errors, and process AI tasks
- Equipping a computing chip with high-reliability memristor devices with self-error correction functions for real-time learning and image processing
Existing computer systems have separate data processing and storage devices, making them inefficient for processing complex data like AI. A KAIST research team has developed a memristor-based integrated system similar to the way our brain processes information. It is now ready for application in various devices including smart security cameras, allowing them to recognize suspicious activity immediately without having to rely on remote cloud servers, and medical devices with which it can help analyze health data in real time.
KAIST (President Kwang Hyung Lee) announced on the 17th of January that the joint research team of Professor Shinhyun Choi and Professor Young-Gyu Yoon of the School of Electrical Engineering has developed a next-generation neuromorphic semiconductor-based ultra-small computing chip that can learn and correct errors on its own.
< Figure 1. Scanning electron microscope (SEM) image of a computing chip equipped with a highly reliable selector-less 32×32 memristor crossbar array (left). Hardware system developed for real-time artificial intelligence implementation (right). >
What is special about this computing chip is that it can learn and correct errors that occur due to non-ideal characteristics that were difficult to solve in existing neuromorphic devices. For example, when processing a video stream, the chip learns to automatically separate a moving object from the background, and it becomes better at this task over time.
This self-learning ability has been proven by achieving accuracy comparable to ideal computer simulations in real-time image processing. The research team's main achievement is that it has completed a system that is both reliable and practical, beyond the development of brain-like components.
The research team has developed the world's first memristor-based integrated system that can adapt to immediate environmental changes, and has presented an innovative solution that overcomes the limitations of existing technology.
< Figure 2. Background and foreground separation results of an image containing non-ideal characteristics of memristor devices (left). Real-time image separation results through on-device learning using the memristor computing chip developed by our research team (right). >
At the heart of this innovation is a next-generation semiconductor device called a memristor*. The variable resistance characteristics of this device can replace the role of synapses in neural networks, and by utilizing it, data storage and computation can be performed simultaneously, just like our brain cells.
*Memristor: A compound word of memory and resistor, next-generation electrical device whose resistance value is determined by the amount and direction of charge that has flowed between the two terminals in the past.
The research team designed a highly reliable memristor that can precisely control resistance changes and developed an efficient system that excludes complex compensation processes through self-learning. This study is significant in that it experimentally verified the commercialization possibility of a next-generation neuromorphic semiconductor-based integrated system that supports real-time learning and inference.
This technology will revolutionize the way artificial intelligence is used in everyday devices, allowing AI tasks to be processed locally without relying on remote cloud servers, making them faster, more privacy-protected, and more energy-efficient.
“This system is like a smart workspace where everything is within arm’s reach instead of having to go back and forth between desks and file cabinets,” explained KAIST researchers Hakcheon Jeong and Seungjae Han, who led the development of this technology. “This is similar to the way our brain processes information, where everything is processed efficiently at once at one spot.”
The research was conducted with Hakcheon Jeong and Seungjae Han, the students of Integrated Master's and Doctoral Program at KAIST School of Electrical Engineering being the co-first authors, the results of which was published online in the international academic journal, Nature Electronics, on January 8, 2025.
*Paper title: Self-supervised video processing with self-calibration on an analogue computing platform based on a selector-less memristor array ( https://doi.org/10.1038/s41928-024-01318-6 )
This research was supported by the Next-Generation Intelligent Semiconductor Technology Development Project, Excellent New Researcher Project and PIM AI Semiconductor Core Technology Development Project of the National Research Foundation of Korea, and the Electronics and Telecommunications Research Institute Research and Development Support Project of the Institute of Information & communications Technology Planning & Evaluation.
2025.01.17 View 16860
KAIST Develops Neuromorphic Semiconductor Chip that Learns and Corrects Itself
< Photo. The research team of the School of Electrical Engineering posed by the newly deveoped processor. (From center to the right) Professor Young-Gyu Yoon, Integrated Master's and Doctoral Program Students Seungjae Han and Hakcheon Jeong and Professor Shinhyun Choi >
- Professor Shinhyun Choi and Professor Young-Gyu Yoon’s Joint Research Team from the School of Electrical Engineering developed a computing chip that can learn, correct errors, and process AI tasks
- Equipping a computing chip with high-reliability memristor devices with self-error correction functions for real-time learning and image processing
Existing computer systems have separate data processing and storage devices, making them inefficient for processing complex data like AI. A KAIST research team has developed a memristor-based integrated system similar to the way our brain processes information. It is now ready for application in various devices including smart security cameras, allowing them to recognize suspicious activity immediately without having to rely on remote cloud servers, and medical devices with which it can help analyze health data in real time.
KAIST (President Kwang Hyung Lee) announced on the 17th of January that the joint research team of Professor Shinhyun Choi and Professor Young-Gyu Yoon of the School of Electrical Engineering has developed a next-generation neuromorphic semiconductor-based ultra-small computing chip that can learn and correct errors on its own.
< Figure 1. Scanning electron microscope (SEM) image of a computing chip equipped with a highly reliable selector-less 32×32 memristor crossbar array (left). Hardware system developed for real-time artificial intelligence implementation (right). >
What is special about this computing chip is that it can learn and correct errors that occur due to non-ideal characteristics that were difficult to solve in existing neuromorphic devices. For example, when processing a video stream, the chip learns to automatically separate a moving object from the background, and it becomes better at this task over time.
This self-learning ability has been proven by achieving accuracy comparable to ideal computer simulations in real-time image processing. The research team's main achievement is that it has completed a system that is both reliable and practical, beyond the development of brain-like components.
The research team has developed the world's first memristor-based integrated system that can adapt to immediate environmental changes, and has presented an innovative solution that overcomes the limitations of existing technology.
< Figure 2. Background and foreground separation results of an image containing non-ideal characteristics of memristor devices (left). Real-time image separation results through on-device learning using the memristor computing chip developed by our research team (right). >
At the heart of this innovation is a next-generation semiconductor device called a memristor*. The variable resistance characteristics of this device can replace the role of synapses in neural networks, and by utilizing it, data storage and computation can be performed simultaneously, just like our brain cells.
*Memristor: A compound word of memory and resistor, next-generation electrical device whose resistance value is determined by the amount and direction of charge that has flowed between the two terminals in the past.
The research team designed a highly reliable memristor that can precisely control resistance changes and developed an efficient system that excludes complex compensation processes through self-learning. This study is significant in that it experimentally verified the commercialization possibility of a next-generation neuromorphic semiconductor-based integrated system that supports real-time learning and inference.
This technology will revolutionize the way artificial intelligence is used in everyday devices, allowing AI tasks to be processed locally without relying on remote cloud servers, making them faster, more privacy-protected, and more energy-efficient.
“This system is like a smart workspace where everything is within arm’s reach instead of having to go back and forth between desks and file cabinets,” explained KAIST researchers Hakcheon Jeong and Seungjae Han, who led the development of this technology. “This is similar to the way our brain processes information, where everything is processed efficiently at once at one spot.”
The research was conducted with Hakcheon Jeong and Seungjae Han, the students of Integrated Master's and Doctoral Program at KAIST School of Electrical Engineering being the co-first authors, the results of which was published online in the international academic journal, Nature Electronics, on January 8, 2025.
*Paper title: Self-supervised video processing with self-calibration on an analogue computing platform based on a selector-less memristor array ( https://doi.org/10.1038/s41928-024-01318-6 )
This research was supported by the Next-Generation Intelligent Semiconductor Technology Development Project, Excellent New Researcher Project and PIM AI Semiconductor Core Technology Development Project of the National Research Foundation of Korea, and the Electronics and Telecommunications Research Institute Research and Development Support Project of the Institute of Information & communications Technology Planning & Evaluation.
2025.01.17 View 16860