TOM
-
 Military Combatants Usher in an Era of Personalized Training with New Materials
< Photo 1. (From left) Professor Steve Park of Materials Science and Engineering, Kyusoon Pak, Ph.D. Candidate (Army Major) >
Traditional military training often relies on standardized methods, which has limited the provision of optimized training tailored to individual combatants' characteristics or specific combat situations. To address this, our research team developed an e-textile platform, securing core technology that can reflect the unique traits of individual combatants and various combat scenarios. This technology has proven robust enough for battlefield use and is economical enough for widespread distribution to a large number of troops.
On June 25th, Professor Steve Park's research team at KAIST's Department of Materials Science and Engineering announced the development of a flexible, wearable electronic textile (E-textile) platform using an innovative technology that 'draws' electronic circuits directly onto fabric.
The wearable e-textile platform developed by the research team combines 3D printing technology with new materials engineering design to directly print flexible and highly durable sensors and electrodes onto textile substrates. This enables the collection of precise movement and human body data from individual combatants, which can then be used to propose customized training models.
Existing e-textile fabrication methods were often complex or limited in their ability to provide personalized customization. To overcome these challenges, the research team adopted an additive manufacturing technology called 'Direct Ink Writing (DIW)' 3D printing.
< Figure 1. Schematic diagram of e-textile manufactured with Direct Ink Writing (DIW) printing technology on various textiles, including combat uniforms >
This technology involves directly dispensing and printing special ink, which functions as sensors and electrodes, onto textile substrates in desired patterns. This allows for flexible implementation of various designs without the complex process of mask fabrication. This is expected to be an effective technology that can be easily supplied to hundreds of thousands of military personnel.
The core of this technology lies in the development of high-performance functional inks based on advanced materials engineering design. The research team combined styrene-butadiene-styrene (SBS) polymer, which provides flexibility, with multi-walled carbon nanotubes (MWCNT) for electrical conductivity. They developed a tensile/bending sensor ink that can stretch up to 102% and maintain stable performance even after 10,000 repetitive tests. This means that accurate data can be consistently obtained even during the strenuous movements of combatants.
< Figure 2. Measurement of human movement and breathing patterns using e-textile >
Furthermore, new material technology was applied to implement 'interconnect electrodes' that electrically connect the upper and lower layers of the fabric. The team developed an electrode ink combining silver (Ag) flakes with rigid polystyrene (PS) polymer, precisely controlling the impregnation level (how much the ink penetrates the fabric) to effectively connect both sides or multiple layers of the fabric. This secures the technology for producing multi-layered wearable electronic systems integrating sensors and electrodes.
< Figure 3. Experimental results of recognizing unknown objects after machine learning six objects using a smart glove >
The research team proved the platform's performance through actual human movement monitoring experiments. They printed the developed e-textile on major joint areas of clothing (shoulders, elbows, knees) and measured movements and posture changes during various exercises such as running, jumping jacks, and push-ups in real-time.
Additionally, they demonstrated the potential for applications such as monitoring breathing patterns using a smart mask and recognizing objects through machine learning and perceiving complex tactile information by printing multiple sensors and electrodes on gloves. These results show that the developed e-textile platform is effective in precisely understanding the movement dynamics of combatants.
This research is an important example demonstrating how cutting-edge new material technology can contribute to the advancement of the defense sector. Major Kyusoon Pak of the Army, who participated in this research, considered required objectives such as military applicability and economic feasibility for practical distribution from the research design stage.
< Figure 4. Experimental results showing that a multi-layered e-textile glove connected with interconnect electrodes can measure tensile/bending signals and pressure signals at a single point >
Major Pak stated, "Our military is currently facing both a crisis and an opportunity due to the decrease in military personnel resources caused by the demographic cliff and the advancement of science and technology. Also, respect for life in the battlefield is emerging as a significant issue. This research aims to secure original technology that can provide customized training according to military branch/duty and type of combat, thereby enhancing the combat power and ensuring the survivability of our soldiers."
He added, "I hope this research will be evaluated as a case that achieved both scientific contribution and military applicability."
This research, where Kyusoon Pak, Ph.D. Candidate (Army Major) from KAIST's Department of Materials Science and Engineering, participated as the first author and Professor Steve Park supervised, was published on May 27, 2025, in `npj Flexible Electronics (top 1.8% in JCR field)', an international academic journal in the electrical, electronic, and materials engineering fields.
* Paper Title: Fabrication of Multifunctional Wearable Interconnect E-textile Platform Using Direct Ink Writing (DIW) 3D Printing
* DOI: https://doi.org/10.1038/s41528-025-00414-7
This research was supported by the Ministry of Trade, Industry and Energy and the National Research Foundation of Korea.
2025.06.25 View 208
Military Combatants Usher in an Era of Personalized Training with New Materials
< Photo 1. (From left) Professor Steve Park of Materials Science and Engineering, Kyusoon Pak, Ph.D. Candidate (Army Major) >
Traditional military training often relies on standardized methods, which has limited the provision of optimized training tailored to individual combatants' characteristics or specific combat situations. To address this, our research team developed an e-textile platform, securing core technology that can reflect the unique traits of individual combatants and various combat scenarios. This technology has proven robust enough for battlefield use and is economical enough for widespread distribution to a large number of troops.
On June 25th, Professor Steve Park's research team at KAIST's Department of Materials Science and Engineering announced the development of a flexible, wearable electronic textile (E-textile) platform using an innovative technology that 'draws' electronic circuits directly onto fabric.
The wearable e-textile platform developed by the research team combines 3D printing technology with new materials engineering design to directly print flexible and highly durable sensors and electrodes onto textile substrates. This enables the collection of precise movement and human body data from individual combatants, which can then be used to propose customized training models.
Existing e-textile fabrication methods were often complex or limited in their ability to provide personalized customization. To overcome these challenges, the research team adopted an additive manufacturing technology called 'Direct Ink Writing (DIW)' 3D printing.
< Figure 1. Schematic diagram of e-textile manufactured with Direct Ink Writing (DIW) printing technology on various textiles, including combat uniforms >
This technology involves directly dispensing and printing special ink, which functions as sensors and electrodes, onto textile substrates in desired patterns. This allows for flexible implementation of various designs without the complex process of mask fabrication. This is expected to be an effective technology that can be easily supplied to hundreds of thousands of military personnel.
The core of this technology lies in the development of high-performance functional inks based on advanced materials engineering design. The research team combined styrene-butadiene-styrene (SBS) polymer, which provides flexibility, with multi-walled carbon nanotubes (MWCNT) for electrical conductivity. They developed a tensile/bending sensor ink that can stretch up to 102% and maintain stable performance even after 10,000 repetitive tests. This means that accurate data can be consistently obtained even during the strenuous movements of combatants.
< Figure 2. Measurement of human movement and breathing patterns using e-textile >
Furthermore, new material technology was applied to implement 'interconnect electrodes' that electrically connect the upper and lower layers of the fabric. The team developed an electrode ink combining silver (Ag) flakes with rigid polystyrene (PS) polymer, precisely controlling the impregnation level (how much the ink penetrates the fabric) to effectively connect both sides or multiple layers of the fabric. This secures the technology for producing multi-layered wearable electronic systems integrating sensors and electrodes.
< Figure 3. Experimental results of recognizing unknown objects after machine learning six objects using a smart glove >
The research team proved the platform's performance through actual human movement monitoring experiments. They printed the developed e-textile on major joint areas of clothing (shoulders, elbows, knees) and measured movements and posture changes during various exercises such as running, jumping jacks, and push-ups in real-time.
Additionally, they demonstrated the potential for applications such as monitoring breathing patterns using a smart mask and recognizing objects through machine learning and perceiving complex tactile information by printing multiple sensors and electrodes on gloves. These results show that the developed e-textile platform is effective in precisely understanding the movement dynamics of combatants.
This research is an important example demonstrating how cutting-edge new material technology can contribute to the advancement of the defense sector. Major Kyusoon Pak of the Army, who participated in this research, considered required objectives such as military applicability and economic feasibility for practical distribution from the research design stage.
< Figure 4. Experimental results showing that a multi-layered e-textile glove connected with interconnect electrodes can measure tensile/bending signals and pressure signals at a single point >
Major Pak stated, "Our military is currently facing both a crisis and an opportunity due to the decrease in military personnel resources caused by the demographic cliff and the advancement of science and technology. Also, respect for life in the battlefield is emerging as a significant issue. This research aims to secure original technology that can provide customized training according to military branch/duty and type of combat, thereby enhancing the combat power and ensuring the survivability of our soldiers."
He added, "I hope this research will be evaluated as a case that achieved both scientific contribution and military applicability."
This research, where Kyusoon Pak, Ph.D. Candidate (Army Major) from KAIST's Department of Materials Science and Engineering, participated as the first author and Professor Steve Park supervised, was published on May 27, 2025, in `npj Flexible Electronics (top 1.8% in JCR field)', an international academic journal in the electrical, electronic, and materials engineering fields.
* Paper Title: Fabrication of Multifunctional Wearable Interconnect E-textile Platform Using Direct Ink Writing (DIW) 3D Printing
* DOI: https://doi.org/10.1038/s41528-025-00414-7
This research was supported by the Ministry of Trade, Industry and Energy and the National Research Foundation of Korea.
2025.06.25 View 208 -
 KAIST Predicts Diseases by Early Detection of Aging Signals in Liver Tissue
- KAIST-KRIBB Develops ‘FiNi-seq’ Technology to Capture Characteristics of Fibrotic Microenvironments Accumulated in Liver Tissue and Dynamic Changes of Early Aging Cells
- Elucidation of the Spatial Ecosystem of Aged Liver Tissue, where Reprogramming of Senescent Cells and Immune Exhaustion Progresses, at the Single-Cell Genome and Epigenome Levels
< (From left) Professor Jong-Eun Park of KAIST Graduate School of Medical Science and Engineering (GSMSE), Dr. Chuna Kim of KRIBB, Dr. Kwon Yong Tak of KAIST GSMSE, Ph.D. Candidate Juyeon Kim of KRIBB, Ph.D. Candidate Myungsun Park of KAIST GSMSE >
Aging and chronic diseases involve the gradual accumulation of subtle tissue changes over a long period. Therefore, there are still limitations in quantitatively understanding these changes within organs and linking them to early signs of disease onset. In response, Korean researchers have successfully developed a platform technology that accurately captures localized changes that first occur within tissue, significantly aiding in faster disease discovery and prediction, and in setting personalized treatment targets.
KAIST (President Kwang Hyung Lee) announced on June 12th that a joint research team led by Professor Jong-Eun Park of the Graduate School of Medical Science and Engineering at KAIST and Dr. Chuna Kim of the Aging Convergence Research Center at the Korea Research Institute of Bioscience and Biotechnology (KRIBB, President Seok-Yoon Kwon) has developed ‘FiNi-seq (Fibrotic Niche enrichment sequencing)’ technology. This technology captures fibrotic microenvironments locally occurring in aged liver tissue and enables precise analysis at the single-cell transcriptome level*.
*Single-cell transcriptome analysis: A method to measure how actively each cell uses which genes, allowing identification and function of individual diseased cells.
The researchers developed a method to selectively enrich early aging microenvironments where regeneration is delayed and fibrosis accumulates, by physically selecting regions with high tissue degradation resistance in aged liver tissue.
In this process, high-resolution identification of fibrosis-related endothelial cells, fibroblasts interacting with the immune system, and immune-exhausted cells such as PD-1 highly expressing CD8 T cells, which were difficult to capture with existing single-cell analysis technologies, was possible.
In particular, the research team confirmed through ‘FiNi-seq’ technology that specific cells observed in fibrotic areas within aged liver tissue secondarily age the surrounding environment through secreted factors, and that this leads to the expansion of the aged environment.
Furthermore, they also elucidated the mechanism by which endothelial cells lose their tissue-specific identity and induce innate immune responses, promoting immune cell infiltration. Through spatial transcriptome analysis, the spatial distribution of fibroblasts interacting with immune cells was quantified, revealing their involvement in tissue regeneration, induction of inflammatory responses, and progression to chronic fibrosis.
The research team performed integrated analysis of multi-omics\* data to obtain transcriptome and epigenome information, precisely interpreting the microenvironment of aged liver tissue and its spatial heterogeneity, and confirming how these changes are connected to the intrahepatic vascular structure.
*Multi-omics: An integrated analysis method for various biological information within an organism, such as genes, proteins, metabolites, and cell information.
The newly developed ‘FiNi-seq’ technology is expected to be a useful platform for high-resolution capture of pathophysiological signals in most chronic liver diseases, including the aging process that causes fibrosis.
< Figure 1. Isolation of fibrotic regions from aged liver tissue, followed by single-cell transcriptome analysis and validation in a fibrosis model. >
The first author, Dr. Kwon Yong Tak of KAIST Graduate School of Medical Science and Engineering (GSMSE), a hepatologist at Seoul St. Mary's Hospital, designed this study to lay the groundwork for early diagnosis and treatment of fibrosis progression, the most important clinical prognostic indicator in chronic liver disease, while pursuing his Ph.D. at KAIST KAIST GSMSE with support from the physician-scientist training program. Co-first author Myungsun Park, a Ph.D. candidate at KAIST KAIST GSMSE, was responsible for the technical implementation of FiNi-seq technology, and Juyeon Kim, a Ph.D. candidate at KRIBB's Aging Convergence Research Center, was responsible for imaging analysis of aged tissue, playing a key role in the research.
Dr. Chuna Kim of KRIBB stated, “Through this study, we were able to precisely elucidate the cellular composition and spatial characteristics of the fibrotic microenvironment observed in aged liver tissue at the single-cell level.”
< Figure 2. Spatially defined stepwise progression patterns of aging-related regions within the liver and identification of regulatory factors inducing them. >
Professor Jong-Eun Park of the Graduate School of Medical Science and Engineering said, “As an analytical technology that can capture subtle changes occurring in the early stages of aging and chronic diseases, it is expected to play a significant role in finding effective treatment targets in the future. Also, we plan to expand this research to chronic diseases in other organs such as the lungs and kidneys, as well as various liver disease models.”
This research was published in the international journal ‘Nature Aging’ on May 5, 2025, with Dr. Kwon Yong Tak of KAIST KAIST GSMSE, Ph.D. Candidate Juyeon Kim of KRIBB, and Ph.D. Candidate Myungsun Park of KAIST as co-first authors.
*Paper Title: Quasi-spatial single-cell transcriptome based on physical tissue properties defines early aging associated niche in liver
*DOI: https://doi.org/10.1038/s43587-025-00857-7
This research was supported by several domestic institutions, including the National Research Foundation of Korea, the Korea Health Industry Development Institute (KHIDI), the Korea Research Institute of Bioscience and Biotechnology (KRIBB), KIST, POSCO Science Fellowship, and the Convergence Medical Scientist Training Program.
2025.06.12 View 1004
KAIST Predicts Diseases by Early Detection of Aging Signals in Liver Tissue
- KAIST-KRIBB Develops ‘FiNi-seq’ Technology to Capture Characteristics of Fibrotic Microenvironments Accumulated in Liver Tissue and Dynamic Changes of Early Aging Cells
- Elucidation of the Spatial Ecosystem of Aged Liver Tissue, where Reprogramming of Senescent Cells and Immune Exhaustion Progresses, at the Single-Cell Genome and Epigenome Levels
< (From left) Professor Jong-Eun Park of KAIST Graduate School of Medical Science and Engineering (GSMSE), Dr. Chuna Kim of KRIBB, Dr. Kwon Yong Tak of KAIST GSMSE, Ph.D. Candidate Juyeon Kim of KRIBB, Ph.D. Candidate Myungsun Park of KAIST GSMSE >
Aging and chronic diseases involve the gradual accumulation of subtle tissue changes over a long period. Therefore, there are still limitations in quantitatively understanding these changes within organs and linking them to early signs of disease onset. In response, Korean researchers have successfully developed a platform technology that accurately captures localized changes that first occur within tissue, significantly aiding in faster disease discovery and prediction, and in setting personalized treatment targets.
KAIST (President Kwang Hyung Lee) announced on June 12th that a joint research team led by Professor Jong-Eun Park of the Graduate School of Medical Science and Engineering at KAIST and Dr. Chuna Kim of the Aging Convergence Research Center at the Korea Research Institute of Bioscience and Biotechnology (KRIBB, President Seok-Yoon Kwon) has developed ‘FiNi-seq (Fibrotic Niche enrichment sequencing)’ technology. This technology captures fibrotic microenvironments locally occurring in aged liver tissue and enables precise analysis at the single-cell transcriptome level*.
*Single-cell transcriptome analysis: A method to measure how actively each cell uses which genes, allowing identification and function of individual diseased cells.
The researchers developed a method to selectively enrich early aging microenvironments where regeneration is delayed and fibrosis accumulates, by physically selecting regions with high tissue degradation resistance in aged liver tissue.
In this process, high-resolution identification of fibrosis-related endothelial cells, fibroblasts interacting with the immune system, and immune-exhausted cells such as PD-1 highly expressing CD8 T cells, which were difficult to capture with existing single-cell analysis technologies, was possible.
In particular, the research team confirmed through ‘FiNi-seq’ technology that specific cells observed in fibrotic areas within aged liver tissue secondarily age the surrounding environment through secreted factors, and that this leads to the expansion of the aged environment.
Furthermore, they also elucidated the mechanism by which endothelial cells lose their tissue-specific identity and induce innate immune responses, promoting immune cell infiltration. Through spatial transcriptome analysis, the spatial distribution of fibroblasts interacting with immune cells was quantified, revealing their involvement in tissue regeneration, induction of inflammatory responses, and progression to chronic fibrosis.
The research team performed integrated analysis of multi-omics\* data to obtain transcriptome and epigenome information, precisely interpreting the microenvironment of aged liver tissue and its spatial heterogeneity, and confirming how these changes are connected to the intrahepatic vascular structure.
*Multi-omics: An integrated analysis method for various biological information within an organism, such as genes, proteins, metabolites, and cell information.
The newly developed ‘FiNi-seq’ technology is expected to be a useful platform for high-resolution capture of pathophysiological signals in most chronic liver diseases, including the aging process that causes fibrosis.
< Figure 1. Isolation of fibrotic regions from aged liver tissue, followed by single-cell transcriptome analysis and validation in a fibrosis model. >
The first author, Dr. Kwon Yong Tak of KAIST Graduate School of Medical Science and Engineering (GSMSE), a hepatologist at Seoul St. Mary's Hospital, designed this study to lay the groundwork for early diagnosis and treatment of fibrosis progression, the most important clinical prognostic indicator in chronic liver disease, while pursuing his Ph.D. at KAIST KAIST GSMSE with support from the physician-scientist training program. Co-first author Myungsun Park, a Ph.D. candidate at KAIST KAIST GSMSE, was responsible for the technical implementation of FiNi-seq technology, and Juyeon Kim, a Ph.D. candidate at KRIBB's Aging Convergence Research Center, was responsible for imaging analysis of aged tissue, playing a key role in the research.
Dr. Chuna Kim of KRIBB stated, “Through this study, we were able to precisely elucidate the cellular composition and spatial characteristics of the fibrotic microenvironment observed in aged liver tissue at the single-cell level.”
< Figure 2. Spatially defined stepwise progression patterns of aging-related regions within the liver and identification of regulatory factors inducing them. >
Professor Jong-Eun Park of the Graduate School of Medical Science and Engineering said, “As an analytical technology that can capture subtle changes occurring in the early stages of aging and chronic diseases, it is expected to play a significant role in finding effective treatment targets in the future. Also, we plan to expand this research to chronic diseases in other organs such as the lungs and kidneys, as well as various liver disease models.”
This research was published in the international journal ‘Nature Aging’ on May 5, 2025, with Dr. Kwon Yong Tak of KAIST KAIST GSMSE, Ph.D. Candidate Juyeon Kim of KRIBB, and Ph.D. Candidate Myungsun Park of KAIST as co-first authors.
*Paper Title: Quasi-spatial single-cell transcriptome based on physical tissue properties defines early aging associated niche in liver
*DOI: https://doi.org/10.1038/s43587-025-00857-7
This research was supported by several domestic institutions, including the National Research Foundation of Korea, the Korea Health Industry Development Institute (KHIDI), the Korea Research Institute of Bioscience and Biotechnology (KRIBB), KIST, POSCO Science Fellowship, and the Convergence Medical Scientist Training Program.
2025.06.12 View 1004 -
 KAIST Professor Jee-Hwan Ryu Receives Global IEEE Robotics Journal Best Paper Award
- Professor Jee-Hwan Ryu of Civil and Environmental Engineering receives the Best Paper Award from the Institute of Electrical and Electronics Engineers (IEEE) Robotics Journal, officially presented at ICRA, a world-renowned robotics conference.
- This is the highest level of international recognition, awarded to only the top 5 papers out of approximately 1,500 published in 2024.
- Securing a new working channel technology for soft growing robots expands the practicality and application possibilities in the field of soft robotics.
< Professor Jee-Hwan Ryu (left), Nam Gyun Kim, Ph.D. Candidate (right) from the KAIST Department of Civil and Environmental Engineering and KAIST Robotics Program >
KAIST (President Kwang-Hyung Lee) announced on the 6th that Professor Jee-Hwan Ryu from the Department of Civil and Environmental Engineering received the 2024 Best Paper Award from the Robotics and Automation Letters (RA-L), a premier journal under the IEEE, at the '2025 IEEE International Conference on Robotics and Automation (ICRA)' held in Atlanta, USA, on May 22nd.
This Best Paper Award is a prestigious honor presented to only the top 5 papers out of approximately 1,500 published in 2024, boasting high international competition and authority.
The award-winning paper by Professor Ryu proposes a novel working channel securing mechanism that significantly expands the practicality and application possibilities of 'Soft Growing Robots,' which are based on soft materials that move or perform tasks through a growing motion similar to plant roots.
< IEEE Robotics Journal Award Ceremony >
Existing soft growing robots move by inflating or contracting their bodies through increasing or decreasing internal pressure, which can lead to blockages in their internal passages. In contrast, the newly developed soft growing robot achieves a growing function while maintaining the internal passage pressure equal to the external atmospheric pressure, thereby successfully securing an internal passage while retaining the robot's flexible and soft characteristics.
This structure allows various materials or tools to be freely delivered through the internal passage (working channel) within the robot and offers the advantage of performing multi-purpose tasks by flexibly replacing equipment according to the working environment.
The research team fabricated a prototype to prove the effectiveness of this technology and verified its performance through various experiments. Specifically, in the slide plate experiment, they confirmed whether materials or equipment could pass through the robot's internal channel without obstruction, and in the pipe pulling experiment, they verified if a long pipe-shaped tool could be pulled through the internal channel.
< Figure 1. Overall hardware structure of the proposed soft growing robot (left) and a cross-sectional view composing the inflatable structure (right) >
Experimental results demonstrated that the internal channel remained stable even while the robot was growing, serving as a key basis for supporting the technology's practicality and scalability.
Professor Jee-Hwan Ryu stated, "This award is very meaningful as it signifies the global recognition of Korea's robotics technology and academic achievements. Especially, it holds great significance in achieving technical progress that can greatly expand the practicality and application fields of soft growing robots. This achievement was possible thanks to the dedication and collaboration of the research team, and I will continue to contribute to the development of robotics technology through innovative research."
< Figure 2. Material supplying mechanism of the Soft Growing Robot >
This research was co-authored by Dongoh Seo, Ph.D. Candidate in Civil and Environmental Engineering, and Nam Gyun Kim, Ph.D. Candidate in Robotics. It was published in IEEE Robotics and Automation Letters on September 1, 2024.
(Paper Title: Inflatable-Structure-Based Working-Channel Securing Mechanism for Soft Growing Robots, DOI: 10.1109/LRA.2024.3426322)
This project was supported simultaneously by the National Research Foundation of Korea's Future Promising Convergence Technology Pioneer Research Project and Mid-career Researcher Project.
2025.06.09 View 1583
KAIST Professor Jee-Hwan Ryu Receives Global IEEE Robotics Journal Best Paper Award
- Professor Jee-Hwan Ryu of Civil and Environmental Engineering receives the Best Paper Award from the Institute of Electrical and Electronics Engineers (IEEE) Robotics Journal, officially presented at ICRA, a world-renowned robotics conference.
- This is the highest level of international recognition, awarded to only the top 5 papers out of approximately 1,500 published in 2024.
- Securing a new working channel technology for soft growing robots expands the practicality and application possibilities in the field of soft robotics.
< Professor Jee-Hwan Ryu (left), Nam Gyun Kim, Ph.D. Candidate (right) from the KAIST Department of Civil and Environmental Engineering and KAIST Robotics Program >
KAIST (President Kwang-Hyung Lee) announced on the 6th that Professor Jee-Hwan Ryu from the Department of Civil and Environmental Engineering received the 2024 Best Paper Award from the Robotics and Automation Letters (RA-L), a premier journal under the IEEE, at the '2025 IEEE International Conference on Robotics and Automation (ICRA)' held in Atlanta, USA, on May 22nd.
This Best Paper Award is a prestigious honor presented to only the top 5 papers out of approximately 1,500 published in 2024, boasting high international competition and authority.
The award-winning paper by Professor Ryu proposes a novel working channel securing mechanism that significantly expands the practicality and application possibilities of 'Soft Growing Robots,' which are based on soft materials that move or perform tasks through a growing motion similar to plant roots.
< IEEE Robotics Journal Award Ceremony >
Existing soft growing robots move by inflating or contracting their bodies through increasing or decreasing internal pressure, which can lead to blockages in their internal passages. In contrast, the newly developed soft growing robot achieves a growing function while maintaining the internal passage pressure equal to the external atmospheric pressure, thereby successfully securing an internal passage while retaining the robot's flexible and soft characteristics.
This structure allows various materials or tools to be freely delivered through the internal passage (working channel) within the robot and offers the advantage of performing multi-purpose tasks by flexibly replacing equipment according to the working environment.
The research team fabricated a prototype to prove the effectiveness of this technology and verified its performance through various experiments. Specifically, in the slide plate experiment, they confirmed whether materials or equipment could pass through the robot's internal channel without obstruction, and in the pipe pulling experiment, they verified if a long pipe-shaped tool could be pulled through the internal channel.
< Figure 1. Overall hardware structure of the proposed soft growing robot (left) and a cross-sectional view composing the inflatable structure (right) >
Experimental results demonstrated that the internal channel remained stable even while the robot was growing, serving as a key basis for supporting the technology's practicality and scalability.
Professor Jee-Hwan Ryu stated, "This award is very meaningful as it signifies the global recognition of Korea's robotics technology and academic achievements. Especially, it holds great significance in achieving technical progress that can greatly expand the practicality and application fields of soft growing robots. This achievement was possible thanks to the dedication and collaboration of the research team, and I will continue to contribute to the development of robotics technology through innovative research."
< Figure 2. Material supplying mechanism of the Soft Growing Robot >
This research was co-authored by Dongoh Seo, Ph.D. Candidate in Civil and Environmental Engineering, and Nam Gyun Kim, Ph.D. Candidate in Robotics. It was published in IEEE Robotics and Automation Letters on September 1, 2024.
(Paper Title: Inflatable-Structure-Based Working-Channel Securing Mechanism for Soft Growing Robots, DOI: 10.1109/LRA.2024.3426322)
This project was supported simultaneously by the National Research Foundation of Korea's Future Promising Convergence Technology Pioneer Research Project and Mid-career Researcher Project.
2025.06.09 View 1583 -
 Professor Hyun Myung's Team Wins First Place in a Challenge at ICRA by IEEE
< Photo 1. (From left) Daebeom Kim (Team Leader, Ph.D. student), Seungjae Lee (Ph.D. student), Seoyeon Jang (Ph.D. student), Jei Kong (Master's student), Professor Hyun Myung >
A team of the Urban Robotics Lab, led by Professor Hyun Myung from the KAIST School of Electrical Engineering, achieved a remarkable first-place overall victory in the Nothing Stands Still Challenge (NSS Challenge) 2025, held at the 2025 IEEE International Conference on Robotics and Automation (ICRA), the world's most prestigious robotics conference, from May 19 to 23 in Atlanta, USA.
The NSS Challenge was co-hosted by HILTI, a global construction company based in Liechtenstein, and Stanford University's Gradient Spaces Group. It is an expanded version of the HILTI SLAM (Simultaneous Localization and Mapping)* Challenge, which has been held since 2021, and is considered one of the most prominent challenges at 2025 IEEE ICRA.*SLAM: Refers to Simultaneous Localization and Mapping, a technology where robots, drones, autonomous vehicles, etc., determine their own position and simultaneously create a map of their surroundings.
< Photo 2. A scene from the oral presentation on the winning team's technology (Speakers: Seungjae Lee and Seoyeon Jang, Ph.D. candidates of KAIST School of Electrical Engineering) >
This challenge primarily evaluates how accurately and robustly LiDAR scan data, collected at various times, can be registered in situations with frequent structural changes, such as construction and industrial environments. In particular, it is regarded as a highly technical competition because it deals with multi-session localization and mapping (Multi-session SLAM) technology that responds to structural changes occurring over multiple timeframes, rather than just single-point registration accuracy.
The Urban Robotics Lab team secured first place overall, surpassing National Taiwan University (3rd place) and Northwestern Polytechnical University of China (2nd place) by a significant margin, with their unique localization and mapping technology that solves the problem of registering LiDAR data collected across multiple times and spaces. The winning team will be awarded a prize of $4,000.
< Figure 1. Example of Multiway-Registration for Registering Multiple Scans >
The Urban Robotics Lab team independently developed a multiway-registration framework that can robustly register multiple scans even without prior connection information. This framework consists of an algorithm for summarizing feature points within scans and finding correspondences (CubicFeat), an algorithm for performing global registration based on the found correspondences (Quatro), and an algorithm for refining results based on change detection (Chamelion). This combination of technologies ensures stable registration performance based on fixed structures, even in highly dynamic industrial environments.
< Figure 2. Example of Change Detection Using the Chamelion Algorithm>
LiDAR scan registration technology is a core component of SLAM (Simultaneous Localization And Mapping) in various autonomous systems such as autonomous vehicles, autonomous robots, autonomous walking systems, and autonomous flying vehicles.
Professor Hyun Myung of the School of Electrical Engineering stated, "This award-winning technology is evaluated as a case that simultaneously proves both academic value and industrial applicability by maximizing the performance of precisely estimating the relative positions between different scans even in complex environments. I am grateful to the students who challenged themselves and never gave up, even when many teams abandoned due to the high difficulty."
< Figure 3. Competition Result Board, Lower RMSE (Root Mean Squared Error) Indicates Higher Score (Unit: meters)>
The Urban Robotics Lab team first participated in the SLAM Challenge in 2022, winning second place among academic teams, and in 2023, they secured first place overall in the LiDAR category and first place among academic teams in the vision category.
2025.05.30 View 2343
Professor Hyun Myung's Team Wins First Place in a Challenge at ICRA by IEEE
< Photo 1. (From left) Daebeom Kim (Team Leader, Ph.D. student), Seungjae Lee (Ph.D. student), Seoyeon Jang (Ph.D. student), Jei Kong (Master's student), Professor Hyun Myung >
A team of the Urban Robotics Lab, led by Professor Hyun Myung from the KAIST School of Electrical Engineering, achieved a remarkable first-place overall victory in the Nothing Stands Still Challenge (NSS Challenge) 2025, held at the 2025 IEEE International Conference on Robotics and Automation (ICRA), the world's most prestigious robotics conference, from May 19 to 23 in Atlanta, USA.
The NSS Challenge was co-hosted by HILTI, a global construction company based in Liechtenstein, and Stanford University's Gradient Spaces Group. It is an expanded version of the HILTI SLAM (Simultaneous Localization and Mapping)* Challenge, which has been held since 2021, and is considered one of the most prominent challenges at 2025 IEEE ICRA.*SLAM: Refers to Simultaneous Localization and Mapping, a technology where robots, drones, autonomous vehicles, etc., determine their own position and simultaneously create a map of their surroundings.
< Photo 2. A scene from the oral presentation on the winning team's technology (Speakers: Seungjae Lee and Seoyeon Jang, Ph.D. candidates of KAIST School of Electrical Engineering) >
This challenge primarily evaluates how accurately and robustly LiDAR scan data, collected at various times, can be registered in situations with frequent structural changes, such as construction and industrial environments. In particular, it is regarded as a highly technical competition because it deals with multi-session localization and mapping (Multi-session SLAM) technology that responds to structural changes occurring over multiple timeframes, rather than just single-point registration accuracy.
The Urban Robotics Lab team secured first place overall, surpassing National Taiwan University (3rd place) and Northwestern Polytechnical University of China (2nd place) by a significant margin, with their unique localization and mapping technology that solves the problem of registering LiDAR data collected across multiple times and spaces. The winning team will be awarded a prize of $4,000.
< Figure 1. Example of Multiway-Registration for Registering Multiple Scans >
The Urban Robotics Lab team independently developed a multiway-registration framework that can robustly register multiple scans even without prior connection information. This framework consists of an algorithm for summarizing feature points within scans and finding correspondences (CubicFeat), an algorithm for performing global registration based on the found correspondences (Quatro), and an algorithm for refining results based on change detection (Chamelion). This combination of technologies ensures stable registration performance based on fixed structures, even in highly dynamic industrial environments.
< Figure 2. Example of Change Detection Using the Chamelion Algorithm>
LiDAR scan registration technology is a core component of SLAM (Simultaneous Localization And Mapping) in various autonomous systems such as autonomous vehicles, autonomous robots, autonomous walking systems, and autonomous flying vehicles.
Professor Hyun Myung of the School of Electrical Engineering stated, "This award-winning technology is evaluated as a case that simultaneously proves both academic value and industrial applicability by maximizing the performance of precisely estimating the relative positions between different scans even in complex environments. I am grateful to the students who challenged themselves and never gave up, even when many teams abandoned due to the high difficulty."
< Figure 3. Competition Result Board, Lower RMSE (Root Mean Squared Error) Indicates Higher Score (Unit: meters)>
The Urban Robotics Lab team first participated in the SLAM Challenge in 2022, winning second place among academic teams, and in 2023, they secured first place overall in the LiDAR category and first place among academic teams in the vision category.
2025.05.30 View 2343 -
 KAIST Develops Virtual Staining Technology for 3D Histopathology
Moving beyond traditional methods of observing thinly sliced and stained cancer tissues, a collaborative international research team led by KAIST has successfully developed a groundbreaking technology. This innovation uses advanced optical techniques combined with an artificial intelligence-based deep learning algorithm to create realistic, virtually stained 3D images of cancer tissue without the need for serial sectioning nor staining. This breakthrough is anticipated to pave the way for next-generation non-invasive pathological diagnosis.
< Photo 1. (From left) Juyeon Park (Ph.D. Candidate, Department of Physics), Professor YongKeun Park (Department of Physics) (Top left) Professor Su-Jin Shin (Gangnam Severance Hospital), Professor Tae Hyun Hwang (Vanderbilt University School of Medicine) >
KAIST (President Kwang Hyung Lee) announced on the 26th that a research team led by Professor YongKeun Park of the Department of Physics, in collaboration with Professor Su-Jin Shin's team at Yonsei University Gangnam Severance Hospital, Professor Tae Hyun Hwang's team at Mayo Clinic, and Tomocube's AI research team, has developed an innovative technology capable of vividly displaying the 3D structure of cancer tissues without separate staining.
For over 200 years, conventional pathology has relied on observing cancer tissues under a microscope, a method that only shows specific cross-sections of the 3D cancer tissue. This has limited the ability to understand the three-dimensional connections and spatial arrangements between cells.
To overcome this, the research team utilized holotomography (HT), an advanced optical technology, to measure the 3D refractive index information of tissues. They then integrated an AI-based deep learning algorithm to successfully generate virtual H&E* images.* H&E (Hematoxylin & Eosin): The most widely used staining method for observing pathological tissues. Hematoxylin stains cell nuclei blue, and eosin stains cytoplasm pink.
The research team quantitatively demonstrated that the images generated by this technology are highly similar to actual stained tissue images. Furthermore, the technology exhibited consistent performance across various organs and tissues, proving its versatility and reliability as a next-generation pathological analysis tool.
< Figure 1. Comparison of conventional 3D tissue pathology procedure and the 3D virtual H&E staining technology proposed in this study. The traditional method requires preparing and staining dozens of tissue slides, while the proposed technology can reduce the number of slides by up to 10 times and quickly generate H&E images without the staining process. >
Moreover, by validating the feasibility of this technology through joint research with hospitals and research institutions in Korea and the United States, utilizing Tomocube's holotomography equipment, the team demonstrated its potential for full-scale adoption in real-world pathological research settings.
Professor YongKeun Park stated, "This research marks a major advancement by transitioning pathological analysis from conventional 2D methods to comprehensive 3D imaging. It will greatly enhance biomedical research and clinical diagnostics, particularly in understanding cancer tumor boundaries and the intricate spatial arrangements of cells within tumor microenvironments."
< Figure 2. Results of AI-based 3D virtual H&E staining and quantitative analysis of pathological tissue. The virtually stained images enabled 3D reconstruction of key pathological features such as cell nuclei and glandular lumens. Based on this, various quantitative indicators, including cell nuclear distribution, volume, and surface area, could be extracted. >
This research, with Juyeon Park, a student of the Integrated Master’s and Ph.D. Program at KAIST, as the first author, was published online in the prestigious journal Nature Communications on May 22.
(Paper title: Revealing 3D microanatomical structures of unlabeled thick cancer tissues using holotomography and virtual H&E staining.
[https://doi.org/10.1038/s41467-025-59820-0]
This study was supported by the Leader Researcher Program of the National Research Foundation of Korea, the Global Industry Technology Cooperation Center Project of the Korea Institute for Advancement of Technology, and the Korea Health Industry Development Institute.
2025.05.26 View 2580
KAIST Develops Virtual Staining Technology for 3D Histopathology
Moving beyond traditional methods of observing thinly sliced and stained cancer tissues, a collaborative international research team led by KAIST has successfully developed a groundbreaking technology. This innovation uses advanced optical techniques combined with an artificial intelligence-based deep learning algorithm to create realistic, virtually stained 3D images of cancer tissue without the need for serial sectioning nor staining. This breakthrough is anticipated to pave the way for next-generation non-invasive pathological diagnosis.
< Photo 1. (From left) Juyeon Park (Ph.D. Candidate, Department of Physics), Professor YongKeun Park (Department of Physics) (Top left) Professor Su-Jin Shin (Gangnam Severance Hospital), Professor Tae Hyun Hwang (Vanderbilt University School of Medicine) >
KAIST (President Kwang Hyung Lee) announced on the 26th that a research team led by Professor YongKeun Park of the Department of Physics, in collaboration with Professor Su-Jin Shin's team at Yonsei University Gangnam Severance Hospital, Professor Tae Hyun Hwang's team at Mayo Clinic, and Tomocube's AI research team, has developed an innovative technology capable of vividly displaying the 3D structure of cancer tissues without separate staining.
For over 200 years, conventional pathology has relied on observing cancer tissues under a microscope, a method that only shows specific cross-sections of the 3D cancer tissue. This has limited the ability to understand the three-dimensional connections and spatial arrangements between cells.
To overcome this, the research team utilized holotomography (HT), an advanced optical technology, to measure the 3D refractive index information of tissues. They then integrated an AI-based deep learning algorithm to successfully generate virtual H&E* images.* H&E (Hematoxylin & Eosin): The most widely used staining method for observing pathological tissues. Hematoxylin stains cell nuclei blue, and eosin stains cytoplasm pink.
The research team quantitatively demonstrated that the images generated by this technology are highly similar to actual stained tissue images. Furthermore, the technology exhibited consistent performance across various organs and tissues, proving its versatility and reliability as a next-generation pathological analysis tool.
< Figure 1. Comparison of conventional 3D tissue pathology procedure and the 3D virtual H&E staining technology proposed in this study. The traditional method requires preparing and staining dozens of tissue slides, while the proposed technology can reduce the number of slides by up to 10 times and quickly generate H&E images without the staining process. >
Moreover, by validating the feasibility of this technology through joint research with hospitals and research institutions in Korea and the United States, utilizing Tomocube's holotomography equipment, the team demonstrated its potential for full-scale adoption in real-world pathological research settings.
Professor YongKeun Park stated, "This research marks a major advancement by transitioning pathological analysis from conventional 2D methods to comprehensive 3D imaging. It will greatly enhance biomedical research and clinical diagnostics, particularly in understanding cancer tumor boundaries and the intricate spatial arrangements of cells within tumor microenvironments."
< Figure 2. Results of AI-based 3D virtual H&E staining and quantitative analysis of pathological tissue. The virtually stained images enabled 3D reconstruction of key pathological features such as cell nuclei and glandular lumens. Based on this, various quantitative indicators, including cell nuclear distribution, volume, and surface area, could be extracted. >
This research, with Juyeon Park, a student of the Integrated Master’s and Ph.D. Program at KAIST, as the first author, was published online in the prestigious journal Nature Communications on May 22.
(Paper title: Revealing 3D microanatomical structures of unlabeled thick cancer tissues using holotomography and virtual H&E staining.
[https://doi.org/10.1038/s41467-025-59820-0]
This study was supported by the Leader Researcher Program of the National Research Foundation of Korea, the Global Industry Technology Cooperation Center Project of the Korea Institute for Advancement of Technology, and the Korea Health Industry Development Institute.
2025.05.26 View 2580 -
 KAIST Discovers Molecular Switch that Reverses Cancerous Transformation at the Critical Moment of Transition
< (From left) PhD student Seoyoon D. Jeong, (bottom) Professor Kwang-Hyun Cho, (top) Dr. Dongkwan Shin, Dr. Jeong-Ryeol Gong >
Professor Kwang-Hyun Cho’s research team has recently been highlighted for their work on developing an original technology for cancer reversal treatment that does not kill cancer cells but only changes their characteristics to reverse them to a state similar to normal cells. This time, they have succeeded in revealing for the first time that a molecular switch that can induce cancer reversal at the moment when normal cells change into cancer cells is hidden in the genetic network.
KAIST (President Kwang-Hyung Lee) announced on the 5th of February that Professor Kwang-Hyun Cho's research team of the Department of Bio and Brain Engineering has succeeded in developing a fundamental technology to capture the critical transition phenomenon at the moment when normal cells change into cancer cells and analyze it to discover a molecular switch that can revert cancer cells back into normal cells.
A critical transition is a phenomenon in which a sudden change in state occurs at a specific point in time, like water changing into steam at 100℃. This critical transition phenomenon also occurs in the process in which normal cells change into cancer cells at a specific point in time due to the accumulation of genetic and epigenetic changes.
The research team discovered that normal cells can enter an unstable critical transition state where normal cells and cancer cells coexist just before they change into cancer cells during tumorigenesis, the production or development of tumors, and analyzed this critical transition state using a systems biology method to develop a cancer reversal molecular switch identification technology that can reverse the cancerization process. They then applied this to colon cancer cells and confirmed through molecular cell experiments that cancer cells can recover the characteristics of normal cells.
This is an original technology that automatically infers a computer model of the genetic network that controls the critical transition of cancer development from single-cell RNA sequencing data, and systematically finds molecular switches for cancer reversion by simulation analysis. It is expected that this technology will be applied to the development of reversion therapies for other cancers in the future.
Professor Kwang-Hyun Cho said, "We have discovered a molecular switch that can revert the fate of cancer cells back to a normal state by capturing the moment of critical transition right before normal cells are changed into an irreversible cancerous state."
< Figure 1. Overall conceptual framework of the technology that automatically constructs a molecular regulatory network from single-cell RNA sequencing data of colon cancer cells to discover molecular switches for cancer reversion through computer simulation analysis. Professor Kwang-Hyun Cho's research team established a fundamental technology for automatic construction of a computer model of a core gene network by analyzing the entire process of tumorigenesis of colon cells turning into cancer cells, and developed an original technology for discovering the molecular switches that can induce cancer cell reversal through attractor landscape analysis. >
He continued, "In particular, this study has revealed in detail, at the genetic network level, what changes occur within cells behind the process of cancer development, which has been considered a mystery until now." He emphasized, "This is the first study to reveal that an important clue that can revert the fate of tumorigenesis is hidden at this very critical moment of change."
< Figure 2. Identification of tumor transition state using single-cell RNA sequencing data from colorectal cancer. Using single-cell RNA sequencing data from colorectal cancer patient-derived organoids for normal and cancerous tissues, a critical transition was identified in which normal and cancerous cells coexist and instability increases (a-d). The critical transition was confirmed to show intermediate levels of major phenotypic features related to cancer or normal tissues that are indicative of the states between the normal and cancerous cells (e). >
The results of this study, conducted by KAIST Dr. Dongkwan Shin (currently at the National Cancer Center), Dr. Jeong-Ryeol Gong, and doctoral student Seoyoon D. Jeong jointly with a research team at Seoul National University that provided the organoids (in vitro cultured tissues) from colon cancer patient, were published as an online paper in the international journal ‘Advanced Science’ published by Wiley on January 22nd.
(Paper title: Attractor landscape analysis reveals a reversion switch in the transition of colorectal tumorigenesis) (DOI: https://doi.org/10.1002/advs.202412503)
< Figure 3. Reconstruction of a dynamic network model for the transition state of colorectal cancer.
A new technology was established to build a gene network computer model that can simulate the dynamic changes between genes by integrating single-cell RNA sequencing data and existing experimental results on gene-to-gene interactions in the critical transition of cancer. (a). Using this technology, a gene network computer model for the critical transition of colorectal cancer was constructed, and the distribution of attractors representing normal and cancer cell phenotypes was investigated through attractor landscape analysis (b-e). >
This study was conducted with the support of the National Research Foundation of Korea under the Ministry of Science and ICT through the Mid-Career Researcher Program and Basic Research Laboratory Program and the Disease-Centered Translational Research Project of the Korea Health Industry Development Institute (KHIDI) of the Ministry of Health and Welfare.
< Figure 4. Quantification of attractor landscapes and discovery of transcription factors for cancer reversibility through perturbation simulation analysis. A methodology for implementing discontinuous attractor landscapes continuously from a computer model of gene networks and quantifying them as cancer scores was introduced (a), and attractor landscapes for the critical transition of colorectal cancer were secured (b-d). By tracking the change patterns of normal and cancer cell attractors through perturbation simulation analysis for each gene, the optimal combination of transcription factors for cancer reversion was discovered (e-h). This was confirmed in various parameter combinations as well (i). >
< Figure 5. Identification and experimental validation of the optimal target gene for cancer reversion. Among the common target genes of the discovered transcription factor combinations, we identified cancer reversing molecular switches that are predicted to suppress cancer cell proliferation and restore the characteristics of normal colon cells (a-d). When inhibitors for the molecular switches were treated to organoids derived from colon cancer patients, it was confirmed that cancer cell proliferation was suppressed and the expression of key genes related to cancer development was inhibited (e-h), and a group of genes related to normal colon epithelium was activated and transformed into a state similar to normal colon cells (i-j). >
< Figure 6. Schematic diagram of the research results. Professor Kwang-Hyun Cho's research team developed an original technology to systematically discover key molecular switches that can induce reversion of colon cancer cells through a systems biology approach using an attractor landscape analysis of a genetic network model for the critical transition at the moment of transformation from normal cells to cancer cells, and verified the reversing effect of actual colon cancer through cellular experiments. >
2025.02.05 View 26440
KAIST Discovers Molecular Switch that Reverses Cancerous Transformation at the Critical Moment of Transition
< (From left) PhD student Seoyoon D. Jeong, (bottom) Professor Kwang-Hyun Cho, (top) Dr. Dongkwan Shin, Dr. Jeong-Ryeol Gong >
Professor Kwang-Hyun Cho’s research team has recently been highlighted for their work on developing an original technology for cancer reversal treatment that does not kill cancer cells but only changes their characteristics to reverse them to a state similar to normal cells. This time, they have succeeded in revealing for the first time that a molecular switch that can induce cancer reversal at the moment when normal cells change into cancer cells is hidden in the genetic network.
KAIST (President Kwang-Hyung Lee) announced on the 5th of February that Professor Kwang-Hyun Cho's research team of the Department of Bio and Brain Engineering has succeeded in developing a fundamental technology to capture the critical transition phenomenon at the moment when normal cells change into cancer cells and analyze it to discover a molecular switch that can revert cancer cells back into normal cells.
A critical transition is a phenomenon in which a sudden change in state occurs at a specific point in time, like water changing into steam at 100℃. This critical transition phenomenon also occurs in the process in which normal cells change into cancer cells at a specific point in time due to the accumulation of genetic and epigenetic changes.
The research team discovered that normal cells can enter an unstable critical transition state where normal cells and cancer cells coexist just before they change into cancer cells during tumorigenesis, the production or development of tumors, and analyzed this critical transition state using a systems biology method to develop a cancer reversal molecular switch identification technology that can reverse the cancerization process. They then applied this to colon cancer cells and confirmed through molecular cell experiments that cancer cells can recover the characteristics of normal cells.
This is an original technology that automatically infers a computer model of the genetic network that controls the critical transition of cancer development from single-cell RNA sequencing data, and systematically finds molecular switches for cancer reversion by simulation analysis. It is expected that this technology will be applied to the development of reversion therapies for other cancers in the future.
Professor Kwang-Hyun Cho said, "We have discovered a molecular switch that can revert the fate of cancer cells back to a normal state by capturing the moment of critical transition right before normal cells are changed into an irreversible cancerous state."
< Figure 1. Overall conceptual framework of the technology that automatically constructs a molecular regulatory network from single-cell RNA sequencing data of colon cancer cells to discover molecular switches for cancer reversion through computer simulation analysis. Professor Kwang-Hyun Cho's research team established a fundamental technology for automatic construction of a computer model of a core gene network by analyzing the entire process of tumorigenesis of colon cells turning into cancer cells, and developed an original technology for discovering the molecular switches that can induce cancer cell reversal through attractor landscape analysis. >
He continued, "In particular, this study has revealed in detail, at the genetic network level, what changes occur within cells behind the process of cancer development, which has been considered a mystery until now." He emphasized, "This is the first study to reveal that an important clue that can revert the fate of tumorigenesis is hidden at this very critical moment of change."
< Figure 2. Identification of tumor transition state using single-cell RNA sequencing data from colorectal cancer. Using single-cell RNA sequencing data from colorectal cancer patient-derived organoids for normal and cancerous tissues, a critical transition was identified in which normal and cancerous cells coexist and instability increases (a-d). The critical transition was confirmed to show intermediate levels of major phenotypic features related to cancer or normal tissues that are indicative of the states between the normal and cancerous cells (e). >
The results of this study, conducted by KAIST Dr. Dongkwan Shin (currently at the National Cancer Center), Dr. Jeong-Ryeol Gong, and doctoral student Seoyoon D. Jeong jointly with a research team at Seoul National University that provided the organoids (in vitro cultured tissues) from colon cancer patient, were published as an online paper in the international journal ‘Advanced Science’ published by Wiley on January 22nd.
(Paper title: Attractor landscape analysis reveals a reversion switch in the transition of colorectal tumorigenesis) (DOI: https://doi.org/10.1002/advs.202412503)
< Figure 3. Reconstruction of a dynamic network model for the transition state of colorectal cancer.
A new technology was established to build a gene network computer model that can simulate the dynamic changes between genes by integrating single-cell RNA sequencing data and existing experimental results on gene-to-gene interactions in the critical transition of cancer. (a). Using this technology, a gene network computer model for the critical transition of colorectal cancer was constructed, and the distribution of attractors representing normal and cancer cell phenotypes was investigated through attractor landscape analysis (b-e). >
This study was conducted with the support of the National Research Foundation of Korea under the Ministry of Science and ICT through the Mid-Career Researcher Program and Basic Research Laboratory Program and the Disease-Centered Translational Research Project of the Korea Health Industry Development Institute (KHIDI) of the Ministry of Health and Welfare.
< Figure 4. Quantification of attractor landscapes and discovery of transcription factors for cancer reversibility through perturbation simulation analysis. A methodology for implementing discontinuous attractor landscapes continuously from a computer model of gene networks and quantifying them as cancer scores was introduced (a), and attractor landscapes for the critical transition of colorectal cancer were secured (b-d). By tracking the change patterns of normal and cancer cell attractors through perturbation simulation analysis for each gene, the optimal combination of transcription factors for cancer reversion was discovered (e-h). This was confirmed in various parameter combinations as well (i). >
< Figure 5. Identification and experimental validation of the optimal target gene for cancer reversion. Among the common target genes of the discovered transcription factor combinations, we identified cancer reversing molecular switches that are predicted to suppress cancer cell proliferation and restore the characteristics of normal colon cells (a-d). When inhibitors for the molecular switches were treated to organoids derived from colon cancer patients, it was confirmed that cancer cell proliferation was suppressed and the expression of key genes related to cancer development was inhibited (e-h), and a group of genes related to normal colon epithelium was activated and transformed into a state similar to normal colon cells (i-j). >
< Figure 6. Schematic diagram of the research results. Professor Kwang-Hyun Cho's research team developed an original technology to systematically discover key molecular switches that can induce reversion of colon cancer cells through a systems biology approach using an attractor landscape analysis of a genetic network model for the critical transition at the moment of transformation from normal cells to cancer cells, and verified the reversing effect of actual colon cancer through cellular experiments. >
2025.02.05 View 26440 -
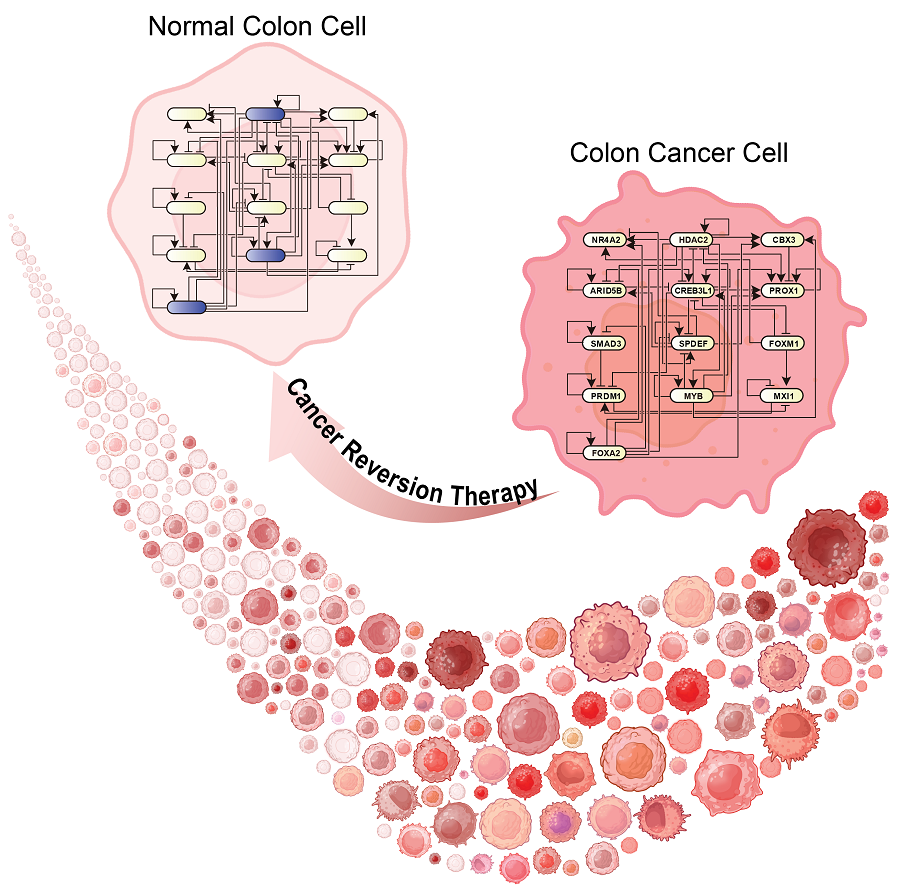 KAIST Develops Foundational Technology to Revert Cancer Cells to Normal Cells
Despite the development of numerous cancer treatment technologies, the common goal of current cancer therapies is to eliminate cancer cells. This approach, however, faces fundamental limitations, including cancer cells developing resistance and returning, as well as severe side effects from the destruction of healthy cells.
< (From top left) Bio and Brain Engineering PhD candidates Juhee Kim, Jeong-Ryeol Gong, Chun-Kyung Lee, and Hoon-Min Kim posed for a group photo with Professor Kwang-Hyun Cho >
KAIST (represented by President Kwang Hyung Lee) announced on the 20th of December that a research team led by Professor Kwang-Hyun Cho from the Department of Bio and Brain Engineering has developed a groundbreaking technology that can treat colon cancer by converting cancer cells into a state resembling normal colon cells without killing them, thus avoiding side effects.
The research team focused on the observation that during the oncogenesis process, normal cells regress along their differentiation trajectory. Building on this insight, they developed a technology to create a digital twin of the gene network associated with the differentiation trajectory of normal cells.
< Figure 1. Technology for creating a digital twin of a gene network from single-cell transcriptome data of a normal cell differentiation trajectory. Professor Kwang-Hyun Cho's research team developed a digital twin creation technology that precisely observes the dynamics of gene regulatory relationships during the process of normal cells differentiating along a differentiation trajectory and analyzes the relationships among key genes to build a mathematical model that can be simulated (A-F). In addition, they developed a technology to discover key regulatory factors that control the differentiation trajectory of normal cells by simulating and analyzing this digital twin. >
< Figure 2. Digital twin simulation simulating the differentiation trajectory of normal colon cells. The dynamics of single-cell transcriptome data for the differentiation trajectory of normal colon cells were analyzed (A) and a digital twin of the gene network was developed representing the regulatory relationships of key genes in this differentiation trajectory (B). The simulation results of the digital twin confirm that it readily reproduces the dynamics of single-cell transcriptome data (C, D). >
Through simulation analysis, the team systematically identified master molecular switches that induce normal cell differentiation. When these switches were applied to colon cancer cells, the cancer cells reverted to a normal-like state, a result confirmed through molecular and cellular experiments as well as animal studies.
< Figure 3. Discovery of top-level key control factors that induce differentiation of normal colon cells. By applying control factor discovery technology to the digital twin model, three genes, HDAC2, FOXA2, and MYB, were discovered as key control factors that induce differentiation of normal colon cells (A, B). The results of simulation analysis of the regulatory effects of the discovered control factors through the digital twin confirmed that they could induce complete differentiation of colon cells (C). >
< Figure 4. Verification of the effect of the key control factors discovered using colon cancer cells and animal experiments on the reversibility of colon cancer. The key control factors of the normal colon cell differentiation trajectory discovered through digital twin simulation analysis were applied to actual colon cancer cells and colon cancer mouse animal models to experimentally verify the effect of cancer reversibility. The key control factors significantly reduced the proliferation of three colon cancer cell lines (A), and this was confirmed in the same way in animal models (B-D). >
This research demonstrates that cancer cell reversion can be systematically achieved by analyzing and utilizing the digital twin of the cancer cell gene network, rather than relying on serendipitous discoveries. The findings hold significant promise for developing reversible cancer therapies that can be applied to various types of cancer.
< Figure 5. The change in overall gene expression was confirmed through the regulation of the identified key regulatory factors, which converted the state of colon cancer cells to that of normal colon cells. The transcriptomes of colon cancer tissues and normal colon tissues from more than 400 colon cancer patients were compared with the transcriptomes of colon cancer cell lines and reversible colon cancer cell lines, respectively. The comparison results confirmed that the regulation of the identified key regulatory factors converted all three colon cancer cell lines to a state similar to the transcriptome expression of normal colon tissues. >
Professor Kwang-Hyun Cho remarked, "The fact that cancer cells can be converted back to normal cells is an astonishing phenomenon. This study proves that such reversion can be systematically induced."
He further emphasized, "This research introduces the novel concept of reversible cancer therapy by reverting cancer cells to normal cells. It also develops foundational technology for identifying targets for cancer reversion through the systematic analysis of normal cell differentiation trajectories."
This research included contributions from Jeong-Ryeol Gong, Chun-Kyung Lee, Hoon-Min Kim, Juhee Kim, and Jaeog Jeon, and was published in the online edition of the international journal Advanced Science by Wiley on December 11. (Title: “Control of Cellular Differentiation Trajectories for Cancer Reversion”) DOI: https://doi.org/10.1002/advs.202402132
< Figure 6. Schematic diagram of the research results. Professor Kwang-Hyun Cho's research team developed a source technology to systematically discover key control factors that can induce reversibility of colon cancer cells through a systems biology approach and a digital twin simulation analysis of the differentiation trajectory of normal colon cells, and verified the effects of reversion on actual colon cancer through molecular cell experiments and animal experiments. >
The study was supported by the Ministry of Science and ICT and the National Research Foundation of Korea through the Mid-Career Researcher Program and Basic Research Laboratory Program. The research findings have been transferred to BioRevert Inc., where they will be used for the development of practical cancer reversion therapies.
2024.12.23 View 101548
KAIST Develops Foundational Technology to Revert Cancer Cells to Normal Cells
Despite the development of numerous cancer treatment technologies, the common goal of current cancer therapies is to eliminate cancer cells. This approach, however, faces fundamental limitations, including cancer cells developing resistance and returning, as well as severe side effects from the destruction of healthy cells.
< (From top left) Bio and Brain Engineering PhD candidates Juhee Kim, Jeong-Ryeol Gong, Chun-Kyung Lee, and Hoon-Min Kim posed for a group photo with Professor Kwang-Hyun Cho >
KAIST (represented by President Kwang Hyung Lee) announced on the 20th of December that a research team led by Professor Kwang-Hyun Cho from the Department of Bio and Brain Engineering has developed a groundbreaking technology that can treat colon cancer by converting cancer cells into a state resembling normal colon cells without killing them, thus avoiding side effects.
The research team focused on the observation that during the oncogenesis process, normal cells regress along their differentiation trajectory. Building on this insight, they developed a technology to create a digital twin of the gene network associated with the differentiation trajectory of normal cells.
< Figure 1. Technology for creating a digital twin of a gene network from single-cell transcriptome data of a normal cell differentiation trajectory. Professor Kwang-Hyun Cho's research team developed a digital twin creation technology that precisely observes the dynamics of gene regulatory relationships during the process of normal cells differentiating along a differentiation trajectory and analyzes the relationships among key genes to build a mathematical model that can be simulated (A-F). In addition, they developed a technology to discover key regulatory factors that control the differentiation trajectory of normal cells by simulating and analyzing this digital twin. >
< Figure 2. Digital twin simulation simulating the differentiation trajectory of normal colon cells. The dynamics of single-cell transcriptome data for the differentiation trajectory of normal colon cells were analyzed (A) and a digital twin of the gene network was developed representing the regulatory relationships of key genes in this differentiation trajectory (B). The simulation results of the digital twin confirm that it readily reproduces the dynamics of single-cell transcriptome data (C, D). >
Through simulation analysis, the team systematically identified master molecular switches that induce normal cell differentiation. When these switches were applied to colon cancer cells, the cancer cells reverted to a normal-like state, a result confirmed through molecular and cellular experiments as well as animal studies.
< Figure 3. Discovery of top-level key control factors that induce differentiation of normal colon cells. By applying control factor discovery technology to the digital twin model, three genes, HDAC2, FOXA2, and MYB, were discovered as key control factors that induce differentiation of normal colon cells (A, B). The results of simulation analysis of the regulatory effects of the discovered control factors through the digital twin confirmed that they could induce complete differentiation of colon cells (C). >
< Figure 4. Verification of the effect of the key control factors discovered using colon cancer cells and animal experiments on the reversibility of colon cancer. The key control factors of the normal colon cell differentiation trajectory discovered through digital twin simulation analysis were applied to actual colon cancer cells and colon cancer mouse animal models to experimentally verify the effect of cancer reversibility. The key control factors significantly reduced the proliferation of three colon cancer cell lines (A), and this was confirmed in the same way in animal models (B-D). >
This research demonstrates that cancer cell reversion can be systematically achieved by analyzing and utilizing the digital twin of the cancer cell gene network, rather than relying on serendipitous discoveries. The findings hold significant promise for developing reversible cancer therapies that can be applied to various types of cancer.
< Figure 5. The change in overall gene expression was confirmed through the regulation of the identified key regulatory factors, which converted the state of colon cancer cells to that of normal colon cells. The transcriptomes of colon cancer tissues and normal colon tissues from more than 400 colon cancer patients were compared with the transcriptomes of colon cancer cell lines and reversible colon cancer cell lines, respectively. The comparison results confirmed that the regulation of the identified key regulatory factors converted all three colon cancer cell lines to a state similar to the transcriptome expression of normal colon tissues. >
Professor Kwang-Hyun Cho remarked, "The fact that cancer cells can be converted back to normal cells is an astonishing phenomenon. This study proves that such reversion can be systematically induced."
He further emphasized, "This research introduces the novel concept of reversible cancer therapy by reverting cancer cells to normal cells. It also develops foundational technology for identifying targets for cancer reversion through the systematic analysis of normal cell differentiation trajectories."
This research included contributions from Jeong-Ryeol Gong, Chun-Kyung Lee, Hoon-Min Kim, Juhee Kim, and Jaeog Jeon, and was published in the online edition of the international journal Advanced Science by Wiley on December 11. (Title: “Control of Cellular Differentiation Trajectories for Cancer Reversion”) DOI: https://doi.org/10.1002/advs.202402132
< Figure 6. Schematic diagram of the research results. Professor Kwang-Hyun Cho's research team developed a source technology to systematically discover key control factors that can induce reversibility of colon cancer cells through a systems biology approach and a digital twin simulation analysis of the differentiation trajectory of normal colon cells, and verified the effects of reversion on actual colon cancer through molecular cell experiments and animal experiments. >
The study was supported by the Ministry of Science and ICT and the National Research Foundation of Korea through the Mid-Career Researcher Program and Basic Research Laboratory Program. The research findings have been transferred to BioRevert Inc., where they will be used for the development of practical cancer reversion therapies.
2024.12.23 View 101548 -
 KAIST Scientifically Identifies a Method to Prevent Dental Erosion from Carbonated Drinks
A Korean research team, which had previously visualized and scientifically proven the harmful effects of carbonated drinks like cola on dental health using nanotechnology, has now identified a mechanism for an effective method to prevent tooth damage caused by these beverages.
KAIST (represented by President Kwang Hyung Lee) announced on the 5th of December that a team led by Professor Seungbum Hong from the Department of Materials Science and Engineering, in collaboration with Seoul National University's School of Dentistry (Departments of Pediatric Dentistry and Oral Microbiology) and Professor Hye Ryung Byon’s research team from the Department of Chemistry, has revealed through nanotechnology that silver diamine fluoride (SDF)* forms a fluoride-containing protective layer on the tooth surface, effectively inhibiting cola-induced erosion.
*SDF (Silver Diamine Fluoride): A dental agent primarily used for the treatment and prevention of tooth decay. SDF strengthens carious lesions, suppresses bacterial growth, and halts the progression of cavities.
The team analyzed the surface morphology and mechanical properties of tooth enamel on a nanoscale using atomic force microscopy (AFM). They also examined the chemical properties of the nano-film formed by SDF treatment using X-ray photoelectron spectroscopy (XPS)* and Fourier-transform infrared spectroscopy (FTIR)*.
*XPS (X-ray Photoelectron Spectroscopy): A powerful surface analysis technique used to investigate the chemical composition and electronic structure of materials.
*FTIR (Fourier-Transform Infrared Spectroscopy): An analytical method that identifies the molecular structure and composition of materials by analyzing how they absorb or transmit infrared light.
The findings showed significant differences in surface roughness and elastic modulus between teeth exposed to cola with and without SDF treatment. Teeth treated with SDF exhibited minimal changes in surface roughness due to erosion (from 64 nm to 70 nm) and maintained a high elastic modulus (from 215 GPa to 205 GPa).
This was attributed to the formation of a fluoroapatite* layer by SDF, which acted as a protective shield.
*Fluoroapatite: A phosphate mineral with the chemical formula Ca₅(PO₄)₃F (calcium fluoro-phosphate). It can occur naturally or be synthesized biologically/artificially and plays a crucial role in strengthening teeth and bones.
< Figure 1. Schematic of the workflow. Surface morphology and mechanical properties of untreated and treated silver diamine fluoride (SDF) treated enamel exposed to cola were analyzed over time using atomic force microscopy (AFM). >
Professor Young J. Kim from Seoul National University's Department of Pediatric Dentistry noted, "This technology could be applied to prevent dental erosion and strengthen teeth for both children and adults. It is a cost-effective and accessible dental treatment."
< Figure 2. Changes in surface roughness and elastic modulus according to time of exposure to cola for SDF untreated and treated teeth. After 1 hour, the surface roughness of the SDF untreated teeth rapidly became rougher from 83 nm to 287 nm and the elastic modulus weakened from 125 GPa to 13 GPa, whereas the surface roughness of the SDF treated teeth changed only slightly from 64 nm to 70 nm and the elastic modulus barely changed from 215 GPa to 205 GPa, maintaining a similar state to the initial state. >
Professor Seungbum Hong emphasized, "Dental health significantly impacts quality of life. This research offers an effective non-invasive method to prevent early dental erosion, moving beyond traditional surgical treatments. By simply applying SDF, dental erosion can be prevented and enamel strengthened, potentially reducing pain and costs associated with treatment."
This study, led by the first author Aditi Saha, a PhD student in KAIST’s Department of Materials Science and Engineering, was published in the international journal Biomaterials Research on November 7 under the title "Nanoscale Study on Noninvasive Prevention of Dental Erosion of Enamel by Silver Diamine Fluoride". The research was supported by the National Research Foundation of Korea.
2024.12.11 View 5001
KAIST Scientifically Identifies a Method to Prevent Dental Erosion from Carbonated Drinks
A Korean research team, which had previously visualized and scientifically proven the harmful effects of carbonated drinks like cola on dental health using nanotechnology, has now identified a mechanism for an effective method to prevent tooth damage caused by these beverages.
KAIST (represented by President Kwang Hyung Lee) announced on the 5th of December that a team led by Professor Seungbum Hong from the Department of Materials Science and Engineering, in collaboration with Seoul National University's School of Dentistry (Departments of Pediatric Dentistry and Oral Microbiology) and Professor Hye Ryung Byon’s research team from the Department of Chemistry, has revealed through nanotechnology that silver diamine fluoride (SDF)* forms a fluoride-containing protective layer on the tooth surface, effectively inhibiting cola-induced erosion.
*SDF (Silver Diamine Fluoride): A dental agent primarily used for the treatment and prevention of tooth decay. SDF strengthens carious lesions, suppresses bacterial growth, and halts the progression of cavities.
The team analyzed the surface morphology and mechanical properties of tooth enamel on a nanoscale using atomic force microscopy (AFM). They also examined the chemical properties of the nano-film formed by SDF treatment using X-ray photoelectron spectroscopy (XPS)* and Fourier-transform infrared spectroscopy (FTIR)*.
*XPS (X-ray Photoelectron Spectroscopy): A powerful surface analysis technique used to investigate the chemical composition and electronic structure of materials.
*FTIR (Fourier-Transform Infrared Spectroscopy): An analytical method that identifies the molecular structure and composition of materials by analyzing how they absorb or transmit infrared light.
The findings showed significant differences in surface roughness and elastic modulus between teeth exposed to cola with and without SDF treatment. Teeth treated with SDF exhibited minimal changes in surface roughness due to erosion (from 64 nm to 70 nm) and maintained a high elastic modulus (from 215 GPa to 205 GPa).
This was attributed to the formation of a fluoroapatite* layer by SDF, which acted as a protective shield.
*Fluoroapatite: A phosphate mineral with the chemical formula Ca₅(PO₄)₃F (calcium fluoro-phosphate). It can occur naturally or be synthesized biologically/artificially and plays a crucial role in strengthening teeth and bones.
< Figure 1. Schematic of the workflow. Surface morphology and mechanical properties of untreated and treated silver diamine fluoride (SDF) treated enamel exposed to cola were analyzed over time using atomic force microscopy (AFM). >
Professor Young J. Kim from Seoul National University's Department of Pediatric Dentistry noted, "This technology could be applied to prevent dental erosion and strengthen teeth for both children and adults. It is a cost-effective and accessible dental treatment."
< Figure 2. Changes in surface roughness and elastic modulus according to time of exposure to cola for SDF untreated and treated teeth. After 1 hour, the surface roughness of the SDF untreated teeth rapidly became rougher from 83 nm to 287 nm and the elastic modulus weakened from 125 GPa to 13 GPa, whereas the surface roughness of the SDF treated teeth changed only slightly from 64 nm to 70 nm and the elastic modulus barely changed from 215 GPa to 205 GPa, maintaining a similar state to the initial state. >
Professor Seungbum Hong emphasized, "Dental health significantly impacts quality of life. This research offers an effective non-invasive method to prevent early dental erosion, moving beyond traditional surgical treatments. By simply applying SDF, dental erosion can be prevented and enamel strengthened, potentially reducing pain and costs associated with treatment."
This study, led by the first author Aditi Saha, a PhD student in KAIST’s Department of Materials Science and Engineering, was published in the international journal Biomaterials Research on November 7 under the title "Nanoscale Study on Noninvasive Prevention of Dental Erosion of Enamel by Silver Diamine Fluoride". The research was supported by the National Research Foundation of Korea.
2024.12.11 View 5001 -
 KAIST Unveils New Possibilities for Treating Intractable Brain Tumors
< Photo 1. (From left) Professor Heung Kyu Lee, KAIST Department of Biological Sciences, and Dr. Keun Bon Ku >
Immunotherapy, which enhances the immune system's T cell response to eliminate cancer cells, has emerged as a key approach in cancer treatment. However, in the case of glioblastoma, an aggressive and treatment-resistant brain tumor, numerous clinical trials have failed to confirm their efficacy. Korean researchers have recently analyzed the mechanisms that cause T cell exhaustion, which is characterized by a loss of function or a weakened response following prolonged exposure to antigens in such intractable cancers, identifying key control factors in T cell activation and clarifying the mechanisms that enhance therapeutic effectiveness.
KAIST (represented by President Kwang Hyung Lee) announced on the 6th of November that Professor Heung Kyu Lee’s team from the Department of Biological Sciences, in collaboration with the Korea Research Institute of Chemical Technology (represented by President Young Kuk Lee), has confirmed improved survival rates in a glioblastoma mouse model. By removing the inhibitory Fc gamma receptor (FcγRIIB), the research team was able to restore the responsiveness of cytotoxic T cells to immune checkpoint inhibitors, leading to enhanced anticancer activity.
The research team examined the effect of FcγRIIB, an inhibitory receptor recently found in cytotoxic T cells, on tumor-infiltrating T cells and the therapeutic effectiveness of the anti-PD-1 immune checkpoint inhibitor.
< Figure 1. Study results on improved survival rate due to increased antitumor activity of anti-PD-1 treatment in inhibitory Fc gamma receptor(Fcgr2b) ablation mice with murine glioblastoma. >
Their findings showed that deleting FcγRIIB induced the increase of tumor antigen-specific memory T cells, which helps to suppress exhaustion, enhances stem-like qualities, and reactivates T cell-mediated antitumor immunity, particularly in response to anti-PD-1 treatment. Furthermore, FcγRIIB deletion led to an increase in antigen-specific memory T cells that maintained continuous infiltration into the tumor tissue.
This study presents a new therapeutic target for tumors unresponsive to immune checkpoint inhibitors and demonstrates that combining FcγRIIB inhibition with anti-PD-1 treatment can produce synergistic effects, potentially improving therapeutic outcomes for tumors like glioblastoma, which typically show resistance to anti-PD-1 therapy.
< Figure 2. Overview of the study on the enhanced response to anti-PD-1 therapy for glioblastoma brain tumors upon deletion of the inhibitory Fc gamma receptor (FcγRIIB) in tumor microenvironment. When the inhibitory Fc gamma receptor (FcγRIIB) of cytotoxic T cells is deleted, an increase in tumor-specific memory T cells (Ttsms) was observed. In addition, this T cell subset is identified as originating from the tumor-draining lymph nodes(TdLNs) and leads to persistent infiltration into the tumor tissue. Anti-PD-1 therapy leads to an increased anti-tumor immune response via Ttsms, which is confirmed by increased tumor cell toxicity and increased cell division and decreased cell de-migration indices. Ultimately, the increased cytotoxic T cell immune response leads to an increase in the survival rate of glioblastoma. >
Professor Heung Kyu Lee explained, "This study offers a way to overcome clinical failures in treating brain tumors with immune checkpoint therapy and opens possibilities for broader applications to other intractable cancers. It also highlights the potential of utilizing cytotoxic T cells for tumor cell therapy."
The study, led by Dr. Keun Bon Ku of KAIST (currently a senior researcher at the Korea Research Institute of Chemical Technology's Center for Infectious Disease Diagnosis and Prevention), along with Chae Won Kim, Yumin Kim, Byeong Hoon Kang, Jeongwoo La, In Kang, Won Hyung Park, Stephen Ahn, and Sung Ki Lee, was published online on October 26 in the Journal for ImmunoTherapy of Cancer, an international journal in tumor immunology and therapy from the Society for Immunotherapy of Cancer. (Paper title: “Inhibitory Fcγ receptor deletion enhances CD8 T cell stemness increasing anti-PD-1 therapy responsiveness against glioblastoma,” http://dx.doi.org/10.1136/jitc-2024-009449).
This research received support from the National Research Foundation of Korea, the Bio & Medical Technology Development Program, and the Samsung Science & Technology Foundation.
2024.11.15 View 5239
KAIST Unveils New Possibilities for Treating Intractable Brain Tumors
< Photo 1. (From left) Professor Heung Kyu Lee, KAIST Department of Biological Sciences, and Dr. Keun Bon Ku >
Immunotherapy, which enhances the immune system's T cell response to eliminate cancer cells, has emerged as a key approach in cancer treatment. However, in the case of glioblastoma, an aggressive and treatment-resistant brain tumor, numerous clinical trials have failed to confirm their efficacy. Korean researchers have recently analyzed the mechanisms that cause T cell exhaustion, which is characterized by a loss of function or a weakened response following prolonged exposure to antigens in such intractable cancers, identifying key control factors in T cell activation and clarifying the mechanisms that enhance therapeutic effectiveness.
KAIST (represented by President Kwang Hyung Lee) announced on the 6th of November that Professor Heung Kyu Lee’s team from the Department of Biological Sciences, in collaboration with the Korea Research Institute of Chemical Technology (represented by President Young Kuk Lee), has confirmed improved survival rates in a glioblastoma mouse model. By removing the inhibitory Fc gamma receptor (FcγRIIB), the research team was able to restore the responsiveness of cytotoxic T cells to immune checkpoint inhibitors, leading to enhanced anticancer activity.
The research team examined the effect of FcγRIIB, an inhibitory receptor recently found in cytotoxic T cells, on tumor-infiltrating T cells and the therapeutic effectiveness of the anti-PD-1 immune checkpoint inhibitor.
< Figure 1. Study results on improved survival rate due to increased antitumor activity of anti-PD-1 treatment in inhibitory Fc gamma receptor(Fcgr2b) ablation mice with murine glioblastoma. >
Their findings showed that deleting FcγRIIB induced the increase of tumor antigen-specific memory T cells, which helps to suppress exhaustion, enhances stem-like qualities, and reactivates T cell-mediated antitumor immunity, particularly in response to anti-PD-1 treatment. Furthermore, FcγRIIB deletion led to an increase in antigen-specific memory T cells that maintained continuous infiltration into the tumor tissue.
This study presents a new therapeutic target for tumors unresponsive to immune checkpoint inhibitors and demonstrates that combining FcγRIIB inhibition with anti-PD-1 treatment can produce synergistic effects, potentially improving therapeutic outcomes for tumors like glioblastoma, which typically show resistance to anti-PD-1 therapy.
< Figure 2. Overview of the study on the enhanced response to anti-PD-1 therapy for glioblastoma brain tumors upon deletion of the inhibitory Fc gamma receptor (FcγRIIB) in tumor microenvironment. When the inhibitory Fc gamma receptor (FcγRIIB) of cytotoxic T cells is deleted, an increase in tumor-specific memory T cells (Ttsms) was observed. In addition, this T cell subset is identified as originating from the tumor-draining lymph nodes(TdLNs) and leads to persistent infiltration into the tumor tissue. Anti-PD-1 therapy leads to an increased anti-tumor immune response via Ttsms, which is confirmed by increased tumor cell toxicity and increased cell division and decreased cell de-migration indices. Ultimately, the increased cytotoxic T cell immune response leads to an increase in the survival rate of glioblastoma. >
Professor Heung Kyu Lee explained, "This study offers a way to overcome clinical failures in treating brain tumors with immune checkpoint therapy and opens possibilities for broader applications to other intractable cancers. It also highlights the potential of utilizing cytotoxic T cells for tumor cell therapy."
The study, led by Dr. Keun Bon Ku of KAIST (currently a senior researcher at the Korea Research Institute of Chemical Technology's Center for Infectious Disease Diagnosis and Prevention), along with Chae Won Kim, Yumin Kim, Byeong Hoon Kang, Jeongwoo La, In Kang, Won Hyung Park, Stephen Ahn, and Sung Ki Lee, was published online on October 26 in the Journal for ImmunoTherapy of Cancer, an international journal in tumor immunology and therapy from the Society for Immunotherapy of Cancer. (Paper title: “Inhibitory Fcγ receptor deletion enhances CD8 T cell stemness increasing anti-PD-1 therapy responsiveness against glioblastoma,” http://dx.doi.org/10.1136/jitc-2024-009449).
This research received support from the National Research Foundation of Korea, the Bio & Medical Technology Development Program, and the Samsung Science & Technology Foundation.
2024.11.15 View 5239 -
 KAIST Succeeds in the Real-time Observation of Organoids using Holotomography
Organoids, which are 3D miniature organs that mimic the structure and function of human organs, play an essential role in disease research and drug development. A Korean research team has overcome the limitations of existing imaging technologies, succeeding in the real-time, high-resolution observation of living organoids.
KAIST (represented by President Kwang Hyung Lee) announced on the 14th of October that Professor YongKeun Park’s research team from the Department of Physics, in collaboration with the Genome Editing Research Center (Director Bon-Kyoung Koo) of the Institute for Basic Science (IBS President Do-Young Noh) and Tomocube Inc., has developed an imaging technology using holotomography to observe live, small intestinal organoids in real time at a high resolution.
Existing imaging techniques have struggled to observe living organoids in high resolution over extended periods and often required additional treatments like fluorescent staining.
< Figure 1. Overview of the low-coherence HT workflow. Using holotomography, 3D morphological restoration and quantitative analysis of organoids can be performed. In order to improve the limited field of view, which is a limitation of the microscope, our research team utilized a large-area field of view combination algorithm and made a 3D restoration by acquiring multi-focus holographic images for 3D measurements. After that, the organoids were compartmentalized to divide the parts necessary for analysis and quantitatively evaluated the protein concentration measurable from the refractive index and the survival rate of the organoids. >
The research team introduced holotomography technology to address these issues, which provides high-resolution images without the need for fluorescent staining and allows for the long-term observation of dynamic changes in real time without causing cell damage.
The team validated this technology using small intestinal organoids from experimental mice and were able to observe various cell structures inside the organoids in detail. They also captured dynamic changes such as growth processes, cell division, and cell death in real time using holotomography.
Additionally, the technology allowed for the precise analysis of the organoids' responses to drug treatments, verifying the survival of the cells.
The researchers believe that this breakthrough will open new horizons in organoid research, enabling the greater utilization of organoids in drug development, personalized medicine, and regenerative medicine.
Future research is expected to more accurately replicate the in vivo environment of organoids, contributing significantly to a more detailed understanding of various life phenomena at the cellular level through more precise 3D imaging.
< Figure 2. Real-time organoid morphology analysis. Using holotomography, it is possible to observe the lumen and villus development process of intestinal organoids in real time, which was difficult to observe with a conventional microscope. In addition, various information about intestinal organoids can be obtained by quantifying the size and protein amount of intestinal organoids through image analysis. >
Dr. Mahn Jae Lee, a graduate of KAIST's Graduate School of Medical Science and Engineering, currently at Chungnam National University Hospital and the first author of the paper, commented, "This research represents a new imaging technology that surpasses previous limitations and is expected to make a major contribution to disease modeling, personalized treatments, and drug development research using organoids."
The research results were published online in the international journal Experimental & Molecular Medicine on October 1, 2024, and the technology has been recognized for its applicability in various fields of life sciences. (Paper title: “Long-term three-dimensional high-resolution imaging of live unlabeled small intestinal organoids via low-coherence holotomography”)
This research was supported by the National Research Foundation of Korea, KAIST Institutes, and the Institute for Basic Science.
2024.10.14 View 5057
KAIST Succeeds in the Real-time Observation of Organoids using Holotomography
Organoids, which are 3D miniature organs that mimic the structure and function of human organs, play an essential role in disease research and drug development. A Korean research team has overcome the limitations of existing imaging technologies, succeeding in the real-time, high-resolution observation of living organoids.
KAIST (represented by President Kwang Hyung Lee) announced on the 14th of October that Professor YongKeun Park’s research team from the Department of Physics, in collaboration with the Genome Editing Research Center (Director Bon-Kyoung Koo) of the Institute for Basic Science (IBS President Do-Young Noh) and Tomocube Inc., has developed an imaging technology using holotomography to observe live, small intestinal organoids in real time at a high resolution.
Existing imaging techniques have struggled to observe living organoids in high resolution over extended periods and often required additional treatments like fluorescent staining.
< Figure 1. Overview of the low-coherence HT workflow. Using holotomography, 3D morphological restoration and quantitative analysis of organoids can be performed. In order to improve the limited field of view, which is a limitation of the microscope, our research team utilized a large-area field of view combination algorithm and made a 3D restoration by acquiring multi-focus holographic images for 3D measurements. After that, the organoids were compartmentalized to divide the parts necessary for analysis and quantitatively evaluated the protein concentration measurable from the refractive index and the survival rate of the organoids. >
The research team introduced holotomography technology to address these issues, which provides high-resolution images without the need for fluorescent staining and allows for the long-term observation of dynamic changes in real time without causing cell damage.
The team validated this technology using small intestinal organoids from experimental mice and were able to observe various cell structures inside the organoids in detail. They also captured dynamic changes such as growth processes, cell division, and cell death in real time using holotomography.
Additionally, the technology allowed for the precise analysis of the organoids' responses to drug treatments, verifying the survival of the cells.
The researchers believe that this breakthrough will open new horizons in organoid research, enabling the greater utilization of organoids in drug development, personalized medicine, and regenerative medicine.
Future research is expected to more accurately replicate the in vivo environment of organoids, contributing significantly to a more detailed understanding of various life phenomena at the cellular level through more precise 3D imaging.
< Figure 2. Real-time organoid morphology analysis. Using holotomography, it is possible to observe the lumen and villus development process of intestinal organoids in real time, which was difficult to observe with a conventional microscope. In addition, various information about intestinal organoids can be obtained by quantifying the size and protein amount of intestinal organoids through image analysis. >
Dr. Mahn Jae Lee, a graduate of KAIST's Graduate School of Medical Science and Engineering, currently at Chungnam National University Hospital and the first author of the paper, commented, "This research represents a new imaging technology that surpasses previous limitations and is expected to make a major contribution to disease modeling, personalized treatments, and drug development research using organoids."
The research results were published online in the international journal Experimental & Molecular Medicine on October 1, 2024, and the technology has been recognized for its applicability in various fields of life sciences. (Paper title: “Long-term three-dimensional high-resolution imaging of live unlabeled small intestinal organoids via low-coherence holotomography”)
This research was supported by the National Research Foundation of Korea, KAIST Institutes, and the Institute for Basic Science.
2024.10.14 View 5057 -
 KAIST Changes the Paradigm of Drug Discovery with World's First Atomic Editing
In pioneering drug development, the new technology that enables the easy and rapid editing of key atoms responsible for drug efficacy has been regarded as a fundamental and "dream" technology, revolutionizing the process of discovering potential drug candidates. KAIST researchers have become the first in the world to successfully develop single-atom editing technology that maximizes drug efficacy.
On October 8th, KAIST (represented by President Kwang-Hyung Lee) announced that Professor Yoonsu Park’s research team from the Department of Chemistry successfully developed technology that enables the easy editing and correction of oxygen atoms in furan compounds into nitrogen atoms, directly converting them into pyrrole frameworks, which are widely used in pharmaceuticals.
< Image. Conceptual image illustrating the main idea of the research >
This research was published in the prestigious scientific journal Science on October 3rd under the title "Photocatalytic Furan-to-Pyrrole Conversion."
Many drugs have complex chemical structures, but their efficacy is often determined by a single critical atom. Atoms like oxygen and nitrogen play a central role in enhancing the pharmacological effects of these drugs, particularly against viruses.
This phenomenon, where the introduction of specific atoms into a drug molecule dramatically affects its efficacy, is known as the "Single Atom Effect." In leading-edge drug development, discovering atoms that maximize drug efficacy is key.
However, evaluating the Single Atom Effect has traditionally required multi-step, costly synthesis processes, as it has been difficult to selectively edit single atoms within stable ring structures containing oxygen or nitrogen.
Professor Park’s team overcame this challenge by introducing a photocatalyst that uses light energy. They developed a photocatalyst that acts as a “molecular scissor,” freely cutting and attaching five-membered rings, enabling single-atom editing at room temperature and atmospheric pressure—a world first.
The team discovered a new reaction mechanism in which the excited molecular scissor removes oxygen from furan via single-electron oxidation and then sequentially adds a nitrogen atom.
Donghyeon Kim and Jaehyun You, the study's first authors and candidates in KAIST’s integrated master's and doctoral program in the Department of Chemistry, explained that this technique offers high versatility by utilizing light energy to replace harsh conditions. They further noted that the technology enables selective editing, even when applied to complex natural products or pharmaceuticals. Professor Yoonsu Park, who led the research, remarked, "This breakthrough, which allows for the selective editing of five-membered organic ring structures, will open new doors for building libraries of drug candidates, a key challenge in pharmaceuticals. I hope this foundational technology will be used to revolutionize the drug development process."
The significance of this research was highlighted in the Perspective section of Science, a feature where a peer scientist of prominence outside of the project group provides commentary on an impactful research.
This research was supported by the National Research Foundation of Korea’s Creative Research Program, the Cross-Generation Collaborative Lab Project at KAIST, and the POSCO Science Fellowship of the POSCO TJ Park Foundation.
2024.10.11 View 5564
KAIST Changes the Paradigm of Drug Discovery with World's First Atomic Editing
In pioneering drug development, the new technology that enables the easy and rapid editing of key atoms responsible for drug efficacy has been regarded as a fundamental and "dream" technology, revolutionizing the process of discovering potential drug candidates. KAIST researchers have become the first in the world to successfully develop single-atom editing technology that maximizes drug efficacy.
On October 8th, KAIST (represented by President Kwang-Hyung Lee) announced that Professor Yoonsu Park’s research team from the Department of Chemistry successfully developed technology that enables the easy editing and correction of oxygen atoms in furan compounds into nitrogen atoms, directly converting them into pyrrole frameworks, which are widely used in pharmaceuticals.
< Image. Conceptual image illustrating the main idea of the research >
This research was published in the prestigious scientific journal Science on October 3rd under the title "Photocatalytic Furan-to-Pyrrole Conversion."
Many drugs have complex chemical structures, but their efficacy is often determined by a single critical atom. Atoms like oxygen and nitrogen play a central role in enhancing the pharmacological effects of these drugs, particularly against viruses.
This phenomenon, where the introduction of specific atoms into a drug molecule dramatically affects its efficacy, is known as the "Single Atom Effect." In leading-edge drug development, discovering atoms that maximize drug efficacy is key.
However, evaluating the Single Atom Effect has traditionally required multi-step, costly synthesis processes, as it has been difficult to selectively edit single atoms within stable ring structures containing oxygen or nitrogen.
Professor Park’s team overcame this challenge by introducing a photocatalyst that uses light energy. They developed a photocatalyst that acts as a “molecular scissor,” freely cutting and attaching five-membered rings, enabling single-atom editing at room temperature and atmospheric pressure—a world first.
The team discovered a new reaction mechanism in which the excited molecular scissor removes oxygen from furan via single-electron oxidation and then sequentially adds a nitrogen atom.
Donghyeon Kim and Jaehyun You, the study's first authors and candidates in KAIST’s integrated master's and doctoral program in the Department of Chemistry, explained that this technique offers high versatility by utilizing light energy to replace harsh conditions. They further noted that the technology enables selective editing, even when applied to complex natural products or pharmaceuticals. Professor Yoonsu Park, who led the research, remarked, "This breakthrough, which allows for the selective editing of five-membered organic ring structures, will open new doors for building libraries of drug candidates, a key challenge in pharmaceuticals. I hope this foundational technology will be used to revolutionize the drug development process."
The significance of this research was highlighted in the Perspective section of Science, a feature where a peer scientist of prominence outside of the project group provides commentary on an impactful research.
This research was supported by the National Research Foundation of Korea’s Creative Research Program, the Cross-Generation Collaborative Lab Project at KAIST, and the POSCO Science Fellowship of the POSCO TJ Park Foundation.
2024.10.11 View 5564 -
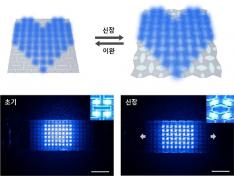 KAIST Develops Stretchable Displays Featuring 25% Expansion Without Image Distortion
Stretchable displays, praised for their spatial efficiency, design flexibility, and human-like flexibility, are seen as the next generation of display technology. A team of Korean researchers has developed a stretchable display that can expand by 25% while maintaining clear image quality without distortion. It can also stretch and contract up to 5,000 times at 15% expansion without any performance degradation, making it the first deformation-free stretchable display with a negative Poisson's ratio* developed in Korea.
*Poisson’s ratio of -1: A ratio where both width and length stretch equally, expressed as a negative value. A positive Poisson's ratio represents the ratio where horizontal stretching leads to vertical contraction, which is the case for most materials.
KAIST (represented by President Kwang-Hyung Lee) announced on the 20th of August that a research team led by Professor Byeong-Soo Bae of the Department of Materials Science and Engineering (Director of the Wearable Platform Materials Technology Center) , in collaboration with the Korea Institute of Machinery & Materials (President Seoghyeon Ryu), successfully developed a stretchable display substrate that suppresses image distortion through omnidirectional stretchability.
Currently, most stretchable displays are made with highly elastic elastomer* materials, but these materials possess a positive Poisson's ratio, causing unavoidable image distortion when the display is stretched.
*Elastomer: A polymer with elasticity similar to rubber.
To address this, the introduction of auxetic* meta-structures has been gaining attention. Unlike conventional materials, auxetic structures have a unique 'negative Poisson's ratio,' expanding in all directions when stretched in just one direction. However, traditional auxetic structures contain many empty spaces, limiting their stability and usability in display substrates.
*Auxetic structure: A special geometric structure that exhibits a negative Poisson's ratio.
To tackle the issue of image distortion, Professor Bae's research team developed a method to create a seamless surface for the auxetic meta-structure, achieving the ideal negative Poisson's ratio of -1 and overcoming the biggest challenge in auxetic meta-structures.
To overcome the second issue of elastic modulus*, the team inserted a textile made of glass fiber bundles with a diameter of just 25 micrometers (a quarter of the thickness of human hair) into the elastomer material. They then filled the empty spaces with the same elastomer, creating a flat and stable integrated film without gaps.
*Elastic Modulus: The ratio that indicates the extent of deformation when force is applied to a material. A higher elastic modulus means that the material is less likely to deform under force.
The research team theoretically identified that the difference in elasticity between the auxetic structure and the elastomer material directly influences the negative Poisson's ratio and successfully achieved an elasticity difference of over 230,000 times, producing a film with a Poisson's ratio of -1, the theoretical limit.
< Figure 2. Deformation of S-AUX film. a) Configurations and visualized principal strain distribution of the optimized S-AUX film at various strain rates. b) Biaxial stretching image. While pristine elastomer shrinks in the directions that were not stretched, S-AUX film developed in this study expands in all directions simultaneously while maintaining its original shape. >
Professor Byeong-Soo Bae, who led the study, explained, "Preventing image distortion using auxetic structures in stretchable displays is a core technology, but it has faced challenges due to the many empty spaces in the surface, making it difficult to use as a substrate. This research outcome is expected to significantly accelerate commercialization through high-resolution, distortion-free stretchable display applications that utilize the entire surface."
This study, co-authored by Dr. Yung Lee from KAIST’s Department of Materials Science and Engineering and Dr. Bongkyun Jang from the Korea Institute of Machinery & Materials, was published on August 20th in the international journal Nature Communications under the title "A seamless auxetic substrate with a negative Poisson's ratio of –1".
The research was supported by the Wearable Platform Materials Technology Center at KAIST, the Korea Institute of Machinery & Materials, and LG Display.
< Figure 3. Structural configuration of the distortion-free display components on the S-AUX film and a contour image of a micro-LED chip transferred onto the S-AUX film. >
< Figure 4. Schematic illustrations and photographic images of the S-AUX film-based image: distortion-free display in its stretched state and released state. >
2024.09.20 View 5843
KAIST Develops Stretchable Displays Featuring 25% Expansion Without Image Distortion
Stretchable displays, praised for their spatial efficiency, design flexibility, and human-like flexibility, are seen as the next generation of display technology. A team of Korean researchers has developed a stretchable display that can expand by 25% while maintaining clear image quality without distortion. It can also stretch and contract up to 5,000 times at 15% expansion without any performance degradation, making it the first deformation-free stretchable display with a negative Poisson's ratio* developed in Korea.
*Poisson’s ratio of -1: A ratio where both width and length stretch equally, expressed as a negative value. A positive Poisson's ratio represents the ratio where horizontal stretching leads to vertical contraction, which is the case for most materials.
KAIST (represented by President Kwang-Hyung Lee) announced on the 20th of August that a research team led by Professor Byeong-Soo Bae of the Department of Materials Science and Engineering (Director of the Wearable Platform Materials Technology Center) , in collaboration with the Korea Institute of Machinery & Materials (President Seoghyeon Ryu), successfully developed a stretchable display substrate that suppresses image distortion through omnidirectional stretchability.
Currently, most stretchable displays are made with highly elastic elastomer* materials, but these materials possess a positive Poisson's ratio, causing unavoidable image distortion when the display is stretched.
*Elastomer: A polymer with elasticity similar to rubber.
To address this, the introduction of auxetic* meta-structures has been gaining attention. Unlike conventional materials, auxetic structures have a unique 'negative Poisson's ratio,' expanding in all directions when stretched in just one direction. However, traditional auxetic structures contain many empty spaces, limiting their stability and usability in display substrates.
*Auxetic structure: A special geometric structure that exhibits a negative Poisson's ratio.
To tackle the issue of image distortion, Professor Bae's research team developed a method to create a seamless surface for the auxetic meta-structure, achieving the ideal negative Poisson's ratio of -1 and overcoming the biggest challenge in auxetic meta-structures.
To overcome the second issue of elastic modulus*, the team inserted a textile made of glass fiber bundles with a diameter of just 25 micrometers (a quarter of the thickness of human hair) into the elastomer material. They then filled the empty spaces with the same elastomer, creating a flat and stable integrated film without gaps.
*Elastic Modulus: The ratio that indicates the extent of deformation when force is applied to a material. A higher elastic modulus means that the material is less likely to deform under force.
The research team theoretically identified that the difference in elasticity between the auxetic structure and the elastomer material directly influences the negative Poisson's ratio and successfully achieved an elasticity difference of over 230,000 times, producing a film with a Poisson's ratio of -1, the theoretical limit.
< Figure 2. Deformation of S-AUX film. a) Configurations and visualized principal strain distribution of the optimized S-AUX film at various strain rates. b) Biaxial stretching image. While pristine elastomer shrinks in the directions that were not stretched, S-AUX film developed in this study expands in all directions simultaneously while maintaining its original shape. >
Professor Byeong-Soo Bae, who led the study, explained, "Preventing image distortion using auxetic structures in stretchable displays is a core technology, but it has faced challenges due to the many empty spaces in the surface, making it difficult to use as a substrate. This research outcome is expected to significantly accelerate commercialization through high-resolution, distortion-free stretchable display applications that utilize the entire surface."
This study, co-authored by Dr. Yung Lee from KAIST’s Department of Materials Science and Engineering and Dr. Bongkyun Jang from the Korea Institute of Machinery & Materials, was published on August 20th in the international journal Nature Communications under the title "A seamless auxetic substrate with a negative Poisson's ratio of –1".
The research was supported by the Wearable Platform Materials Technology Center at KAIST, the Korea Institute of Machinery & Materials, and LG Display.
< Figure 3. Structural configuration of the distortion-free display components on the S-AUX film and a contour image of a micro-LED chip transferred onto the S-AUX film. >
< Figure 4. Schematic illustrations and photographic images of the S-AUX film-based image: distortion-free display in its stretched state and released state. >
2024.09.20 View 5843