Department+of+Biological+Science
-
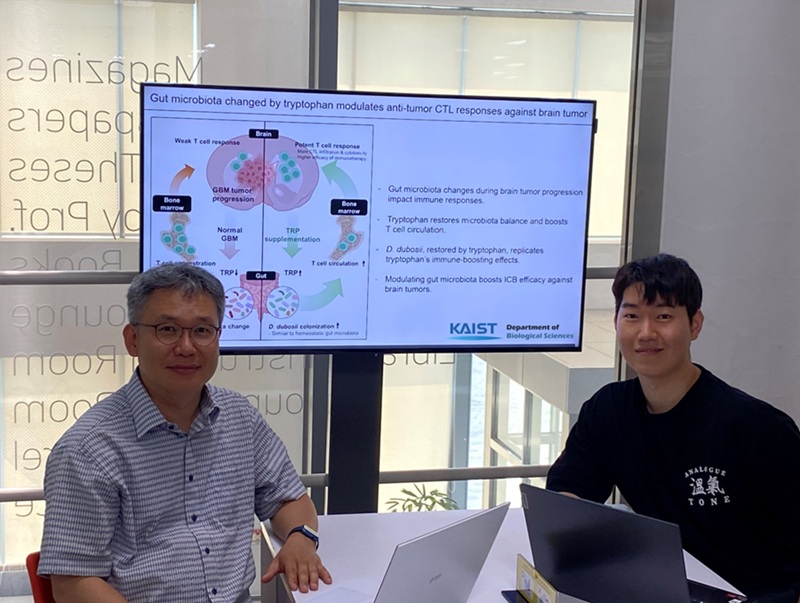 KAIST Enhances Immunotherapy for Difficult-to-Treat Brain Tumors with Gut Microbiota
< Photo 1.(From left) Prof. Heung Kyu Lee, Department of Biological Sciences,
and Dr. Hyeon Cheol Kim>
Advanced treatments, known as immunotherapies that activate T cells—our body's immune cells—to eliminate cancer cells, have shown limited efficacy as standalone therapies for glioblastoma, the most lethal form of brain tumor. This is due to their minimal response to glioblastoma and high resistance to treatment.
Now, a KAIST research team has now demonstrated a new therapeutic strategy that can enhance the efficacy of immunotherapy for brain tumors by utilizing gut microbes and their metabolites. This also opens up possibilities for developing microbiome-based immunotherapy supplements in the future.
KAIST (President Kwang Hyung Lee) announced on July 1 that a research team led by Professor Heung Kyu Lee of the Department of Biological Sciences discovered and demonstrated a method to significantly improve the efficiency of glioblastoma immunotherapy by focusing on changes in the gut microbial ecosystem.
The research team noted that as glioblastoma progresses, the concentration of ‘tryptophan’, an important amino acid in the gut, sharply decreases, leading to changes in the gut microbial ecosystem. They discovered that by supplementing tryptophan to restore microbial diversity, specific beneficial strains activate CD8 T cells (a type of immune cell) and induce their infiltration into tumor tissues. Through a mouse model of glioblastoma, the research team confirmed that tryptophan supplementation enhanced the response of cancer-attacking T cells (especially CD8 T cells), leading to their increased migration to tumor sites such as lymph nodes and the brain.
In this process, they also revealed that ‘Duncaniella dubosii’, a beneficial commensal bacterium present in the gut, plays a crucial role. This bacterium helped T cells effectively redistribute within the body, and survival rates significantly improved when used in combination with immunotherapy (anti-PD-1).
Furthermore, it was demonstrated that even when this commensal bacterium was administered alone to germ-free mice (mice without any commensal microbes), the survival rate for glioblastoma increased. This is because the bacterium utilizes tryptophan to regulate the gut environment, and the metabolites produced in this process strengthen the ability of CD8 T cells to attack cancer cells.
Professor Heung Kyu Lee explained, "This research is a meaningful achievement, showing that even in intractable brain tumors where immune checkpoint inhibitors had no effect, a combined strategy utilizing gut microbes can significantly enhance treatment response."
Dr. Hyeon Cheol Kim of KAIST (currently a postdoctoral researcher at the Institute for Biological Sciences) participated as the first author. The research findings were published online in Cell Reports, an international journal in the life sciences, on June 26.
This research was conducted as part of the Basic Research Program and Bio & Medical Technology Development Program supported by the Ministry of Science and ICT and the National Research Foundation of Korea.
※Paper Title: Gut microbiota dysbiosis induced by brain tumor modulates the efficacy of immunotherapy
※DOI: https://doi.org/10.1016/j.celrep.2025.115825
2025.07.02 View 46
KAIST Enhances Immunotherapy for Difficult-to-Treat Brain Tumors with Gut Microbiota
< Photo 1.(From left) Prof. Heung Kyu Lee, Department of Biological Sciences,
and Dr. Hyeon Cheol Kim>
Advanced treatments, known as immunotherapies that activate T cells—our body's immune cells—to eliminate cancer cells, have shown limited efficacy as standalone therapies for glioblastoma, the most lethal form of brain tumor. This is due to their minimal response to glioblastoma and high resistance to treatment.
Now, a KAIST research team has now demonstrated a new therapeutic strategy that can enhance the efficacy of immunotherapy for brain tumors by utilizing gut microbes and their metabolites. This also opens up possibilities for developing microbiome-based immunotherapy supplements in the future.
KAIST (President Kwang Hyung Lee) announced on July 1 that a research team led by Professor Heung Kyu Lee of the Department of Biological Sciences discovered and demonstrated a method to significantly improve the efficiency of glioblastoma immunotherapy by focusing on changes in the gut microbial ecosystem.
The research team noted that as glioblastoma progresses, the concentration of ‘tryptophan’, an important amino acid in the gut, sharply decreases, leading to changes in the gut microbial ecosystem. They discovered that by supplementing tryptophan to restore microbial diversity, specific beneficial strains activate CD8 T cells (a type of immune cell) and induce their infiltration into tumor tissues. Through a mouse model of glioblastoma, the research team confirmed that tryptophan supplementation enhanced the response of cancer-attacking T cells (especially CD8 T cells), leading to their increased migration to tumor sites such as lymph nodes and the brain.
In this process, they also revealed that ‘Duncaniella dubosii’, a beneficial commensal bacterium present in the gut, plays a crucial role. This bacterium helped T cells effectively redistribute within the body, and survival rates significantly improved when used in combination with immunotherapy (anti-PD-1).
Furthermore, it was demonstrated that even when this commensal bacterium was administered alone to germ-free mice (mice without any commensal microbes), the survival rate for glioblastoma increased. This is because the bacterium utilizes tryptophan to regulate the gut environment, and the metabolites produced in this process strengthen the ability of CD8 T cells to attack cancer cells.
Professor Heung Kyu Lee explained, "This research is a meaningful achievement, showing that even in intractable brain tumors where immune checkpoint inhibitors had no effect, a combined strategy utilizing gut microbes can significantly enhance treatment response."
Dr. Hyeon Cheol Kim of KAIST (currently a postdoctoral researcher at the Institute for Biological Sciences) participated as the first author. The research findings were published online in Cell Reports, an international journal in the life sciences, on June 26.
This research was conducted as part of the Basic Research Program and Bio & Medical Technology Development Program supported by the Ministry of Science and ICT and the National Research Foundation of Korea.
※Paper Title: Gut microbiota dysbiosis induced by brain tumor modulates the efficacy of immunotherapy
※DOI: https://doi.org/10.1016/j.celrep.2025.115825
2025.07.02 View 46 -
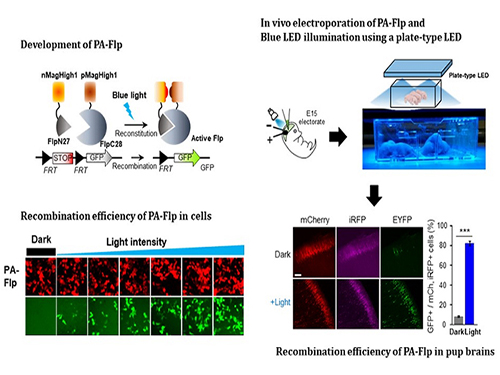 Noninvasive Light-Sensitive Recombinase for Deep Brain Genetic Manipulation
A KAIST team presented a noninvasive light-sensitive photoactivatable recombinase suitable for genetic manipulation in vivo. The highly light-sensitive property of photoactivatable Flp recombinase will be ideal for controlling genetic manipulation in deep mouse brain regions by illumination with a noninvasive light-emitting diode. This easy-to-use optogenetic module made by Professor Won Do Heo and his team will provide a side-effect free and expandable genetic manipulation tool for neuroscience research.
Spatiotemporal control of gene expression has been acclaimed as a valuable strategy for identifying functions of genes with complex neural circuits. Studies of complex brain functions require highly sophisticated and robust technologies that enable specific labeling and rapid genetic modification in live animals. A number of approaches for controlling the activity of proteins or expression of genes in a spatiotemporal manner using light, small molecules, hormones, and peptides have been developed for manipulating intact circuits or functions.
Among them, recombination-employing, chemically inducible systems are the most commonly used in vivo gene-modification systems. Other approaches include selective or conditional Cre-activation systems within subsets of green fluorescent protein-expressing cells or dual-promoter-driven intersectional populations of cells.
However, these methods are limited by the considerable time and effort required to establish knock-in mouse lines and by constraints on spatiotemporal control, which relies on a limited set of available genetic promoters and transgenic mouse resources.
Beyond these constraints, optogenetic approaches allow the activity of genetically defined neurons in the mouse brain to be controlled with high spatiotemporal resolution. However, an optogenetic module for gene-manipulation capable of revealing the spatiotemporal functions of specific target genes in the mouse brain has remained a challenge.
In the study published at Nature Communication on Jan. 18, the team featured photoactivatable Flp recombinase by searching out split sites of Flp recombinase that were not previously identified, being capable of reconstitution to be active. The team validated the highly light-sensitive, efficient performance of photoactivatable Flp recombinase through precise light targeting by showing transgene expression within anatomically confined mouse brain regions.
The concept of local genetic labeling presented here suggests a new approach for genetically identifying subpopulations of cells defined by the spatial and temporal characteristics of light delivery. To date, an optogenetic module for gene-manipulation capable of revealing spatiotemporal functions of specific target genes in the mouse brain has remained out of reach and no such light-inducible Flp system has been developed. Accordingly, the team sought to develop a photoactivatable Flp recombinase that takes full advantage of the high spatiotemporal control offered by light stimulation.
This activation through noninvasive light illumination deep inside the brain is advantageous in that it avoids chemical or optic fiber implantation-mediated side effects, such as off-target cytotoxicity or physical lesions that might influence animal physiology or behaviors. The technique provides expandable utilities for transgene expression systems upon Flp recombinase activity in vivo, by designing a viral vector for minimal leaky expression influenced by viral nascent promoters.
The team demonstrated the utility of PA-Flp as a noninvasive in vivo optogenetic manipulation tool for use in the mouse brain, even applicable for deep brain structures as it can reach the hippocampus or medial septum using external LED light illumination.
The study is the result of five years of research by Professor Heo, who has led the bio-imaging and optogenetics fields by developing his own bio-imaging and optogenetics technologies. “It will be a great advantage to control specific gene expression desired by LEDs with little physical and chemical stimulation that can affect the physiological phenomenon in living animals,” he explained.
2019.01.22 View 7974
Noninvasive Light-Sensitive Recombinase for Deep Brain Genetic Manipulation
A KAIST team presented a noninvasive light-sensitive photoactivatable recombinase suitable for genetic manipulation in vivo. The highly light-sensitive property of photoactivatable Flp recombinase will be ideal for controlling genetic manipulation in deep mouse brain regions by illumination with a noninvasive light-emitting diode. This easy-to-use optogenetic module made by Professor Won Do Heo and his team will provide a side-effect free and expandable genetic manipulation tool for neuroscience research.
Spatiotemporal control of gene expression has been acclaimed as a valuable strategy for identifying functions of genes with complex neural circuits. Studies of complex brain functions require highly sophisticated and robust technologies that enable specific labeling and rapid genetic modification in live animals. A number of approaches for controlling the activity of proteins or expression of genes in a spatiotemporal manner using light, small molecules, hormones, and peptides have been developed for manipulating intact circuits or functions.
Among them, recombination-employing, chemically inducible systems are the most commonly used in vivo gene-modification systems. Other approaches include selective or conditional Cre-activation systems within subsets of green fluorescent protein-expressing cells or dual-promoter-driven intersectional populations of cells.
However, these methods are limited by the considerable time and effort required to establish knock-in mouse lines and by constraints on spatiotemporal control, which relies on a limited set of available genetic promoters and transgenic mouse resources.
Beyond these constraints, optogenetic approaches allow the activity of genetically defined neurons in the mouse brain to be controlled with high spatiotemporal resolution. However, an optogenetic module for gene-manipulation capable of revealing the spatiotemporal functions of specific target genes in the mouse brain has remained a challenge.
In the study published at Nature Communication on Jan. 18, the team featured photoactivatable Flp recombinase by searching out split sites of Flp recombinase that were not previously identified, being capable of reconstitution to be active. The team validated the highly light-sensitive, efficient performance of photoactivatable Flp recombinase through precise light targeting by showing transgene expression within anatomically confined mouse brain regions.
The concept of local genetic labeling presented here suggests a new approach for genetically identifying subpopulations of cells defined by the spatial and temporal characteristics of light delivery. To date, an optogenetic module for gene-manipulation capable of revealing spatiotemporal functions of specific target genes in the mouse brain has remained out of reach and no such light-inducible Flp system has been developed. Accordingly, the team sought to develop a photoactivatable Flp recombinase that takes full advantage of the high spatiotemporal control offered by light stimulation.
This activation through noninvasive light illumination deep inside the brain is advantageous in that it avoids chemical or optic fiber implantation-mediated side effects, such as off-target cytotoxicity or physical lesions that might influence animal physiology or behaviors. The technique provides expandable utilities for transgene expression systems upon Flp recombinase activity in vivo, by designing a viral vector for minimal leaky expression influenced by viral nascent promoters.
The team demonstrated the utility of PA-Flp as a noninvasive in vivo optogenetic manipulation tool for use in the mouse brain, even applicable for deep brain structures as it can reach the hippocampus or medial septum using external LED light illumination.
The study is the result of five years of research by Professor Heo, who has led the bio-imaging and optogenetics fields by developing his own bio-imaging and optogenetics technologies. “It will be a great advantage to control specific gene expression desired by LEDs with little physical and chemical stimulation that can affect the physiological phenomenon in living animals,” he explained.
2019.01.22 View 7974 -
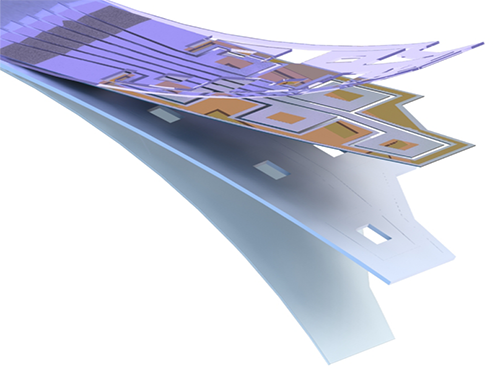 Flexible Drug Delivery Microdevice to Advance Precision Medicine
(Schematic view of flexible microdevice: The flexible drug delivery device for controlled release fabricated via inorganic laser lift off.)
A KAIST research team has developed a flexible drug delivery device with controlled release for personalized medicine, blazing the path toward theragnosis.
Theragnosis, an emerging medical technology, is gaining attention as key factor to advance precision medicine for its featuring simultaneous diagnosis and therapeutics. Theragnosis devices including smart contact lenses and microneedle patches integrate physiological data sensors and drug delivery devices. The controlled drug delivery boasts fewer side-effects, uniform therapeutic results, and minimal dosages compared to oral ingestion. Recently, some research groups conducted in-human applications of controlled-release bulky microchips for osteoporosis treatment. However they failed to demonstrate successful human-friendly flexible drug delivery systems for controlled release.
For this microdevice, the team under Professor Daesoo Kim from the Department of Biological Science and Professor Keon Jae Lee from the Department of Materials Science and Engineering, fabricated a device on a rigid substrate and transferred a 50 µm-thick active drug delivery layer to the flexible substrate via inorganic laser lift off. The fabricated device shows mechanical flexibility while maintaining the capability of precise administration of exact dosages at desired times. The core technology is to produce a freestanding gold capping layer directly on top of the microreservoir with the drugs inside, which had been regarded as impossible in conventional microfabrication.
The developed flexible drug delivery system can be applied to smart contact lenses or the brain disease treatments by implanting them into cramped and corrugated organs. In addition, when powered wirelessly, it will represent a novel platform for personalized medicine. The team already proved through animal experimentation that treatment for brain epilepsy made progress by releasing anti-epileptic medication through the device.
Professor Lee believes the flexible microdevice will further expand the applications of smart contact lenses, therapeutic treatments for brain disease, and subcutaneous implantations for daily healthcare system. This study “Flexible Wireless Powered Drug Delivery System for Targeted Administration on Cerebral Cortex” was described in the June online issue of Nano Energy.
(Photo: The flexible drug delivery device for contolled relase attached on a glass rod.)
2018.08.13 View 9857
Flexible Drug Delivery Microdevice to Advance Precision Medicine
(Schematic view of flexible microdevice: The flexible drug delivery device for controlled release fabricated via inorganic laser lift off.)
A KAIST research team has developed a flexible drug delivery device with controlled release for personalized medicine, blazing the path toward theragnosis.
Theragnosis, an emerging medical technology, is gaining attention as key factor to advance precision medicine for its featuring simultaneous diagnosis and therapeutics. Theragnosis devices including smart contact lenses and microneedle patches integrate physiological data sensors and drug delivery devices. The controlled drug delivery boasts fewer side-effects, uniform therapeutic results, and minimal dosages compared to oral ingestion. Recently, some research groups conducted in-human applications of controlled-release bulky microchips for osteoporosis treatment. However they failed to demonstrate successful human-friendly flexible drug delivery systems for controlled release.
For this microdevice, the team under Professor Daesoo Kim from the Department of Biological Science and Professor Keon Jae Lee from the Department of Materials Science and Engineering, fabricated a device on a rigid substrate and transferred a 50 µm-thick active drug delivery layer to the flexible substrate via inorganic laser lift off. The fabricated device shows mechanical flexibility while maintaining the capability of precise administration of exact dosages at desired times. The core technology is to produce a freestanding gold capping layer directly on top of the microreservoir with the drugs inside, which had been regarded as impossible in conventional microfabrication.
The developed flexible drug delivery system can be applied to smart contact lenses or the brain disease treatments by implanting them into cramped and corrugated organs. In addition, when powered wirelessly, it will represent a novel platform for personalized medicine. The team already proved through animal experimentation that treatment for brain epilepsy made progress by releasing anti-epileptic medication through the device.
Professor Lee believes the flexible microdevice will further expand the applications of smart contact lenses, therapeutic treatments for brain disease, and subcutaneous implantations for daily healthcare system. This study “Flexible Wireless Powered Drug Delivery System for Targeted Administration on Cerebral Cortex” was described in the June online issue of Nano Energy.
(Photo: The flexible drug delivery device for contolled relase attached on a glass rod.)
2018.08.13 View 9857 -
 How to Trigger Innate Fear Response?
(Figure:This illustration describes how ACC-BLA circuit controls innate freezing response depending on its activity level.)
When animals encounter danger, they usually respond to the situation in one of two ways: to freeze or to flee. How do they make this quick decision in a life or death moment?
According to KAIST neuroscientists, there are two types of fear: learned versus innate. The latter is known to be induced without any prior experience and is thus naturally encoded in the brain. A research team under Professor Jin-Hee Han in the Department of Biological Sciences identified the brain circuit responsible for regulating the innate fear response.
The study, which appeared in the July 24 issue of Nature Communications represents a significant step toward understanding how the neural circuits in the prefrontal cortex create behavioral responses to external threats. This also represents a new paradigm in therapeutic development for fear-related mental disorders.
Responses of freezing or fleeing when facing external threats reflect behavioral and physiological changes in an instinctive move to adapt to the new environment for survival. These responses are controlled by the emotional circuit systems of the brain and the malfunction of this circuit leads to fear-related disorders.
The anterior cingulate cortex (ACC) is a sub-region within the prefrontal cortex, comprising a part of the brain circuitry that regulates behavioral and physiological fear responses. This area is capable of high-order processing of the perceived sensory information and conveys ‘top-down’ information toward the amygdala and brainstem areas, known as the response outlet.
Many studies have already demonstrated that the brain regions in the prefrontal cortex regulate the response against learned threats. However, it has been unknown how innate responses against fear are encoded in the neural circuits in the prefrontal cortex.
Dr. Jinho Jhang, the lead author of the study explains how the team achieved their key idea. “Many overseas studies have already proved that the prefrontal cortex circuit works to regulate the fear response. However, researchers have paid little attention to the innate response against predators. Professor Han suggested we do research on the instinctive fear response instead of the learned response. We particularly focused on the anterior cingulate region, which has been connected with memory, pain, and sympathy, but not the fear response itself. Since we turned in this new direction, we have accumulated some significant data,” said Dr. Jhang.
For this study, Professor Han’s team investigated how mice react when exposed to the olfactory stimuli of predators. Based on the results of optogenetic manipulation, neural circuit tracing, and ex vivo slice electrophysiology experiments, the team demonstrated that the anterior cingulate cortex and its projection input to the basolateral amygdala play a role in the inhibitory regulation of innate fear responses to predators’ odors in mice.
Professor Han believes these results will extend the understanding of how instinctive fear responses can be encoded in our brain circuits. “Our findings will help to develop therapeutic treatments for mental disorders aroused from fear such as panic disorders and post-traumatic stress disorder,” said Professor Han.
2018.08.08 View 8364
How to Trigger Innate Fear Response?
(Figure:This illustration describes how ACC-BLA circuit controls innate freezing response depending on its activity level.)
When animals encounter danger, they usually respond to the situation in one of two ways: to freeze or to flee. How do they make this quick decision in a life or death moment?
According to KAIST neuroscientists, there are two types of fear: learned versus innate. The latter is known to be induced without any prior experience and is thus naturally encoded in the brain. A research team under Professor Jin-Hee Han in the Department of Biological Sciences identified the brain circuit responsible for regulating the innate fear response.
The study, which appeared in the July 24 issue of Nature Communications represents a significant step toward understanding how the neural circuits in the prefrontal cortex create behavioral responses to external threats. This also represents a new paradigm in therapeutic development for fear-related mental disorders.
Responses of freezing or fleeing when facing external threats reflect behavioral and physiological changes in an instinctive move to adapt to the new environment for survival. These responses are controlled by the emotional circuit systems of the brain and the malfunction of this circuit leads to fear-related disorders.
The anterior cingulate cortex (ACC) is a sub-region within the prefrontal cortex, comprising a part of the brain circuitry that regulates behavioral and physiological fear responses. This area is capable of high-order processing of the perceived sensory information and conveys ‘top-down’ information toward the amygdala and brainstem areas, known as the response outlet.
Many studies have already demonstrated that the brain regions in the prefrontal cortex regulate the response against learned threats. However, it has been unknown how innate responses against fear are encoded in the neural circuits in the prefrontal cortex.
Dr. Jinho Jhang, the lead author of the study explains how the team achieved their key idea. “Many overseas studies have already proved that the prefrontal cortex circuit works to regulate the fear response. However, researchers have paid little attention to the innate response against predators. Professor Han suggested we do research on the instinctive fear response instead of the learned response. We particularly focused on the anterior cingulate region, which has been connected with memory, pain, and sympathy, but not the fear response itself. Since we turned in this new direction, we have accumulated some significant data,” said Dr. Jhang.
For this study, Professor Han’s team investigated how mice react when exposed to the olfactory stimuli of predators. Based on the results of optogenetic manipulation, neural circuit tracing, and ex vivo slice electrophysiology experiments, the team demonstrated that the anterior cingulate cortex and its projection input to the basolateral amygdala play a role in the inhibitory regulation of innate fear responses to predators’ odors in mice.
Professor Han believes these results will extend the understanding of how instinctive fear responses can be encoded in our brain circuits. “Our findings will help to develop therapeutic treatments for mental disorders aroused from fear such as panic disorders and post-traumatic stress disorder,” said Professor Han.
2018.08.08 View 8364 -
 Animal Cyborg: Behavioral Control by 'Toy' Craving Circuit
Children love to get toys from parents for their birthday present. This craving toward items also involves object hoarding disorders and shopping addiction. However, the biological meaning of why the brain pursues objects or items has remained unknown. Part of the answer may lie with a neural circuit in the hypothalamus associated with “object craving,” says neuroscientist Daesoo Kim from the Department of Biological Sciences at KAIST.
His research team found that some neurons in the hypothalamus are activated during playing with toys in mice. Thanks to optogenetics, they proved that these neurons in the hypothalamus actually governs obsessive behavior toward non-food objects in mice.
“When we stimulate a neuron in the hypothalamus of mice, they anxiously chased target objects. We found evidence that the neural circuits in the medial preoptic area (MPA) modulate “object craving,” the appetite for possessing objects” said Professor Kim.
Researchers also proved that the MPA circuit facilitate hunting behavior in response to crickets, a natural prey to mice, showing the role of this circuit for catching prey. Further, the MPA nerves send excitatory signals to the periaqueductal gray (PAG), located around the cerebral aqueduct, to create such behavior. The team named this circuit the ‘MPA-PAG’ circuit.
The team showed that they could control mammalian behavior for the first time with this scheme of MPA-Induced Drive Assisted Steering (MIDAS), in which a mouse chase the target objects in the front of head during stimulation of the MPA-PAG circuit. MIDAS allows mice to overcome obstacles to move in a desired path using optogenetics.
(Professor Daesoo Kim)
Professor Kim, who teamed up with Professor Phill Seung Lee in the Department of Mechanical Engineering, explained the significance of the research, “This study provides evidence to treat brain disorders such as compulsive hoarding and kleptomania. It also contributes to the development of technology to control the behavior of animals and humans using strong innate motivation, and thus could impact neuro-economics, defense, and disaster relief.”
He said the team would like to complete the neural circuit map governing behaviors of possession and hunting in the near future by exploring correlations with other neural behaviors controlling possessing and hunting activities.
This research was funded by the Samsung Science and Technology Foundation and published in Nature Neuroscience in March 2018.
(Figure 1: Schematics showing possessive behavior induced by the MPA neural circuit)
(Figure 2: Schematics of the MIDAS system that controls mammals behavior using the desire to possess. A MIDAS mouse is following the bait object controlled wirelessly.)
2018.04.23 View 10524
Animal Cyborg: Behavioral Control by 'Toy' Craving Circuit
Children love to get toys from parents for their birthday present. This craving toward items also involves object hoarding disorders and shopping addiction. However, the biological meaning of why the brain pursues objects or items has remained unknown. Part of the answer may lie with a neural circuit in the hypothalamus associated with “object craving,” says neuroscientist Daesoo Kim from the Department of Biological Sciences at KAIST.
His research team found that some neurons in the hypothalamus are activated during playing with toys in mice. Thanks to optogenetics, they proved that these neurons in the hypothalamus actually governs obsessive behavior toward non-food objects in mice.
“When we stimulate a neuron in the hypothalamus of mice, they anxiously chased target objects. We found evidence that the neural circuits in the medial preoptic area (MPA) modulate “object craving,” the appetite for possessing objects” said Professor Kim.
Researchers also proved that the MPA circuit facilitate hunting behavior in response to crickets, a natural prey to mice, showing the role of this circuit for catching prey. Further, the MPA nerves send excitatory signals to the periaqueductal gray (PAG), located around the cerebral aqueduct, to create such behavior. The team named this circuit the ‘MPA-PAG’ circuit.
The team showed that they could control mammalian behavior for the first time with this scheme of MPA-Induced Drive Assisted Steering (MIDAS), in which a mouse chase the target objects in the front of head during stimulation of the MPA-PAG circuit. MIDAS allows mice to overcome obstacles to move in a desired path using optogenetics.
(Professor Daesoo Kim)
Professor Kim, who teamed up with Professor Phill Seung Lee in the Department of Mechanical Engineering, explained the significance of the research, “This study provides evidence to treat brain disorders such as compulsive hoarding and kleptomania. It also contributes to the development of technology to control the behavior of animals and humans using strong innate motivation, and thus could impact neuro-economics, defense, and disaster relief.”
He said the team would like to complete the neural circuit map governing behaviors of possession and hunting in the near future by exploring correlations with other neural behaviors controlling possessing and hunting activities.
This research was funded by the Samsung Science and Technology Foundation and published in Nature Neuroscience in March 2018.
(Figure 1: Schematics showing possessive behavior induced by the MPA neural circuit)
(Figure 2: Schematics of the MIDAS system that controls mammals behavior using the desire to possess. A MIDAS mouse is following the bait object controlled wirelessly.)
2018.04.23 View 10524 -
 Photoacoustic Imaging and Photothermal Cancer Therapy Using BR Nanoparticles
(Professor Sangyong Jon and PhD Candidate Dong Yun Lee)
Sangyong Jon, a professor in the Department of Biological Sciences at KAIST, and his team developed combined photoacoustic imaging and photothermal therapy for cancer by using Bilirubin (BR) nanoparticles.
The research team applied the properties of a bile pigment called BR, which exerts potent antioxidant and anti-inflammatory effects, to this research.
The team expects this research, which shows high biocompatibility as well as outstanding photoacoustic imaging and photothermal therapy, to be an appropriate system in the field of treatment for cancer.
In the past, the research team developed a PEGylated bilirubin-based nanoparticle system by combining water-insoluble BR with water-soluble Polyethylene Glycol (PEG).
This technology facilitated BR exerting antioxidants yet prevented them from being accumulated in the body. Its efficiency and safety was identified in an animal disease model, for conditions such as inflammatory bowel disease, islet cell transportation, and asthma.
Differing from previous research methods, this research applied the different physicochemical properties of BR to cancer treatment.
When the causative agent of jaundice, yellow BR, is exposed to a certain wavelength of blue light, the agent becomes a photonic nanomaterial as it responses to the light. This light-responsive nanomaterial can be used to cure jaundice because it allows for active excretion in infants.
Secondly, the team identified that BR is a major component of black pigment gallstones which can be often found in gall bladders or bile ducts under certain pathological conditions. The findings show that BR forms black pigment gallstones without the role of an intermediate or cation, such as calcium and copper. The research team combined cisplatin, a platinum metal-based anticancer drug, with BR so that BR nanoparticles changed the solution color from yellow to purple.
The team also examined the possibility of cisplatin-chelated BR nanoparticles as a probe for photoacoustic images. They found that considerable photoacoustic activity was shown when it was exposed to near infrared light. In fact, the photoacoustic signal was increased significantly in tumors of animals with colorectal cancer when the nanoparticles were administered to it intravenously. The team expects a more accurate diagnosis of tumors through this technology.
Moreover, the team assessed the photothermal effects of cisplatin-chelated BR nanoparticles. The research showed that the temperature of tumors increased by 25 degrees Celsius within five minutes when they were exposed to near infrared light, due to the photothermal effect. After two weeks, their size was reduced compared to that of other groups, and sometimes the tumors were even necrotized.
Professor Jon said, “Existing substances have a low biocompatibility and limitation for clinical therapy because they are artificially oriented; therefore, they might have toxicity. I am hoping that these cisplatin-chelated BR-based nanoparticles will provide a new platform for preclinical, translational research and clinical adaptation of the photoacoustic imaging and photothermal therapy.”
The paper (Dong Yun Lee as a first author) was published online in the renowned journal in the field of applied chemistry, Angewandte Chemi International Edition, on September 4. This research was sponsored by the National Research Foundation of Korea.
(Schematic diagram of the research)
(From left: Bilirubin nanoparticles, cisplatin-chelated Bilirubin nanoparticles)
2017.09.26 View 9679
Photoacoustic Imaging and Photothermal Cancer Therapy Using BR Nanoparticles
(Professor Sangyong Jon and PhD Candidate Dong Yun Lee)
Sangyong Jon, a professor in the Department of Biological Sciences at KAIST, and his team developed combined photoacoustic imaging and photothermal therapy for cancer by using Bilirubin (BR) nanoparticles.
The research team applied the properties of a bile pigment called BR, which exerts potent antioxidant and anti-inflammatory effects, to this research.
The team expects this research, which shows high biocompatibility as well as outstanding photoacoustic imaging and photothermal therapy, to be an appropriate system in the field of treatment for cancer.
In the past, the research team developed a PEGylated bilirubin-based nanoparticle system by combining water-insoluble BR with water-soluble Polyethylene Glycol (PEG).
This technology facilitated BR exerting antioxidants yet prevented them from being accumulated in the body. Its efficiency and safety was identified in an animal disease model, for conditions such as inflammatory bowel disease, islet cell transportation, and asthma.
Differing from previous research methods, this research applied the different physicochemical properties of BR to cancer treatment.
When the causative agent of jaundice, yellow BR, is exposed to a certain wavelength of blue light, the agent becomes a photonic nanomaterial as it responses to the light. This light-responsive nanomaterial can be used to cure jaundice because it allows for active excretion in infants.
Secondly, the team identified that BR is a major component of black pigment gallstones which can be often found in gall bladders or bile ducts under certain pathological conditions. The findings show that BR forms black pigment gallstones without the role of an intermediate or cation, such as calcium and copper. The research team combined cisplatin, a platinum metal-based anticancer drug, with BR so that BR nanoparticles changed the solution color from yellow to purple.
The team also examined the possibility of cisplatin-chelated BR nanoparticles as a probe for photoacoustic images. They found that considerable photoacoustic activity was shown when it was exposed to near infrared light. In fact, the photoacoustic signal was increased significantly in tumors of animals with colorectal cancer when the nanoparticles were administered to it intravenously. The team expects a more accurate diagnosis of tumors through this technology.
Moreover, the team assessed the photothermal effects of cisplatin-chelated BR nanoparticles. The research showed that the temperature of tumors increased by 25 degrees Celsius within five minutes when they were exposed to near infrared light, due to the photothermal effect. After two weeks, their size was reduced compared to that of other groups, and sometimes the tumors were even necrotized.
Professor Jon said, “Existing substances have a low biocompatibility and limitation for clinical therapy because they are artificially oriented; therefore, they might have toxicity. I am hoping that these cisplatin-chelated BR-based nanoparticles will provide a new platform for preclinical, translational research and clinical adaptation of the photoacoustic imaging and photothermal therapy.”
The paper (Dong Yun Lee as a first author) was published online in the renowned journal in the field of applied chemistry, Angewandte Chemi International Edition, on September 4. This research was sponsored by the National Research Foundation of Korea.
(Schematic diagram of the research)
(From left: Bilirubin nanoparticles, cisplatin-chelated Bilirubin nanoparticles)
2017.09.26 View 9679 -
 Unlocking the Keys to Parkinson's Disease
A KAIST research team has identified a new mechanism that causes the hallmark symptoms of Parkinson’s disease, namely tremors, rigidity, and loss of voluntary movement.
The discovery, made in collaboration with Nanyang Technological University in Singapore, presents a new perspective to three decades of conventional wisdom in Parkinson’s disease research. It also opens up new avenues that can help alleviate the motor problems suffered by patients of the disease, which reportedly number more than 10 million worldwide. The research was published in Neuron on August 30.
The research team was led by Professor Daesoo Kim from the Department of Biological Sciences at KAIST and Professor George Augustine from the Lee Kong Chian School of Medicine at NTU. Dr. Jeongjin Kim, a former postdoctoral fellow at KAIST who now works at the Korea Institute of Science and Technology (KIST), is the lead author.
It is known that Parkinson’s disease is caused by a lack of dopamine, a chemical in the brain that transmits neural signals. However, it remains unknown how the disease causes the motor
Smooth, voluntary movements, such as reaching for a cup of coffee, are controlled by the basal ganglia, which issue instructions via neurons (nerve cells that process and transmit information in the brain) in the thalamus to the cortex. These instructions come in two types: one that triggers a response (excitatory signals) and the other that suppresses a response (inhibitory signals). Proper balance between the two controls movement.
A low level of dopamine causes the basal ganglia to severely inhibit target neurons in the thalamus, called an inhibition. Scientists have long assumed that this stronger inhibition causes the motor problems of Parkinson’s disease patients.
To test this assumption, the research team used optogenetic technology in an animal model to study the effects of this increased inhibition of the thalamus and ultimately movement. Optogenetics is the use of light to control the activity of specific types of neurons within the brain.
They found that when signals from the basal ganglia are more strongly activated by light, the target neurons in the thalamus paradoxically became hyperactive. Called rebound excitation, this hyperactivity produced abnormal muscular stiffness and tremor. Such motor problems are very similar to the symptoms of Parkinson’s disease patients. When this hyperactivity of thalamic neurons is suppressed by light, mice show normal movments without Parkinson’s disease symptoms. Reducing the levels of activity back to normal caused the motor symptoms to stop, proving that the hyperactivity caused the motor problems experienced by Parkinson’s disease patients.
Professor Kim at KAIST said, “This study overturns three decades of consensus on the provenance of Parkinsonian symptoms.” The lead author, Dr Jeongjin Kim said, “The therapeutic implications of this study for the treatment of Parkinsonian symptoms are profound. It may soon become possible to remedy movement disorders without using L-DOPA, a pre-cursor to dopamine.”
Professor Augustine at NTU added, “Our findings are a breakthrough, both for understanding how the brain normally controls the movement of our body and how this control goes awry during Parkinson’s disease and related dopamine-deficiency disorders.”
The study took five years to complete, and includes researchers from the Department of Bio & Brain Engineering at KAIST.
The research team will move forward by investigating how hyperactivity in neurons in the thalamus leads to abnormal movement, as well as developing therapeutic strategies for the disease by targeting this neural mechanism.
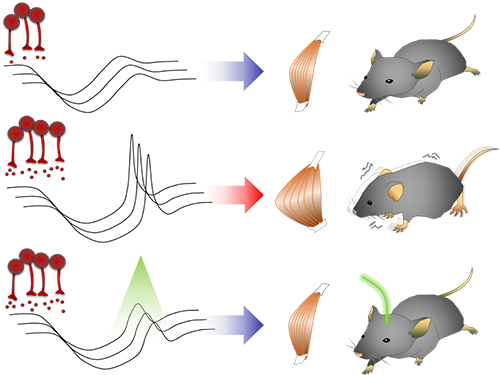
Figure abstract: Inhibitory inputs from the basal ganglia inhibit thalamic neurons (upper). In low-dopamine states, like PD, rebound firing follows inhibition and causes movement disorders (middle). The inhibition of rebound firing alleviates PD-like symptoms in a mouse model of PD.
2017.09.22 View 11326
Unlocking the Keys to Parkinson's Disease
A KAIST research team has identified a new mechanism that causes the hallmark symptoms of Parkinson’s disease, namely tremors, rigidity, and loss of voluntary movement.
The discovery, made in collaboration with Nanyang Technological University in Singapore, presents a new perspective to three decades of conventional wisdom in Parkinson’s disease research. It also opens up new avenues that can help alleviate the motor problems suffered by patients of the disease, which reportedly number more than 10 million worldwide. The research was published in Neuron on August 30.
The research team was led by Professor Daesoo Kim from the Department of Biological Sciences at KAIST and Professor George Augustine from the Lee Kong Chian School of Medicine at NTU. Dr. Jeongjin Kim, a former postdoctoral fellow at KAIST who now works at the Korea Institute of Science and Technology (KIST), is the lead author.
It is known that Parkinson’s disease is caused by a lack of dopamine, a chemical in the brain that transmits neural signals. However, it remains unknown how the disease causes the motor
Smooth, voluntary movements, such as reaching for a cup of coffee, are controlled by the basal ganglia, which issue instructions via neurons (nerve cells that process and transmit information in the brain) in the thalamus to the cortex. These instructions come in two types: one that triggers a response (excitatory signals) and the other that suppresses a response (inhibitory signals). Proper balance between the two controls movement.
A low level of dopamine causes the basal ganglia to severely inhibit target neurons in the thalamus, called an inhibition. Scientists have long assumed that this stronger inhibition causes the motor problems of Parkinson’s disease patients.
To test this assumption, the research team used optogenetic technology in an animal model to study the effects of this increased inhibition of the thalamus and ultimately movement. Optogenetics is the use of light to control the activity of specific types of neurons within the brain.
They found that when signals from the basal ganglia are more strongly activated by light, the target neurons in the thalamus paradoxically became hyperactive. Called rebound excitation, this hyperactivity produced abnormal muscular stiffness and tremor. Such motor problems are very similar to the symptoms of Parkinson’s disease patients. When this hyperactivity of thalamic neurons is suppressed by light, mice show normal movments without Parkinson’s disease symptoms. Reducing the levels of activity back to normal caused the motor symptoms to stop, proving that the hyperactivity caused the motor problems experienced by Parkinson’s disease patients.
Professor Kim at KAIST said, “This study overturns three decades of consensus on the provenance of Parkinsonian symptoms.” The lead author, Dr Jeongjin Kim said, “The therapeutic implications of this study for the treatment of Parkinsonian symptoms are profound. It may soon become possible to remedy movement disorders without using L-DOPA, a pre-cursor to dopamine.”
Professor Augustine at NTU added, “Our findings are a breakthrough, both for understanding how the brain normally controls the movement of our body and how this control goes awry during Parkinson’s disease and related dopamine-deficiency disorders.”
The study took five years to complete, and includes researchers from the Department of Bio & Brain Engineering at KAIST.
The research team will move forward by investigating how hyperactivity in neurons in the thalamus leads to abnormal movement, as well as developing therapeutic strategies for the disease by targeting this neural mechanism.
Figure abstract: Inhibitory inputs from the basal ganglia inhibit thalamic neurons (upper). In low-dopamine states, like PD, rebound firing follows inhibition and causes movement disorders (middle). The inhibition of rebound firing alleviates PD-like symptoms in a mouse model of PD.
2017.09.22 View 11326 -
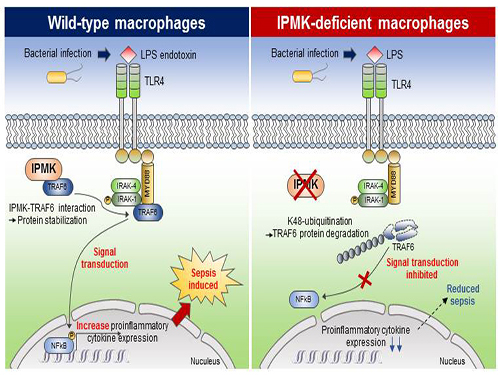 Study Identifies the Novel Molecular Signal for Triggering Septic Shock
Professor Seyun Kim’s team at the Department of Biological Sciences reported the mechanism by which cellular signaling transduction networks are precisely controlled in mediating innate immune responses, such as sepsis, by the enzyme IPMK (Inositol polyphosphate multikinase) which is essential for inositol biosynthesis metabolism.
In collaboration with Professor Hyun Seong Roh at Seoul National University, the study’s first author, Eunha Kim, a Ph.D. candidate in Department of Biological Sciences, performed a series of cellular, biochemical, and physiological experiments searching for the new function of IPMK enzymes in macrophages. The research findings were published in Science Advances on April 21.
Professor Kim’s team has been investigating various inositol metabolites and their biosynthesis metabolism for several years and has multilaterally identified the signaling actions of IPMK for controlling cellular growth and energy homeostasis.
This research showed that the specific deletion of IPMK enzymes in macrophages could significantly reduce levels of inflammation and increase survival rates in mice when they were challenged by microbial septic shock and endotoxins. This suggests a role for IPMK enzymes in mediating innate inflammatory responses that are directly related to a host’s defense against pathogenic bacterial infection.
The team further discovered that IPMK enzymes directly bind to TRAF6 proteins, a key player in immune signaling, thus protecting TRAF6 proteins from ubiquitination reactions that are involved in protein degradation. In addition, Kim and his colleagues successfully verified this IPMK-dependent immune control by employing short peptides which can specifically interfere with the binding between IPMK enzymes and TRAF6 proteins in macrophage cells.
This research revealed a novel function of IPMK enzymes in the fine tuning of innate immune signaling networks, suggesting a new direction for developing therapeutics targeting serious medical conditions such as neuroinflammation, type 2 diabetes, as well as polymicrobial sepsis that are developed from uncontrolled host immune responses. This research was funded by the Ministry of Science, ICT and Future Planning.
(Figure: Deletion of IPMK (inositol polyphosphate multikinase) in macrophages reduces the stability of TRAF6 protein which is the key to innate immune signaling, thereby blocking excessive inflammation in response to pathological bacterial infection.)
2017.05.11 View 9995
Study Identifies the Novel Molecular Signal for Triggering Septic Shock
Professor Seyun Kim’s team at the Department of Biological Sciences reported the mechanism by which cellular signaling transduction networks are precisely controlled in mediating innate immune responses, such as sepsis, by the enzyme IPMK (Inositol polyphosphate multikinase) which is essential for inositol biosynthesis metabolism.
In collaboration with Professor Hyun Seong Roh at Seoul National University, the study’s first author, Eunha Kim, a Ph.D. candidate in Department of Biological Sciences, performed a series of cellular, biochemical, and physiological experiments searching for the new function of IPMK enzymes in macrophages. The research findings were published in Science Advances on April 21.
Professor Kim’s team has been investigating various inositol metabolites and their biosynthesis metabolism for several years and has multilaterally identified the signaling actions of IPMK for controlling cellular growth and energy homeostasis.
This research showed that the specific deletion of IPMK enzymes in macrophages could significantly reduce levels of inflammation and increase survival rates in mice when they were challenged by microbial septic shock and endotoxins. This suggests a role for IPMK enzymes in mediating innate inflammatory responses that are directly related to a host’s defense against pathogenic bacterial infection.
The team further discovered that IPMK enzymes directly bind to TRAF6 proteins, a key player in immune signaling, thus protecting TRAF6 proteins from ubiquitination reactions that are involved in protein degradation. In addition, Kim and his colleagues successfully verified this IPMK-dependent immune control by employing short peptides which can specifically interfere with the binding between IPMK enzymes and TRAF6 proteins in macrophage cells.
This research revealed a novel function of IPMK enzymes in the fine tuning of innate immune signaling networks, suggesting a new direction for developing therapeutics targeting serious medical conditions such as neuroinflammation, type 2 diabetes, as well as polymicrobial sepsis that are developed from uncontrolled host immune responses. This research was funded by the Ministry of Science, ICT and Future Planning.
(Figure: Deletion of IPMK (inositol polyphosphate multikinase) in macrophages reduces the stability of TRAF6 protein which is the key to innate immune signaling, thereby blocking excessive inflammation in response to pathological bacterial infection.)
2017.05.11 View 9995 -
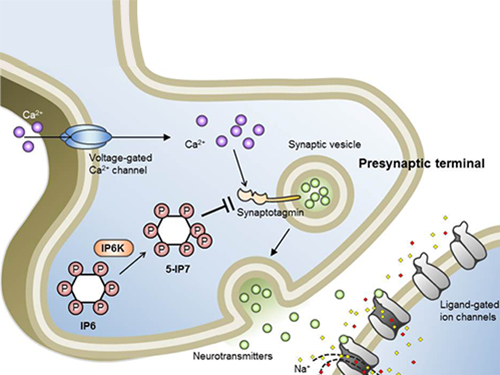 Professor Seyun Kim Identifies a Neuron Signal Controlling Molecule
A research team led by Professor Seyun Kim of the Department of Biological Sciences at KAIST has identified inositol pyrophosphates as the molecule that strongly controls neuron signaling via synaptotagmin.
Professors Tae-Young Yoon of Yonsei University’s Y-IBS and Sung-Hyun Kim of Kyung Hee University’s Department of Biomedical Science also joined the team.
The results were published in the Proceedings of the National Academy of Sciences of the United States of America (PNAS) on June 30, 2016.
This interdisciplinary research project was conducted by six research teams from four different countries and covered a wide scope of academic fields, from neurobiology to super resolution optic imaging.
Inositol pyrophosphates such as 5-diphosphoinositol pentakisphos-phate (5-IP7), which naturally occur in corns and beans, are essential metabolites in the body. In particular, inositol hexakisphosphate (IP6) has anti-cancer properties and is thought to have an important role in cell signaling.
Inositol pentakisphosphate (IP7) differs from IP6 by having an additional phosphate group, which was first discovered 20 years ago. IP7 has recently been identified as playing a key role in diabetes and obesity.
Psychopathy and neurodegenerative diseases are known to result from the disrupted balance of inositol pyrophosphates. However, the role and the mechanism of action of IP7 in brain neurons and nerve transmission remained unknown.
Professor Kim’s team has worked on inositol pyrophosphates for several years and discovered that very small quantities of IP7 control cell-signaling transduction. Professor Yoon of Yonsei University identified IP7 as a much stronger inhibitor of neuron signaling compared to IP6. In particular, IP7 directly suppresses synaptotagmin, one of the key proteins in neuron signaling. Moreover, Professor Kim of Kyung Hee University observed IP7 inhibition in sea horse neurons.
Together, the joint research team identified inositol pyrophosphates as the key switch metabolite of brain-signaling transduction.
The researchers hope that future research on synaptotagmin and IP7 will reveal the mechanism of neuron-signal transduction and thus enable the treatment of neurological disorders.
These research findings were the result of cooperation of various science and technology institutes: KAIST, Yonsei-IBS (Institute for Basic Science), Kyung Hee University, Sungkyunkwan University, KIST, University of Zurich in Switzerland, and Albert-Ludwigs-University Freiburg in Germany.
Schematic Image of Controlling the Synaptic Exocytotic Pathway by 5-IP7 , Helping the Understanding of the Signaling Mechanisms of Inositol Pyrophosphates
2016.07.21 View 12898
Professor Seyun Kim Identifies a Neuron Signal Controlling Molecule
A research team led by Professor Seyun Kim of the Department of Biological Sciences at KAIST has identified inositol pyrophosphates as the molecule that strongly controls neuron signaling via synaptotagmin.
Professors Tae-Young Yoon of Yonsei University’s Y-IBS and Sung-Hyun Kim of Kyung Hee University’s Department of Biomedical Science also joined the team.
The results were published in the Proceedings of the National Academy of Sciences of the United States of America (PNAS) on June 30, 2016.
This interdisciplinary research project was conducted by six research teams from four different countries and covered a wide scope of academic fields, from neurobiology to super resolution optic imaging.
Inositol pyrophosphates such as 5-diphosphoinositol pentakisphos-phate (5-IP7), which naturally occur in corns and beans, are essential metabolites in the body. In particular, inositol hexakisphosphate (IP6) has anti-cancer properties and is thought to have an important role in cell signaling.
Inositol pentakisphosphate (IP7) differs from IP6 by having an additional phosphate group, which was first discovered 20 years ago. IP7 has recently been identified as playing a key role in diabetes and obesity.
Psychopathy and neurodegenerative diseases are known to result from the disrupted balance of inositol pyrophosphates. However, the role and the mechanism of action of IP7 in brain neurons and nerve transmission remained unknown.
Professor Kim’s team has worked on inositol pyrophosphates for several years and discovered that very small quantities of IP7 control cell-signaling transduction. Professor Yoon of Yonsei University identified IP7 as a much stronger inhibitor of neuron signaling compared to IP6. In particular, IP7 directly suppresses synaptotagmin, one of the key proteins in neuron signaling. Moreover, Professor Kim of Kyung Hee University observed IP7 inhibition in sea horse neurons.
Together, the joint research team identified inositol pyrophosphates as the key switch metabolite of brain-signaling transduction.
The researchers hope that future research on synaptotagmin and IP7 will reveal the mechanism of neuron-signal transduction and thus enable the treatment of neurological disorders.
These research findings were the result of cooperation of various science and technology institutes: KAIST, Yonsei-IBS (Institute for Basic Science), Kyung Hee University, Sungkyunkwan University, KIST, University of Zurich in Switzerland, and Albert-Ludwigs-University Freiburg in Germany.
Schematic Image of Controlling the Synaptic Exocytotic Pathway by 5-IP7 , Helping the Understanding of the Signaling Mechanisms of Inositol Pyrophosphates
2016.07.21 View 12898 -
 Nikon Instruments Korea Donates a Fund to KAIST's Department of Biological Sciences
Representatives from Nikon Instruments Korea Co., Ltd., a producer of microscopes and measuring instruments, visited the KAIST campus on September 25, 2015, and donated USD 9,000 to the Department of Biological Sciences at KAIST.
A small ceremony to mark the donation took place at the department’s conference room.
In the picture from left to right were Professor Won-Do Heo, Department Head Byung-Ha Oh, Professor Sangyong Jon, President Sam-Sup Jang of Korea Instech, and Director Ik-Soo Yoo of Nikon Instruments Korea.
The department announced that the fund would be used to build its new research center to house the state-of-the-art research equipment and tools for the development of new medicine.
2015.10.03 View 5909
Nikon Instruments Korea Donates a Fund to KAIST's Department of Biological Sciences
Representatives from Nikon Instruments Korea Co., Ltd., a producer of microscopes and measuring instruments, visited the KAIST campus on September 25, 2015, and donated USD 9,000 to the Department of Biological Sciences at KAIST.
A small ceremony to mark the donation took place at the department’s conference room.
In the picture from left to right were Professor Won-Do Heo, Department Head Byung-Ha Oh, Professor Sangyong Jon, President Sam-Sup Jang of Korea Instech, and Director Ik-Soo Yoo of Nikon Instruments Korea.
The department announced that the fund would be used to build its new research center to house the state-of-the-art research equipment and tools for the development of new medicine.
2015.10.03 View 5909 -
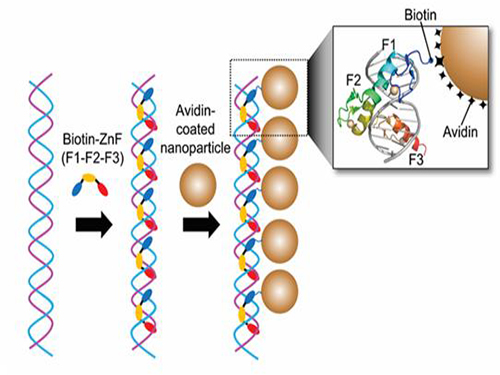 Nanoparticle Cluster Manufacturing Technique Using DNA Binding Protein Developed
Professor Hak-Sung Kim of the Department of Biological Sciences at KAIST and Yiseul Ryu, a doctoral candidate, used the Zinc Finger protein that specifically binds to target DNA sequence to develop a new manufacturing technique for size-controllable magnetic Nanoparticle Clusters (NPCs). Their research results were published in Angewandte Chemie International Edition online on 25 November 2014.
NPCs are structures consisting of magnetic nanoparticles, gold nanoparticles, and quantum dots, each of which are smaller than 100 nm (10-9m). NPCs have a distinctive property of collectivity not seen in single nanoparticles.
Specifically NPCS differ in physical and optical properties such as Plasmon coupling absorbance, energy transfers between particles, electron transfers, and conductivity. Therefore, NPCs can be employed in biological and medical research as well as the development of nanoelectric and nanoplasmon devices.
To make use of these novel properties, the size and the composition of the cluster must be exquisitely controlled. However, previous techniques relied on chemical binding which required complex steps, making it difficult to control the size and composition of NPCs.
Professor Kim’s team used Zinc Finger, a DNA binding protein, to develop a NPCs manufacturing technique to create clusters of the desired size easily. The Zinc Finger protein contains a zinc ion and specifically recognizes DNA sequence upon binding, which allows the exquisite control of the size and the cluster composition. The technique is also bio-friendly.
Professor Kim’s team created linear structure of different sizes of NPCs using Zinc Finger proteins and three DNA sequences of different lengths. The NPCs they produced confirmed their ability to control the size and structure of the cluster by using different DNA lengths.
The NPCs showed tripled T2 relaxation rates compared to the existing MRI contrast media (Feridex) and effectively transported to targeted cells. The research findings show the potential use of NPCs in biological and medical fields such as MRI contrast media, fluorescence imaging, and drug transport.
The research used the specific binding property of protein and DNA to develop a new method to create an inorganic nanoparticle’s supramolecular assembly. The technique can be used and applied extensively in other nanoparticles for future research in diagnosis, imaging, and drug and gene delivery.
Figure 1. A Mimetic Diagram of NPCs Manufacturing Technique Using DNA Binding Protein Zinc Finger
Figure 2. Transmission Electron Microscopy Images showing different sizes of NPCs depending on the length of the DNA
2014.12.04 View 14520
Nanoparticle Cluster Manufacturing Technique Using DNA Binding Protein Developed
Professor Hak-Sung Kim of the Department of Biological Sciences at KAIST and Yiseul Ryu, a doctoral candidate, used the Zinc Finger protein that specifically binds to target DNA sequence to develop a new manufacturing technique for size-controllable magnetic Nanoparticle Clusters (NPCs). Their research results were published in Angewandte Chemie International Edition online on 25 November 2014.
NPCs are structures consisting of magnetic nanoparticles, gold nanoparticles, and quantum dots, each of which are smaller than 100 nm (10-9m). NPCs have a distinctive property of collectivity not seen in single nanoparticles.
Specifically NPCS differ in physical and optical properties such as Plasmon coupling absorbance, energy transfers between particles, electron transfers, and conductivity. Therefore, NPCs can be employed in biological and medical research as well as the development of nanoelectric and nanoplasmon devices.
To make use of these novel properties, the size and the composition of the cluster must be exquisitely controlled. However, previous techniques relied on chemical binding which required complex steps, making it difficult to control the size and composition of NPCs.
Professor Kim’s team used Zinc Finger, a DNA binding protein, to develop a NPCs manufacturing technique to create clusters of the desired size easily. The Zinc Finger protein contains a zinc ion and specifically recognizes DNA sequence upon binding, which allows the exquisite control of the size and the cluster composition. The technique is also bio-friendly.
Professor Kim’s team created linear structure of different sizes of NPCs using Zinc Finger proteins and three DNA sequences of different lengths. The NPCs they produced confirmed their ability to control the size and structure of the cluster by using different DNA lengths.
The NPCs showed tripled T2 relaxation rates compared to the existing MRI contrast media (Feridex) and effectively transported to targeted cells. The research findings show the potential use of NPCs in biological and medical fields such as MRI contrast media, fluorescence imaging, and drug transport.
The research used the specific binding property of protein and DNA to develop a new method to create an inorganic nanoparticle’s supramolecular assembly. The technique can be used and applied extensively in other nanoparticles for future research in diagnosis, imaging, and drug and gene delivery.
Figure 1. A Mimetic Diagram of NPCs Manufacturing Technique Using DNA Binding Protein Zinc Finger
Figure 2. Transmission Electron Microscopy Images showing different sizes of NPCs depending on the length of the DNA
2014.12.04 View 14520 -
 Discovery of New Therapeutic Targets for Alzheimer's Disease
A Korean research team headed by Professor Dae-Soo Kim of Biological Sciences at KAIST and Dr. Chang-Jun Lee from the Korea Institute of Science and Technology (KIST) successfully identified that reactive astrocytes, commonly observed in brains affected by Alzheimer’s disease, produce abnormal amounts of inhibitory neurotransmitter gamma-Aminobutyric acid (GABA) in reaction to the enzyme Monoamine oxidase B (Mao-B) and release GABA through the Bestrophin-1 channel to suppress the normal signal transmission of brain nerve cells.
By suppressing the GABA production or release from reactive astrocytes, the research team was able to restore the model mice's memory and learning impairment caused by Alzheimer’s disease. This discovery will allow the development of new drugs to treat Alzheimer’s and other related diseases.
The research result was published in the June 29, 2014 edition of Nature Medicine (Title: GABA from Reactive Astrocytes Impairs Memory in Mouse Models of Alzheimer’s Disease).
For details, please read the article below:
Technology News, July 10, 2014
"Discovery of New Drug Targets for Memory Impairment in Alzheimer’s Disease"
http://technews.tmcnet.com/news/2014/07/10/7917811.htm
2014.07.16 View 10304
Discovery of New Therapeutic Targets for Alzheimer's Disease
A Korean research team headed by Professor Dae-Soo Kim of Biological Sciences at KAIST and Dr. Chang-Jun Lee from the Korea Institute of Science and Technology (KIST) successfully identified that reactive astrocytes, commonly observed in brains affected by Alzheimer’s disease, produce abnormal amounts of inhibitory neurotransmitter gamma-Aminobutyric acid (GABA) in reaction to the enzyme Monoamine oxidase B (Mao-B) and release GABA through the Bestrophin-1 channel to suppress the normal signal transmission of brain nerve cells.
By suppressing the GABA production or release from reactive astrocytes, the research team was able to restore the model mice's memory and learning impairment caused by Alzheimer’s disease. This discovery will allow the development of new drugs to treat Alzheimer’s and other related diseases.
The research result was published in the June 29, 2014 edition of Nature Medicine (Title: GABA from Reactive Astrocytes Impairs Memory in Mouse Models of Alzheimer’s Disease).
For details, please read the article below:
Technology News, July 10, 2014
"Discovery of New Drug Targets for Memory Impairment in Alzheimer’s Disease"
http://technews.tmcnet.com/news/2014/07/10/7917811.htm
2014.07.16 View 10304