research
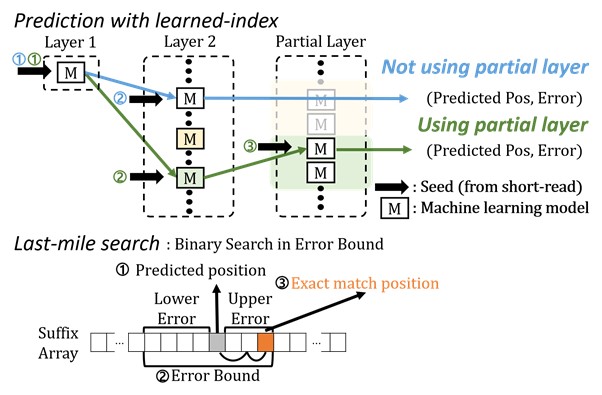
The algorithm presents the first full-fledged, short-read alignment software that leverages learned indices for solving the exact match search problem for efficient seeding
< Image:Scientists from KAIST develops new machine-learning-based approach to speed up DNA sequencing. >
The human genome consists of a complete set of DNA, which is about 6.4 billion letters long. Because of its size, reading the whole genome sequence at once is challenging. So scientists use DNA sequencers to produce hundreds of millions of DNA sequence fragments, or short reads, up to 300 letters long. Then the DNA sequencer assembles all the short reads like a giant jigsaw puzzle to reconstruct the entire genome sequence. Even with very fast computers, this job can take hours to complete.
A research team at KAIST has achieved up to 3.45x faster speeds by developing the first short-read alignment software that uses a recent advance in machine-learning called a learned index.
The research team reported their findings on March 7, 2022 in the journal Bioinformatics. The software has been released as open source and can be found on github (https://github.com/kaist-ina/BWA-MEME).
Next-generation sequencing (NGS) is a state-of-the-art DNA sequencing method. Projects are underway with the goal of producing genome sequencing at population scale. Modern NGS hardware is capable of generating billions of short reads in a single run. Then the short reads have to be aligned with the reference DNA sequence. With large-scale DNA sequencing operations running hundreds of next-generation sequences, the need for an efficient short read alignment tool has become even more critical. Accelerating the DNA sequence alignment would be a step toward achieving the goal of population-scale sequencing. However, existing algorithms are limited in their performance because of their frequent memory accesses.
BWA-MEM2 is a popular short-read alignment software package currently used to sequence the DNA. However, it has its limitations. The state-of-the-art alignment has two phases – seeding and extending. During the seeding phase, searches find exact matches of short reads in the reference DNA sequence. During the extending phase, the short reads from the seeding phase are extended. In the current process, bottlenecks occur in the seeding phase. Finding the exact matches slows the process.
The researchers set out to solve the problem of accelerating the DNA sequence alignment. To speed the process, they applied machine learning techniques to create an algorithmic improvement. Their algorithm, BWA-MEME (BWA-MEM emulated) leverages learned indices to solve the exact match search problem. The original software compared one character at a time for an exact match search. The team’s new algorithm achieves up to 3.45x faster speeds in seeding throughput over BWA-MEM2 by reducing the number of instructions by 4.60x and memory accesses by 8.77x. “Through this study, it has been shown that full genome big data analysis can be performed faster and less costly than conventional methods by applying machine learning technology,” said Professor Dongsu Han from the School of Electrical Engineering at KAIST.
The researchers’ ultimate goal was to develop efficient software that scientists from academia and industry could use on a daily basis for analyzing big data in genomics. “With the recent advances in artificial intelligence and machine learning, we see so many opportunities for designing better software for genomic data analysis. The potential is there for accelerating existing analysis as well as enabling new types of analysis, and our goal is to develop such software,” added Han.
Whole genome sequencing has traditionally been used for discovering genomic mutations and identifying the root causes of diseases, which leads to the discovery and development of new drugs and cures. There could be many potential applications. Whole genome sequencing is used not only for research, but also for clinical purposes. “The science and technology for analyzing genomic data is making rapid progress to make it more accessible for scientists and patients. This will enhance our understanding about diseases and develop a better cure for patients of various diseases.”
The research was funded by the National Research Foundation of the Korean government’s Ministry of Science and ICT.
-Publication
Youngmok Jung, Dongsu Han, “BWA-MEME:BWA-MEM emulated with a machine learning approach,” Bioinformatics, Volume 38, Issue 9, May 2022
(https://doi.org/10.1093/bioinformatics/btac137)
-Profile
Professor Dongsu Han
School of Electrical Engineering
KAIST
-
research Genome Sequencing Unveils Mutational Impacts of Radiation on Mammalian Cells
Recent release of the waste water from Japan's Fukushima nuclear disaster stirred apprehension regarding the health implications of radiation exposure. Classified as a Group 1 carcinogen, ionizing radiation has long been associated with various cancers and genetic disorders, as evidenced by survivors and descendants of atomic bombings and the Chernobyl disaster. Despite much smaller amount, we remain consistently exposed to low levels of radiation in everyday life and medical procedures. Radi
2024-02-15 -
research KAIST research team develops a forgery prevention technique using salmon DNA
The authenticity scandal that plagued the artwork “Beautiful Woman” by Kyung-ja Chun for 30 years shows how concerns about replicas can become a burden to artists, as most of them are not experts in the field of anti-counterfeiting. To solve this problem, artist-friendly physical unclonable functions (PUFs) based on optical techniques instead of electronic ones, which can be applied immediately onto artwork through brushstrokes are needed. On May 23, a KAIST research team led by P
2023-06-08 -
research Overview of the 30-year history of metabolic engineering
< Distinguished Professor Sang Yup Lee from the Department of Chemical and Biomolecular Engineering at KAIST > A research team comprised of Gi Bae Kim, Dr. So Young Choi, Dr. In Jin Cho, Da-Hee Ahn, and Distinguished Professor Sang Yup Lee from the Department of Chemical and Biomolecular Engineering at KAIST reported the 30-year history of metabolic engineering, highlighting examples of recent progress in the field and contributions to sustainability and health. Their paper “Met
2023-01-25 -
research Phage resistant Escherichia coli strains developed to reduce fermentation failure
A genome engineering-based systematic strategy for developing phage resistant Escherichia coli strains has been successfully developed through the collaborative efforts of a team led by Professor Sang Yup Lee, Professor Shi Chen, and Professor Lianrong Wang. This study by Xuan Zou et al. was published in Nature Communications in August 2022 and featured in Nature Communications Editors’ Highlights. The collaboration by the School of Pharmaceutical Sciences at Wuhan University, the First Af
2022-08-23 -
research Now You Can See Floral Scents!
Optical interferometry visualizes how often lilies emit volatile organic compounds Have you ever thought about when flowers emit their scents? KAIST mechanical engineers and biological scientists directly visualized how often a lily releases a floral scent using a laser interferometry method. These measurement results can provide new insights for understanding and further exploring the biosynthesis and emission mechanisms of floral volatiles. Why is it important to know this? It is well kno
2022-05-25