BIO
-
 Professor Jong Chul Ye Appointed as Distinguished Lecturer of IEEE EMBS
Professor Jong Chul Ye from the Department of Bio and Brain Engineering was appointed as a distinguished lecturer by the International Association of Electrical and Electronic Engineers (IEEE) Engineering in Medicine and Biology Society (EMBS). Professor Ye was invited to deliver a lecture on his leading research on artificial intelligence (AI) technology in medical video restoration. He will serve a term of two years beginning in 2020.
IEEE EMBS's distinguished lecturer program is designed to educate researchers around the world on the latest trends and technology in biomedical engineering. Sponsored by IEEE, its members can attend lectures on the distinguished professor's research subject.
Professor Ye said, "We are at a time where the importance of AI in medical imaging is increasing.” He added, “I am proud to be appointed as a distinguished lecturer of the IEEE EMBS in recognition of my contributions to this field.”
(END)
2020.02.27 View 12639
Professor Jong Chul Ye Appointed as Distinguished Lecturer of IEEE EMBS
Professor Jong Chul Ye from the Department of Bio and Brain Engineering was appointed as a distinguished lecturer by the International Association of Electrical and Electronic Engineers (IEEE) Engineering in Medicine and Biology Society (EMBS). Professor Ye was invited to deliver a lecture on his leading research on artificial intelligence (AI) technology in medical video restoration. He will serve a term of two years beginning in 2020.
IEEE EMBS's distinguished lecturer program is designed to educate researchers around the world on the latest trends and technology in biomedical engineering. Sponsored by IEEE, its members can attend lectures on the distinguished professor's research subject.
Professor Ye said, "We are at a time where the importance of AI in medical imaging is increasing.” He added, “I am proud to be appointed as a distinguished lecturer of the IEEE EMBS in recognition of my contributions to this field.”
(END)
2020.02.27 View 12639 -
 New Catalyst Recycles Greenhouse Gases into Fuel and Hydrogen Gas
< Professor Cafer T. Yavuz (left), PhD Candidate Youngdong Song (center), and Researcher Sreerangappa Ramesh (right) >
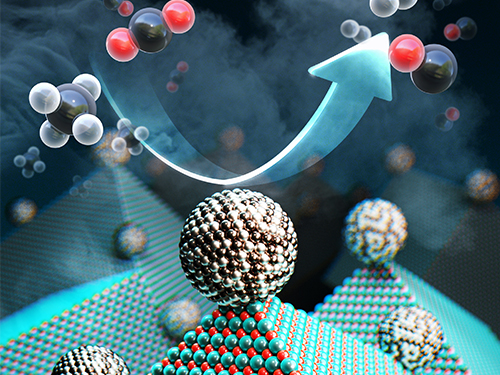
Scientists have taken a major step toward a circular carbon economy by developing a long-lasting, economical catalyst that recycles greenhouse gases into ingredients that can be used in fuel, hydrogen gas, and other chemicals. The results could be revolutionary in the effort to reverse global warming, according to the researchers. The study was published on February 14 in Science.
“We set out to develop an effective catalyst that can convert large amounts of the greenhouse gases carbon dioxide and methane without failure,” said Cafer T. Yavuz, paper author and associate professor of chemical and biomolecular engineering and of chemistry at KAIST.
The catalyst, made from inexpensive and abundant nickel, magnesium, and molybdenum, initiates and speeds up the rate of reaction that converts carbon dioxide and methane into hydrogen gas. It can work efficiently for more than a month.
This conversion is called ‘dry reforming’, where harmful gases, such as carbon dioxide, are processed to produce more useful chemicals that could be refined for use in fuel, plastics, or even pharmaceuticals. It is an effective process, but it previously required rare and expensive metals such as platinum and rhodium to induce a brief and inefficient chemical reaction.
Other researchers had previously proposed nickel as a more economical solution, but carbon byproducts would build up and the surface nanoparticles would bind together on the cheaper metal, fundamentally changing the composition and geometry of the catalyst and rendering it useless.
“The difficulty arises from the lack of control on scores of active sites over the bulky catalysts surfaces because any refinement procedures attempted also change the nature of the catalyst itself,” Yavuz said.
The researchers produced nickel-molybdenum nanoparticles under a reductive environment in the presence of a single crystalline magnesium oxide. As the ingredients were heated under reactive gas, the nanoparticles moved on the pristine crystal surface seeking anchoring points. The resulting activated catalyst sealed its own high-energy active sites and permanently fixed the location of the nanoparticles — meaning that the nickel-based catalyst will not have a carbon build up, nor will the surface particles bind to one another.
“It took us almost a year to understand the underlying mechanism,” said first author Youngdong Song, a graduate student in the Department of Chemical and Biomolecular Engineering at KAIST. “Once we studied all the chemical events in detail, we were shocked.”
The researchers dubbed the catalyst Nanocatalysts on Single Crystal Edges (NOSCE). The magnesium-oxide nanopowder comes from a finely structured form of magnesium oxide, where the molecules bind continuously to the edge. There are no breaks or defects in the surface, allowing for uniform and predictable reactions.
“Our study solves a number of challenges the catalyst community faces,” Yavuz said. “We believe the NOSCE mechanism will improve other inefficient catalytic reactions and provide even further savings of greenhouse gas emissions.”
This work was supported, in part, by the Saudi-Aramco-KAIST CO2 Management Center and the National Research Foundation of Korea.
Other contributors include Ercan Ozdemir, Sreerangappa Ramesh, Aldiar Adishev, and Saravanan Subramanian, all of whom are affiliated with the Graduate School of Energy, Environment, Water and Sustainability at KAIST; Aadesh Harale, Mohammed Albuali, Bandar Abdullah Fadhel, and Aqil Jamal, all of whom are with the Research and Development Center in Saudi Arabia; and Dohyun Moon and Sun Hee Choi, both of whom are with the Pohang Accelerator Laboratory in Korea. Ozdemir is also affiliated with the Institute of Nanotechnology at the Gebze Technical University in Turkey; Fadhel and Jamal are also affiliated with the Saudi-Armco-KAIST CO2 Management Center in Korea.
<Newly developed catalyst that recycles greenhouse gases into ingredients that can be used in fuel, hydrogen gas and other chemicals.>
Publication:
Song et al. (2020) Dry reforming of methane by stable Ni–Mo nanocatalysts on single-crystalline MgO. Science, Vol. 367, Issue 6479, pp. 777-781. Available online at http://dx.doi.org/10.1126/science.aav2412
Profile: Prof. Cafer T. Yavuz, MA, PhD
yavuz@kaist.ac.kr
http://yavuz.kaist.ac.kr/
Associate Professor
Oxide and Organic Nanomaterials for the Environment (ONE) Laboratory
Graduate School of Energy, Environment, Water and Sustainability (EEWS)
Korea Advanced Institute of Science and Technology (KAIST)
http://kaist.ac.kr
Daejeon, Republic of Korea
Profile: Youngdong Song ydsong88@kaist.ac.kr
Ph.D. Candidate
Department of Chemical and Biomolecular Engineering
Korea Advanced Institute of Science and Technology (KAIST)
http://kaist.ac.kr
Daejeon, Republic of Korea
(END)
2020.02.17 View 19075
New Catalyst Recycles Greenhouse Gases into Fuel and Hydrogen Gas
< Professor Cafer T. Yavuz (left), PhD Candidate Youngdong Song (center), and Researcher Sreerangappa Ramesh (right) >
Scientists have taken a major step toward a circular carbon economy by developing a long-lasting, economical catalyst that recycles greenhouse gases into ingredients that can be used in fuel, hydrogen gas, and other chemicals. The results could be revolutionary in the effort to reverse global warming, according to the researchers. The study was published on February 14 in Science.
“We set out to develop an effective catalyst that can convert large amounts of the greenhouse gases carbon dioxide and methane without failure,” said Cafer T. Yavuz, paper author and associate professor of chemical and biomolecular engineering and of chemistry at KAIST.
The catalyst, made from inexpensive and abundant nickel, magnesium, and molybdenum, initiates and speeds up the rate of reaction that converts carbon dioxide and methane into hydrogen gas. It can work efficiently for more than a month.
This conversion is called ‘dry reforming’, where harmful gases, such as carbon dioxide, are processed to produce more useful chemicals that could be refined for use in fuel, plastics, or even pharmaceuticals. It is an effective process, but it previously required rare and expensive metals such as platinum and rhodium to induce a brief and inefficient chemical reaction.
Other researchers had previously proposed nickel as a more economical solution, but carbon byproducts would build up and the surface nanoparticles would bind together on the cheaper metal, fundamentally changing the composition and geometry of the catalyst and rendering it useless.
“The difficulty arises from the lack of control on scores of active sites over the bulky catalysts surfaces because any refinement procedures attempted also change the nature of the catalyst itself,” Yavuz said.
The researchers produced nickel-molybdenum nanoparticles under a reductive environment in the presence of a single crystalline magnesium oxide. As the ingredients were heated under reactive gas, the nanoparticles moved on the pristine crystal surface seeking anchoring points. The resulting activated catalyst sealed its own high-energy active sites and permanently fixed the location of the nanoparticles — meaning that the nickel-based catalyst will not have a carbon build up, nor will the surface particles bind to one another.
“It took us almost a year to understand the underlying mechanism,” said first author Youngdong Song, a graduate student in the Department of Chemical and Biomolecular Engineering at KAIST. “Once we studied all the chemical events in detail, we were shocked.”
The researchers dubbed the catalyst Nanocatalysts on Single Crystal Edges (NOSCE). The magnesium-oxide nanopowder comes from a finely structured form of magnesium oxide, where the molecules bind continuously to the edge. There are no breaks or defects in the surface, allowing for uniform and predictable reactions.
“Our study solves a number of challenges the catalyst community faces,” Yavuz said. “We believe the NOSCE mechanism will improve other inefficient catalytic reactions and provide even further savings of greenhouse gas emissions.”
This work was supported, in part, by the Saudi-Aramco-KAIST CO2 Management Center and the National Research Foundation of Korea.
Other contributors include Ercan Ozdemir, Sreerangappa Ramesh, Aldiar Adishev, and Saravanan Subramanian, all of whom are affiliated with the Graduate School of Energy, Environment, Water and Sustainability at KAIST; Aadesh Harale, Mohammed Albuali, Bandar Abdullah Fadhel, and Aqil Jamal, all of whom are with the Research and Development Center in Saudi Arabia; and Dohyun Moon and Sun Hee Choi, both of whom are with the Pohang Accelerator Laboratory in Korea. Ozdemir is also affiliated with the Institute of Nanotechnology at the Gebze Technical University in Turkey; Fadhel and Jamal are also affiliated with the Saudi-Armco-KAIST CO2 Management Center in Korea.
<Newly developed catalyst that recycles greenhouse gases into ingredients that can be used in fuel, hydrogen gas and other chemicals.>
Publication:
Song et al. (2020) Dry reforming of methane by stable Ni–Mo nanocatalysts on single-crystalline MgO. Science, Vol. 367, Issue 6479, pp. 777-781. Available online at http://dx.doi.org/10.1126/science.aav2412
Profile: Prof. Cafer T. Yavuz, MA, PhD
yavuz@kaist.ac.kr
http://yavuz.kaist.ac.kr/
Associate Professor
Oxide and Organic Nanomaterials for the Environment (ONE) Laboratory
Graduate School of Energy, Environment, Water and Sustainability (EEWS)
Korea Advanced Institute of Science and Technology (KAIST)
http://kaist.ac.kr
Daejeon, Republic of Korea
Profile: Youngdong Song ydsong88@kaist.ac.kr
Ph.D. Candidate
Department of Chemical and Biomolecular Engineering
Korea Advanced Institute of Science and Technology (KAIST)
http://kaist.ac.kr
Daejeon, Republic of Korea
(END)
2020.02.17 View 19075 -
 What Fuels a “Domino Effect” in Cancer Drug Resistance?
KAIST researchers have identified mechanisms that relay prior acquired resistance to the first-line chemotherapy to the second-line targeted therapy, fueling a “domino effect” in cancer drug resistance. Their study featured in the February 7 edition of Science Advances suggests a new strategy for improving the second-line setting of cancer treatment for patients who showed resistance to anti-cancer drugs.
Resistance to cancer drugs is often managed in the clinic by chemotherapy and targeted therapy. Unlike chemotherapy that works by repressing fast-proliferating cells, targeted therapy blocks a single oncogenic pathway to halt tumor growth. In many cases, targeted therapy is engaged as a maintenance therapy or employed in the second-line after front-line chemotherapy.
A team of researchers led by Professor Yoosik Kim from the Department of Chemical and Biomolecular Engineering and the KAIST Institute for Health Science and Technology (KIHST) has discovered an unexpected resistance signature that occurs between chemotherapy and targeted therapy. The team further identified a set of integrated mechanisms that promotes this kind of sequential therapy resistance.
“There have been multiple clinical accounts reflecting that targeted therapies tend to be least successful in patients who have exhausted all standard treatments,” said the first author of the paper Mark Borris D. Aldonza. He continued, “These accounts ignited our hypothesis that failed responses to some chemotherapies might speed up the evolution of resistance to other drugs, particularly those with specific targets.”
Aldonza and his colleagues extracted large amounts of drug-resistance information from the open-source database the Genomics of Drug Sensitivity in Cancer (GDSC), which contains thousands of drug response data entries from various human cancer cell lines. Their big data analysis revealed that cancer cell lines resistant to chemotherapies classified as anti-mitotic drugs (AMDs), toxins that inhibit overacting cell division, are also resistant to a class of targeted therapies called epidermal growth factor receptor-tyrosine kinase inhibitors (EGFR-TKIs).
In all of the cancer types analyzed, more than 84 percent of those resistant to AMDs, representatively ‘paclitaxel’, were also resistant to at least nine EGFR-TKIs. In lung, pancreatic, and breast cancers where paclitaxel is often used as a first-line, standard-of-care regimen, greater than 92 percent showed resistance to EGFR-TKIs. Professor Kim said, “It is surprising to see that such collateral resistance can occur specifically between two chemically different classes of drugs.”
To figure out how failed responses to paclitaxel leads to resistance to EGFR-TKIs, the team validated co-resistance signatures that they found in the database by generating and analyzing a subset of slow-doubling, paclitaxel-resistant cancer models called ‘persisters’.
The results demonstrated that paclitaxel-resistant cancers remodel their stress response by first becoming more stem cell-like, evolving the ability to self-renew to adapt to more stressful conditions like drug exposures. More surprisingly, when the researchers characterized the metabolic state of the cells, EGFR-TKI persisters derived from paclitaxel-resistant cancer cells showed high dependencies to energy-producing processes such as glycolysis and glutaminolysis.
“We found that, without an energy stimulus like glucose, these cells transform to becoming more senescent, a characteristic of cells that have arrested cell division. However, this senescence is controlled by stem cell factors, which the paclitaxel-resistant cancers use to escape from this arrested state given a favorable condition to re-grow,” said Aldonza.
Professor Kim explained, “Before this research, there was no reason to expect that acquiring the cancer stem cell phenotype that dramatically leads to a cascade of changes in cellular states affecting metabolism and cell death is linked with drug-specific sequential resistance between two classes of therapies.”
He added, “The expansion of our work to other working models of drug resistance in a much more clinically-relevant setting, perhaps in clinical trials, will take on increasing importance, as sequential treatment strategies will continue to be adapted to various forms of anti-cancer therapy regimens.”
This study was supported by the Basic Science Research Program of the National Research Foundation of Korea (NRF-2016R1C1B2009886), and the KAIST Future Systems Healthcare Project (KAISTHEALTHCARE42) funded by the Korean Ministry of Science and ICT (MSIT). Undergraduate student Aldonza participated in this research project and presented the findings as the lead author as part of the Undergraduate Research Participation (URP) Program at KAIST.
< Figure 1. Schematic overview of the study. >
< Figure 2. Big data analysis revealing co-resistance signatures between classes of anti-cancer drugs. >
Publication:
Aldonza et al. (2020) Prior acquired resistance to paclitaxel relays diverse EGFR-targeted therapy persistence mechanisms. Science Advances, Vol. 6, No. 6, eaav7416. Available online at http://dx.doi.org/10.1126/sciadv.aav7416
Profile: Prof. Yoosik Kim, MA, PhD
ysyoosik@kaist.ac.kr
https://qcbio.kaist.ac.kr/
Assistant Professor
Bio Network Analysis Laboratory
Department of Chemical and Biomolecular Engineering
Korea Advanced Institute of Science and Technology (KAIST)
http://kaist.ac.kr
Daejeon, Republic of Korea
Profile: Mark Borris D. Aldonza
borris@kaist.ac.kr
Undergraduate Student
Department of Biological Sciences
Korea Advanced Institute of Science and Technology (KAIST)
http://kaist.ac.kr
Daejeon, Republic of Korea
(END)
2020.02.10 View 16677
What Fuels a “Domino Effect” in Cancer Drug Resistance?
KAIST researchers have identified mechanisms that relay prior acquired resistance to the first-line chemotherapy to the second-line targeted therapy, fueling a “domino effect” in cancer drug resistance. Their study featured in the February 7 edition of Science Advances suggests a new strategy for improving the second-line setting of cancer treatment for patients who showed resistance to anti-cancer drugs.
Resistance to cancer drugs is often managed in the clinic by chemotherapy and targeted therapy. Unlike chemotherapy that works by repressing fast-proliferating cells, targeted therapy blocks a single oncogenic pathway to halt tumor growth. In many cases, targeted therapy is engaged as a maintenance therapy or employed in the second-line after front-line chemotherapy.
A team of researchers led by Professor Yoosik Kim from the Department of Chemical and Biomolecular Engineering and the KAIST Institute for Health Science and Technology (KIHST) has discovered an unexpected resistance signature that occurs between chemotherapy and targeted therapy. The team further identified a set of integrated mechanisms that promotes this kind of sequential therapy resistance.
“There have been multiple clinical accounts reflecting that targeted therapies tend to be least successful in patients who have exhausted all standard treatments,” said the first author of the paper Mark Borris D. Aldonza. He continued, “These accounts ignited our hypothesis that failed responses to some chemotherapies might speed up the evolution of resistance to other drugs, particularly those with specific targets.”
Aldonza and his colleagues extracted large amounts of drug-resistance information from the open-source database the Genomics of Drug Sensitivity in Cancer (GDSC), which contains thousands of drug response data entries from various human cancer cell lines. Their big data analysis revealed that cancer cell lines resistant to chemotherapies classified as anti-mitotic drugs (AMDs), toxins that inhibit overacting cell division, are also resistant to a class of targeted therapies called epidermal growth factor receptor-tyrosine kinase inhibitors (EGFR-TKIs).
In all of the cancer types analyzed, more than 84 percent of those resistant to AMDs, representatively ‘paclitaxel’, were also resistant to at least nine EGFR-TKIs. In lung, pancreatic, and breast cancers where paclitaxel is often used as a first-line, standard-of-care regimen, greater than 92 percent showed resistance to EGFR-TKIs. Professor Kim said, “It is surprising to see that such collateral resistance can occur specifically between two chemically different classes of drugs.”
To figure out how failed responses to paclitaxel leads to resistance to EGFR-TKIs, the team validated co-resistance signatures that they found in the database by generating and analyzing a subset of slow-doubling, paclitaxel-resistant cancer models called ‘persisters’.
The results demonstrated that paclitaxel-resistant cancers remodel their stress response by first becoming more stem cell-like, evolving the ability to self-renew to adapt to more stressful conditions like drug exposures. More surprisingly, when the researchers characterized the metabolic state of the cells, EGFR-TKI persisters derived from paclitaxel-resistant cancer cells showed high dependencies to energy-producing processes such as glycolysis and glutaminolysis.
“We found that, without an energy stimulus like glucose, these cells transform to becoming more senescent, a characteristic of cells that have arrested cell division. However, this senescence is controlled by stem cell factors, which the paclitaxel-resistant cancers use to escape from this arrested state given a favorable condition to re-grow,” said Aldonza.
Professor Kim explained, “Before this research, there was no reason to expect that acquiring the cancer stem cell phenotype that dramatically leads to a cascade of changes in cellular states affecting metabolism and cell death is linked with drug-specific sequential resistance between two classes of therapies.”
He added, “The expansion of our work to other working models of drug resistance in a much more clinically-relevant setting, perhaps in clinical trials, will take on increasing importance, as sequential treatment strategies will continue to be adapted to various forms of anti-cancer therapy regimens.”
This study was supported by the Basic Science Research Program of the National Research Foundation of Korea (NRF-2016R1C1B2009886), and the KAIST Future Systems Healthcare Project (KAISTHEALTHCARE42) funded by the Korean Ministry of Science and ICT (MSIT). Undergraduate student Aldonza participated in this research project and presented the findings as the lead author as part of the Undergraduate Research Participation (URP) Program at KAIST.
< Figure 1. Schematic overview of the study. >
< Figure 2. Big data analysis revealing co-resistance signatures between classes of anti-cancer drugs. >
Publication:
Aldonza et al. (2020) Prior acquired resistance to paclitaxel relays diverse EGFR-targeted therapy persistence mechanisms. Science Advances, Vol. 6, No. 6, eaav7416. Available online at http://dx.doi.org/10.1126/sciadv.aav7416
Profile: Prof. Yoosik Kim, MA, PhD
ysyoosik@kaist.ac.kr
https://qcbio.kaist.ac.kr/
Assistant Professor
Bio Network Analysis Laboratory
Department of Chemical and Biomolecular Engineering
Korea Advanced Institute of Science and Technology (KAIST)
http://kaist.ac.kr
Daejeon, Republic of Korea
Profile: Mark Borris D. Aldonza
borris@kaist.ac.kr
Undergraduate Student
Department of Biological Sciences
Korea Advanced Institute of Science and Technology (KAIST)
http://kaist.ac.kr
Daejeon, Republic of Korea
(END)
2020.02.10 View 16677 -
 Blood-Based Multiplexed Diagnostic Sensor Helps to Accurately Detect Alzheimer’s Disease
A research team at KAIST reported clinically accurate multiplexed electrical biosensor for detecting Alzheimer’s disease by measuring its core biomarkers using densely aligned carbon nanotubes.
Alzheimer’s disease is the most prevalent neurodegenerative disorder, affecting one in ten aged over 65 years. Early diagnosis can reduce the risk of suffering the disease by one-third, according to recent reports. However, its early diagnosis remains challenging due to the low accuracy but high cost of diagnosis.
Research team led by Professors Chan Beum Park and Steve Park described an ultrasensitive detection of multiple Alzheimer's disease core biomarker in human plasma. The team have designed the sensor array by employing a densely aligned single-walled carbon nanotube thin films as a transducer.
The representative biomarkers of Alzheimer's disease are beta-amyloid42, beta-amyloid40, total tau protein, phosphorylated tau protein and the concentrations of these biomarkers in human plasma are directly correlated with the pathology of Alzheimer’s disease.
The research team developed a highly sensitive resistive biosensor based on densely aligned carbon nanotubes fabricated by Langmuir-Blodgett method with a low manufacturing cost.
Aligned carbon nanotubes with high density minimizes the tube-to-tube junction resistance compared with randomly distributed carbon nanotubes, which leads to the improvement of sensor sensitivity. To be more specific, this resistive sensor with densely aligned carbon nanotubes exhibits a sensitivity over 100 times higher than that of conventional carbon nanotube-based biosensors.
By measuring the concentrations of four Alzheimer’s disease biomarkers simultaneously Alzheimer patients can be discriminated from health controls with an average sensitivity of 90.0%, a selectivity of 90.0% and an average accuracy of 88.6%.
This work, titled “Clinically accurate diagnosis of Alzheimer’s disease via multiplexed sensing of core biomarkers in human plasma”, were published in Nature Communications on January 8th 2020. The authors include PhD candidate Kayoung Kim and MS candidate Min-Ji Kim.
Professor Steve Park said, “This study was conducted on patients who are already confirmed with Alzheimer’s Disease. For further use in practical setting, it is necessary to test the patients with mild cognitive impairment.” He also emphasized that, “It is essential to establish a nationwide infrastructure, such as mild cognitive impairment cohort study and a dementia cohort study. This would enable the establishment of world-wide research network, and will help various private and public institutions.”
This research was supported by the Ministry of Science and ICT, Human Resource Bank of Chungnam National University Hospital and Chungbuk National University Hospital.
< A schematic diagram of a high-density aligned carbon nanotube-based resistive sensor that distinguishes patients with Alzheimer’s Disease by measuring the concentration of four biomarkers in the blood. >
Profile:
Professor Steve Park
stevepark@kaist.ac.kr
Department of Materials Science and Engineering
http://steveparklab.kaist.ac.kr/
KAIST
Profile:
Professor Chan Beum Park
parkcb at kaist.ac.kr
Department of Materials Science and Engineering
http://biomaterials.kaist.ac.kr/
KAIST
2020.02.07 View 12274
Blood-Based Multiplexed Diagnostic Sensor Helps to Accurately Detect Alzheimer’s Disease
A research team at KAIST reported clinically accurate multiplexed electrical biosensor for detecting Alzheimer’s disease by measuring its core biomarkers using densely aligned carbon nanotubes.
Alzheimer’s disease is the most prevalent neurodegenerative disorder, affecting one in ten aged over 65 years. Early diagnosis can reduce the risk of suffering the disease by one-third, according to recent reports. However, its early diagnosis remains challenging due to the low accuracy but high cost of diagnosis.
Research team led by Professors Chan Beum Park and Steve Park described an ultrasensitive detection of multiple Alzheimer's disease core biomarker in human plasma. The team have designed the sensor array by employing a densely aligned single-walled carbon nanotube thin films as a transducer.
The representative biomarkers of Alzheimer's disease are beta-amyloid42, beta-amyloid40, total tau protein, phosphorylated tau protein and the concentrations of these biomarkers in human plasma are directly correlated with the pathology of Alzheimer’s disease.
The research team developed a highly sensitive resistive biosensor based on densely aligned carbon nanotubes fabricated by Langmuir-Blodgett method with a low manufacturing cost.
Aligned carbon nanotubes with high density minimizes the tube-to-tube junction resistance compared with randomly distributed carbon nanotubes, which leads to the improvement of sensor sensitivity. To be more specific, this resistive sensor with densely aligned carbon nanotubes exhibits a sensitivity over 100 times higher than that of conventional carbon nanotube-based biosensors.
By measuring the concentrations of four Alzheimer’s disease biomarkers simultaneously Alzheimer patients can be discriminated from health controls with an average sensitivity of 90.0%, a selectivity of 90.0% and an average accuracy of 88.6%.
This work, titled “Clinically accurate diagnosis of Alzheimer’s disease via multiplexed sensing of core biomarkers in human plasma”, were published in Nature Communications on January 8th 2020. The authors include PhD candidate Kayoung Kim and MS candidate Min-Ji Kim.
Professor Steve Park said, “This study was conducted on patients who are already confirmed with Alzheimer’s Disease. For further use in practical setting, it is necessary to test the patients with mild cognitive impairment.” He also emphasized that, “It is essential to establish a nationwide infrastructure, such as mild cognitive impairment cohort study and a dementia cohort study. This would enable the establishment of world-wide research network, and will help various private and public institutions.”
This research was supported by the Ministry of Science and ICT, Human Resource Bank of Chungnam National University Hospital and Chungbuk National University Hospital.
< A schematic diagram of a high-density aligned carbon nanotube-based resistive sensor that distinguishes patients with Alzheimer’s Disease by measuring the concentration of four biomarkers in the blood. >
Profile:
Professor Steve Park
stevepark@kaist.ac.kr
Department of Materials Science and Engineering
http://steveparklab.kaist.ac.kr/
KAIST
Profile:
Professor Chan Beum Park
parkcb at kaist.ac.kr
Department of Materials Science and Engineering
http://biomaterials.kaist.ac.kr/
KAIST
2020.02.07 View 12274 -
 Scientists Discover the Mechanism of DNA High-Order Structure Formation
(Molecular structures of Abo1 in different energy states (left), Demonstration of an Abo1-assisted histone loading onto DNA by the DNA curtain assay. )
The genetic material of our cells—DNA—exists in a high-order structure called “chromatin”. Chromatin consists of DNA wrapped around histone proteins and efficiently packs DNA into a small volume. Moreover, using a spool and thread analogy, chromatin allows DNA to be locally wound or unwound, thus enabling genes to be enclosed or exposed. The misregulation of chromatin structures results in aberrant gene expression and can ultimately lead to developmental disorders or cancers. Despite the importance of DNA high-order structures, the complexity of the underlying machinery has circumvented molecular dissection.
For the first time, molecular biologists have uncovered how one particular mechanism uses energy to ensure proper histone placement onto DNA to form chromatin. They published their results on Dec. 17 in Nature Communications.
The study focused on proteins called histone chaperones. Histone chaperones are responsible for adding and removing specific histones at specific times during the DNA packaging process. The wrong histone at the wrong time and place could result in the misregulation of gene expression or aberrant DNA replication. Thus, histone chaperones are key players in the assembly and disassembly of chromatin.
“In order to carefully control the assembly and disassembly of chromatin units, histone chaperones act as molecular escorts that prevent histone aggregation and undesired interactions,” said Professor Ji-Joon Song in the Department of Biological Sciences at KAIST. “We set out to understand how a unique histone chaperone uses chemical energy to assemble or disassemble chromatin.”
Song and his team looked to Abo1, the only known histone chaperone that utilizes cellular energy (ATP). While Abo1 is found in yeast, it has an analogous partner in other organisms, including humans, called ATAD2. Both use ATP, which is produced through a cellular process where enzymes break down a molecule’s phosphate bond. ATP energy is typically used to power other cellular processes, but it is a rare partner for histone chaperones.
“This was an interesting problem in the field because all other histone chaperones studied to date do not use ATP,” Song said.
By imaging Abo1 with a single-molecule fluorescence imaging technique known as the DNA curtain assay, the researchers could examine the protein interactions at the single-molecule level. The technique allows scientists to arrange the DNA molecules and proteins on a single layer of a microfluidic chamber and examine the layer with fluorescence microscopy.
The researchers found through real-time observation that Abo1 is ring-shaped and changes its structure to accommodate a specific histone and deposit it on DNA. Moreover, they found that the accommodating structural changes are powered by ADP.
“We discovered a mechanism by which Abo1 accommodates histone substrates, ultimately allowing it to function as a unique energy-dependent histone chaperone,” Song said. “We also found that despite looking like a protein disassembly machine, Abo1 actually loads histone substrates onto DNA to facilitate chromatin assembly.”
The researchers plan to continue exploring how energy-dependent histone chaperones bind and release histones, with the ultimate goal of developing therapeutics that can target cancer-causing misbehavior by Abo1’s analogous human counterpart, ATAD2.
-Profile
Professor Ji-Joon Song
Department of Biological Sciences KI for the BioCentury (https://kis.kaist.ac.kr/index.php?mid=KIB_O) KAIST
2020.01.07 View 11788
Scientists Discover the Mechanism of DNA High-Order Structure Formation
(Molecular structures of Abo1 in different energy states (left), Demonstration of an Abo1-assisted histone loading onto DNA by the DNA curtain assay. )
The genetic material of our cells—DNA—exists in a high-order structure called “chromatin”. Chromatin consists of DNA wrapped around histone proteins and efficiently packs DNA into a small volume. Moreover, using a spool and thread analogy, chromatin allows DNA to be locally wound or unwound, thus enabling genes to be enclosed or exposed. The misregulation of chromatin structures results in aberrant gene expression and can ultimately lead to developmental disorders or cancers. Despite the importance of DNA high-order structures, the complexity of the underlying machinery has circumvented molecular dissection.
For the first time, molecular biologists have uncovered how one particular mechanism uses energy to ensure proper histone placement onto DNA to form chromatin. They published their results on Dec. 17 in Nature Communications.
The study focused on proteins called histone chaperones. Histone chaperones are responsible for adding and removing specific histones at specific times during the DNA packaging process. The wrong histone at the wrong time and place could result in the misregulation of gene expression or aberrant DNA replication. Thus, histone chaperones are key players in the assembly and disassembly of chromatin.
“In order to carefully control the assembly and disassembly of chromatin units, histone chaperones act as molecular escorts that prevent histone aggregation and undesired interactions,” said Professor Ji-Joon Song in the Department of Biological Sciences at KAIST. “We set out to understand how a unique histone chaperone uses chemical energy to assemble or disassemble chromatin.”
Song and his team looked to Abo1, the only known histone chaperone that utilizes cellular energy (ATP). While Abo1 is found in yeast, it has an analogous partner in other organisms, including humans, called ATAD2. Both use ATP, which is produced through a cellular process where enzymes break down a molecule’s phosphate bond. ATP energy is typically used to power other cellular processes, but it is a rare partner for histone chaperones.
“This was an interesting problem in the field because all other histone chaperones studied to date do not use ATP,” Song said.
By imaging Abo1 with a single-molecule fluorescence imaging technique known as the DNA curtain assay, the researchers could examine the protein interactions at the single-molecule level. The technique allows scientists to arrange the DNA molecules and proteins on a single layer of a microfluidic chamber and examine the layer with fluorescence microscopy.
The researchers found through real-time observation that Abo1 is ring-shaped and changes its structure to accommodate a specific histone and deposit it on DNA. Moreover, they found that the accommodating structural changes are powered by ADP.
“We discovered a mechanism by which Abo1 accommodates histone substrates, ultimately allowing it to function as a unique energy-dependent histone chaperone,” Song said. “We also found that despite looking like a protein disassembly machine, Abo1 actually loads histone substrates onto DNA to facilitate chromatin assembly.”
The researchers plan to continue exploring how energy-dependent histone chaperones bind and release histones, with the ultimate goal of developing therapeutics that can target cancer-causing misbehavior by Abo1’s analogous human counterpart, ATAD2.
-Profile
Professor Ji-Joon Song
Department of Biological Sciences KI for the BioCentury (https://kis.kaist.ac.kr/index.php?mid=KIB_O) KAIST
2020.01.07 View 11788 -
 New IEEE Fellow, Professor Jong Chul Ye
Professor Jong Chul Ye from the Department of Bio and Brain Engineering was named a new fellow of the Institute of Electrical and Electronics Engineers (IEEE). IEEE announced this on December 1 in recognition of Professor Ye’s contributions to the development of signal processing and artificial intelligence (AI) technology in the field of biomedical imaging.
As the world’s largest society in the electrical and electronics field, IEEE names the top 0.1% of their members as fellows based on their research achievements.Professor Ye has published more than 100 research papers in world-leading journals in the biomedical imaging field, including those affiliated with IEEE.
He also gave a keynote talk at the yearly conference of the International Society for Magnetic Resonance Imaging (ISMRM) on medical AI technology. In addition, Professor Ye has been appointed to serve as the next chair of the Computational Imaging Technical Committee of the IEEE Signal Processing Society, and the chair of the IEEE Symposium on Biomedical Imaging (ISBI) 2020 to be held in April in Iowa, USA.
Professor Ye said, “The importance of AI technology is developing in the biomedical imaging field. I feel proud that my contributions have been internationally recognized and allowed me to be named an IEEE fellow.”
2019.12.18 View 11543
New IEEE Fellow, Professor Jong Chul Ye
Professor Jong Chul Ye from the Department of Bio and Brain Engineering was named a new fellow of the Institute of Electrical and Electronics Engineers (IEEE). IEEE announced this on December 1 in recognition of Professor Ye’s contributions to the development of signal processing and artificial intelligence (AI) technology in the field of biomedical imaging.
As the world’s largest society in the electrical and electronics field, IEEE names the top 0.1% of their members as fellows based on their research achievements.Professor Ye has published more than 100 research papers in world-leading journals in the biomedical imaging field, including those affiliated with IEEE.
He also gave a keynote talk at the yearly conference of the International Society for Magnetic Resonance Imaging (ISMRM) on medical AI technology. In addition, Professor Ye has been appointed to serve as the next chair of the Computational Imaging Technical Committee of the IEEE Signal Processing Society, and the chair of the IEEE Symposium on Biomedical Imaging (ISBI) 2020 to be held in April in Iowa, USA.
Professor Ye said, “The importance of AI technology is developing in the biomedical imaging field. I feel proud that my contributions have been internationally recognized and allowed me to be named an IEEE fellow.”
2019.12.18 View 11543 -
 Professor Il-Doo Kim Named Scientist of the Year by the Journalists
Professor Il-Doo Kim from the Department of Materials Science and Engineering was named the 2019 Scientist of the Year by Korean science journalists. The award was conferred at the 2019 Science Press Night ceremony of the Korea Science Journalists Association (KSJA) on November 29.
Professor Kim focuses on developing nanofiber gas sensors for diagnosing diseases in advance by analyzing exhaled biomarkers with electrospinning technology. His outstanding research was praised and selected as one of the top 10 nanotechnology of 2019 by the Korea Nano Technology Research Society (KoNTRS), the Ministry of Science and ICT (MSIT), and the Ministry of Trade, Industry and Energy (MOTIE).
Professor Kim was honored with the QIAN Baojun Fiber Award, which is awarded every two years by Donghua University in Shanghai, China to recognize outstanding contributions in fiber science and technology. Professor Kim was also elected as an academician of the Asia Pacific Academy of Materials (APAM) on November 21 in Guangzhou, China.
In May, Professor Kim was appointed as an associate editor of ACS Nano, a leading international research journal in the field of nanoscience. In his editorial published in the May issue of ACS Nano, Professor Kim introduced and shared the history of KAIST and its vision for the future with other members of the journal. He hopes this will help with promoting a closer relationship between the members of the journal and KAIST moving forward.
“Above all,” he said in his acceptance speech, “the greatest news for me as an educator is that the first PhD graduate from our lab, Dr. Seonjin Choi, was appointed as the youngest professor in the Division of Materials Science and Engineering at Hanyang University on September 1.”
2019.12.17 View 12825
Professor Il-Doo Kim Named Scientist of the Year by the Journalists
Professor Il-Doo Kim from the Department of Materials Science and Engineering was named the 2019 Scientist of the Year by Korean science journalists. The award was conferred at the 2019 Science Press Night ceremony of the Korea Science Journalists Association (KSJA) on November 29.
Professor Kim focuses on developing nanofiber gas sensors for diagnosing diseases in advance by analyzing exhaled biomarkers with electrospinning technology. His outstanding research was praised and selected as one of the top 10 nanotechnology of 2019 by the Korea Nano Technology Research Society (KoNTRS), the Ministry of Science and ICT (MSIT), and the Ministry of Trade, Industry and Energy (MOTIE).
Professor Kim was honored with the QIAN Baojun Fiber Award, which is awarded every two years by Donghua University in Shanghai, China to recognize outstanding contributions in fiber science and technology. Professor Kim was also elected as an academician of the Asia Pacific Academy of Materials (APAM) on November 21 in Guangzhou, China.
In May, Professor Kim was appointed as an associate editor of ACS Nano, a leading international research journal in the field of nanoscience. In his editorial published in the May issue of ACS Nano, Professor Kim introduced and shared the history of KAIST and its vision for the future with other members of the journal. He hopes this will help with promoting a closer relationship between the members of the journal and KAIST moving forward.
“Above all,” he said in his acceptance speech, “the greatest news for me as an educator is that the first PhD graduate from our lab, Dr. Seonjin Choi, was appointed as the youngest professor in the Division of Materials Science and Engineering at Hanyang University on September 1.”
2019.12.17 View 12825 -
 Tungsten Suboxide Improves the Efficiency of Platinum in Hydrogen Production
< PhD Candidate Jinkyu Park and Professor Jinwoo Lee >
Researchers presented a new strategy for enhancing catalytic activity using tungsten suboxide as a single-atom catalyst (SAC). This strategy, which significantly improves hydrogen evolution reaction (HER) in metal platinum (pt) by 16.3 times, sheds light on the development of new electrochemical catalyst technologies.
Hydrogen has been touted as a promising alternative to fossil fuels. However, most of the conventional industrial hydrogen production methods come with environmental issues, releasing significant amounts of carbon dioxide and greenhouse gases.
Electrochemical water splitting is considered a potential approach for clean hydrogen production. Pt is one of the most commonly used catalysts to improve HER performance in electrochemical water splitting, but the high cost and scarcity of Pt remain key obstacles to mass commercial applications.
SACs, where all metal species are individually dispersed on a desired support material, have been identified as one way to reduce the amount of Pt usage, as they offer the maximum number of surface exposed Pt atoms.
Inspired by earlier studies, which mainly focused on SACs supported by carbon-based materials, a KAIST research team led by Professor Jinwoo Lee from the Department of Chemical and Biomolecular Engineering investigated the influence of support materials on the performance of SACs.
Professor Lee and his researchers suggested mesoporous tungsten suboxide as a new support material for atomically dispersed Pt, as this was expected to provide high electronic conductivity and have a synergetic effect with Pt.
They compared the performance of single-atom Pt supported by carbon and tungsten suboxide respectively. The results revealed that the support effect occurred with tungsten suboxide, in which the mass activity of a single-atom Pt supported by tungsten suboxide was 2.1 times greater than that of single-atom Pt supported by carbon, and 16.3 times higher than that of Pt nanoparticles supported by carbon.
The team indicated a change in the electronic structure of Pt via charge transfer from tungsten suboxide to Pt. This phenomenon was reported as a result of strong metal-support interaction between Pt and tungsten suboxide.
HER performance can be improved not only by changing the electronic structure of the supported metal, but also by inducing another support effect, the spillover effect, the research group reported. Hydrogen spillover is a phenomenon where adsorbed hydrogen migrates from one surface to another, and it occurs more easily as the Pt size becomes smaller.
The researchers compared the performance of single-atom Pt and Pt nanoparticles supported by tungsten suboxide. The single-atom Pt supported by tungsten suboxide exhibited a higher degree of hydrogen spillover phenomenon, which enhanced the Pt mass activity for hydrogen evolution up to 10.7 times compared to Pt nanoparticles supported by tungsten suboxide.
Professor Lee said, “Choosing the right support material is important for improving electrocatalysis in hydrogen production. The tungsten suboxide catalyst we used to support Pt in our study implies that interactions between the well-matched metal and support can drastically enhance the efficiency of the process.”
This research was supported by the Ministry of Science and ICT and introduced in the International Edition of the German journal Angewandte Chemie.
Figure. Schematic representation of hydrogen evolution reaction (HER) of pseudo single-atom Pt supported by tungsten suboxide
-Publication
Jinkyu Park, Dr. Seonggyu Lee, Hee-Eun Kim, Ara Cho, Seongbeen Kim, Dr. Youngjin Ye, Prof. Jeong Woo Han, Prof. Hyunjoo Lee, Dr. Jong Hyun Jang, and Prof. Jinwoo Lee. 2019. Investigation of the Support Effect in Atomically Dispersed Pt on WO3−x for Utilization of Pt in the Hydrogen Evolution Reaction. International Edition of Angewandte Chemie. Volume No. 58. Issue No. 45. 6 pages. https://doi.org/10.1002/anie.201908122
-ProfileProfessor Jinwoo LeeConvergence of Energy and Nano Science Laboratoryhttp://cens.kaist.ac.kr
Department of Chemical and Biomolecular EngineeringKAIST
2019.10.28 View 22913
Tungsten Suboxide Improves the Efficiency of Platinum in Hydrogen Production
< PhD Candidate Jinkyu Park and Professor Jinwoo Lee >
Researchers presented a new strategy for enhancing catalytic activity using tungsten suboxide as a single-atom catalyst (SAC). This strategy, which significantly improves hydrogen evolution reaction (HER) in metal platinum (pt) by 16.3 times, sheds light on the development of new electrochemical catalyst technologies.
Hydrogen has been touted as a promising alternative to fossil fuels. However, most of the conventional industrial hydrogen production methods come with environmental issues, releasing significant amounts of carbon dioxide and greenhouse gases.
Electrochemical water splitting is considered a potential approach for clean hydrogen production. Pt is one of the most commonly used catalysts to improve HER performance in electrochemical water splitting, but the high cost and scarcity of Pt remain key obstacles to mass commercial applications.
SACs, where all metal species are individually dispersed on a desired support material, have been identified as one way to reduce the amount of Pt usage, as they offer the maximum number of surface exposed Pt atoms.
Inspired by earlier studies, which mainly focused on SACs supported by carbon-based materials, a KAIST research team led by Professor Jinwoo Lee from the Department of Chemical and Biomolecular Engineering investigated the influence of support materials on the performance of SACs.
Professor Lee and his researchers suggested mesoporous tungsten suboxide as a new support material for atomically dispersed Pt, as this was expected to provide high electronic conductivity and have a synergetic effect with Pt.
They compared the performance of single-atom Pt supported by carbon and tungsten suboxide respectively. The results revealed that the support effect occurred with tungsten suboxide, in which the mass activity of a single-atom Pt supported by tungsten suboxide was 2.1 times greater than that of single-atom Pt supported by carbon, and 16.3 times higher than that of Pt nanoparticles supported by carbon.
The team indicated a change in the electronic structure of Pt via charge transfer from tungsten suboxide to Pt. This phenomenon was reported as a result of strong metal-support interaction between Pt and tungsten suboxide.
HER performance can be improved not only by changing the electronic structure of the supported metal, but also by inducing another support effect, the spillover effect, the research group reported. Hydrogen spillover is a phenomenon where adsorbed hydrogen migrates from one surface to another, and it occurs more easily as the Pt size becomes smaller.
The researchers compared the performance of single-atom Pt and Pt nanoparticles supported by tungsten suboxide. The single-atom Pt supported by tungsten suboxide exhibited a higher degree of hydrogen spillover phenomenon, which enhanced the Pt mass activity for hydrogen evolution up to 10.7 times compared to Pt nanoparticles supported by tungsten suboxide.
Professor Lee said, “Choosing the right support material is important for improving electrocatalysis in hydrogen production. The tungsten suboxide catalyst we used to support Pt in our study implies that interactions between the well-matched metal and support can drastically enhance the efficiency of the process.”
This research was supported by the Ministry of Science and ICT and introduced in the International Edition of the German journal Angewandte Chemie.
Figure. Schematic representation of hydrogen evolution reaction (HER) of pseudo single-atom Pt supported by tungsten suboxide
-Publication
Jinkyu Park, Dr. Seonggyu Lee, Hee-Eun Kim, Ara Cho, Seongbeen Kim, Dr. Youngjin Ye, Prof. Jeong Woo Han, Prof. Hyunjoo Lee, Dr. Jong Hyun Jang, and Prof. Jinwoo Lee. 2019. Investigation of the Support Effect in Atomically Dispersed Pt on WO3−x for Utilization of Pt in the Hydrogen Evolution Reaction. International Edition of Angewandte Chemie. Volume No. 58. Issue No. 45. 6 pages. https://doi.org/10.1002/anie.201908122
-ProfileProfessor Jinwoo LeeConvergence of Energy and Nano Science Laboratoryhttp://cens.kaist.ac.kr
Department of Chemical and Biomolecular EngineeringKAIST
2019.10.28 View 22913 -
 A Single, Master Switch for Sugar Levels?
When a fly eats sugar, a single brain cell sends simultaneous messages to stimulate one hormone and inhibit another to control glucose levels in the body. Further research into this control system with remarkable precision could shed light on the neural mechanisms of diabetes and obesity in humans .
A single neuron appears to monitor and control sugar levels in the fly body, according to research published this week in Nature. This new insight into the mechanisms in the fly brain that maintain a balance of two key hormones controlling glucose levels, insulin and glucagon, can provide a framework for understanding diabetes and obesity in humans.
Neurons that sense and respond to glucose were identified more than 50 years ago, but what they do in our body has remained unclear. Researchers at the Korea Advanced Institute of Science and Technology (KAIST) and New York University School of Medicine have now found a single “glucose-sensing neuron” that appears to be the master controller in Drosophila, the vinegar fly, for maintaining an ideal glucose balance, called homeostasis.
Professor Greg Seong-Bae Suh, Dr. Yangkyun Oh and colleagues identified a key neuron that is excited by glucose, which they called CN neuron. This CN neuron has a unique shape – it has an axon (which is used to transmit information to downstream cells) that is bifurcated. One branch projects to insulin-producing cells, and sends a signal triggering the secretion of the insulin equivalent in flies. The other branch projects to glucagon-producing cells and sends a signal inhibiting the secretion of the glucagon equivalent.
When flies consume food, the levels of glucose in their body increase; this excites the CN neuron, which fires the simultaneous signals to stimulate insulin and inhibit glucagon secretion, thereby maintaining the appropriate balance between the hormones and sugar in the blood. The researchers were able to see this happening in the brain in real time by using a combination of cutting-edge fluorescent calcium imaging technology, as well as measuring hormone and sugar levels and applying highly sophisticated molecular genetic techniques.
When flies were not fed, however, the researchers observed a reduction in the activity of CN neuron, a reduction in insulin secretion and an increase in glucagon secretion. These findings indicate that these key hormones are under the direct control of the glucose-sensing neuron. Furthermore, when they silenced the CN neuron rendering dysfunctional CN neuron in flies, these animals experienced an imbalance, resulting in hyperglycemia – high levels of sugars in the blood, similar to what is observed in diabetes in humans. This further suggests that the CN neuron is critical to maintaining glucose homeostasis in animals.
While further research is required to investigate this process in humans, Suh notes this is a significant step forward in the fields of both neurobiology and endocrinology.
“This work lays the foundation for translational research to better understand how this delicate regulatory process is affected by diabetes, obesity, excessive nutrition and diets high in sugar,” Suh said.
Profile: Greg Seong-Bae Suh
seongbaesuh@kaist.ac.kr
Professor Department of Biological Sciences
KAIST
(Figure: A single glucose-excited CN neuron extends bifurcated axonal branches,
one of which innervates insulin producing cells and stimulates their activity an the other axonal branch projects to glucagon producing cells and inhibits their activity.)
2019.10.24 View 18733
A Single, Master Switch for Sugar Levels?
When a fly eats sugar, a single brain cell sends simultaneous messages to stimulate one hormone and inhibit another to control glucose levels in the body. Further research into this control system with remarkable precision could shed light on the neural mechanisms of diabetes and obesity in humans .
A single neuron appears to monitor and control sugar levels in the fly body, according to research published this week in Nature. This new insight into the mechanisms in the fly brain that maintain a balance of two key hormones controlling glucose levels, insulin and glucagon, can provide a framework for understanding diabetes and obesity in humans.
Neurons that sense and respond to glucose were identified more than 50 years ago, but what they do in our body has remained unclear. Researchers at the Korea Advanced Institute of Science and Technology (KAIST) and New York University School of Medicine have now found a single “glucose-sensing neuron” that appears to be the master controller in Drosophila, the vinegar fly, for maintaining an ideal glucose balance, called homeostasis.
Professor Greg Seong-Bae Suh, Dr. Yangkyun Oh and colleagues identified a key neuron that is excited by glucose, which they called CN neuron. This CN neuron has a unique shape – it has an axon (which is used to transmit information to downstream cells) that is bifurcated. One branch projects to insulin-producing cells, and sends a signal triggering the secretion of the insulin equivalent in flies. The other branch projects to glucagon-producing cells and sends a signal inhibiting the secretion of the glucagon equivalent.
When flies consume food, the levels of glucose in their body increase; this excites the CN neuron, which fires the simultaneous signals to stimulate insulin and inhibit glucagon secretion, thereby maintaining the appropriate balance between the hormones and sugar in the blood. The researchers were able to see this happening in the brain in real time by using a combination of cutting-edge fluorescent calcium imaging technology, as well as measuring hormone and sugar levels and applying highly sophisticated molecular genetic techniques.
When flies were not fed, however, the researchers observed a reduction in the activity of CN neuron, a reduction in insulin secretion and an increase in glucagon secretion. These findings indicate that these key hormones are under the direct control of the glucose-sensing neuron. Furthermore, when they silenced the CN neuron rendering dysfunctional CN neuron in flies, these animals experienced an imbalance, resulting in hyperglycemia – high levels of sugars in the blood, similar to what is observed in diabetes in humans. This further suggests that the CN neuron is critical to maintaining glucose homeostasis in animals.
While further research is required to investigate this process in humans, Suh notes this is a significant step forward in the fields of both neurobiology and endocrinology.
“This work lays the foundation for translational research to better understand how this delicate regulatory process is affected by diabetes, obesity, excessive nutrition and diets high in sugar,” Suh said.
Profile: Greg Seong-Bae Suh
seongbaesuh@kaist.ac.kr
Professor Department of Biological Sciences
KAIST
(Figure: A single glucose-excited CN neuron extends bifurcated axonal branches,
one of which innervates insulin producing cells and stimulates their activity an the other axonal branch projects to glucagon producing cells and inhibits their activity.)
2019.10.24 View 18733 -
 A Mathematical Model Reveals Long-Distance Cell Communication Mechanism
How can tens of thousands of people in a large football stadium all clap together with the same beat even though they can only hear the people near them clapping?
A combination of a partial differential equation and a synthetic circuit in microbes answers this question. An interdisciplinary collaborative team of Professor Jae Kyoung Kim at KAIST, Professor Krešimir Josić at the University of Houston, and Professor Matt Bennett at Rice University has identified how a large community can communicate with each other almost simultaneously even with very short distance signaling. The research was reported at Nature Chemical Biology.
Cells often communicate using signaling molecules, which can travel only a short distance. Nevertheless, the cells can also communicate over large distances to spur collective action. The team revealed a cell communication mechanism that quickly forms a network of local interactions to spur collective action, even in large communities.
The research team used an engineered transcriptional circuit of combined positive and negative feedback loops in E. coli, which can periodically release two types of signaling molecules: activator and repressor. As the signaling molecules travel over a short distance, cells can only talk to their nearest neighbors. However, cell communities synchronize oscillatory gene expression in spatially extended systems as long as the transcriptional circuit contains a positive feedback loop for the activator.
Professor Kim said that analyzing and understanding such high-dimensional dynamics was extremely difficult. He explained, “That’s why we used high-dimensional partial differential equation to describe the system based on the interactions among various types of molecules.” Surprisingly, the mathematical model accurately simulates the synthesis of the signaling molecules in the cell and their spatial diffusion throughout the chamber and their effect on neighboring cells.
The team simplified the high-dimensional system into a one-dimensional orbit, noting that the system repeats periodically. This allowed them to discover that cells can make one voice when they lowered their own voice and listened to the others. “It turns out the positive feedback loop reduces the distance between moving points and finally makes them move all together. That’s why you clap louder when you hear applause from nearby neighbors and everyone eventually claps together at almost the same time,” said Professor Kim.
Professor Kim added, “Math is a powerful as it simplifies complex thing so that we can find an essential underlying property. This finding would not have been possible without the simplification of complex systems using mathematics."
The National Institutes of Health, the National Science Foundation, the Robert A. Welch Foundation, the Hamill Foundation, the National Research Foundation of Korea, and the T.J. Park Science Fellowship of POSCO supported the research.
(Figure: Complex molecular interactions among microbial consortia is simplified as interactions among points on a limit cycle (right).)
2019.10.15 View 29575
A Mathematical Model Reveals Long-Distance Cell Communication Mechanism
How can tens of thousands of people in a large football stadium all clap together with the same beat even though they can only hear the people near them clapping?
A combination of a partial differential equation and a synthetic circuit in microbes answers this question. An interdisciplinary collaborative team of Professor Jae Kyoung Kim at KAIST, Professor Krešimir Josić at the University of Houston, and Professor Matt Bennett at Rice University has identified how a large community can communicate with each other almost simultaneously even with very short distance signaling. The research was reported at Nature Chemical Biology.
Cells often communicate using signaling molecules, which can travel only a short distance. Nevertheless, the cells can also communicate over large distances to spur collective action. The team revealed a cell communication mechanism that quickly forms a network of local interactions to spur collective action, even in large communities.
The research team used an engineered transcriptional circuit of combined positive and negative feedback loops in E. coli, which can periodically release two types of signaling molecules: activator and repressor. As the signaling molecules travel over a short distance, cells can only talk to their nearest neighbors. However, cell communities synchronize oscillatory gene expression in spatially extended systems as long as the transcriptional circuit contains a positive feedback loop for the activator.
Professor Kim said that analyzing and understanding such high-dimensional dynamics was extremely difficult. He explained, “That’s why we used high-dimensional partial differential equation to describe the system based on the interactions among various types of molecules.” Surprisingly, the mathematical model accurately simulates the synthesis of the signaling molecules in the cell and their spatial diffusion throughout the chamber and their effect on neighboring cells.
The team simplified the high-dimensional system into a one-dimensional orbit, noting that the system repeats periodically. This allowed them to discover that cells can make one voice when they lowered their own voice and listened to the others. “It turns out the positive feedback loop reduces the distance between moving points and finally makes them move all together. That’s why you clap louder when you hear applause from nearby neighbors and everyone eventually claps together at almost the same time,” said Professor Kim.
Professor Kim added, “Math is a powerful as it simplifies complex thing so that we can find an essential underlying property. This finding would not have been possible without the simplification of complex systems using mathematics."
The National Institutes of Health, the National Science Foundation, the Robert A. Welch Foundation, the Hamill Foundation, the National Research Foundation of Korea, and the T.J. Park Science Fellowship of POSCO supported the research.
(Figure: Complex molecular interactions among microbial consortia is simplified as interactions among points on a limit cycle (right).)
2019.10.15 View 29575 -
 Two Professors Recognized for the National R&D Excellence 100
< Professor Haeng-Ki Lee (left) and Professor Jeong-Ho Lee (right) >
Two KAIST professors were listed among the 2019 National R&D Excellence 100 announced by the Ministry of Science and ICT and the Korea Institute of S&T Evaluation and Planning.
Professor Haeng-Ki Lee from the Department of Civil and Environmental Engineering was recognized in the field of mechanics and materials for his research on developing new construction materials through the convergence of nano- and biotechnologies.
In the field of life and marine science, Professor Jeong-Ho Lee from the Graduate School of Medical Science and Engineering was lauded for his research of diagnostic tools and therapies for glioblastoma and pediatric brain tumors.
A certificate from the Minister of Ministry of Science and ICT will be conferred to these two professors, and their names will be inscribed on a special 2019 National R&D Excellence 100 plaque to celebrate their achievements. The professors will also be given privileges during the process of new R&D project selection.
(END)
2019.10.15 View 13707
Two Professors Recognized for the National R&D Excellence 100
< Professor Haeng-Ki Lee (left) and Professor Jeong-Ho Lee (right) >
Two KAIST professors were listed among the 2019 National R&D Excellence 100 announced by the Ministry of Science and ICT and the Korea Institute of S&T Evaluation and Planning.
Professor Haeng-Ki Lee from the Department of Civil and Environmental Engineering was recognized in the field of mechanics and materials for his research on developing new construction materials through the convergence of nano- and biotechnologies.
In the field of life and marine science, Professor Jeong-Ho Lee from the Graduate School of Medical Science and Engineering was lauded for his research of diagnostic tools and therapies for glioblastoma and pediatric brain tumors.
A certificate from the Minister of Ministry of Science and ICT will be conferred to these two professors, and their names will be inscribed on a special 2019 National R&D Excellence 100 plaque to celebrate their achievements. The professors will also be given privileges during the process of new R&D project selection.
(END)
2019.10.15 View 13707 -
 Professor Hyun Gyu Park Appointed as Associate Editor for Biosensors and Bioelectronics
Professor Hyun Gyu Park from the Department of Chemical and Biomolecular Engineering was appointed as an associate editor for Biosensors and Bioelectronics, an international journal published by Elsevier.
Biosensors and Bioelectronics is one of the top SCI journals in the fields of chemistry and analytical science (IF 9.518 as of 2018). Professor Park was recognized and appointed as the associate editor for this journal due to his outstanding research achievements in the fields of nucleic acid engineering, biosensors, and nanobiotechnology.
Professor Park will serve as the associate editor from this October until December 2021.
(END)
2019.10.01 View 8704
Professor Hyun Gyu Park Appointed as Associate Editor for Biosensors and Bioelectronics
Professor Hyun Gyu Park from the Department of Chemical and Biomolecular Engineering was appointed as an associate editor for Biosensors and Bioelectronics, an international journal published by Elsevier.
Biosensors and Bioelectronics is one of the top SCI journals in the fields of chemistry and analytical science (IF 9.518 as of 2018). Professor Park was recognized and appointed as the associate editor for this journal due to his outstanding research achievements in the fields of nucleic acid engineering, biosensors, and nanobiotechnology.
Professor Park will serve as the associate editor from this October until December 2021.
(END)
2019.10.01 View 8704