AND
-
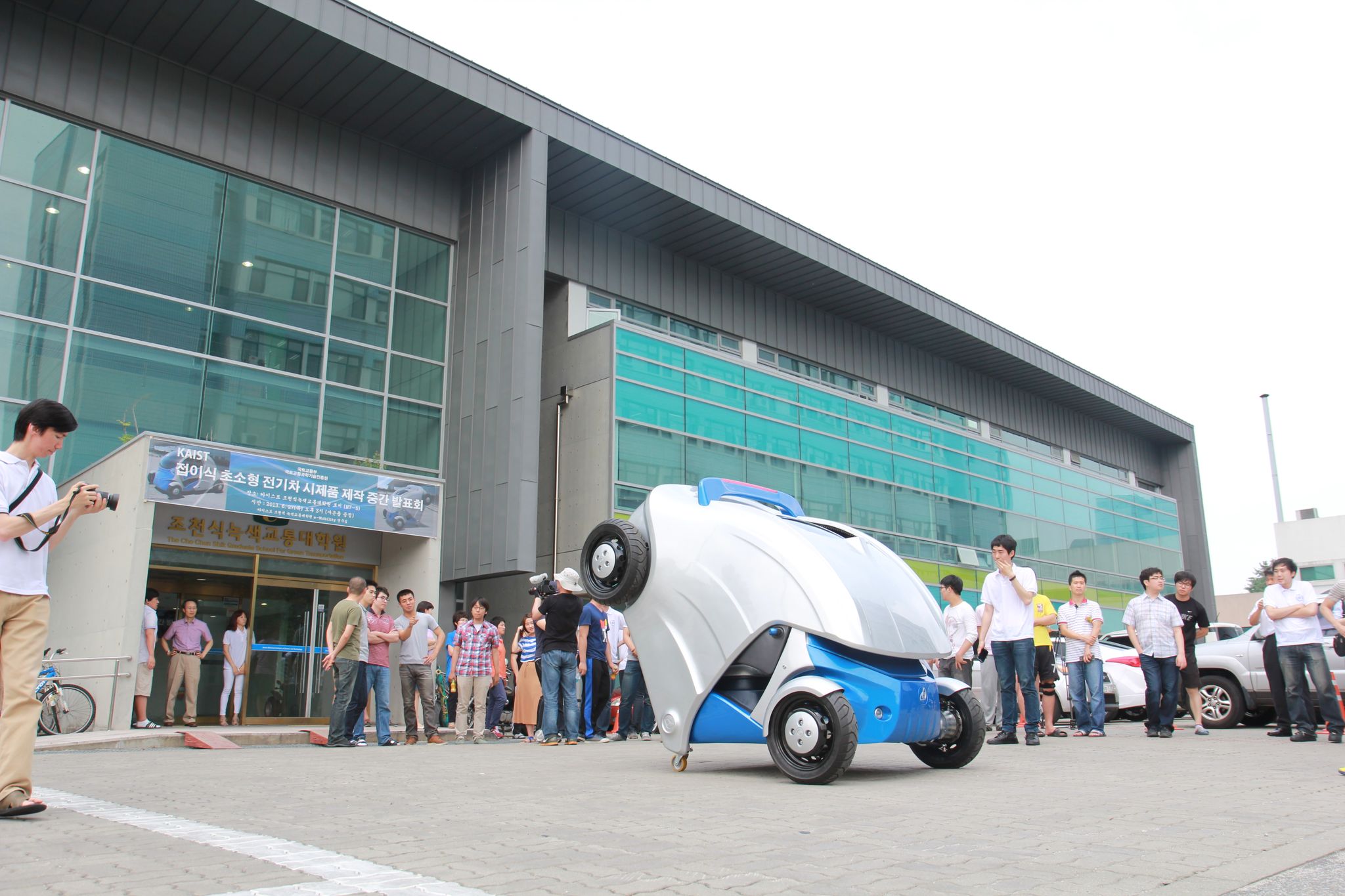 KAIST unveils foldable micro electric car, Armadillo-T
The small and light electric car completely folds in half when parking, making it a perfect fit for public or private transportation in an urban environment.
Looking for a parking space for hours at a busy shopping mall or being stuck on roads jammed with cars releasing large amounts of carbon dioxide are all-too-familiar scenes for city dwellers.
A group of researchers at the Korea Advanced Institute of Science and Technology (KAIST) recently developed a possible solution to such problems: a foldable, compact electric vehicle that can be utilized either as a personal car or part of the public transit system to connect major transportation routes within a city.
In-Soo Suh, associate professor of the Graduate School for Green Transportation at KAIST, and his research team introduced a prototype micro electric car called "Armadillo-T," whose design is based on a native animal of South America, the armadillo, a placental mammal with a leathery armor shell.
The research team imitated the animal"s distinctive protection characteristic of rolling up into a ball when facing with threat from predators. Just as armadillos hide themselves inside the shell, Armadillo-T tucks its rear body away, shrinking its original size of 2.8 meters (110 inches) down to almost half, 1.65 meters (65 inches), when folding.
Armadillo-T is a four-wheel-drive, all-electric car with two seats and four in-wheel motors. Since the motors are installed inside the wheels, and the 13.6 kWh capacity of lithium-ion battery pack is housed on the front side, the battery and motors do not have to change their positions when the car folds. This not only optimizes the energy efficiency but also provides stability and ample room to drivers and passengers.
Once folded, the small and light (weighs 450 kg) electric vehicle takes up only one-third of a 5-meter parking space, the standard parking size in Korea, allowing three of its kind to be parked. With a smartphone-interfaced remote control on the wheels, the vehicle can turn 360 degrees, enhancing drivers" convenience to park the car, even in an odd space in a parking lot, the corner of a building, for example.
Professor In-Soo Suh said, "I expect that people living in cities will eventually shift their preferences from bulky, petro-engine cars to smaller and lighter electric cars. Armadillo-T can be one of the alternatives city drivers can opt for. Particularly, this car is ideal for urban travels, including car-sharing and transit transfer, to offer major transportation links in a city. In addition to the urban application, local near-distance travels such as tourist zones or large buildings can be another example of application."
The concept car has loads of smart features on board, too: the cameras installed inside the car eliminate the need for side mirrors and increase the driver"s ability to see the car"s right and left side, thereby reducing blind spots. With a smartphone, the driver can control Armadillo-T and enable remote folding control. The car has a maximum speed of 60 km/h, and with a ten-minute fast charge, it can run up to 100 km.
Professor Suh explained that the concept of Armadillo-T was originally initiated in 2011 as he focused his research interest on the sub-A segment of personal mobility vehicles (PMVs), which are smaller and lighter than the current compact cars, as a new personalized transport mode.
"In coming years, we will see more mega-size cities established and face more serious environmental problems. Throughout the world, the aging population is rapidly growing as well. To cope with climate, energy, and limited petroleum resources, we really need to think outside the box, once again, to find more convenient and eco-friendly transportation, just as the Ford Model T did in the early 1920s. A further level of R&D, technical standards, and regulatory reviews are required to have these types of micro vehicles or PMVs on the market through test-bed evaluations, but we believe that Armadillo-T is an icon toward the future transport system with technology innovation."
The research project has been supported by the Korean government, the Ministry of Land, Infrastructure and Transport and the Korea Agency for Infrastructure Technology Advancement, since December 2012.Youtube Link: http://www.youtube.com/watch?v=8DoZH7Y-sR0
2013.08.21 View 16724
KAIST unveils foldable micro electric car, Armadillo-T
The small and light electric car completely folds in half when parking, making it a perfect fit for public or private transportation in an urban environment.
Looking for a parking space for hours at a busy shopping mall or being stuck on roads jammed with cars releasing large amounts of carbon dioxide are all-too-familiar scenes for city dwellers.
A group of researchers at the Korea Advanced Institute of Science and Technology (KAIST) recently developed a possible solution to such problems: a foldable, compact electric vehicle that can be utilized either as a personal car or part of the public transit system to connect major transportation routes within a city.
In-Soo Suh, associate professor of the Graduate School for Green Transportation at KAIST, and his research team introduced a prototype micro electric car called "Armadillo-T," whose design is based on a native animal of South America, the armadillo, a placental mammal with a leathery armor shell.
The research team imitated the animal"s distinctive protection characteristic of rolling up into a ball when facing with threat from predators. Just as armadillos hide themselves inside the shell, Armadillo-T tucks its rear body away, shrinking its original size of 2.8 meters (110 inches) down to almost half, 1.65 meters (65 inches), when folding.
Armadillo-T is a four-wheel-drive, all-electric car with two seats and four in-wheel motors. Since the motors are installed inside the wheels, and the 13.6 kWh capacity of lithium-ion battery pack is housed on the front side, the battery and motors do not have to change their positions when the car folds. This not only optimizes the energy efficiency but also provides stability and ample room to drivers and passengers.
Once folded, the small and light (weighs 450 kg) electric vehicle takes up only one-third of a 5-meter parking space, the standard parking size in Korea, allowing three of its kind to be parked. With a smartphone-interfaced remote control on the wheels, the vehicle can turn 360 degrees, enhancing drivers" convenience to park the car, even in an odd space in a parking lot, the corner of a building, for example.
Professor In-Soo Suh said, "I expect that people living in cities will eventually shift their preferences from bulky, petro-engine cars to smaller and lighter electric cars. Armadillo-T can be one of the alternatives city drivers can opt for. Particularly, this car is ideal for urban travels, including car-sharing and transit transfer, to offer major transportation links in a city. In addition to the urban application, local near-distance travels such as tourist zones or large buildings can be another example of application."
The concept car has loads of smart features on board, too: the cameras installed inside the car eliminate the need for side mirrors and increase the driver"s ability to see the car"s right and left side, thereby reducing blind spots. With a smartphone, the driver can control Armadillo-T and enable remote folding control. The car has a maximum speed of 60 km/h, and with a ten-minute fast charge, it can run up to 100 km.
Professor Suh explained that the concept of Armadillo-T was originally initiated in 2011 as he focused his research interest on the sub-A segment of personal mobility vehicles (PMVs), which are smaller and lighter than the current compact cars, as a new personalized transport mode.
"In coming years, we will see more mega-size cities established and face more serious environmental problems. Throughout the world, the aging population is rapidly growing as well. To cope with climate, energy, and limited petroleum resources, we really need to think outside the box, once again, to find more convenient and eco-friendly transportation, just as the Ford Model T did in the early 1920s. A further level of R&D, technical standards, and regulatory reviews are required to have these types of micro vehicles or PMVs on the market through test-bed evaluations, but we believe that Armadillo-T is an icon toward the future transport system with technology innovation."
The research project has been supported by the Korean government, the Ministry of Land, Infrastructure and Transport and the Korea Agency for Infrastructure Technology Advancement, since December 2012.Youtube Link: http://www.youtube.com/watch?v=8DoZH7Y-sR0
2013.08.21 View 16724 -
 2013 International Conference for the Integration of Science, Technology, and Society at KAIST (ICISTS-KAIST)
The International Conference for the Integration of Science, Technology, and Society at KAIST (ICISTS-KAIST) is a global forum organized by KAIST undergraduate students to promote the exchange of ideas and facilitate the discussion of issues that are important to science, technology, society, and higher education. The ICISTS-KAIST conference has been held annually every summer since 2005, inviting distinguished speakers and guests from all around the world to share their insights and expertise with students gathered from Korea and abroad. Last year alone, more than 300 students from 22 nations and 40 speakers participated in the event.
Originally, the ICISTS-KAIST was established by KAIST students who were inspired by the Harvard Project for Asian and International Relations (HPAIR), which is one of the Harvard’s largest annual student conferences in Asia.
This year, 335 students from 103 universities in 22 countries joined the conference that was held on August 5th-9th in Daejeon, making the 2013 ICISTS-KAIST the biggest science and engineering gathering hosted by university students in Asia. About 36% of the participants were international students.
The theme of the conference was “Perfect Alliance: Coexistence for Human Society,” in which students and speakers addressed issues on how to harmonize the speed of scientific progress with the development of important values in society, as well as to explore solutions to overcome the chasm, if any, between the boundaries of science and society.
In his opening remarks, President Steve Kang said, “Creativity and innovation are born out of openness. Therefore, it is essential for young scientists and engineers to communicate with people from different cultural and political backgrounds. Through this kind of global interaction and exchange of ideas and views, students will have an opportunity to deepen their understanding of the world and to better examine the purpose of their intellectual exploration in science and technology.” At the 2013 ICISTS-KAIST, 25 distinguished speakers participated including Walter Bender, a former director of the Media Lab at MIT and David Christian, a professor of Macquarie University in Australia.
2013.08.08 View 11947
2013 International Conference for the Integration of Science, Technology, and Society at KAIST (ICISTS-KAIST)
The International Conference for the Integration of Science, Technology, and Society at KAIST (ICISTS-KAIST) is a global forum organized by KAIST undergraduate students to promote the exchange of ideas and facilitate the discussion of issues that are important to science, technology, society, and higher education. The ICISTS-KAIST conference has been held annually every summer since 2005, inviting distinguished speakers and guests from all around the world to share their insights and expertise with students gathered from Korea and abroad. Last year alone, more than 300 students from 22 nations and 40 speakers participated in the event.
Originally, the ICISTS-KAIST was established by KAIST students who were inspired by the Harvard Project for Asian and International Relations (HPAIR), which is one of the Harvard’s largest annual student conferences in Asia.
This year, 335 students from 103 universities in 22 countries joined the conference that was held on August 5th-9th in Daejeon, making the 2013 ICISTS-KAIST the biggest science and engineering gathering hosted by university students in Asia. About 36% of the participants were international students.
The theme of the conference was “Perfect Alliance: Coexistence for Human Society,” in which students and speakers addressed issues on how to harmonize the speed of scientific progress with the development of important values in society, as well as to explore solutions to overcome the chasm, if any, between the boundaries of science and society.
In his opening remarks, President Steve Kang said, “Creativity and innovation are born out of openness. Therefore, it is essential for young scientists and engineers to communicate with people from different cultural and political backgrounds. Through this kind of global interaction and exchange of ideas and views, students will have an opportunity to deepen their understanding of the world and to better examine the purpose of their intellectual exploration in science and technology.” At the 2013 ICISTS-KAIST, 25 distinguished speakers participated including Walter Bender, a former director of the Media Lab at MIT and David Christian, a professor of Macquarie University in Australia.
2013.08.08 View 11947 -
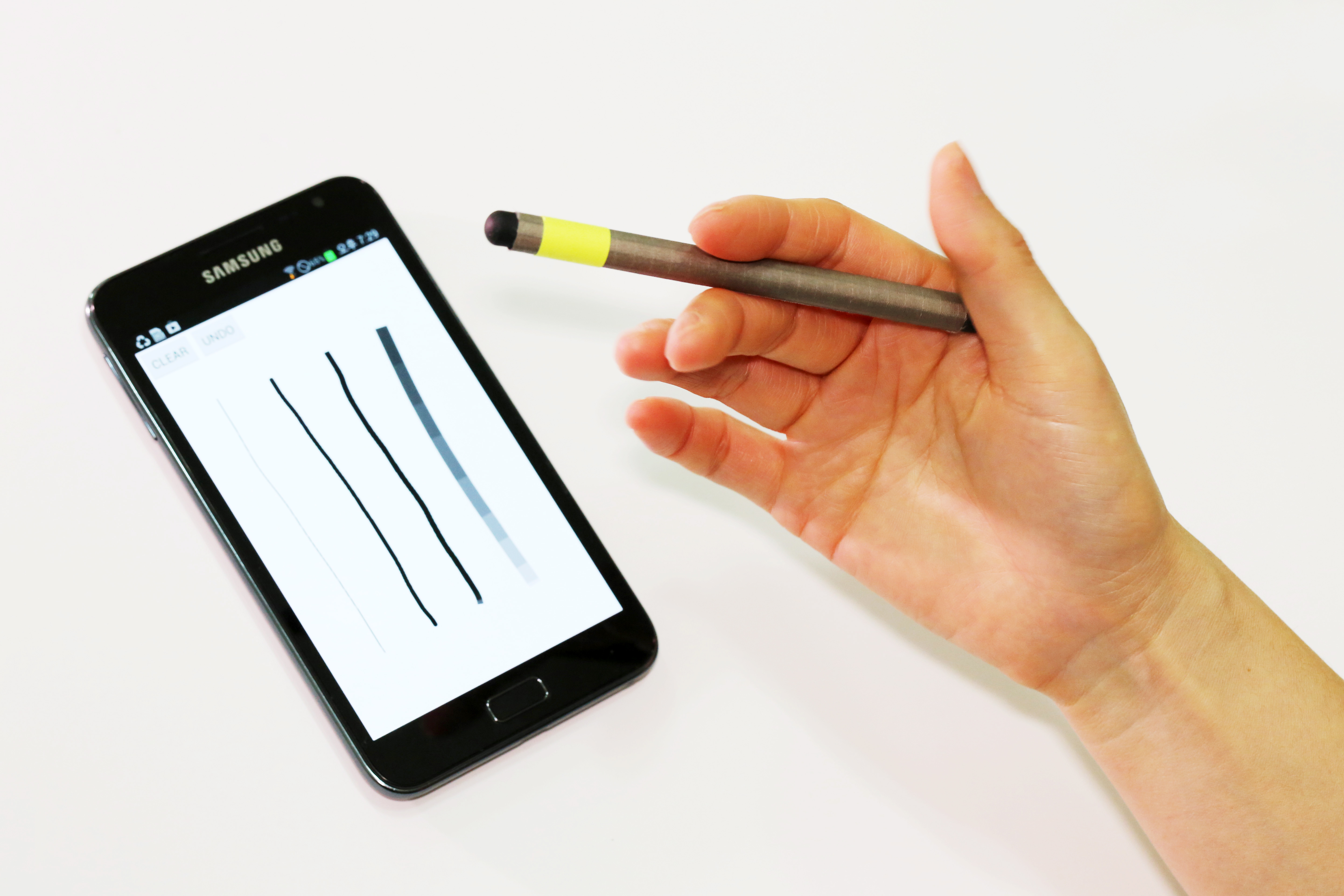 A magnetic pen for smartphones adds another level of conveniences
Utilizing existing features on smartphones, the MagPen provides users with a compatible and simple input tool regardless of the type of phones they are using.
A doctoral candidate at the Korea Advanced Institute of Science and Technology (KAIST) developed a magnetically driven pen interface that works both on and around mobile devices. This interface, called the MagPen, can be used for any type of smartphones and tablet computers so long as they have magnetometers embedded in.
Advised by Professor Kwang-yun Wohn of the Graduate School of Culture Technology (GSCT) at KAIST, Sungjae Hwang, a Ph.D. student, created the MagPen in collaboration with Myung-Wook Ahn, a master"s student at the GSCT of KAIST, and Andrea Bianchi, a professor at Sungkyunkwan University.
Almost all mobile devices today provide location-based services, and magnetometers are incorporated in the integrated circuits of smartphones or tablet PCs, functioning as compasses. Taking advantage of built-in magnetometers, Hwang"s team came up with a technology that enabled an input tool for mobile devices such as a capacitive stylus pen to interact more sensitively and effectively with the devices" touch screen. Text and command entered by a stylus pen are expressed better on the screen of mobile devices than those done by human fingers.
The MagPen utilizes magnetometers equipped with smartphones, thus there is no need to build an additional sensing panel for a touchscreen as well as circuits, communication modules, or batteries for the pen. With an application installed on smartphones, it senses and analyzes the magnetic field produced by a permanent magnet embedded in a standard capacitive stylus pen.
Sungjae Hwang said, "Our technology is eco-friendly and very affordable because we are able to improve the expressiveness of the stylus pen without requiring additional hardware beyond those already installed on the current mobile devices. The technology allows smartphone users to enjoy added convenience while no wastes generated."
The MagPen detects the direction at which a stylus pen is pointing; selects colors by dragging the pen across smartphone bezel; identifies pens with different magnetic properties; recognizes pen-spinning gestures; and estimates the finger pressure applied to the pen.
Notably, with its spinning motion, the MagPen expands the scope of input gestures recognized by a stylus pen beyond its existing vocabularies of gestures and techniques such as titling, hovering, and varying pressures. The tip of the pen switches from a pointer to an eraser and vice versa when spinning. Or, it can choose the thickness of the lines drawn on a screen by spinning.
"It"s quite remarkable to see that the MagPen can understand spinning motion. It"s like the pen changes its living environment from two dimensions to three dimensions. This is the most creative characteristic of our technology," added Sungjae Hwang.
Hwang"s initial research result was first presented at the International Conference on Intelligent User Interfaces organized by the Association for Computing Machinery and held on March 19-22 in Santa Monica, the US.
In the next month of August, the research team will present a paper on the MagPen technology, entitled "MagPen: Magnetically Driven Pen Interaction On and Around Conventional Smartphones" and receive an Honorable Mention Award at the 15th International Conference on Human-Computer Interaction with Mobile Devices and Services (MobileHCI 2013) to be held in Germany.
In addition to the MagPen, Hwang and his team are conducting other projects to develop different types of magnetic gadgets (collectively called "MagGetz") that include the Magnetic Marionette, a magnetic cover for a smartphone, which offers augmented interactions with the phone, as well as magnetic widgets such as buttons and toggle interface.
Hwang has filed ten patents for the MagGetz technology.
Youtube Links: http://www.youtube.com/watch?v=NkPo2las7wc, http://www.youtube.com/watch?v=J9GtgyzoZmM
2013.07.25 View 12870
A magnetic pen for smartphones adds another level of conveniences
Utilizing existing features on smartphones, the MagPen provides users with a compatible and simple input tool regardless of the type of phones they are using.
A doctoral candidate at the Korea Advanced Institute of Science and Technology (KAIST) developed a magnetically driven pen interface that works both on and around mobile devices. This interface, called the MagPen, can be used for any type of smartphones and tablet computers so long as they have magnetometers embedded in.
Advised by Professor Kwang-yun Wohn of the Graduate School of Culture Technology (GSCT) at KAIST, Sungjae Hwang, a Ph.D. student, created the MagPen in collaboration with Myung-Wook Ahn, a master"s student at the GSCT of KAIST, and Andrea Bianchi, a professor at Sungkyunkwan University.
Almost all mobile devices today provide location-based services, and magnetometers are incorporated in the integrated circuits of smartphones or tablet PCs, functioning as compasses. Taking advantage of built-in magnetometers, Hwang"s team came up with a technology that enabled an input tool for mobile devices such as a capacitive stylus pen to interact more sensitively and effectively with the devices" touch screen. Text and command entered by a stylus pen are expressed better on the screen of mobile devices than those done by human fingers.
The MagPen utilizes magnetometers equipped with smartphones, thus there is no need to build an additional sensing panel for a touchscreen as well as circuits, communication modules, or batteries for the pen. With an application installed on smartphones, it senses and analyzes the magnetic field produced by a permanent magnet embedded in a standard capacitive stylus pen.
Sungjae Hwang said, "Our technology is eco-friendly and very affordable because we are able to improve the expressiveness of the stylus pen without requiring additional hardware beyond those already installed on the current mobile devices. The technology allows smartphone users to enjoy added convenience while no wastes generated."
The MagPen detects the direction at which a stylus pen is pointing; selects colors by dragging the pen across smartphone bezel; identifies pens with different magnetic properties; recognizes pen-spinning gestures; and estimates the finger pressure applied to the pen.
Notably, with its spinning motion, the MagPen expands the scope of input gestures recognized by a stylus pen beyond its existing vocabularies of gestures and techniques such as titling, hovering, and varying pressures. The tip of the pen switches from a pointer to an eraser and vice versa when spinning. Or, it can choose the thickness of the lines drawn on a screen by spinning.
"It"s quite remarkable to see that the MagPen can understand spinning motion. It"s like the pen changes its living environment from two dimensions to three dimensions. This is the most creative characteristic of our technology," added Sungjae Hwang.
Hwang"s initial research result was first presented at the International Conference on Intelligent User Interfaces organized by the Association for Computing Machinery and held on March 19-22 in Santa Monica, the US.
In the next month of August, the research team will present a paper on the MagPen technology, entitled "MagPen: Magnetically Driven Pen Interaction On and Around Conventional Smartphones" and receive an Honorable Mention Award at the 15th International Conference on Human-Computer Interaction with Mobile Devices and Services (MobileHCI 2013) to be held in Germany.
In addition to the MagPen, Hwang and his team are conducting other projects to develop different types of magnetic gadgets (collectively called "MagGetz") that include the Magnetic Marionette, a magnetic cover for a smartphone, which offers augmented interactions with the phone, as well as magnetic widgets such as buttons and toggle interface.
Hwang has filed ten patents for the MagGetz technology.
Youtube Links: http://www.youtube.com/watch?v=NkPo2las7wc, http://www.youtube.com/watch?v=J9GtgyzoZmM
2013.07.25 View 12870 -
 Professor Jay H. Lee to receive the 2013 AIChE CAST Computing in Chemical Engineering Award
Professor Jay H. Lee of Chemical and Biomolecular Engineering Department at KAIST has won the 2013 Computing in Chemical Engineering Award of AIChE"s CAST Division (AIChE, American Institute of Chemical Engineers and CAST, Computing & Systems Technology Division).
The CAST Computing in Chemical Engineering Award, sponsored by The Dow Chemical Company, is annually given to an individual who has made outstanding contributions in the application of computing and systems technology to chemical engineering.Professor Lee has been recognized for his pioneering research contributions for “novel paradigms for much improved and robust model predictive control in industrial processes.” He is currently the Head of Chemical and Biomolecular Engineering Department and Director of Brain Korea (BK) 21 Program at the department. BK21 is the Korean government’s initiative to support the growth of research universities in the nation and foster highly trained master’s and doctoral students as well as researchers.
The CAST Computing in Chemical Engineering Award will be presented to Professor Jay H. Lee at the CAST Division dinner to be held at the AIChE Annual Meeting this November in San Francisco, where he will also deliver the after dinner lecture associated with this award.
2013.06.12 View 11787
Professor Jay H. Lee to receive the 2013 AIChE CAST Computing in Chemical Engineering Award
Professor Jay H. Lee of Chemical and Biomolecular Engineering Department at KAIST has won the 2013 Computing in Chemical Engineering Award of AIChE"s CAST Division (AIChE, American Institute of Chemical Engineers and CAST, Computing & Systems Technology Division).
The CAST Computing in Chemical Engineering Award, sponsored by The Dow Chemical Company, is annually given to an individual who has made outstanding contributions in the application of computing and systems technology to chemical engineering.Professor Lee has been recognized for his pioneering research contributions for “novel paradigms for much improved and robust model predictive control in industrial processes.” He is currently the Head of Chemical and Biomolecular Engineering Department and Director of Brain Korea (BK) 21 Program at the department. BK21 is the Korean government’s initiative to support the growth of research universities in the nation and foster highly trained master’s and doctoral students as well as researchers.
The CAST Computing in Chemical Engineering Award will be presented to Professor Jay H. Lee at the CAST Division dinner to be held at the AIChE Annual Meeting this November in San Francisco, where he will also deliver the after dinner lecture associated with this award.
2013.06.12 View 11787 -
 International Student Conference (ICISTS-KAIST) to be Held in August
- 300 participants including university students worldwide and renowned speakers expected to gather
- Ideal coexistence of science & technology and society explored under the theme of “Perfect Alliance”
Science & technology and society are at the core of 21st century’s development. ICISTS-KAIST 2013, international conference for university students, seeks ways for the two to coexist harmoniously and is to be held from August 5 to 9 on KAIST campus as well as at Daejeon Convention Center.
ICISTS stands for International Conference for the Integration of Science, Technology and Society. ICISTS-KAIST is a non-profit organization run by KAIST students who are directly engaged in the coordination, planning, finance, public relations, and management of this academic event. The upcoming ninth annual event of ICISTS (www.icists.org) 2013 is centered around the theme, “Perfect Alliance: Coexistence for Human Society.” The conference will last for four nights and five days; scholars and students across various academic backgrounds gather to narrow the gap between fields of study and discuss possible solutions to the problems in today’s society.
The annual conference, ICISTS-KAIST attracts hundreds of participants from all over the world to KAIST, Daejeon and its most recent event last year witnessed discussions among some 300 students from 22 countries hearing the lectures from 40 academics and scholars. This year’s event will welcome the 16-year old inventor, scientist, and cancer researcher Jack Thomas Andraka, the founder of the “One Laptop Per Child” project Walter Bender, Chemistry Nobel Prize laureate Harold Walter Kroto, and many more.
The application period for ICISTS-KAIST 2013 runs from May 20 to July 12, and applications are received through the website at www.icists.org.
ICISTS-KAIST 2013 Promgram Summary
Event Title: International Conference for the Integration of Science, Technology and Society 2013 (ICISTS-KAIST 2013)
Theme: Perfect Alliance: Coexistence for Human Society
Date and Venue: 2013 Aug. 5 (Mon.) ~ Aug. 9 (Fri.), KAIST Campus and Daejeon Convention Center
Host and Organizer: ICISTS KAIST
Sponsor: Korean National Commission for UNESCO, Korea Tourism Organization, Korea Ministry of Education, Science & Technology, KOFST
Session Description:
Keynote Speech - Keynote address on fundamental approach to coexistence
Parallel Session - Multiple simultaneous lecture of delegates’ choice
Group Discussion - Small group discussions among delegates and speakers
Panel Discussion - In-depth and thought-revealing discussion among speakers
Experience Session - First-person experience on relevant technology
Team Project & Poster Fair - Team mission, poster exhibition and evaluation
Subtopics:
- New Values from Coexistence of Science & Technology and Society
- Synergetic Resolution via Coexistence of Science & Technology and Society
- Essential Communication for Coexistence of Science & Technology and Society
Notable Speakers:
- Gretchen Kalonji: Assistant to Director-General at UNESCO
- Sheila Jasanoff: Director of STS Program at Harvard Kennedy School
- Walter Bender: Former Director of MIT Media Lab and One Laptop Per Child- Jack Andraka: 16-year old Cancer Resesarcher
2013.05.31 View 10318
International Student Conference (ICISTS-KAIST) to be Held in August
- 300 participants including university students worldwide and renowned speakers expected to gather
- Ideal coexistence of science & technology and society explored under the theme of “Perfect Alliance”
Science & technology and society are at the core of 21st century’s development. ICISTS-KAIST 2013, international conference for university students, seeks ways for the two to coexist harmoniously and is to be held from August 5 to 9 on KAIST campus as well as at Daejeon Convention Center.
ICISTS stands for International Conference for the Integration of Science, Technology and Society. ICISTS-KAIST is a non-profit organization run by KAIST students who are directly engaged in the coordination, planning, finance, public relations, and management of this academic event. The upcoming ninth annual event of ICISTS (www.icists.org) 2013 is centered around the theme, “Perfect Alliance: Coexistence for Human Society.” The conference will last for four nights and five days; scholars and students across various academic backgrounds gather to narrow the gap between fields of study and discuss possible solutions to the problems in today’s society.
The annual conference, ICISTS-KAIST attracts hundreds of participants from all over the world to KAIST, Daejeon and its most recent event last year witnessed discussions among some 300 students from 22 countries hearing the lectures from 40 academics and scholars. This year’s event will welcome the 16-year old inventor, scientist, and cancer researcher Jack Thomas Andraka, the founder of the “One Laptop Per Child” project Walter Bender, Chemistry Nobel Prize laureate Harold Walter Kroto, and many more.
The application period for ICISTS-KAIST 2013 runs from May 20 to July 12, and applications are received through the website at www.icists.org.
ICISTS-KAIST 2013 Promgram Summary
Event Title: International Conference for the Integration of Science, Technology and Society 2013 (ICISTS-KAIST 2013)
Theme: Perfect Alliance: Coexistence for Human Society
Date and Venue: 2013 Aug. 5 (Mon.) ~ Aug. 9 (Fri.), KAIST Campus and Daejeon Convention Center
Host and Organizer: ICISTS KAIST
Sponsor: Korean National Commission for UNESCO, Korea Tourism Organization, Korea Ministry of Education, Science & Technology, KOFST
Session Description:
Keynote Speech - Keynote address on fundamental approach to coexistence
Parallel Session - Multiple simultaneous lecture of delegates’ choice
Group Discussion - Small group discussions among delegates and speakers
Panel Discussion - In-depth and thought-revealing discussion among speakers
Experience Session - First-person experience on relevant technology
Team Project & Poster Fair - Team mission, poster exhibition and evaluation
Subtopics:
- New Values from Coexistence of Science & Technology and Society
- Synergetic Resolution via Coexistence of Science & Technology and Society
- Essential Communication for Coexistence of Science & Technology and Society
Notable Speakers:
- Gretchen Kalonji: Assistant to Director-General at UNESCO
- Sheila Jasanoff: Director of STS Program at Harvard Kennedy School
- Walter Bender: Former Director of MIT Media Lab and One Laptop Per Child- Jack Andraka: 16-year old Cancer Resesarcher
2013.05.31 View 10318 -
 Cooperation Agreement signed between KAIST and University of Oxford
On April 6th, a Cooperation Memorandum of Understanding (MOU), including the exchange of students between two universities, was signed by KAIST and University of Oxford on April 6th.
In the MOU, KAIST and Oxford agreed to achieve mutual development through cooperation, such as joint research programs and the exchange of professors and students.
President Sung-Mo Kang stated, “This agreement will be the start of close cooperation between the two universities, and I will continue to endeavor to expand the relationship.”
Andrew Hamilton, the president of Oxford University replied, “I am well aware of the excellent quality of teaching and research of Korean students and KAIST. I hope the cooperation between KAIST and Oxford will achieve exchange at diverse levels.”
President Hamilton taught chemistry at the University of Pittsburgh and Princeton University and also served as the vice-president of Yale University. Having experience of teaching a KAIST student before, he has high praise for the ability of KAIST students.
The President of the University of Oxford, Andrew Hamilton (left), and the President of KAIST, Sung-Mo "Steve" Kang, shake hands in acknowledgement of signing the Cooperation MOU.
2013.05.20 View 7343
Cooperation Agreement signed between KAIST and University of Oxford
On April 6th, a Cooperation Memorandum of Understanding (MOU), including the exchange of students between two universities, was signed by KAIST and University of Oxford on April 6th.
In the MOU, KAIST and Oxford agreed to achieve mutual development through cooperation, such as joint research programs and the exchange of professors and students.
President Sung-Mo Kang stated, “This agreement will be the start of close cooperation between the two universities, and I will continue to endeavor to expand the relationship.”
Andrew Hamilton, the president of Oxford University replied, “I am well aware of the excellent quality of teaching and research of Korean students and KAIST. I hope the cooperation between KAIST and Oxford will achieve exchange at diverse levels.”
President Hamilton taught chemistry at the University of Pittsburgh and Princeton University and also served as the vice-president of Yale University. Having experience of teaching a KAIST student before, he has high praise for the ability of KAIST students.
The President of the University of Oxford, Andrew Hamilton (left), and the President of KAIST, Sung-Mo "Steve" Kang, shake hands in acknowledgement of signing the Cooperation MOU.
2013.05.20 View 7343 -
 The new era of personalized cancer diagnosis and treatment
Professor Tae-Young Yoon
- Succeeded in observing carcinogenic protein at the molecular level
- “Paved the way to customized cancer treatment through accurate analysis of carcinogenic protein”
The joint KAIST research team of Professor Tae Young Yoon of the Department of Physics and Professor Won Do Huh of the Department of Biological Sciences have developed the technology to monitor characteristics of carcinogenic protein in cancer tissue – for the first time in the world.
The technology makes it possible to analyse the mechanism of cancer development through a small amount of carcinogenic protein from a cancer patient. Therefore, a personalised approach to diagnosis and treatment using the knowledge of the specific mechanism of cancer development in the patient may be possible in the future.
Until recently, modern medicine could only speculate on the cause of cancer through statistics. Although developed countries, such as the United States, are known to use a large sequencing technology that analyses the patient’s DNA, identification of the interactions between proteins responsible for causing cancer remained an unanswered question for a long time in medicine.
Firstly, Professor Yoon’s research team has developed a fluorescent microscope that can observe even a single molecule. Then, the “Immunoprecipitation method”, a technology to extract a specific protein exploiting the high affinity between antigens and antibodies was developed. Using this technology and the microscope, “Real-Time Single Molecule co-Immunoprecipitation Method” was created. In this way, the team succeeded in observing the interactions between carcinogenic and other proteins at a molecular level, in real time.
To validate the developed technology, the team investigated Ras, a carcinogenic protein; its mutation statistically is known to cause around 30% of cancers.
The experimental results confirmed that 30-50% of Ras protein was expressed in mouse tumour and human cancer cells. In normal cells, less than 5% of Ras protein was expressed. Thus, the experiment showed that unusual increase in activation of Ras protein induces cancer.
The increase in the ratio of active Ras protein can be inferred from existing research data but the measurement of specific numerical data has never been done before.
The team suggested a new molecular level diagnosis technique of identifying the progress of cancer in patients through measuring the percentage of activated carcinogenic protein in cancer tissue.
Professor Yoon Tae-young said, “This newly developed technology does not require a separate procedure of protein expression or refining, hence the existing proteins in real biological tissues or cancer cells can be observed directly.” He also said, “Since carcinogenic protein can be analyzed accurately, it has opened up the path to customized cancer treatment in the future.”
“Since the observation is possible on a molecular level, the technology confers the advantage that researchers can carry out various examinations on a small sample of the cancer patient.” He added, “The clinical trial will start in December 2012 and in a few years customized cancer diagnosis and treatment will be possible.”
Meanwhile, the research has been published in Nature Communications (February 19). Many researchers from various fields have participated, regardless of the differences in their speciality, and successfully produced interdisciplinary research. Professor Tae Young Yoon of the Department of Physics and Professors Dae Sik Lim and Won Do Huh of Biological Sciences at KAIST, and Professor Chang Bong Hyun of Computational Science of KIAS contributed to developing the technique.
Figure 1: Schematic diagram of observed interactions at the molecular level in real time using fluorescent microscope. The carcinogenic protein from a mouse tumour is fixed on the microchip, and its molecular characteristics are observed live.
Figure 2: Molecular interaction data using a molecular level fluorescent microscope. A signal in the form of spike is shown when two proteins combine. This is monitored live using an Electron Multiplying Charge Coupled Device (EMCCD). It shows signal results in bright dots.
An organism has an immune system as a defence mechanism to foreign intruders. The immune system is activated when unwanted pathogens or foreign protein are in the body. Antibodies form in recognition of the specific antigen to protect itself. Organisms evolved to form antibodies with high specificity to a certain antigen. Antibodies only react to its complementary antigens. The field of molecular biology uses the affinity between antigens and antibodies to extract specific proteins; a technology called immunoprecipitation. Even in a mixture of many proteins, the protein sought can be extracted using antibodies. Thus immunoprecipitation is widely used to detect pathogens or to extract specific proteins.
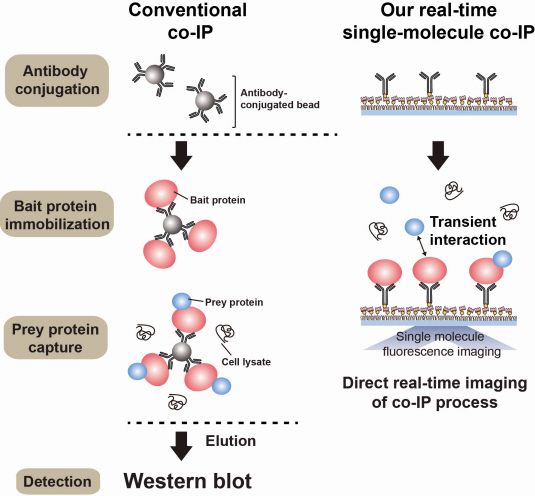
Technology co-IP is a well-known example that uses immunoprecipitation. The research on interactions between proteins uses co-IP in general. The basis of fixing the antigen on the antibody to extract antigen protein is the same as immunoprecipitation. Then, researchers inject and observe its reaction with the partner protein to observe the interactions and precipitate the antibodies. If the reaction occurs, the partner protein will be found with the antibodies in the precipitations. If not, then the partner protein will not be found. This shows that the two proteins interact.
However, the traditional co-IP can be used to infer the interactions between the two proteins although the information of the dynamics on how the reaction occurs is lost. To overcome these shortcomings, the Real-Time Single Molecule co-IP Method enables observation on individual protein level in real time. Therefore, the significance of the new technique is in making observation of interactions more direct and quantitative.
Additional Figure 1: Comparison between Conventional co-IP and Real-Time Single Molecule co-IP
2013.04.01 View 21035
The new era of personalized cancer diagnosis and treatment
Professor Tae-Young Yoon
- Succeeded in observing carcinogenic protein at the molecular level
- “Paved the way to customized cancer treatment through accurate analysis of carcinogenic protein”
The joint KAIST research team of Professor Tae Young Yoon of the Department of Physics and Professor Won Do Huh of the Department of Biological Sciences have developed the technology to monitor characteristics of carcinogenic protein in cancer tissue – for the first time in the world.
The technology makes it possible to analyse the mechanism of cancer development through a small amount of carcinogenic protein from a cancer patient. Therefore, a personalised approach to diagnosis and treatment using the knowledge of the specific mechanism of cancer development in the patient may be possible in the future.
Until recently, modern medicine could only speculate on the cause of cancer through statistics. Although developed countries, such as the United States, are known to use a large sequencing technology that analyses the patient’s DNA, identification of the interactions between proteins responsible for causing cancer remained an unanswered question for a long time in medicine.
Firstly, Professor Yoon’s research team has developed a fluorescent microscope that can observe even a single molecule. Then, the “Immunoprecipitation method”, a technology to extract a specific protein exploiting the high affinity between antigens and antibodies was developed. Using this technology and the microscope, “Real-Time Single Molecule co-Immunoprecipitation Method” was created. In this way, the team succeeded in observing the interactions between carcinogenic and other proteins at a molecular level, in real time.
To validate the developed technology, the team investigated Ras, a carcinogenic protein; its mutation statistically is known to cause around 30% of cancers.
The experimental results confirmed that 30-50% of Ras protein was expressed in mouse tumour and human cancer cells. In normal cells, less than 5% of Ras protein was expressed. Thus, the experiment showed that unusual increase in activation of Ras protein induces cancer.
The increase in the ratio of active Ras protein can be inferred from existing research data but the measurement of specific numerical data has never been done before.
The team suggested a new molecular level diagnosis technique of identifying the progress of cancer in patients through measuring the percentage of activated carcinogenic protein in cancer tissue.
Professor Yoon Tae-young said, “This newly developed technology does not require a separate procedure of protein expression or refining, hence the existing proteins in real biological tissues or cancer cells can be observed directly.” He also said, “Since carcinogenic protein can be analyzed accurately, it has opened up the path to customized cancer treatment in the future.”
“Since the observation is possible on a molecular level, the technology confers the advantage that researchers can carry out various examinations on a small sample of the cancer patient.” He added, “The clinical trial will start in December 2012 and in a few years customized cancer diagnosis and treatment will be possible.”
Meanwhile, the research has been published in Nature Communications (February 19). Many researchers from various fields have participated, regardless of the differences in their speciality, and successfully produced interdisciplinary research. Professor Tae Young Yoon of the Department of Physics and Professors Dae Sik Lim and Won Do Huh of Biological Sciences at KAIST, and Professor Chang Bong Hyun of Computational Science of KIAS contributed to developing the technique.
Figure 1: Schematic diagram of observed interactions at the molecular level in real time using fluorescent microscope. The carcinogenic protein from a mouse tumour is fixed on the microchip, and its molecular characteristics are observed live.
Figure 2: Molecular interaction data using a molecular level fluorescent microscope. A signal in the form of spike is shown when two proteins combine. This is monitored live using an Electron Multiplying Charge Coupled Device (EMCCD). It shows signal results in bright dots.
An organism has an immune system as a defence mechanism to foreign intruders. The immune system is activated when unwanted pathogens or foreign protein are in the body. Antibodies form in recognition of the specific antigen to protect itself. Organisms evolved to form antibodies with high specificity to a certain antigen. Antibodies only react to its complementary antigens. The field of molecular biology uses the affinity between antigens and antibodies to extract specific proteins; a technology called immunoprecipitation. Even in a mixture of many proteins, the protein sought can be extracted using antibodies. Thus immunoprecipitation is widely used to detect pathogens or to extract specific proteins.
Technology co-IP is a well-known example that uses immunoprecipitation. The research on interactions between proteins uses co-IP in general. The basis of fixing the antigen on the antibody to extract antigen protein is the same as immunoprecipitation. Then, researchers inject and observe its reaction with the partner protein to observe the interactions and precipitate the antibodies. If the reaction occurs, the partner protein will be found with the antibodies in the precipitations. If not, then the partner protein will not be found. This shows that the two proteins interact.
However, the traditional co-IP can be used to infer the interactions between the two proteins although the information of the dynamics on how the reaction occurs is lost. To overcome these shortcomings, the Real-Time Single Molecule co-IP Method enables observation on individual protein level in real time. Therefore, the significance of the new technique is in making observation of interactions more direct and quantitative.
Additional Figure 1: Comparison between Conventional co-IP and Real-Time Single Molecule co-IP
2013.04.01 View 21035 -
 Ligand Recognition Mechanism of Protein Identified
Professor Hak-Sung Kim
-“Solved the 50 year old mystery of how protein recognises and binds to ligands”
- Exciting potential for understanding life phenomena and the further development of highly effective therapeutic agent development
KAIST’s Biological Science Department’s Professor Hak-Sung Kim, working in collaboration with Professor Sung-Chul Hong of Department of Physics, Seoul National University, has identified the mechanism of how the protein recognizes and binds to ligands within the human body.
The research findings were published in the online edition of Nature Chemical Biology (March 18), which is the most prestigious journal in the field of life science.
Since the research identified the mechanism, of which protein recognises and binds to ligands, it will take an essential role in understanding complex life phenomenon by understanding regulatory function of protein.
Also, ligand recognition of proteins is closely related to the cause of various diseases. Therefore the research team hopes to contribute to the development of highly effective treatments.
Ligands, well-known examples include nucleic acid and proteins, form the structure of an organism or are essential constituents with special functions such as information signalling.
In particular, the most important role of protein is recognising and binding to a particular ligand and hence regulating and maintaining life phenomena. The abnormal occurrence of an error in recognition of ligands may lead to various diseases.
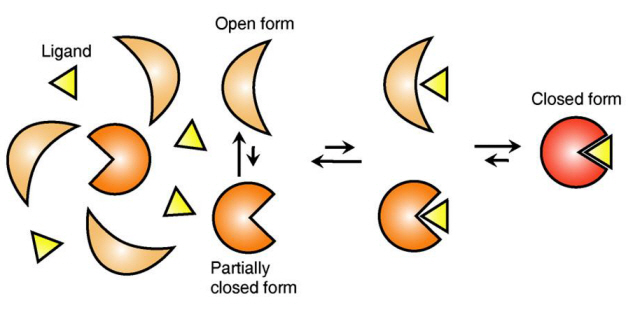
The research team focused on the repetition of change in protein structure from the most stable “open form” to a relatively unstable “partially closed form”.
Professor Kim’s team analysed the change in protein structure when binding to a ligand on a molecular level in real time to explain the ligand recognition mechanism.
The research findings showed that ligands prefer the most stable protein structure. The team was the first in the world to identify that ligands alter protein structure to the most stable, the lowest energy level, when it binds to the protein.
In addition, the team found that ligands bind to unstable partially-closed forms to change protein structure.
The existing models to explain ligand recognition mechanism of protein are “Induced Custom Model”, which involves change in protein structure in binding to ligands, and the “Structure Selection Model”, which argues that ligands select and recognise only the best protein structure out of many. The academic world considers that the team’s research findings have perfectly proved the models through experiments for the first time in the world.
Professor Kim explained, “In the presence of ligands, there exists a phenomenon where the speed of altering protein structure is changed. This phenomenon is analysed on a molecular level to prove ligand recognition mechanism of protein for the first time”. He also said, “The 50-year old mystery, that existed only as a hypothesis on biology textbooks and was thought never to be solved, has been confirmed through experiments for the first time.”
Figure 1: Proteins, with open and partially open form, recognising and binding to ligands.
Figure 2: Ligands temporarily bind to a stable protein structure, open form, which changes into the most stable structure, closed form. In addition, binding to partially closed form also changes protein structure to closed form.
2013.04.01 View 13365
Ligand Recognition Mechanism of Protein Identified
Professor Hak-Sung Kim
-“Solved the 50 year old mystery of how protein recognises and binds to ligands”
- Exciting potential for understanding life phenomena and the further development of highly effective therapeutic agent development
KAIST’s Biological Science Department’s Professor Hak-Sung Kim, working in collaboration with Professor Sung-Chul Hong of Department of Physics, Seoul National University, has identified the mechanism of how the protein recognizes and binds to ligands within the human body.
The research findings were published in the online edition of Nature Chemical Biology (March 18), which is the most prestigious journal in the field of life science.
Since the research identified the mechanism, of which protein recognises and binds to ligands, it will take an essential role in understanding complex life phenomenon by understanding regulatory function of protein.
Also, ligand recognition of proteins is closely related to the cause of various diseases. Therefore the research team hopes to contribute to the development of highly effective treatments.
Ligands, well-known examples include nucleic acid and proteins, form the structure of an organism or are essential constituents with special functions such as information signalling.
In particular, the most important role of protein is recognising and binding to a particular ligand and hence regulating and maintaining life phenomena. The abnormal occurrence of an error in recognition of ligands may lead to various diseases.
The research team focused on the repetition of change in protein structure from the most stable “open form” to a relatively unstable “partially closed form”.
Professor Kim’s team analysed the change in protein structure when binding to a ligand on a molecular level in real time to explain the ligand recognition mechanism.
The research findings showed that ligands prefer the most stable protein structure. The team was the first in the world to identify that ligands alter protein structure to the most stable, the lowest energy level, when it binds to the protein.
In addition, the team found that ligands bind to unstable partially-closed forms to change protein structure.
The existing models to explain ligand recognition mechanism of protein are “Induced Custom Model”, which involves change in protein structure in binding to ligands, and the “Structure Selection Model”, which argues that ligands select and recognise only the best protein structure out of many. The academic world considers that the team’s research findings have perfectly proved the models through experiments for the first time in the world.
Professor Kim explained, “In the presence of ligands, there exists a phenomenon where the speed of altering protein structure is changed. This phenomenon is analysed on a molecular level to prove ligand recognition mechanism of protein for the first time”. He also said, “The 50-year old mystery, that existed only as a hypothesis on biology textbooks and was thought never to be solved, has been confirmed through experiments for the first time.”
Figure 1: Proteins, with open and partially open form, recognising and binding to ligands.
Figure 2: Ligands temporarily bind to a stable protein structure, open form, which changes into the most stable structure, closed form. In addition, binding to partially closed form also changes protein structure to closed form.
2013.04.01 View 13365 -
 KAIST and Saudi Aramco agreed to establish a joint CO2 research center in Korea
The Korea Advanced Institute of Science and Technology (KAIST) and Saudi Aramco, a global energy and petrochemicals enterprise, signed a memorandum of understanding (MOU) on January 6, 2013 in Dhahran, Saudi Arabia and pledged to jointly collaborate in research and development of innovative technologies and solutions to address the world"s energy challenges.
Under the MOU, the two entities agreed to establish a research center, Saudi Aramco-KAIST CO2 Research Center, near KAIST"s main campus in Daejeon, Korea. The research center, to be jointly managed by KAIST and Saudi Aramco, will foster and facilitate research collaborations in areas such as tackling carbon dioxide (CO2) emissions by removal or capture of CO2, conversing CO2 into useful products, developing efficiency improvements in energy production, sharing carbon management technologies, establishing exchange programs, and conducting joint projects.
According to Saudi Aramco, the company"s collaboration with KAIST is the first partnership established in Asia. Khalid A. Al-Falih, President and CEO of Saudi Aramco, said,
"The CO2 Research Center represents a major step in Saudi Aramco"s research and technology strategy to partner with top global institutions to help address and find sustainable solutions to the world’s energy challenge both domestically and internationally."
2013.03.19 View 11527
KAIST and Saudi Aramco agreed to establish a joint CO2 research center in Korea
The Korea Advanced Institute of Science and Technology (KAIST) and Saudi Aramco, a global energy and petrochemicals enterprise, signed a memorandum of understanding (MOU) on January 6, 2013 in Dhahran, Saudi Arabia and pledged to jointly collaborate in research and development of innovative technologies and solutions to address the world"s energy challenges.
Under the MOU, the two entities agreed to establish a research center, Saudi Aramco-KAIST CO2 Research Center, near KAIST"s main campus in Daejeon, Korea. The research center, to be jointly managed by KAIST and Saudi Aramco, will foster and facilitate research collaborations in areas such as tackling carbon dioxide (CO2) emissions by removal or capture of CO2, conversing CO2 into useful products, developing efficiency improvements in energy production, sharing carbon management technologies, establishing exchange programs, and conducting joint projects.
According to Saudi Aramco, the company"s collaboration with KAIST is the first partnership established in Asia. Khalid A. Al-Falih, President and CEO of Saudi Aramco, said,
"The CO2 Research Center represents a major step in Saudi Aramco"s research and technology strategy to partner with top global institutions to help address and find sustainable solutions to the world’s energy challenge both domestically and internationally."
2013.03.19 View 11527 -
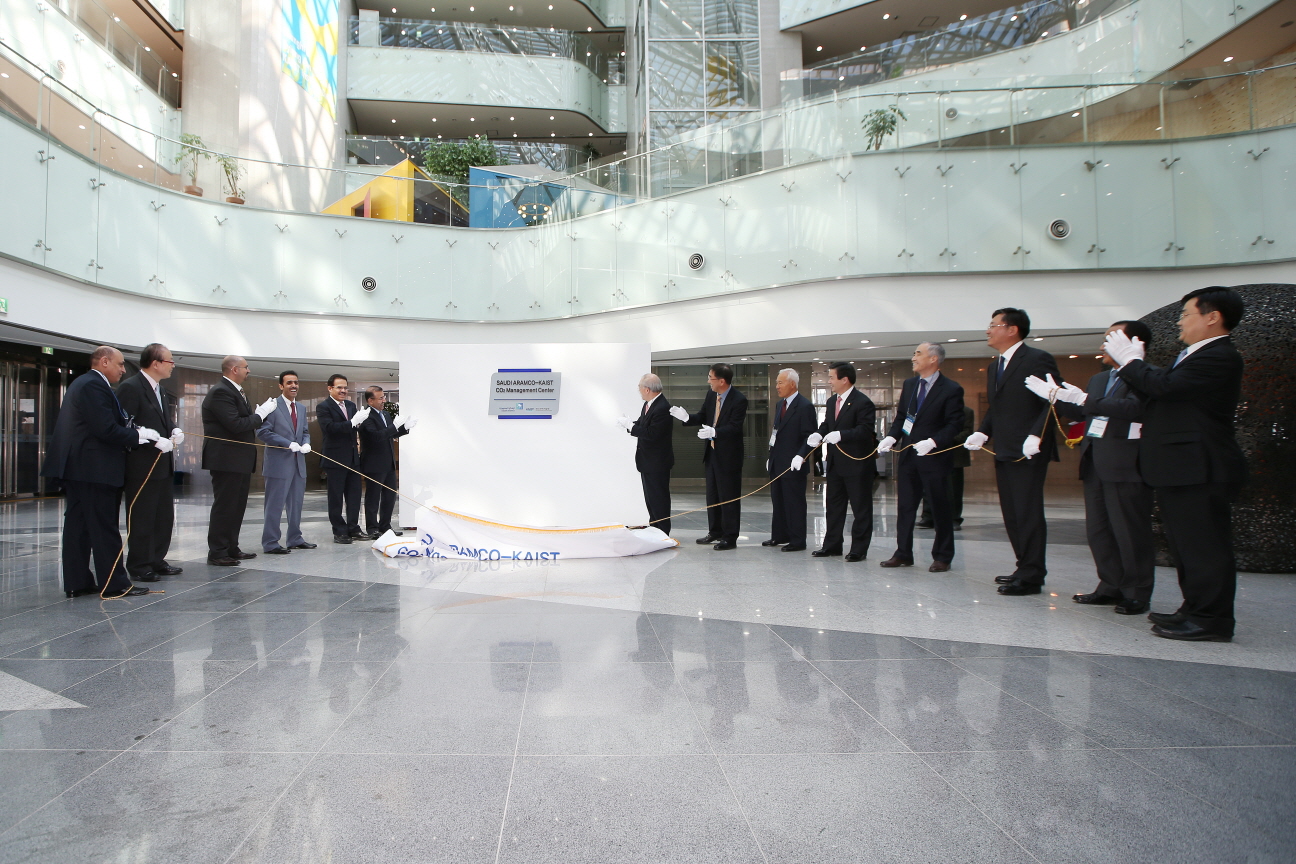 Launched the Saudi Aramco-KAIST CO2 Management Center in Korea
KAIST and Saudi Aramco, a global energy and petrochemicals enterprise, signed on February 20, 2013 the Master Research and Collaboration Agreement (the Agreement) on joint collaborations in research and development of carbon management between the two entities.
The Agreement was subsequently concluded upon the signing of the Memorandum of Understanding (MOU) between KAIST and Saudi Aramco, dated January 7th, 2013. In the Agreement, the two organizations specified terms and conditions necessary to conduct joint research projects and stipulated governing body for the operation of the Saudi Aramco-KAIST CO2 Management Center.
KAIST and Saudi Aramco, a national oil company for Saudi Arabia, entered into the MOU, in which the two parties shared a common interest in addressing the issue of CO2 capture, CO2storage, CO2 avoidance using efficiency improvements, and converting CO2 into useful chemicals and other materials, and agreed to “create a major research center for CO2” in Korea.
As envisioned by the MOU and its subsequent agreement, KAIST and Saudi Aramco decided to operate an interim office of the Saudi Aramco-KAIST CO2 Management Center at KAIST campus in Daejeon, Korea, pending the establishment of the research center. The full-fledged, independent research facility will be built at a location and during a period to be agreed between the two parties.
Following the signing of the Agreement, there was a celebration event taken place, including a signboard hanging ceremony for the interim research office. A 10-member delegation from Saudi Aramco, which was headed by Vice President of Engineering Services Samir Al-Tubayyeb, Dr. Nam-Pyo Suh, former president of KAIST, Vice President of Research at KAIST Kyung-Wook Paik, and senior representatives from Korean oil and petrochemical companies such as S-Oil, Lotte Chemicals, SK Innovation, and STX attended the event.
Kyung-Wook Paik, Vice President of Research at KAIST, said,
“In order to help find solutions to carbon management, KAIST and Saudi Aramco will facilitate to exchange each party’s complementary technical expertise, gain insight into new research fields, and have access to key sources of talent, while promoting innovation for technology solutions and contributing to the lifelong learning agenda of both organizations.”
Samir Al-Tubayyeb, Vice President of Engineering Services at Saudi Aramco, added that “As a world-leading oil and gas company, Saudi Aramco’s mission is to promote the continued use of safe, environmentally-friendly petroleum products with a vision to becoming a global leader in research and technology. Building a strong and cooperative relationship with KAIST in our endeavor to search for alternative ways to better utilization of fossil fuels will expedite the creation of opportunities to make the world environmentally safer and sustainable.”
KAIST and Saudi Aramco will each chip in a maximum of USD 5 million annually for the establishment and operation of the Saudi Aramco-KAIST CO2 Management Center during the initial term of the Master Research and Collaboration Agreement, which starts in 2013 and continues through 2018.
2013.03.19 View 15468
Launched the Saudi Aramco-KAIST CO2 Management Center in Korea
KAIST and Saudi Aramco, a global energy and petrochemicals enterprise, signed on February 20, 2013 the Master Research and Collaboration Agreement (the Agreement) on joint collaborations in research and development of carbon management between the two entities.
The Agreement was subsequently concluded upon the signing of the Memorandum of Understanding (MOU) between KAIST and Saudi Aramco, dated January 7th, 2013. In the Agreement, the two organizations specified terms and conditions necessary to conduct joint research projects and stipulated governing body for the operation of the Saudi Aramco-KAIST CO2 Management Center.
KAIST and Saudi Aramco, a national oil company for Saudi Arabia, entered into the MOU, in which the two parties shared a common interest in addressing the issue of CO2 capture, CO2storage, CO2 avoidance using efficiency improvements, and converting CO2 into useful chemicals and other materials, and agreed to “create a major research center for CO2” in Korea.
As envisioned by the MOU and its subsequent agreement, KAIST and Saudi Aramco decided to operate an interim office of the Saudi Aramco-KAIST CO2 Management Center at KAIST campus in Daejeon, Korea, pending the establishment of the research center. The full-fledged, independent research facility will be built at a location and during a period to be agreed between the two parties.
Following the signing of the Agreement, there was a celebration event taken place, including a signboard hanging ceremony for the interim research office. A 10-member delegation from Saudi Aramco, which was headed by Vice President of Engineering Services Samir Al-Tubayyeb, Dr. Nam-Pyo Suh, former president of KAIST, Vice President of Research at KAIST Kyung-Wook Paik, and senior representatives from Korean oil and petrochemical companies such as S-Oil, Lotte Chemicals, SK Innovation, and STX attended the event.
Kyung-Wook Paik, Vice President of Research at KAIST, said,
“In order to help find solutions to carbon management, KAIST and Saudi Aramco will facilitate to exchange each party’s complementary technical expertise, gain insight into new research fields, and have access to key sources of talent, while promoting innovation for technology solutions and contributing to the lifelong learning agenda of both organizations.”
Samir Al-Tubayyeb, Vice President of Engineering Services at Saudi Aramco, added that “As a world-leading oil and gas company, Saudi Aramco’s mission is to promote the continued use of safe, environmentally-friendly petroleum products with a vision to becoming a global leader in research and technology. Building a strong and cooperative relationship with KAIST in our endeavor to search for alternative ways to better utilization of fossil fuels will expedite the creation of opportunities to make the world environmentally safer and sustainable.”
KAIST and Saudi Aramco will each chip in a maximum of USD 5 million annually for the establishment and operation of the Saudi Aramco-KAIST CO2 Management Center during the initial term of the Master Research and Collaboration Agreement, which starts in 2013 and continues through 2018.
2013.03.19 View 15468 -
 Synthesis of a New Organic Supermolecule Succeeded
From left to right: Prof.Stoddart, Prof.Goddard and Prof.Jang Wook Choi
KAIST EEWS graduate school’s research team led by Prof. Stoddart, Prof. Goddard and Prof. Jang Wook Choi has succeeded the synthesis of a new organic supermolecule that is stable in a radical condition under room temperature.
Prof. Stoddart, who mainly led this research, is the world’s great scholar on orgaic molecular structure especially on catenane with an interconnection of several ring structures. Catenane is originated from Latin “catenane” referring to “chain”. The brief structure of the synthesized catenane is as following:
Usually radicals are known to be unstable since they are electronically neutral and have very high reactivity. However, the radicals from this research showed air- and water- stability. It also showed a reversible change in oxidation number from o to +8 through chemical/electrochemical oxidation-reduction reaction. The phenomenon where paramagnetic and diamagnetic characteristics change according to the oxidation number has also been observed.
Thus, the research like this - on the molecules showing various characteristics with stable radical - is expected to give a new direction to the next-generation electromemory system, semiconductor and energy storage system research.
Meanwhile, this research, led by Prof.Stoddart team with Prof.Goddard and Prof. Jang Wook Choi’s team, is conducted under the support of Science and Technology’s World Class University project by Ministry of Education and published in ‘Science’ on 25th of Jan.
2013.02.24 View 12702
Synthesis of a New Organic Supermolecule Succeeded
From left to right: Prof.Stoddart, Prof.Goddard and Prof.Jang Wook Choi
KAIST EEWS graduate school’s research team led by Prof. Stoddart, Prof. Goddard and Prof. Jang Wook Choi has succeeded the synthesis of a new organic supermolecule that is stable in a radical condition under room temperature.
Prof. Stoddart, who mainly led this research, is the world’s great scholar on orgaic molecular structure especially on catenane with an interconnection of several ring structures. Catenane is originated from Latin “catenane” referring to “chain”. The brief structure of the synthesized catenane is as following:
Usually radicals are known to be unstable since they are electronically neutral and have very high reactivity. However, the radicals from this research showed air- and water- stability. It also showed a reversible change in oxidation number from o to +8 through chemical/electrochemical oxidation-reduction reaction. The phenomenon where paramagnetic and diamagnetic characteristics change according to the oxidation number has also been observed.
Thus, the research like this - on the molecules showing various characteristics with stable radical - is expected to give a new direction to the next-generation electromemory system, semiconductor and energy storage system research.
Meanwhile, this research, led by Prof.Stoddart team with Prof.Goddard and Prof. Jang Wook Choi’s team, is conducted under the support of Science and Technology’s World Class University project by Ministry of Education and published in ‘Science’ on 25th of Jan.
2013.02.24 View 12702 -
 Online Article on President Sung-Mo 'Steve' Kang by California Council on Science and Technology (CCST)
The California Council on Science and Technology (CCST), an independent, not-for-profit organization established by the mandate of California Legislature in 1988, is designed to offer expert advice to the California state government and recommend solutions to science and technology-related policy issues.
Over the past three years, President Sung-Mo “Steve” Kang has served as a member of CCST Council, an assembly of corporate CEOs, academicians, scientists, and scholars of the highest distinction.
On February 21, 2013, CCST posted on its website the announcement of Council Member Sung-Mo “Steve” Kang as President of KAIST along with his personal comments on his move to KAIST and its presidency. For the online article, please visit: http://www.ccst.us/news/2013/0221KAIST.php
2013.02.23 View 9153
Online Article on President Sung-Mo 'Steve' Kang by California Council on Science and Technology (CCST)
The California Council on Science and Technology (CCST), an independent, not-for-profit organization established by the mandate of California Legislature in 1988, is designed to offer expert advice to the California state government and recommend solutions to science and technology-related policy issues.
Over the past three years, President Sung-Mo “Steve” Kang has served as a member of CCST Council, an assembly of corporate CEOs, academicians, scientists, and scholars of the highest distinction.
On February 21, 2013, CCST posted on its website the announcement of Council Member Sung-Mo “Steve” Kang as President of KAIST along with his personal comments on his move to KAIST and its presidency. For the online article, please visit: http://www.ccst.us/news/2013/0221KAIST.php
2013.02.23 View 9153