system
-
 A KAIST Research Team Produces Eco-Friendly Nylon with Engineered Bacterium
With worsening climate change and environmental issues, in recent years, there has been increased interest in the eco-friendly production of polymers like nylon.
On August 10, Dr. Taehee Han from a KAIST research team led by Distinguished Professor Sang Yup Lee in the Department of Chemical and Biomolecular Engineering revealed the successful development of a microbial strain that produces valerolactam, a monomer of nylon-5.
Valerolactam is an important monomer that constitutes nylon-5 and nylon-6,5. Nylon is the oldest synthetic polymer, and nylon-5 is one of its derivatives composed of monomers with five carbons, while nylon-5,6 is composed of two types of monomers with either five or six carbons. They not only have excellent processability, but are also light and tough, which allows them to be applied in a wide range of industrial sectors including clothing, badminton rackets, fishing nets, tents, and gear parts. Monomers are materials that can be built into polymers, and synthetic processes are what connects them into a polymer.
The chemical production of valerolactam, however, is based on petrochemistry, where extreme reaction conditions are required and toxic waste is produced. To solve these problems, efforts are being made to develop environmentally friendly and highly efficient microbial cell factories for lactam production. Systems metabolic engineering, a key strategy for effective microbial strain development, is a research field pioneered by Professor Sang Yup Lee.
Professor Lee’s team used metabolic engineering, a technique for manipulating microbial metabolic pathways, to construct a synthetic metabolic pathway for valerolactam production in Corynebacteriam glutamicum, a bacterium commonly used for amino acid production. With this, they successfully developed a microbial strain that utilizes biomass-derived glucose as a carbon source to produce high-value valerolactam.
In 2017, the team suggested a novel method that metabolically manipulates Escherichia coli to produce valerolactam. However, there were several limitations at the time including low producibility and the generation of harmful byproducts.
< Figure 1. Schematic graphical representation of the development of microorganisms that produce valerolactam, a nylon-5 monomer >
In this research, the team improved valerolactam producibility and incorporated an additional systems metabolic strategy to the developed microbial strain while eliminating the harmful byproducts. By removing the gene involved in the production of the main byproduct and through gene screening, the team successfully converted 5-aminovaleric acid, a byproduct and a precursor, into valerolactam.
Furthermore, by employing a strategy where the 5-aminovaleric acid-converting gene is inserted multiple times into the genome, the team strengthened the metabolic flux for valerolactam production. As a result, they reached a world-record concentration of 76.1 g/L, which is 6.17 times greater than what was previously reported.
This study was published in Metabolic Engineering on July 12, under the title, “Metabolic engineering of Corynebacterium glutamicum for the high-level production of valerolactam, a nylon-5 monomer”.
Dr. Taehee Han, the first author of the paper, said, “The significance of this research lies in our development of an environmentally friendly technology that efficiently produces monomer lactam for nylon production using microorganisms.” She added, “Through this technology, we will be able to take a step forward in replacing the petrochemical industry with a microorganism-based biopolymer industry.”
This work was supported by the “Development of Next-Generation Biofinery Platform Technologies for Leading Bio-based Chemicals Industry Project” funded by the Korean Ministry of Science and ICT.
2023.08.24 View 7679
A KAIST Research Team Produces Eco-Friendly Nylon with Engineered Bacterium
With worsening climate change and environmental issues, in recent years, there has been increased interest in the eco-friendly production of polymers like nylon.
On August 10, Dr. Taehee Han from a KAIST research team led by Distinguished Professor Sang Yup Lee in the Department of Chemical and Biomolecular Engineering revealed the successful development of a microbial strain that produces valerolactam, a monomer of nylon-5.
Valerolactam is an important monomer that constitutes nylon-5 and nylon-6,5. Nylon is the oldest synthetic polymer, and nylon-5 is one of its derivatives composed of monomers with five carbons, while nylon-5,6 is composed of two types of monomers with either five or six carbons. They not only have excellent processability, but are also light and tough, which allows them to be applied in a wide range of industrial sectors including clothing, badminton rackets, fishing nets, tents, and gear parts. Monomers are materials that can be built into polymers, and synthetic processes are what connects them into a polymer.
The chemical production of valerolactam, however, is based on petrochemistry, where extreme reaction conditions are required and toxic waste is produced. To solve these problems, efforts are being made to develop environmentally friendly and highly efficient microbial cell factories for lactam production. Systems metabolic engineering, a key strategy for effective microbial strain development, is a research field pioneered by Professor Sang Yup Lee.
Professor Lee’s team used metabolic engineering, a technique for manipulating microbial metabolic pathways, to construct a synthetic metabolic pathway for valerolactam production in Corynebacteriam glutamicum, a bacterium commonly used for amino acid production. With this, they successfully developed a microbial strain that utilizes biomass-derived glucose as a carbon source to produce high-value valerolactam.
In 2017, the team suggested a novel method that metabolically manipulates Escherichia coli to produce valerolactam. However, there were several limitations at the time including low producibility and the generation of harmful byproducts.
< Figure 1. Schematic graphical representation of the development of microorganisms that produce valerolactam, a nylon-5 monomer >
In this research, the team improved valerolactam producibility and incorporated an additional systems metabolic strategy to the developed microbial strain while eliminating the harmful byproducts. By removing the gene involved in the production of the main byproduct and through gene screening, the team successfully converted 5-aminovaleric acid, a byproduct and a precursor, into valerolactam.
Furthermore, by employing a strategy where the 5-aminovaleric acid-converting gene is inserted multiple times into the genome, the team strengthened the metabolic flux for valerolactam production. As a result, they reached a world-record concentration of 76.1 g/L, which is 6.17 times greater than what was previously reported.
This study was published in Metabolic Engineering on July 12, under the title, “Metabolic engineering of Corynebacterium glutamicum for the high-level production of valerolactam, a nylon-5 monomer”.
Dr. Taehee Han, the first author of the paper, said, “The significance of this research lies in our development of an environmentally friendly technology that efficiently produces monomer lactam for nylon production using microorganisms.” She added, “Through this technology, we will be able to take a step forward in replacing the petrochemical industry with a microorganism-based biopolymer industry.”
This work was supported by the “Development of Next-Generation Biofinery Platform Technologies for Leading Bio-based Chemicals Industry Project” funded by the Korean Ministry of Science and ICT.
2023.08.24 View 7679 -
 Professor Joseph J. Lim of KAIST receives the Best System Paper Award from RSS 2023, First in Korea
- Professor Joseph J. Lim from the Kim Jaechul Graduate School of AI at KAIST and his team receive an award for the most outstanding paper in the implementation of robot systems.
- Professor Lim works on AI-based perception, reasoning, and sequential decision-making to develop systems capable of intelligent decision-making, including robot learning
< Photo 1. RSS2023 Best System Paper Award Presentation >
The team of Professor Joseph J. Lim from the Kim Jaechul Graduate School of AI at KAIST has been honored with the 'Best System Paper Award' at "Robotics: Science and Systems (RSS) 2023".
The RSS conference is globally recognized as a leading event for showcasing the latest discoveries and advancements in the field of robotics. It is a venue where the greatest minds in robotics engineering and robot learning come together to share their research breakthroughs. The RSS Best System Paper Award is a prestigious honor granted to a paper that excels in presenting real-world robot system implementation and experimental results.
< Photo 2. Professor Joseph J. Lim of Kim Jaechul Graduate School of AI at KAIST >
The team led by Professor Lim, including two Master's students and an alumnus (soon to be appointed at Yonsei University), received the prestigious RSS Best System Paper Award, making it the first-ever achievement for a Korean and for a domestic institution.
< Photo 3. Certificate of the Best System Paper Award presented at RSS 2023 >
This award is especially meaningful considering the broader challenges in the field. Although recent progress in artificial intelligence and deep learning algorithms has resulted in numerous breakthroughs in robotics, most of these achievements have been confined to relatively simple and short tasks, like walking or pick-and-place. Moreover, tasks are typically performed in simulated environments rather than dealing with more complex, long-horizon real-world tasks such as factory operations or household chores. These limitations primarily stem from the considerable challenge of acquiring data required to develop and validate learning-based AI techniques, particularly in real-world complex tasks.
In light of these challenges, this paper introduced a benchmark that employs 3D printing to simplify the reproduction of furniture assembly tasks in real-world environments. Furthermore, it proposed a standard benchmark for the development and comparison of algorithms for complex and long-horizon tasks, supported by teleoperation data. Ultimately, the paper suggests a new research direction of addressing complex and long-horizon tasks and encourages diverse advancements in research by facilitating reproducible experiments in real-world environments.
Professor Lim underscored the growing potential for integrating robots into daily life, driven by an aging population and an increase in single-person households. As robots become part of everyday life, testing their performance in real-world scenarios becomes increasingly crucial. He hoped this research would serve as a cornerstone for future studies in this field.
The Master's students, Minho Heo and Doohyun Lee, from the Kim Jaechul Graduate School of AI at KAIST, also shared their aspirations to become global researchers in the domain of robot learning. Meanwhile, the alumnus of Professor Lim's research lab, Dr. Youngwoon Lee, is set to be appointed to the Graduate School of AI at Yonsei University and will continue pursuing research in robot learning.
Paper title: Furniture Bench: Reproducible Real-World Benchmark for Long-Horizon Complex Manipulation. Robotics: Science and Systems.
< Image. Conceptual Summary of the 3D Printing Technology >
2023.07.31 View 12372
Professor Joseph J. Lim of KAIST receives the Best System Paper Award from RSS 2023, First in Korea
- Professor Joseph J. Lim from the Kim Jaechul Graduate School of AI at KAIST and his team receive an award for the most outstanding paper in the implementation of robot systems.
- Professor Lim works on AI-based perception, reasoning, and sequential decision-making to develop systems capable of intelligent decision-making, including robot learning
< Photo 1. RSS2023 Best System Paper Award Presentation >
The team of Professor Joseph J. Lim from the Kim Jaechul Graduate School of AI at KAIST has been honored with the 'Best System Paper Award' at "Robotics: Science and Systems (RSS) 2023".
The RSS conference is globally recognized as a leading event for showcasing the latest discoveries and advancements in the field of robotics. It is a venue where the greatest minds in robotics engineering and robot learning come together to share their research breakthroughs. The RSS Best System Paper Award is a prestigious honor granted to a paper that excels in presenting real-world robot system implementation and experimental results.
< Photo 2. Professor Joseph J. Lim of Kim Jaechul Graduate School of AI at KAIST >
The team led by Professor Lim, including two Master's students and an alumnus (soon to be appointed at Yonsei University), received the prestigious RSS Best System Paper Award, making it the first-ever achievement for a Korean and for a domestic institution.
< Photo 3. Certificate of the Best System Paper Award presented at RSS 2023 >
This award is especially meaningful considering the broader challenges in the field. Although recent progress in artificial intelligence and deep learning algorithms has resulted in numerous breakthroughs in robotics, most of these achievements have been confined to relatively simple and short tasks, like walking or pick-and-place. Moreover, tasks are typically performed in simulated environments rather than dealing with more complex, long-horizon real-world tasks such as factory operations or household chores. These limitations primarily stem from the considerable challenge of acquiring data required to develop and validate learning-based AI techniques, particularly in real-world complex tasks.
In light of these challenges, this paper introduced a benchmark that employs 3D printing to simplify the reproduction of furniture assembly tasks in real-world environments. Furthermore, it proposed a standard benchmark for the development and comparison of algorithms for complex and long-horizon tasks, supported by teleoperation data. Ultimately, the paper suggests a new research direction of addressing complex and long-horizon tasks and encourages diverse advancements in research by facilitating reproducible experiments in real-world environments.
Professor Lim underscored the growing potential for integrating robots into daily life, driven by an aging population and an increase in single-person households. As robots become part of everyday life, testing their performance in real-world scenarios becomes increasingly crucial. He hoped this research would serve as a cornerstone for future studies in this field.
The Master's students, Minho Heo and Doohyun Lee, from the Kim Jaechul Graduate School of AI at KAIST, also shared their aspirations to become global researchers in the domain of robot learning. Meanwhile, the alumnus of Professor Lim's research lab, Dr. Youngwoon Lee, is set to be appointed to the Graduate School of AI at Yonsei University and will continue pursuing research in robot learning.
Paper title: Furniture Bench: Reproducible Real-World Benchmark for Long-Horizon Complex Manipulation. Robotics: Science and Systems.
< Image. Conceptual Summary of the 3D Printing Technology >
2023.07.31 View 12372 -
 KAIST presents a microbial cell factory as a source of eco-friendly food and cosmetic coloring
Despite decades of global population growth, global food crisis seems to be at hand yet again because the food productivity is cut severely due to prolonged presence of abnormal weather from intensifying climate change and global food supply chain is deteriorated due to international conflicts such as wars exacerbating food shortages and nutritional inequality around the globe. At the same time, however, as awareness of the environment and sustainability rises, an increase in demand for more eco-friendly and high-quality food and beauty products is being observed not without a sense of irony. At a time like this, microorganisms are attracting attention as a key that can handle this couple of seemingly distant problems.
KAIST (President Kwang-Hyung Lee) announced on the 26th that Kyeong Rok Choi, a research professor of the Bioprocess Research Center and Sang Yup Lee, a Distinguished Professor of the Department of Chemical and Biomolecular Engineering, published a paper titled “Metabolic Engineering of Microorganisms for Food and Cosmetics Production” upon invitation by “Nature Reviews Bioengineering” to be published online published by Nature after peer review.
※ Paper title: Systems metabolic engineering of microorganisms for food and cosmetics production
※ Author information: Kyeong Rok Choi (first author) and Sang Yup Lee (corresponding author)
Systems metabolic engineering is a research field founded by Distinguished Professor Sang Yup Lee of KAIST to more effectively develop microbial cell factories, the core factor of the next-generation bio industry to replace the existing chemical industry that relies heavily on petroleum. By applying a systemic metabolic engineering strategy, the researchers have developed a number of high-performance microbial cell factories that produce a variety of food and cosmetic compounds including natural substances like heme and zinc protoporphyrin IX compounds which can improve the flavor and color of synthetic meat, lycopene and β-carotene which are functional natural pigments that can be widely used in food and cosmetics, and methyl anthranilate, a grape-derived compound widely used to impart grape flavor in food and beverage manufacturing.
In this paper written upon invitation by Nature, the research team covered remarkable cases of microbial cell factory that can produce amino acids, proteins, fats and fatty acids, vitamins, flavors, pigments, alcohols, functional compounds and other food additives used in various foods and cosmetics and the companies that have successfully commercialized these microbial-derived materials Furthermore, the paper organized and presents systems metabolic engineering strategies that can spur the development of industrial microbial cell factories that can produce more diverse food and cosmetic compounds in an eco-friendly way with economic feasibility.
< Figure 1. Examples of production of food and cosmetic compounds using microbial cell factories >
For example, by producing proteins or amino acids with high nutritional value through non-edible biomass used as animal feed or fertilizer through the microbial fermentation process, it will contribute to the increase in production and stable supply of food around the world. Furthermore, by contributing to developing more viable alternative meat, further reducing dependence on animal protein, it can also contribute to reducing greenhouse gases and environmental pollution generated through livestock breeding or fish farming.
In addition, vanillin or methyl anthranilate, which give off vanilla or grape flavor, are widely added to various foods, but natural products isolated and refined from plants are low in production and high in production cost, so in most cases, petrochemicals substances derived from vanillin and methylanthranilic acid are added to food. These materials can also be produced through an eco-friendly and human-friendly method by borrowing the power of microorganisms.
Ethical and resource problems that arise in producing compounds like Calmin (cochineal pigment), a coloring added to various cosmetics and foods such as red lipstick and strawberry-flavored milk, which must be extracted from cochineal insects that live only in certain cacti. and Hyaluronic acid, which is widely consumed as a health supplement, but is only present in omega-3 fatty acids extracted from shark or fish livers, can also be resolved when they can be produced in an eco-friendly way using microorganisms.
KAIST Research Professor Kyeong Rok Choi, the first author of this paper, said, “In addition to traditional fermented foods such as kimchi and yogurt, foods produced with the help of microorganisms like cocoa butter, a base ingredient for chocolate that can only be obtained from fermented cacao beans, and monosodium glutamate, a seasoning produced through microbial fermentation are already familiar to us”. “In the future, we will be able to acquire a wider variety of foods and cosmetics even more easily produced in an eco-friendly and sustainable way in our daily lives through microbial cell factories.” he added.
< Figure 2. Systems metabolic engineering strategy to improve metabolic flow in microbial cell factories >
Distinguished Professor Sang Yup Lee said, “It is engineers’ mission to make the world a better place utilizing science and technology.” and added, “Continuous advancement and active use of systems metabolic engineering will contribute greatly to easing and resolving the problems arising from both the food crisis and the climate change."
This research was carried out as a part of the “Development of Protein Production Technology from Inorganic Substances through Control of Microbial Metabolism System Project” (Project Leader: Kyeong Rok Choi, KAIST Research Professor) of the the Center for Agricultural Microorganism and Enzyme (Director Pahn-Shick Chang) supported by the Rural Development Administration and the “Development of Platform Technologies of Microbial Cell Factories for the Next-generation Biorefineries Project” (Project Leader: Sang Yup Lee, KAIST Distinguished Professor) of the Petroleum-Substitute Eco-friendly Chemical Technology Development Program supported by the Ministry of Science and ICT.
2023.07.28 View 10975
KAIST presents a microbial cell factory as a source of eco-friendly food and cosmetic coloring
Despite decades of global population growth, global food crisis seems to be at hand yet again because the food productivity is cut severely due to prolonged presence of abnormal weather from intensifying climate change and global food supply chain is deteriorated due to international conflicts such as wars exacerbating food shortages and nutritional inequality around the globe. At the same time, however, as awareness of the environment and sustainability rises, an increase in demand for more eco-friendly and high-quality food and beauty products is being observed not without a sense of irony. At a time like this, microorganisms are attracting attention as a key that can handle this couple of seemingly distant problems.
KAIST (President Kwang-Hyung Lee) announced on the 26th that Kyeong Rok Choi, a research professor of the Bioprocess Research Center and Sang Yup Lee, a Distinguished Professor of the Department of Chemical and Biomolecular Engineering, published a paper titled “Metabolic Engineering of Microorganisms for Food and Cosmetics Production” upon invitation by “Nature Reviews Bioengineering” to be published online published by Nature after peer review.
※ Paper title: Systems metabolic engineering of microorganisms for food and cosmetics production
※ Author information: Kyeong Rok Choi (first author) and Sang Yup Lee (corresponding author)
Systems metabolic engineering is a research field founded by Distinguished Professor Sang Yup Lee of KAIST to more effectively develop microbial cell factories, the core factor of the next-generation bio industry to replace the existing chemical industry that relies heavily on petroleum. By applying a systemic metabolic engineering strategy, the researchers have developed a number of high-performance microbial cell factories that produce a variety of food and cosmetic compounds including natural substances like heme and zinc protoporphyrin IX compounds which can improve the flavor and color of synthetic meat, lycopene and β-carotene which are functional natural pigments that can be widely used in food and cosmetics, and methyl anthranilate, a grape-derived compound widely used to impart grape flavor in food and beverage manufacturing.
In this paper written upon invitation by Nature, the research team covered remarkable cases of microbial cell factory that can produce amino acids, proteins, fats and fatty acids, vitamins, flavors, pigments, alcohols, functional compounds and other food additives used in various foods and cosmetics and the companies that have successfully commercialized these microbial-derived materials Furthermore, the paper organized and presents systems metabolic engineering strategies that can spur the development of industrial microbial cell factories that can produce more diverse food and cosmetic compounds in an eco-friendly way with economic feasibility.
< Figure 1. Examples of production of food and cosmetic compounds using microbial cell factories >
For example, by producing proteins or amino acids with high nutritional value through non-edible biomass used as animal feed or fertilizer through the microbial fermentation process, it will contribute to the increase in production and stable supply of food around the world. Furthermore, by contributing to developing more viable alternative meat, further reducing dependence on animal protein, it can also contribute to reducing greenhouse gases and environmental pollution generated through livestock breeding or fish farming.
In addition, vanillin or methyl anthranilate, which give off vanilla or grape flavor, are widely added to various foods, but natural products isolated and refined from plants are low in production and high in production cost, so in most cases, petrochemicals substances derived from vanillin and methylanthranilic acid are added to food. These materials can also be produced through an eco-friendly and human-friendly method by borrowing the power of microorganisms.
Ethical and resource problems that arise in producing compounds like Calmin (cochineal pigment), a coloring added to various cosmetics and foods such as red lipstick and strawberry-flavored milk, which must be extracted from cochineal insects that live only in certain cacti. and Hyaluronic acid, which is widely consumed as a health supplement, but is only present in omega-3 fatty acids extracted from shark or fish livers, can also be resolved when they can be produced in an eco-friendly way using microorganisms.
KAIST Research Professor Kyeong Rok Choi, the first author of this paper, said, “In addition to traditional fermented foods such as kimchi and yogurt, foods produced with the help of microorganisms like cocoa butter, a base ingredient for chocolate that can only be obtained from fermented cacao beans, and monosodium glutamate, a seasoning produced through microbial fermentation are already familiar to us”. “In the future, we will be able to acquire a wider variety of foods and cosmetics even more easily produced in an eco-friendly and sustainable way in our daily lives through microbial cell factories.” he added.
< Figure 2. Systems metabolic engineering strategy to improve metabolic flow in microbial cell factories >
Distinguished Professor Sang Yup Lee said, “It is engineers’ mission to make the world a better place utilizing science and technology.” and added, “Continuous advancement and active use of systems metabolic engineering will contribute greatly to easing and resolving the problems arising from both the food crisis and the climate change."
This research was carried out as a part of the “Development of Protein Production Technology from Inorganic Substances through Control of Microbial Metabolism System Project” (Project Leader: Kyeong Rok Choi, KAIST Research Professor) of the the Center for Agricultural Microorganism and Enzyme (Director Pahn-Shick Chang) supported by the Rural Development Administration and the “Development of Platform Technologies of Microbial Cell Factories for the Next-generation Biorefineries Project” (Project Leader: Sang Yup Lee, KAIST Distinguished Professor) of the Petroleum-Substitute Eco-friendly Chemical Technology Development Program supported by the Ministry of Science and ICT.
2023.07.28 View 10975 -
 A KAIST research team develops a high-performance modular SSD system semiconductor
In recent years, there has been a rise in demand for large amounts of data to train AI models and, thus, data size has become increasingly important over time. Accordingly, solid state drives (SSDs, storage devices that use a semiconductor memory unit), which are core storage devices for data centers and cloud services, have also seen an increase in demand. However, the internal components of higher performing SSDs have become more tightly coupled, and this tightly-coupled structure limits SSD from maximized performance.
On June 15, a KAIST research team led by Professor Dongjun Kim (John Kim) from the School of Electrical Engineering (EE) announced the development of the first SSD system semiconductor structure that can increase the reading/writing performance of next generation SSDs and extend their lifespan through high-performance modular SSD systems.
Professor Kim’s team identified the limitations of the tightly-coupled structures in existing SSD designs and proposed a de-coupled structure that can maximize SSD performance by configuring an internal on-chip network specialized for flash memory. This technique utilizes on-chip network technology, which can freely send packet-based data within the chip and is often used to design non-memory system semiconductors like CPUs and GPUs. Through this, the team developed a ‘modular SSD’, which shows reduced interdependence between front-end and back-end designs, and allows their independent design and assembly.
*on-chip network: a packet-based connection structure for the internal components of system semiconductors like CPUs/GPUs. On-chip networks are one of the most critical design components for high-performing system semiconductors, and their importance grows with the size of the semiconductor chip.
Professor Kim’s team refers to the components nearer to the CPU as the front-end and the parts closer to the flash memory as back-end. They newly constructed an on-chip network specific to flash memory in order to allow data transmission between the back-end’s flash controller, proposing a de-coupled structure that can minimize performance drop.
The SSD can accelerate some functions of the flash translation layer, a critical element to drive the SSD, in order to allow flash memory to actively overcome its limitations. Another advantage of the de-coupled, modular structure is that the flash translation layer is not limited to the characteristics of specific flash memories. Instead, their front-end and back-end designs can be carried out independently. Through this, the team could produce 21-times faster response times compared to existing systems and extend SSD lifespan by 23% by also applying the DDS defect detection technique.
< Figure 1. Schematic diagram of the structure of a high-performance modular SSD system developed by Professor Dong-Jun Kim's team >
This research, conducted by first author and Ph.D. candidate Jiho Kim from the KAIST School of EE and co-author Professor Myoungsoo Jung, was presented on the 19th of June at the 50th IEEE/ACM International Symposium on Computer Architecture, the most prestigious academic conference in the field of computer architecture, held in Orlando, Florida. (Paper Title: Decoupled SSD: Rethinking SSD Architecture through Network-based Flash Controllers)
< Figure 2. Conceptual diagram of hardware acceleration through high-performance modular SSD system >
Professor Dongjun Kim, who led the research, said, “This research is significant in that it identified the structural limitations of existing SSDs, and showed that on-chip network technology based on system memory semiconductors like CPUs can drive the hardware to actively carry out the necessary actions. We expect this to contribute greatly to the next-generation high-performance SSD market.” He added, “The de-coupled architecture is a structure that can actively operate to extend devices’ lifespan. In other words, its significance is not limited to the level of performance and can, therefore, be used for various applications.”
KAIST commented that this research is also meaningful in that the results were reaped through a collaborative study between two world-renowned researchers: Professor Myeongsoo Jung, recognized in the field of computer system storage devices, and Professor Dongjun Kim, a leading researcher in computer architecture and interconnection networks.
This research was funded by the National Research Foundation of Korea, Samsung Electronics, the IC Design Education Center, and Next Generation Semiconductor Technology and Development granted by the Institute of Information & Communications Technology, Planning & Evaluation.
2023.06.23 View 9300
A KAIST research team develops a high-performance modular SSD system semiconductor
In recent years, there has been a rise in demand for large amounts of data to train AI models and, thus, data size has become increasingly important over time. Accordingly, solid state drives (SSDs, storage devices that use a semiconductor memory unit), which are core storage devices for data centers and cloud services, have also seen an increase in demand. However, the internal components of higher performing SSDs have become more tightly coupled, and this tightly-coupled structure limits SSD from maximized performance.
On June 15, a KAIST research team led by Professor Dongjun Kim (John Kim) from the School of Electrical Engineering (EE) announced the development of the first SSD system semiconductor structure that can increase the reading/writing performance of next generation SSDs and extend their lifespan through high-performance modular SSD systems.
Professor Kim’s team identified the limitations of the tightly-coupled structures in existing SSD designs and proposed a de-coupled structure that can maximize SSD performance by configuring an internal on-chip network specialized for flash memory. This technique utilizes on-chip network technology, which can freely send packet-based data within the chip and is often used to design non-memory system semiconductors like CPUs and GPUs. Through this, the team developed a ‘modular SSD’, which shows reduced interdependence between front-end and back-end designs, and allows their independent design and assembly.
*on-chip network: a packet-based connection structure for the internal components of system semiconductors like CPUs/GPUs. On-chip networks are one of the most critical design components for high-performing system semiconductors, and their importance grows with the size of the semiconductor chip.
Professor Kim’s team refers to the components nearer to the CPU as the front-end and the parts closer to the flash memory as back-end. They newly constructed an on-chip network specific to flash memory in order to allow data transmission between the back-end’s flash controller, proposing a de-coupled structure that can minimize performance drop.
The SSD can accelerate some functions of the flash translation layer, a critical element to drive the SSD, in order to allow flash memory to actively overcome its limitations. Another advantage of the de-coupled, modular structure is that the flash translation layer is not limited to the characteristics of specific flash memories. Instead, their front-end and back-end designs can be carried out independently. Through this, the team could produce 21-times faster response times compared to existing systems and extend SSD lifespan by 23% by also applying the DDS defect detection technique.
< Figure 1. Schematic diagram of the structure of a high-performance modular SSD system developed by Professor Dong-Jun Kim's team >
This research, conducted by first author and Ph.D. candidate Jiho Kim from the KAIST School of EE and co-author Professor Myoungsoo Jung, was presented on the 19th of June at the 50th IEEE/ACM International Symposium on Computer Architecture, the most prestigious academic conference in the field of computer architecture, held in Orlando, Florida. (Paper Title: Decoupled SSD: Rethinking SSD Architecture through Network-based Flash Controllers)
< Figure 2. Conceptual diagram of hardware acceleration through high-performance modular SSD system >
Professor Dongjun Kim, who led the research, said, “This research is significant in that it identified the structural limitations of existing SSDs, and showed that on-chip network technology based on system memory semiconductors like CPUs can drive the hardware to actively carry out the necessary actions. We expect this to contribute greatly to the next-generation high-performance SSD market.” He added, “The de-coupled architecture is a structure that can actively operate to extend devices’ lifespan. In other words, its significance is not limited to the level of performance and can, therefore, be used for various applications.”
KAIST commented that this research is also meaningful in that the results were reaped through a collaborative study between two world-renowned researchers: Professor Myeongsoo Jung, recognized in the field of computer system storage devices, and Professor Dongjun Kim, a leading researcher in computer architecture and interconnection networks.
This research was funded by the National Research Foundation of Korea, Samsung Electronics, the IC Design Education Center, and Next Generation Semiconductor Technology and Development granted by the Institute of Information & Communications Technology, Planning & Evaluation.
2023.06.23 View 9300 -
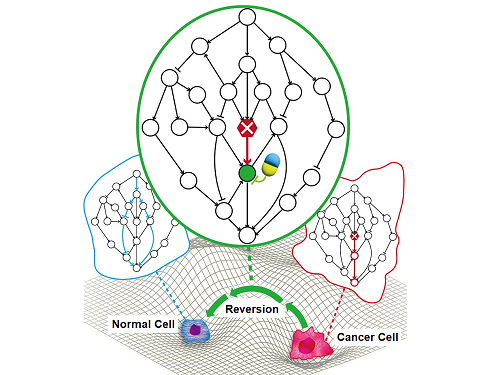 A KAIST Research Team Identifies a Cancer Reversion Mechanism
Despite decades of intensive cancer research by numerous biomedical scientists, cancer still holds its place as the number one cause of death in Korea. The fundamental reason behind the limitations of current cancer treatment methods is the fact that they all aim to completely destroy cancer cells, which eventually allows the cancer cells to acquire immunity. In other words, recurrences and side-effects caused by the destruction of healthy cells are inevitable. To this end, some have suggested anticancer treatment methods based on cancer reversion, which can revert cancer cells back to normal or near-normal cells under certain conditions. However, the practical development of this idea has not yet been attempted.
On June 8, a KAIST research team led by Professor Kwang-Hyun Cho from the Department of Bio and Brain Engineering reported to have successfully identified the fundamental principle of a process that can revert cancer cells back to normal cells without killing the cells.
Professor Cho’s team focused on the fact that unlike normal cells, which react according to external stimuli, cancer cells tend to ignore such stimuli and only undergo uncontrolled cell division. Through computer simulation analysis, the team discovered that the input-output (I/O) relationships that were distorted by genetic mutations could be reverted back to normal I/O relationships under certain conditions. The team then demonstrated through molecular cell experiments that such I/O relationship recovery also occurred in real cancer cells.
The results of this study, written by Dr. Jae Il Joo and Dr. Hwa-Jeong Park, were published in Wiley’s Advanced Science online on June 2 under the title, "Normalizing input-output relationships of cancer networks for reversion therapy."
< Image 1. Input-output (I/O) relationships in gene regulatory networks >
Professor Kwang-Hyun Cho's research team classified genes into four types by simulation-analyzing the effect of gene mutations on the I/O relationship of gene regulatory networks. (Figure A-J) In addition, by analyzing 18 genes of the cancer-related gene regulatory network, it was confirmed that when mutations occur in more than half of the genes constituting each network, reversibility is possible through appropriate control. (Figure K)
Professor Cho’s team uncovered that the reason the distorted I/O relationships of cancer cells could be reverted back to normal ones was the robustness and redundancy of intracellular gene control networks that developed over the course of evolution. In addition, they found that some genes were more promising as targets for cancer reversion than others, and showed through molecular cell experiments that controlling such genes could revert the distorted I/O relationships of cancer cells back to normal ones.
< Image 2. Simulation results of restoration of bladder cancer gene regulation network and I/O relationship of bladder cancer cells. >
The research team classified the effects of gene mutations on the I/O relationship in the bladder cancer gene regulation network by simulation analysis and classified them into 4 types. (Figure A) Through this, it was found that the distorted input-output relationship between bladder cancer cell lines KU-1919 and HCT-1197 could be restored to normal. (Figure B)
< Image 3. Analysis of survival of bladder cancer patients according to reversible gene mutation and I/O recovery experiment of bladder cancer cells. >
As predicted through network simulation analysis, Professor Kwang-Hyun Cho's research team confirmed through molecular cell experiments that the response to TGF-b was normally restored when AKT and MAP3K1 were inhibited in the bladder cancer cell line KU-1919. (Figure A-G) In addition, it was confirmed that there is a difference in the survival rate of bladder cancer patients depending on the presence or absence of a reversible gene mutation. (Figure H)
The results of this research show that the reversion of real cancer cells does not happen by chance, and that it is possible to systematically explore targets that can induce this phenomenon, thereby creating the potential for the development of innovative anticancer drugs that can control such target genes.
< Image 4. Cancer cell reversibility principle >
The research team analyzed the reversibility, redundancy, and robustness of various networks and found that there was a positive correlation between them. From this, it was found that reversibility was additionally inherent in the process of evolution in which the gene regulatory network acquired redundancy and consistency.
Professor Cho said, “By uncovering the fundamental principles of a new cancer reversion treatment strategy that may overcome the unresolved limitations of existing chemotherapy, we have increased the possibility of developing new and innovative drugs that can improve both the prognosis and quality of life of cancer patients.”
< Image 5. Conceptual diagram of research results >
The research team identified the fundamental control principle of cancer cell reversibility through systems biology research. When the I/O relationship of the intracellular gene regulatory network is distorted by mutation, the distorted I/O relationship can be restored to a normal state by identifying and adjusting the reversible gene target based on the redundancy of the molecular circuit inherent in the complex network.
After Professor Cho’s team first suggested the concept of reversion treatment, they published their results for reverting colorectal cancer in January 2020, and in January 2022 they successfully re-programmed malignant breast cancer cells back into hormone-treatable ones. In January 2023, the team successfully removed the metastasis ability from lung cancer cells and reverted them back to a state that allowed improved drug reactivity. However, these results were case studies of specific types of cancer and did not reveal what common principle allowed cancer reversion across all cancer types, making this the first revelation of the general principle of cancer reversion and its evolutionary origins.
This research was funded by the Ministry of Science and ICT of the Republic of Korea and the National Research Foundation of Korea.
2023.06.20 View 13220
A KAIST Research Team Identifies a Cancer Reversion Mechanism
Despite decades of intensive cancer research by numerous biomedical scientists, cancer still holds its place as the number one cause of death in Korea. The fundamental reason behind the limitations of current cancer treatment methods is the fact that they all aim to completely destroy cancer cells, which eventually allows the cancer cells to acquire immunity. In other words, recurrences and side-effects caused by the destruction of healthy cells are inevitable. To this end, some have suggested anticancer treatment methods based on cancer reversion, which can revert cancer cells back to normal or near-normal cells under certain conditions. However, the practical development of this idea has not yet been attempted.
On June 8, a KAIST research team led by Professor Kwang-Hyun Cho from the Department of Bio and Brain Engineering reported to have successfully identified the fundamental principle of a process that can revert cancer cells back to normal cells without killing the cells.
Professor Cho’s team focused on the fact that unlike normal cells, which react according to external stimuli, cancer cells tend to ignore such stimuli and only undergo uncontrolled cell division. Through computer simulation analysis, the team discovered that the input-output (I/O) relationships that were distorted by genetic mutations could be reverted back to normal I/O relationships under certain conditions. The team then demonstrated through molecular cell experiments that such I/O relationship recovery also occurred in real cancer cells.
The results of this study, written by Dr. Jae Il Joo and Dr. Hwa-Jeong Park, were published in Wiley’s Advanced Science online on June 2 under the title, "Normalizing input-output relationships of cancer networks for reversion therapy."
< Image 1. Input-output (I/O) relationships in gene regulatory networks >
Professor Kwang-Hyun Cho's research team classified genes into four types by simulation-analyzing the effect of gene mutations on the I/O relationship of gene regulatory networks. (Figure A-J) In addition, by analyzing 18 genes of the cancer-related gene regulatory network, it was confirmed that when mutations occur in more than half of the genes constituting each network, reversibility is possible through appropriate control. (Figure K)
Professor Cho’s team uncovered that the reason the distorted I/O relationships of cancer cells could be reverted back to normal ones was the robustness and redundancy of intracellular gene control networks that developed over the course of evolution. In addition, they found that some genes were more promising as targets for cancer reversion than others, and showed through molecular cell experiments that controlling such genes could revert the distorted I/O relationships of cancer cells back to normal ones.
< Image 2. Simulation results of restoration of bladder cancer gene regulation network and I/O relationship of bladder cancer cells. >
The research team classified the effects of gene mutations on the I/O relationship in the bladder cancer gene regulation network by simulation analysis and classified them into 4 types. (Figure A) Through this, it was found that the distorted input-output relationship between bladder cancer cell lines KU-1919 and HCT-1197 could be restored to normal. (Figure B)
< Image 3. Analysis of survival of bladder cancer patients according to reversible gene mutation and I/O recovery experiment of bladder cancer cells. >
As predicted through network simulation analysis, Professor Kwang-Hyun Cho's research team confirmed through molecular cell experiments that the response to TGF-b was normally restored when AKT and MAP3K1 were inhibited in the bladder cancer cell line KU-1919. (Figure A-G) In addition, it was confirmed that there is a difference in the survival rate of bladder cancer patients depending on the presence or absence of a reversible gene mutation. (Figure H)
The results of this research show that the reversion of real cancer cells does not happen by chance, and that it is possible to systematically explore targets that can induce this phenomenon, thereby creating the potential for the development of innovative anticancer drugs that can control such target genes.
< Image 4. Cancer cell reversibility principle >
The research team analyzed the reversibility, redundancy, and robustness of various networks and found that there was a positive correlation between them. From this, it was found that reversibility was additionally inherent in the process of evolution in which the gene regulatory network acquired redundancy and consistency.
Professor Cho said, “By uncovering the fundamental principles of a new cancer reversion treatment strategy that may overcome the unresolved limitations of existing chemotherapy, we have increased the possibility of developing new and innovative drugs that can improve both the prognosis and quality of life of cancer patients.”
< Image 5. Conceptual diagram of research results >
The research team identified the fundamental control principle of cancer cell reversibility through systems biology research. When the I/O relationship of the intracellular gene regulatory network is distorted by mutation, the distorted I/O relationship can be restored to a normal state by identifying and adjusting the reversible gene target based on the redundancy of the molecular circuit inherent in the complex network.
After Professor Cho’s team first suggested the concept of reversion treatment, they published their results for reverting colorectal cancer in January 2020, and in January 2022 they successfully re-programmed malignant breast cancer cells back into hormone-treatable ones. In January 2023, the team successfully removed the metastasis ability from lung cancer cells and reverted them back to a state that allowed improved drug reactivity. However, these results were case studies of specific types of cancer and did not reveal what common principle allowed cancer reversion across all cancer types, making this the first revelation of the general principle of cancer reversion and its evolutionary origins.
This research was funded by the Ministry of Science and ICT of the Republic of Korea and the National Research Foundation of Korea.
2023.06.20 View 13220 -
 KAIST Holds 2023 Commencement Ceremony
< Photo 1. On the 17th, KAIST held the 2023 Commencement Ceremony for a total of 2,870 students, including 691 doctors. >
KAIST held its 2023 commencement ceremony at the Sports Complex of its main campus in Daejeon at 2 p.m. on February 27. It was the first commencement ceremony to invite all its graduates since the start of COVID-19 quarantine measures.
KAIST awarded a total of 2,870 degrees including 691 PhD degrees, 1,464 master’s degrees, and 715 bachelor’s degrees, which adds to the total of 74,999 degrees KAIST has conferred since its foundation in 1971, which includes 15,772 PhD, 38,360 master’s and 20,867 bachelor’s degrees.
This year’s Cum Laude, Gabin Ryu, from the Department of Mechanical Engineering received the Minister of Science and ICT Award. Seung-ju Lee from the School of Computing received the Chairman of the KAIST Board of Trustees Award, while Jantakan Nedsaengtip, an international student from Thailand received the KAIST Presidential Award, and Jaeyong Hwang from the Department of Physics and Junmo Lee from the Department of Industrial and Systems Engineering each received the President of the Alumni Association Award and the Chairman of the KAIST Development Foundation Award, respectively.
Minister Jong-ho Lee of the Ministry of Science and ICT awarded the recipients of the academic awards and delivered a congratulatory speech.
Yujin Cha from the Department of Bio and Brain Engineering, who received a PhD degree after 19 years since his entrance to KAIST as an undergraduate student in 2004 gave a speech on behalf of the graduates to move and inspire the graduates and the guests.
After Cha received a bachelor’s degree from the Department of Nuclear and Quantum Engineering, he entered a medical graduate school and became a radiation oncology specialist. But after experiencing the death of a young patient who suffered from osteosarcoma, he returned to his alma mater to become a scientist. As he believes that science and technology is the ultimate solution to the limitations of modern medicine, he started as a PhD student at the Department of Bio and Brain Engineering in 2018, hoping to find such solutions.
During his course, he identified the characteristics of the decision-making process of doctors during diagnosis, and developed a brain-inspired AI algorithm. It is an original and challenging study that attempted to develop a fundamental machine learning theory from the data he collected from 200 doctors of different specialties.
Cha said, “Humans and AI can cooperate by humans utilizing the unique learning abilities of AI to develop our expertise, while AIs can mimic us humans’ learning abilities to improve.” He added, “My ultimate goal is to develop technology to a level at which humans and machines influence each other and ‘coevolve’, and applying it not only to medicine, but in all areas.”
Cha, who is currently an assistant professor at the KAIST Biomedical Research Center, has also written Artificial Intelligence for Doctors in 2017 to help medical personnel use AI in clinical fields, and the book was selected as one of the 2018 Sejong Books in the academic category.
During his speech at this year’s commencement ceremony, he shared that “there are so many things in the world that are difficult to solve and many things to solve them with, but I believe the things that can really broaden the horizons of the world and find fundamental solutions to the problems at hand are science and technology.”
Meanwhile, singer-songwriter Sae Byul Park who studied at the KAIST Graduate School of Culture Technology will also receive her PhD degree.
Natural language processing (NLP) is a field in AI that teaches a computer to understand and analyze human language that is actively being studied. An example of NLP is ChatGTP, which recently received a lot of attention. For her research, Park analyzed music rather than language using NLP technology.
To analyze music, which is in the form of sound, using the methods for NLP, it is necessary to rebuild notes and beats into a form of words or sentences as in a language. For this, Park designed an algorithm called Mel2Word and applied it to her research.
She also suggested that by converting melodies into texts for analysis, one would be able to quantitatively express music as sentences or words with meaning and context rather than as simple sounds representing a certain note.
Park said, “music has always been considered as a product of subjective emotion, but this research provides a framework that can calculate and analyze music.”
Park’s study can later be developed into a tool to measure the similarities between musical work, as well as a piece’s originality, artistry and popularity, and it can be used as a clue to explore the fundamental principles of how humans respond to music from a cognitive science perspective.
Park began her Ph.D. program in 2014, while carrying on with her musical activities as well as public and university lectures alongside, and dealing with personally major events including marriage and childbirth during the course of years. She already met the requirements to receive her degree in 2019, but delayed her graduation in order to improve the level of completion of her research, and finally graduated with her current achievements after nine years.
Professor Juhan Nam, who supervised Park’s research, said, “Park, who has a bachelor’s degree in psychology, later learned to code for graduate school, and has complete high-quality research in the field of artificial intelligence.” He added, “Though it took a long time, her attitude of not giving up until the end as a researcher is also excellent.”
Sae Byul Park is currently lecturing courses entitled Culture Technology and Music Information Retrieval at the Underwood International College of Yonsei University.
Park said, “the 10 or so years I’ve spent at KAIST as a graduate student was a time I could learn and prosper not only academically but from all angles of life.” She added, “having received a doctorate degree is not the end, but a ‘commencement’. Therefore, I will start to root deeper from the seeds I sowed and work harder as a both a scholar and an artist.”
< Photo 2. From left) Yujin Cha (Valedictorian, Medical-Scientist Program Ph.D. graduate), Saebyeol Park (a singer-songwriter, Ph.D. graduate from the Graduate School of Culture and Technology), Junseok Moon and Inah Seo (the two highlighted CEO graduates from the Department of Management Engineering's master’s program) >
Young entrepreneurs who dream of solving social problems will also be wearing their graduation caps. Two such graduates are Jun-seok Moon and Inah Seo, receiving their master’s degrees in social entrepreneurship MBA from the KAIST College of Business.
Before entrance, Moon ran a café helping African refugees stand on their own feet. Then, he entered KAIST to later expand his business and learn social entrepreneurship in order to sustainably help refugees in the blind spots of human rights and welfare.
During his master’s course, Moon realized that he could achieve active carbon reduction by changing the coffee alone, and switched his business field and founded Equal Table. The amount of carbon an individual can reduce by refraining from using a single paper cup is 10g, while changing the coffee itself can reduce it by 300g.
1kg of coffee emits 15kg of carbon over the course of its production, distribution, processing, and consumption, but Moon produces nearly carbon-neutral coffee beans by having innovated the entire process. In particular, the company-to-company ESG business solution is Moon’s new start-up area. It provides companies with carbon-reduced coffee made by roasting raw beans from carbon-neutral certified farms with 100% renewable energy, and shows how much carbon has been reduced in its making. Equal Table will launch the service this month in collaboration with SK Telecom, its first partner.
Inah Seo, who also graduated with Moon, founded Conscious Wear to start a fashion business reducing environmental pollution. In order to realize her mission, she felt the need to gain the appropriate expertise in management, and enrolled for the social entrepreneurship MBA.
Out of the various fashion industries, Seo focused on the leather market, which is worth 80 trillion won. Due to thickness or contamination issues, only about 60% of animal skin fabric is used, and the rest is discarded. Heavy metals are used during such processes, which also directly affects the environment.
During the social entrepreneurship MBA course, Seo collaborated with SK Chemicals, which had links through the program, and launched eco-friendly leather bags. The bags used discarded leather that was recycled by grinding and reprocessing into a biomaterial called PO3G. It was the first case in which PO3G that is over 90% biodegradable was applied to regenerated leather. In other words, it can reduce environmental pollution in the processing and disposal stages, while also reducing carbon emissions and water usage by one-tenth compared to existing cowhide products.
The social entrepreneurship MBA course, from which Moon and Seo graduated, will run in integration with the Graduate School of Green Growth as an Impact MBA program starting this year. KAIST plans to steadily foster entrepreneurs who will lead meaningful changes in the environment and society as well as economic values through innovative technologies and ideas.
< Photo 3. NYU President Emeritus John Sexton (left), who received this year's honorary doctorate of science, poses with President Kwang Hyung Lee >
Meanwhile, during this day’s commencement ceremony, KAIST also presented President Emeritus John Sexton of New York University with an honorary doctorate in science. He was recognized for laying the foundation for the cooperation between KAIST and New York University, such as promoting joint campuses.
< Photo 4. At the commencement ceremony of KAIST held on the 17th, President Kwang Hyung Lee is encouraging the graduates with his commencement address. >
President Kwang Hyung Lee emphasized in his commencement speech that, “if you can draw up the future and work hard toward your goal, the future can become a work of art that you create with your own hands,” and added, “Never stop on the journey toward your dreams, and do not give up even when you are met with failure. Failure happens to everyone, all the time. The important thing is to know 'why you failed', and to use those elements of failure as the driving force for the next try.”
2023.02.20 View 24564
KAIST Holds 2023 Commencement Ceremony
< Photo 1. On the 17th, KAIST held the 2023 Commencement Ceremony for a total of 2,870 students, including 691 doctors. >
KAIST held its 2023 commencement ceremony at the Sports Complex of its main campus in Daejeon at 2 p.m. on February 27. It was the first commencement ceremony to invite all its graduates since the start of COVID-19 quarantine measures.
KAIST awarded a total of 2,870 degrees including 691 PhD degrees, 1,464 master’s degrees, and 715 bachelor’s degrees, which adds to the total of 74,999 degrees KAIST has conferred since its foundation in 1971, which includes 15,772 PhD, 38,360 master’s and 20,867 bachelor’s degrees.
This year’s Cum Laude, Gabin Ryu, from the Department of Mechanical Engineering received the Minister of Science and ICT Award. Seung-ju Lee from the School of Computing received the Chairman of the KAIST Board of Trustees Award, while Jantakan Nedsaengtip, an international student from Thailand received the KAIST Presidential Award, and Jaeyong Hwang from the Department of Physics and Junmo Lee from the Department of Industrial and Systems Engineering each received the President of the Alumni Association Award and the Chairman of the KAIST Development Foundation Award, respectively.
Minister Jong-ho Lee of the Ministry of Science and ICT awarded the recipients of the academic awards and delivered a congratulatory speech.
Yujin Cha from the Department of Bio and Brain Engineering, who received a PhD degree after 19 years since his entrance to KAIST as an undergraduate student in 2004 gave a speech on behalf of the graduates to move and inspire the graduates and the guests.
After Cha received a bachelor’s degree from the Department of Nuclear and Quantum Engineering, he entered a medical graduate school and became a radiation oncology specialist. But after experiencing the death of a young patient who suffered from osteosarcoma, he returned to his alma mater to become a scientist. As he believes that science and technology is the ultimate solution to the limitations of modern medicine, he started as a PhD student at the Department of Bio and Brain Engineering in 2018, hoping to find such solutions.
During his course, he identified the characteristics of the decision-making process of doctors during diagnosis, and developed a brain-inspired AI algorithm. It is an original and challenging study that attempted to develop a fundamental machine learning theory from the data he collected from 200 doctors of different specialties.
Cha said, “Humans and AI can cooperate by humans utilizing the unique learning abilities of AI to develop our expertise, while AIs can mimic us humans’ learning abilities to improve.” He added, “My ultimate goal is to develop technology to a level at which humans and machines influence each other and ‘coevolve’, and applying it not only to medicine, but in all areas.”
Cha, who is currently an assistant professor at the KAIST Biomedical Research Center, has also written Artificial Intelligence for Doctors in 2017 to help medical personnel use AI in clinical fields, and the book was selected as one of the 2018 Sejong Books in the academic category.
During his speech at this year’s commencement ceremony, he shared that “there are so many things in the world that are difficult to solve and many things to solve them with, but I believe the things that can really broaden the horizons of the world and find fundamental solutions to the problems at hand are science and technology.”
Meanwhile, singer-songwriter Sae Byul Park who studied at the KAIST Graduate School of Culture Technology will also receive her PhD degree.
Natural language processing (NLP) is a field in AI that teaches a computer to understand and analyze human language that is actively being studied. An example of NLP is ChatGTP, which recently received a lot of attention. For her research, Park analyzed music rather than language using NLP technology.
To analyze music, which is in the form of sound, using the methods for NLP, it is necessary to rebuild notes and beats into a form of words or sentences as in a language. For this, Park designed an algorithm called Mel2Word and applied it to her research.
She also suggested that by converting melodies into texts for analysis, one would be able to quantitatively express music as sentences or words with meaning and context rather than as simple sounds representing a certain note.
Park said, “music has always been considered as a product of subjective emotion, but this research provides a framework that can calculate and analyze music.”
Park’s study can later be developed into a tool to measure the similarities between musical work, as well as a piece’s originality, artistry and popularity, and it can be used as a clue to explore the fundamental principles of how humans respond to music from a cognitive science perspective.
Park began her Ph.D. program in 2014, while carrying on with her musical activities as well as public and university lectures alongside, and dealing with personally major events including marriage and childbirth during the course of years. She already met the requirements to receive her degree in 2019, but delayed her graduation in order to improve the level of completion of her research, and finally graduated with her current achievements after nine years.
Professor Juhan Nam, who supervised Park’s research, said, “Park, who has a bachelor’s degree in psychology, later learned to code for graduate school, and has complete high-quality research in the field of artificial intelligence.” He added, “Though it took a long time, her attitude of not giving up until the end as a researcher is also excellent.”
Sae Byul Park is currently lecturing courses entitled Culture Technology and Music Information Retrieval at the Underwood International College of Yonsei University.
Park said, “the 10 or so years I’ve spent at KAIST as a graduate student was a time I could learn and prosper not only academically but from all angles of life.” She added, “having received a doctorate degree is not the end, but a ‘commencement’. Therefore, I will start to root deeper from the seeds I sowed and work harder as a both a scholar and an artist.”
< Photo 2. From left) Yujin Cha (Valedictorian, Medical-Scientist Program Ph.D. graduate), Saebyeol Park (a singer-songwriter, Ph.D. graduate from the Graduate School of Culture and Technology), Junseok Moon and Inah Seo (the two highlighted CEO graduates from the Department of Management Engineering's master’s program) >
Young entrepreneurs who dream of solving social problems will also be wearing their graduation caps. Two such graduates are Jun-seok Moon and Inah Seo, receiving their master’s degrees in social entrepreneurship MBA from the KAIST College of Business.
Before entrance, Moon ran a café helping African refugees stand on their own feet. Then, he entered KAIST to later expand his business and learn social entrepreneurship in order to sustainably help refugees in the blind spots of human rights and welfare.
During his master’s course, Moon realized that he could achieve active carbon reduction by changing the coffee alone, and switched his business field and founded Equal Table. The amount of carbon an individual can reduce by refraining from using a single paper cup is 10g, while changing the coffee itself can reduce it by 300g.
1kg of coffee emits 15kg of carbon over the course of its production, distribution, processing, and consumption, but Moon produces nearly carbon-neutral coffee beans by having innovated the entire process. In particular, the company-to-company ESG business solution is Moon’s new start-up area. It provides companies with carbon-reduced coffee made by roasting raw beans from carbon-neutral certified farms with 100% renewable energy, and shows how much carbon has been reduced in its making. Equal Table will launch the service this month in collaboration with SK Telecom, its first partner.
Inah Seo, who also graduated with Moon, founded Conscious Wear to start a fashion business reducing environmental pollution. In order to realize her mission, she felt the need to gain the appropriate expertise in management, and enrolled for the social entrepreneurship MBA.
Out of the various fashion industries, Seo focused on the leather market, which is worth 80 trillion won. Due to thickness or contamination issues, only about 60% of animal skin fabric is used, and the rest is discarded. Heavy metals are used during such processes, which also directly affects the environment.
During the social entrepreneurship MBA course, Seo collaborated with SK Chemicals, which had links through the program, and launched eco-friendly leather bags. The bags used discarded leather that was recycled by grinding and reprocessing into a biomaterial called PO3G. It was the first case in which PO3G that is over 90% biodegradable was applied to regenerated leather. In other words, it can reduce environmental pollution in the processing and disposal stages, while also reducing carbon emissions and water usage by one-tenth compared to existing cowhide products.
The social entrepreneurship MBA course, from which Moon and Seo graduated, will run in integration with the Graduate School of Green Growth as an Impact MBA program starting this year. KAIST plans to steadily foster entrepreneurs who will lead meaningful changes in the environment and society as well as economic values through innovative technologies and ideas.
< Photo 3. NYU President Emeritus John Sexton (left), who received this year's honorary doctorate of science, poses with President Kwang Hyung Lee >
Meanwhile, during this day’s commencement ceremony, KAIST also presented President Emeritus John Sexton of New York University with an honorary doctorate in science. He was recognized for laying the foundation for the cooperation between KAIST and New York University, such as promoting joint campuses.
< Photo 4. At the commencement ceremony of KAIST held on the 17th, President Kwang Hyung Lee is encouraging the graduates with his commencement address. >
President Kwang Hyung Lee emphasized in his commencement speech that, “if you can draw up the future and work hard toward your goal, the future can become a work of art that you create with your own hands,” and added, “Never stop on the journey toward your dreams, and do not give up even when you are met with failure. Failure happens to everyone, all the time. The important thing is to know 'why you failed', and to use those elements of failure as the driving force for the next try.”
2023.02.20 View 24564 -
 KAIST’s unmanned racing car to race in the Indy Autonomous Challenge @ CES 2023 as the only contender representing Asia
- Professor David Hyunchul Shim of the School of Electrical Engineering, is at the Las Vegas Motor Speedway in Las Vegas, Nevada with his students of the Unmanned Systems Research Group (USRG), participating in the Indy Autonomous Challenge (IAC) @ CES as the only Asian team in the race.
Photo 1. Nine teams that competed at the first Indy Autonomous Challenge on October 23, 2021. (KAIST team is the right most team in the front row)
- The EE USRG team won the slot to race in the IAC @ CES 2023 rightly as the semifinals entree of the IAC @ CES 2022’ held in January of last year
- Through the partnership with Hyundai Motor Company, USRG received support to participate in the competition, and is to share the latest developments and trends of the technology with the company researchers
- With upgrades from last year, USRG is to race with a high-speed Indy racing car capable of driving up to 300 km/h and the technology developed in the process is to be used in further advancement of the high-speed autonomous vehicle technology of the future.
KAIST (President Kwang Hyung Lee) announced on the 5th that it will participate in the “Indy Autonomous Challenge (IAC) @ CES 2023”, an official event of the world's largest electronics and information technology exhibition held every year in Las Vegas, Nevada, of the United States from January 5th to 8th.
Photo 2. KAIST Racing Team participating in the Indy Autonomous Challenge @ CES 2023 (Team Leader: Sungwon Na, Team Members: Seongwoo Moon, Hyunwoo Nam, Chanhoe Ryu, Jaeyoung Kang)
“IAC @ CES 2023”, which is to be held at the Las Vegas Motor Speedway (LVMS) on January 7, seeks to advance technology developed as the result of last year's competition to share the results of such advanced high-speed autonomous vehicle technology with the public.
This competition is the 4th competition following the “Indy Autonomous Challenge (IAC)” held for the first time in Indianapolis, USA on October 23, 2021. At the IAC @ CES 2022 following the first IAC competition, the Unmmaned Systems Research Group (USRG) team led by Professor David Hyunchul Shim advanced to the semifinals out of a total of nine teams and won a spot to participate in CES 2023. As a result, the USRG comes into the challenge as the only Asian team to compete with other teams comprised of students and researchers of American and European backgrounds where the culture of motorsports is more deep-rooted.
For CES 2022, Professor David Hyunchul Shim’s research team was able to successfully develop a software that controlled the racing car to comply with the race flags and regulations while going up to 240 km/h all on its on.
Photo 3. KAIST Team’s vehicle on Las Vegas Motor Speedway during the IAC @ CES 2022
In the IAC @ CES 2023, the official racing vehicle AV-23, is a converted version of IL-15, the official racing car for Indy 500, fully automated while maintaining the optimal design for high-speed racing, and was upgraded from the last year’s competition taking up the highest speed up to 300 km/h.
This year’s competition, will develop on last year’s head-to-head autonomous racing and take the form of the single elimination tournament to have the cars overtake the others without any restrictions on the driving course, which would have the team that constantly drives at the fastest speed will win the competition.
Photo 4. KAIST Team’s vehicle overtaking the Italian team, PoliMOVE’s vehicle during one of the race in the IAC @ CES 2022
Professor Shim's team further developed on the CES 2022 certified software to fine tune the external recognition mechanisms and is now focused on precise positioning and driving control technology that factors into maintaining stability even when driving at high speed.
Professor Shim's research team won the Autonomous Driving Competition hosted by Hyundai Motor Company in 2021. Starting with this CES 2023 competition, they signed a partnership contract with Hyundai to receive financial support to participate in the CES competition and share the latest developments and trends of autonomous driving technology with Hyundai Motor's research team.
During CES 2023, the research team will also participate in other events such as the exhibition by the KAIST racing team at the IAC’s official booth located in the West Hall.
Professor David Hyunchul Shim said, “With these competitions being held overseas, there were many difficulties having to keep coming back, but the students took part in it diligently, for which I am deeply grateful. Thanks to their efforts, we were able to continue in this competition, which will be a way to verify the autonomous driving technology that we developed ourselves over the past 13 years, and I highly appreciate that.”
“While high-speed autonomous driving technology is a technology that is not yet sought out in Korea, but it can be applied most effectively for long-distance travel in the Korea,” he went on to add. “It has huge advantages in that it does not require constructions for massive infrastructure that costs enormous amount of money such as high-speed rail or urban aviation and with our design, it is minimally affected by weather conditions.” he emphasized.
On a different note, the IAC @ CES 2023 is co-hosted by the Consumer Technology Association (CTA) and Energy Systems Network (ESN), the organizers of CES. Last year’s IAC winner, Technische Universität München of Germany, and MIT-PITT-RW, a team of Massachusetts Institute of Technology (Massachusetts), University of Pittsburgh (Pennsylvania), Rochester Institute of Technology (New York), University of Waterloo (Canada), with and the University of Waterloo, along with TII EuroRacing - University of Modena and Reggio Emilia (Italy), Technology Innovation Institute (United Arab Emirates), and five other teams are in the race for the win against KAIST.
Photo 5. KAIST Team’s vehicle on the track during the IAC @ CES 2022
The Indy Autonomous Challenge is scheduled to hold its fifth competition at the Monza track in Italy in June 2023 and the sixth competition at CES 2024.
2023.01.05 View 12530
KAIST’s unmanned racing car to race in the Indy Autonomous Challenge @ CES 2023 as the only contender representing Asia
- Professor David Hyunchul Shim of the School of Electrical Engineering, is at the Las Vegas Motor Speedway in Las Vegas, Nevada with his students of the Unmanned Systems Research Group (USRG), participating in the Indy Autonomous Challenge (IAC) @ CES as the only Asian team in the race.
Photo 1. Nine teams that competed at the first Indy Autonomous Challenge on October 23, 2021. (KAIST team is the right most team in the front row)
- The EE USRG team won the slot to race in the IAC @ CES 2023 rightly as the semifinals entree of the IAC @ CES 2022’ held in January of last year
- Through the partnership with Hyundai Motor Company, USRG received support to participate in the competition, and is to share the latest developments and trends of the technology with the company researchers
- With upgrades from last year, USRG is to race with a high-speed Indy racing car capable of driving up to 300 km/h and the technology developed in the process is to be used in further advancement of the high-speed autonomous vehicle technology of the future.
KAIST (President Kwang Hyung Lee) announced on the 5th that it will participate in the “Indy Autonomous Challenge (IAC) @ CES 2023”, an official event of the world's largest electronics and information technology exhibition held every year in Las Vegas, Nevada, of the United States from January 5th to 8th.
Photo 2. KAIST Racing Team participating in the Indy Autonomous Challenge @ CES 2023 (Team Leader: Sungwon Na, Team Members: Seongwoo Moon, Hyunwoo Nam, Chanhoe Ryu, Jaeyoung Kang)
“IAC @ CES 2023”, which is to be held at the Las Vegas Motor Speedway (LVMS) on January 7, seeks to advance technology developed as the result of last year's competition to share the results of such advanced high-speed autonomous vehicle technology with the public.
This competition is the 4th competition following the “Indy Autonomous Challenge (IAC)” held for the first time in Indianapolis, USA on October 23, 2021. At the IAC @ CES 2022 following the first IAC competition, the Unmmaned Systems Research Group (USRG) team led by Professor David Hyunchul Shim advanced to the semifinals out of a total of nine teams and won a spot to participate in CES 2023. As a result, the USRG comes into the challenge as the only Asian team to compete with other teams comprised of students and researchers of American and European backgrounds where the culture of motorsports is more deep-rooted.
For CES 2022, Professor David Hyunchul Shim’s research team was able to successfully develop a software that controlled the racing car to comply with the race flags and regulations while going up to 240 km/h all on its on.
Photo 3. KAIST Team’s vehicle on Las Vegas Motor Speedway during the IAC @ CES 2022
In the IAC @ CES 2023, the official racing vehicle AV-23, is a converted version of IL-15, the official racing car for Indy 500, fully automated while maintaining the optimal design for high-speed racing, and was upgraded from the last year’s competition taking up the highest speed up to 300 km/h.
This year’s competition, will develop on last year’s head-to-head autonomous racing and take the form of the single elimination tournament to have the cars overtake the others without any restrictions on the driving course, which would have the team that constantly drives at the fastest speed will win the competition.
Photo 4. KAIST Team’s vehicle overtaking the Italian team, PoliMOVE’s vehicle during one of the race in the IAC @ CES 2022
Professor Shim's team further developed on the CES 2022 certified software to fine tune the external recognition mechanisms and is now focused on precise positioning and driving control technology that factors into maintaining stability even when driving at high speed.
Professor Shim's research team won the Autonomous Driving Competition hosted by Hyundai Motor Company in 2021. Starting with this CES 2023 competition, they signed a partnership contract with Hyundai to receive financial support to participate in the CES competition and share the latest developments and trends of autonomous driving technology with Hyundai Motor's research team.
During CES 2023, the research team will also participate in other events such as the exhibition by the KAIST racing team at the IAC’s official booth located in the West Hall.
Professor David Hyunchul Shim said, “With these competitions being held overseas, there were many difficulties having to keep coming back, but the students took part in it diligently, for which I am deeply grateful. Thanks to their efforts, we were able to continue in this competition, which will be a way to verify the autonomous driving technology that we developed ourselves over the past 13 years, and I highly appreciate that.”
“While high-speed autonomous driving technology is a technology that is not yet sought out in Korea, but it can be applied most effectively for long-distance travel in the Korea,” he went on to add. “It has huge advantages in that it does not require constructions for massive infrastructure that costs enormous amount of money such as high-speed rail or urban aviation and with our design, it is minimally affected by weather conditions.” he emphasized.
On a different note, the IAC @ CES 2023 is co-hosted by the Consumer Technology Association (CTA) and Energy Systems Network (ESN), the organizers of CES. Last year’s IAC winner, Technische Universität München of Germany, and MIT-PITT-RW, a team of Massachusetts Institute of Technology (Massachusetts), University of Pittsburgh (Pennsylvania), Rochester Institute of Technology (New York), University of Waterloo (Canada), with and the University of Waterloo, along with TII EuroRacing - University of Modena and Reggio Emilia (Italy), Technology Innovation Institute (United Arab Emirates), and five other teams are in the race for the win against KAIST.
Photo 5. KAIST Team’s vehicle on the track during the IAC @ CES 2022
The Indy Autonomous Challenge is scheduled to hold its fifth competition at the Monza track in Italy in June 2023 and the sixth competition at CES 2024.
2023.01.05 View 12530 -
 Phage resistant Escherichia coli strains developed to reduce fermentation failure
A genome engineering-based systematic strategy for developing phage resistant Escherichia coli strains has been successfully developed through the collaborative efforts of a team led by Professor Sang Yup Lee, Professor Shi Chen, and Professor Lianrong Wang. This study by Xuan Zou et al. was published in Nature Communications in August 2022 and featured in Nature Communications Editors’ Highlights. The collaboration by the School of Pharmaceutical Sciences at Wuhan University, the First Affiliated Hospital of Shenzhen University, and the KAIST Department of Chemical and Biomolecular Engineering has made an important advance in the metabolic engineering and fermentation industry as it solves a big problem of phage infection causing fermentation failure.
Systems metabolic engineering is a highly interdisciplinary field that has made the development of microbial cell factories to produce various bioproducts including chemicals, fuels, and materials possible in a sustainable and environmentally friendly way, mitigating the impact of worldwide resource depletion and climate change. Escherichia coli is one of the most important chassis microbial strains, given its wide applications in the bio-based production of a diverse range of chemicals and materials. With the development of tools and strategies for systems metabolic engineering using E. coli, a highly optimized and well-characterized cell factory will play a crucial role in converting cheap and readily available raw materials into products of great economic and industrial value.
However, the consistent problem of phage contamination in fermentation imposes a devastating impact on host cells and threatens the productivity of bacterial bioprocesses in biotechnology facilities, which can lead to widespread fermentation failure and immeasurable economic loss. Host-controlled defense systems can be developed into effective genetic engineering solutions to address bacteriophage contamination in industrial-scale fermentation; however, most of the resistance mechanisms only narrowly restrict phages and their effect on phage contamination will be limited.
There have been attempts to develop diverse abilities/systems for environmental adaptation or antiviral defense. The team’s collaborative efforts developed a new type II single-stranded DNA phosphorothioation (Ssp) defense system derived from E. coli 3234/A, which can be used in multiple industrial E. coli strains (e.g., E. coli K-12, B and W) to provide broad protection against various types of dsDNA coliphages. Furthermore, they developed a systematic genome engineering strategy involving the simultaneous genomic integration of the Ssp defense module and mutations in components that are essential to the phage life cycle. This strategy can be used to transform E. coli hosts that are highly susceptible to phage attack into strains with powerful restriction effects on the tested bacteriophages. This endows hosts with strong resistance against a wide spectrum of phage infections without affecting bacterial growth and normal physiological function. More importantly, the resulting engineered phage-resistant strains maintained the capabilities of producing the desired chemicals and recombinant proteins even under high levels of phage cocktail challenge, which provides crucial protection against phage attacks.
This is a major step forward, as it provides a systematic solution for engineering phage-resistant bacterial strains, especially industrial bioproduction strains, to protect cells from a wide range of bacteriophages. Considering the functionality of this engineering strategy with diverse E. coli strains, the strategy reported in this study can be widely extended to other bacterial species and industrial applications, which will be of great interest to researchers in academia and industry alike.
Fig. A schematic model of the systematic strategy for engineering phage-sensitive industrial E. coli strains into strains with broad antiphage activities. Through the simultaneous genomic integration of a DNA phosphorothioation-based Ssp defense module and mutations of components essential for the phage life cycle, the engineered E. coli strains show strong resistance against diverse phages tested and maintain the capabilities of producing example recombinant proteins, even under high levels of phage cocktail challenge.
2022.08.23 View 17144
Phage resistant Escherichia coli strains developed to reduce fermentation failure
A genome engineering-based systematic strategy for developing phage resistant Escherichia coli strains has been successfully developed through the collaborative efforts of a team led by Professor Sang Yup Lee, Professor Shi Chen, and Professor Lianrong Wang. This study by Xuan Zou et al. was published in Nature Communications in August 2022 and featured in Nature Communications Editors’ Highlights. The collaboration by the School of Pharmaceutical Sciences at Wuhan University, the First Affiliated Hospital of Shenzhen University, and the KAIST Department of Chemical and Biomolecular Engineering has made an important advance in the metabolic engineering and fermentation industry as it solves a big problem of phage infection causing fermentation failure.
Systems metabolic engineering is a highly interdisciplinary field that has made the development of microbial cell factories to produce various bioproducts including chemicals, fuels, and materials possible in a sustainable and environmentally friendly way, mitigating the impact of worldwide resource depletion and climate change. Escherichia coli is one of the most important chassis microbial strains, given its wide applications in the bio-based production of a diverse range of chemicals and materials. With the development of tools and strategies for systems metabolic engineering using E. coli, a highly optimized and well-characterized cell factory will play a crucial role in converting cheap and readily available raw materials into products of great economic and industrial value.
However, the consistent problem of phage contamination in fermentation imposes a devastating impact on host cells and threatens the productivity of bacterial bioprocesses in biotechnology facilities, which can lead to widespread fermentation failure and immeasurable economic loss. Host-controlled defense systems can be developed into effective genetic engineering solutions to address bacteriophage contamination in industrial-scale fermentation; however, most of the resistance mechanisms only narrowly restrict phages and their effect on phage contamination will be limited.
There have been attempts to develop diverse abilities/systems for environmental adaptation or antiviral defense. The team’s collaborative efforts developed a new type II single-stranded DNA phosphorothioation (Ssp) defense system derived from E. coli 3234/A, which can be used in multiple industrial E. coli strains (e.g., E. coli K-12, B and W) to provide broad protection against various types of dsDNA coliphages. Furthermore, they developed a systematic genome engineering strategy involving the simultaneous genomic integration of the Ssp defense module and mutations in components that are essential to the phage life cycle. This strategy can be used to transform E. coli hosts that are highly susceptible to phage attack into strains with powerful restriction effects on the tested bacteriophages. This endows hosts with strong resistance against a wide spectrum of phage infections without affecting bacterial growth and normal physiological function. More importantly, the resulting engineered phage-resistant strains maintained the capabilities of producing the desired chemicals and recombinant proteins even under high levels of phage cocktail challenge, which provides crucial protection against phage attacks.
This is a major step forward, as it provides a systematic solution for engineering phage-resistant bacterial strains, especially industrial bioproduction strains, to protect cells from a wide range of bacteriophages. Considering the functionality of this engineering strategy with diverse E. coli strains, the strategy reported in this study can be widely extended to other bacterial species and industrial applications, which will be of great interest to researchers in academia and industry alike.
Fig. A schematic model of the systematic strategy for engineering phage-sensitive industrial E. coli strains into strains with broad antiphage activities. Through the simultaneous genomic integration of a DNA phosphorothioation-based Ssp defense module and mutations of components essential for the phage life cycle, the engineered E. coli strains show strong resistance against diverse phages tested and maintain the capabilities of producing example recombinant proteins, even under high levels of phage cocktail challenge.
2022.08.23 View 17144 -
 Interactive Map of Metabolical Synthesis of Chemicals
An interactive map that compiled the chemicals produced by biological, chemical and combined reactions has been distributed on the web
- A team led by Distinguished Professor Sang Yup Lee of the Department of Chemical and Biomolecular Engineering, organized and distributed an all-inclusive listing of chemical substances that can be synthesized using microorganisms
- It is expected to be used by researchers around the world as it enables easy assessment of the synthetic pathway through the web.
A research team comprised of Woo Dae Jang, Gi Bae Kim, and Distinguished Professor Sang Yup Lee of the Department of Chemical and Biomolecular Engineering at KAIST reported an interactive metabolic map of bio-based chemicals. Their research paper “An interactive metabolic map of bio-based chemicals” was published online in Trends in Biotechnology on August 10, 2022.
As a response to rapid climate change and environmental pollution, research on the production of petrochemical products using microorganisms is receiving attention as a sustainable alternative to existing methods of productions. In order to synthesize various chemical substances, materials, and fuel using microorganisms, it is necessary to first construct the biosynthetic pathway toward desired product by exploration and discovery and introduce them into microorganisms. In addition, in order to efficiently synthesize various chemical substances, it is sometimes necessary to employ chemical methods along with bioengineering methods using microorganisms at the same time. For the production of non-native chemicals, novel pathways are designed by recruiting enzymes from heterologous sources or employing enzymes designed though rational engineering, directed evolution, or ab initio design.
The research team had completed a map of chemicals which compiled all available pathways of biological and/or chemical reactions that lead to the production of various bio-based chemicals back in 2019 and published the map in Nature Catalysis. The map was distributed in the form of a poster to industries and academia so that the synthesis paths of bio-based chemicals could be checked at a glance.
The research team has expanded the bio-based chemicals map this time in the form of an interactive map on the web so that anyone with internet access can quickly explore efficient paths to synthesize desired products. The web-based map provides interactive visual tools to allow interactive visualization, exploration, and analysis of complex networks of biological and/or chemical reactions toward the desired products. In addition, the reported paper also discusses the production of natural compounds that are used for diverse purposes such as food and medicine, which will help designing novel pathways through similar approaches or by exploiting the promiscuity of enzymes described in the map. The published bio-based chemicals map is also available at http://systemsbiotech.co.kr.
The co-first authors, Dr. Woo Dae Jang and Ph.D. student Gi Bae Kim, said, “We conducted this study to address the demand for updating the previously distributed chemicals map and enhancing its versatility.” “The map is expected to be utilized in a variety of research and in efforts to set strategies and prospects for chemical production incorporating bio and chemical methods that are detailed in the map.”
Distinguished Professor Sang Yup Lee said, “The interactive bio-based chemicals map is expected to help design and optimization of the metabolic pathways for the biosynthesis of target chemicals together with the strategies of chemical conversions, serving as a blueprint for developing further ideas on the production of desired chemicals through biological and/or chemical reactions.”
The interactive metabolic map of bio-based chemicals.
2022.08.11 View 18604
Interactive Map of Metabolical Synthesis of Chemicals
An interactive map that compiled the chemicals produced by biological, chemical and combined reactions has been distributed on the web
- A team led by Distinguished Professor Sang Yup Lee of the Department of Chemical and Biomolecular Engineering, organized and distributed an all-inclusive listing of chemical substances that can be synthesized using microorganisms
- It is expected to be used by researchers around the world as it enables easy assessment of the synthetic pathway through the web.
A research team comprised of Woo Dae Jang, Gi Bae Kim, and Distinguished Professor Sang Yup Lee of the Department of Chemical and Biomolecular Engineering at KAIST reported an interactive metabolic map of bio-based chemicals. Their research paper “An interactive metabolic map of bio-based chemicals” was published online in Trends in Biotechnology on August 10, 2022.
As a response to rapid climate change and environmental pollution, research on the production of petrochemical products using microorganisms is receiving attention as a sustainable alternative to existing methods of productions. In order to synthesize various chemical substances, materials, and fuel using microorganisms, it is necessary to first construct the biosynthetic pathway toward desired product by exploration and discovery and introduce them into microorganisms. In addition, in order to efficiently synthesize various chemical substances, it is sometimes necessary to employ chemical methods along with bioengineering methods using microorganisms at the same time. For the production of non-native chemicals, novel pathways are designed by recruiting enzymes from heterologous sources or employing enzymes designed though rational engineering, directed evolution, or ab initio design.
The research team had completed a map of chemicals which compiled all available pathways of biological and/or chemical reactions that lead to the production of various bio-based chemicals back in 2019 and published the map in Nature Catalysis. The map was distributed in the form of a poster to industries and academia so that the synthesis paths of bio-based chemicals could be checked at a glance.
The research team has expanded the bio-based chemicals map this time in the form of an interactive map on the web so that anyone with internet access can quickly explore efficient paths to synthesize desired products. The web-based map provides interactive visual tools to allow interactive visualization, exploration, and analysis of complex networks of biological and/or chemical reactions toward the desired products. In addition, the reported paper also discusses the production of natural compounds that are used for diverse purposes such as food and medicine, which will help designing novel pathways through similar approaches or by exploiting the promiscuity of enzymes described in the map. The published bio-based chemicals map is also available at http://systemsbiotech.co.kr.
The co-first authors, Dr. Woo Dae Jang and Ph.D. student Gi Bae Kim, said, “We conducted this study to address the demand for updating the previously distributed chemicals map and enhancing its versatility.” “The map is expected to be utilized in a variety of research and in efforts to set strategies and prospects for chemical production incorporating bio and chemical methods that are detailed in the map.”
Distinguished Professor Sang Yup Lee said, “The interactive bio-based chemicals map is expected to help design and optimization of the metabolic pathways for the biosynthesis of target chemicals together with the strategies of chemical conversions, serving as a blueprint for developing further ideas on the production of desired chemicals through biological and/or chemical reactions.”
The interactive metabolic map of bio-based chemicals.
2022.08.11 View 18604 -
 Metabolically Engineered Bacterium Produces Lutein
A research group at KAIST has engineered a bacterial strain capable of producing lutein. The research team applied systems metabolic engineering strategies, including substrate channeling and electron channeling, to enhance the production of lutein in an engineered Escherichia coli strain. The strategies will be also useful for the efficient production of other industrially important natural products used in the food, pharmaceutical, and cosmetic industries.
Figure: Systems metabolic engineering was employed to construct and optimize the metabolic pathways for lutein production, and substrate channeling and electron channeling strategies were additionally employed to increase the production of the lutein with high productivity.
Lutein is classified as a xanthophyll chemical that is abundant in egg yolk, fruits, and vegetables. It protects the eye from oxidative damage from radiation and reduces the risk of eye diseases including macular degeneration and cataracts. Commercialized products featuring lutein are derived from the extracts of the marigold flower, which is known to harbor abundant amounts of lutein. However, the drawback of lutein production from nature is that it takes a long time to grow and harvest marigold flowers. Furthermore, it requires additional physical and chemical-based extractions with a low yield, which makes it economically unfeasible in terms of productivity. The high cost and low yield of these bioprocesses has made it difficult to readily meet the demand for lutein.
These challenges inspired the metabolic engineers at KAIST, including researchers Dr. Seon Young Park, Ph.D. Candidate Hyunmin Eun, and Distinguished Professor Sang Yup Lee from the Department of Chemical and Biomolecular Engineering. The team’s study entitled “Metabolic engineering of Escherichia coli with electron channeling for the production of natural products” was published in Nature Catalysis on August 5, 2022.
This research details the ability to produce lutein from E. coli with a high yield using a cheap carbon source, glycerol, via systems metabolic engineering. The research group focused on solving the bottlenecks of the biosynthetic pathway for lutein production constructed within an individual cell. First, using systems metabolic engineering, which is an integrated technology to engineer the metabolism of a microorganism, lutein was produced when the lutein biosynthesis pathway was introduced, albeit in very small amounts.
To improve the productivity of lutein production, the bottleneck enzymes within the metabolic pathway were first identified. It turned out that metabolic reactions that involve a promiscuous enzyme, an enzyme that is involved in two or more metabolic reactions, and electron-requiring cytochrome P450 enzymes are the main bottleneck steps of the pathway inhibiting lutein biosynthesis.
To overcome these challenges, substrate channeling, a strategy to artificially recruit enzymes in physical proximity within the cell in order to increase the local concentrations of substrates that can be converted into products, was employed to channel more metabolic flux towards the target chemical while reducing the formation of unwanted byproducts.
Furthermore, electron channeling, a strategy similar to substrate channeling but differing in terms of increasing the local concentrations of electrons required for oxidoreduction reactions mediated by P450 and its reductase partners, was applied to further streamline the metabolic flux towards lutein biosynthesis, which led to the highest titer of lutein production achieved in a bacterial host ever reported. The same electron channeling strategy was successfully applied for the production of other natural products including nootkatone and apigenin in E. coli, showcasing the general applicability of the strategy in the research field.
“It is expected that this microbial cell factory-based production of lutein will be able to replace the current plant extraction-based process,” said Dr. Seon Young Park, the first author of the paper. She explained that another important point of the research is that integrated metabolic engineering strategies developed from this study can be generally applicable for the efficient production of other natural products useful as pharmaceuticals or nutraceuticals.
“As maintaining good health in an aging society is becoming increasingly important, we expect that the technology and strategies developed here will play pivotal roles in producing other valuable natural products of medical or nutritional importance,” explained Distinguished Professor Sang Yup Lee.
This work was supported by the Cooperative Research Program for Agriculture Science & Technology Development funded by the Rural Development Administration of Korea, with further support from the Development of Next-generation Biorefinery Platform Technologies for Leading Bio-based Chemicals Industry Project and by the Development of Platform Technologies of Microbial Cell Factories for the Next-generation Biorefineries Project of the National Research Foundation funded by the Ministry of Science and ICT of Korea.
2022.08.05 View 12370
Metabolically Engineered Bacterium Produces Lutein
A research group at KAIST has engineered a bacterial strain capable of producing lutein. The research team applied systems metabolic engineering strategies, including substrate channeling and electron channeling, to enhance the production of lutein in an engineered Escherichia coli strain. The strategies will be also useful for the efficient production of other industrially important natural products used in the food, pharmaceutical, and cosmetic industries.
Figure: Systems metabolic engineering was employed to construct and optimize the metabolic pathways for lutein production, and substrate channeling and electron channeling strategies were additionally employed to increase the production of the lutein with high productivity.
Lutein is classified as a xanthophyll chemical that is abundant in egg yolk, fruits, and vegetables. It protects the eye from oxidative damage from radiation and reduces the risk of eye diseases including macular degeneration and cataracts. Commercialized products featuring lutein are derived from the extracts of the marigold flower, which is known to harbor abundant amounts of lutein. However, the drawback of lutein production from nature is that it takes a long time to grow and harvest marigold flowers. Furthermore, it requires additional physical and chemical-based extractions with a low yield, which makes it economically unfeasible in terms of productivity. The high cost and low yield of these bioprocesses has made it difficult to readily meet the demand for lutein.
These challenges inspired the metabolic engineers at KAIST, including researchers Dr. Seon Young Park, Ph.D. Candidate Hyunmin Eun, and Distinguished Professor Sang Yup Lee from the Department of Chemical and Biomolecular Engineering. The team’s study entitled “Metabolic engineering of Escherichia coli with electron channeling for the production of natural products” was published in Nature Catalysis on August 5, 2022.
This research details the ability to produce lutein from E. coli with a high yield using a cheap carbon source, glycerol, via systems metabolic engineering. The research group focused on solving the bottlenecks of the biosynthetic pathway for lutein production constructed within an individual cell. First, using systems metabolic engineering, which is an integrated technology to engineer the metabolism of a microorganism, lutein was produced when the lutein biosynthesis pathway was introduced, albeit in very small amounts.
To improve the productivity of lutein production, the bottleneck enzymes within the metabolic pathway were first identified. It turned out that metabolic reactions that involve a promiscuous enzyme, an enzyme that is involved in two or more metabolic reactions, and electron-requiring cytochrome P450 enzymes are the main bottleneck steps of the pathway inhibiting lutein biosynthesis.
To overcome these challenges, substrate channeling, a strategy to artificially recruit enzymes in physical proximity within the cell in order to increase the local concentrations of substrates that can be converted into products, was employed to channel more metabolic flux towards the target chemical while reducing the formation of unwanted byproducts.
Furthermore, electron channeling, a strategy similar to substrate channeling but differing in terms of increasing the local concentrations of electrons required for oxidoreduction reactions mediated by P450 and its reductase partners, was applied to further streamline the metabolic flux towards lutein biosynthesis, which led to the highest titer of lutein production achieved in a bacterial host ever reported. The same electron channeling strategy was successfully applied for the production of other natural products including nootkatone and apigenin in E. coli, showcasing the general applicability of the strategy in the research field.
“It is expected that this microbial cell factory-based production of lutein will be able to replace the current plant extraction-based process,” said Dr. Seon Young Park, the first author of the paper. She explained that another important point of the research is that integrated metabolic engineering strategies developed from this study can be generally applicable for the efficient production of other natural products useful as pharmaceuticals or nutraceuticals.
“As maintaining good health in an aging society is becoming increasingly important, we expect that the technology and strategies developed here will play pivotal roles in producing other valuable natural products of medical or nutritional importance,” explained Distinguished Professor Sang Yup Lee.
This work was supported by the Cooperative Research Program for Agriculture Science & Technology Development funded by the Rural Development Administration of Korea, with further support from the Development of Next-generation Biorefinery Platform Technologies for Leading Bio-based Chemicals Industry Project and by the Development of Platform Technologies of Microbial Cell Factories for the Next-generation Biorefineries Project of the National Research Foundation funded by the Ministry of Science and ICT of Korea.
2022.08.05 View 12370 -
 Professor Jae-Woong Jeong Receives Hyonwoo KAIST Academic Award
Professor Jae-Woong Jeong from the School of Electrical Engineering was selected for the Hyonwoo KAIST Academic Award, funded by the HyonWoo Cultural Foundation (Chairman Soo-il Kwak, honorary professor at Seoul National University Business School).
The Hyonwoo KAIST Academic Award, presented for the first time in 2021, is an award newly founded by the donations of Chairman Soo-il Kwak of the HyonWoo Cultural Foundation, who aims to reward excellent KAIST scholars who have made outstanding academic achievements.
Every year, through the strict evaluations of the selection committee of the HyonWoo Cultural Foundation and the faculty reward recommendation board, KAIST will choose one faculty member that may represent the school with their excellent academic achievement, and reward them with a plaque and 100 million won.
Professor Jae-Woong Jeong, the winner of this year’s award, developed the first IoT-based wireless remote brain neural network control system to overcome brain diseases, and has been leading the field. The research was published in 2021 in Nature Biomedical Engineering, one of world’s best scientific journals, and has been recognized as a novel technology that suggested a new vision for the automation of brain research and disease treatment. This study, led by Professor Jeong’s research team, was part of the KAIST College of Engineering Global Initiative Interdisciplinary Research Project, and was jointly studied by Washington University School of Medicine through an international research collaboration. The technology was introduced more than 60 times through both domestic and international media, including Medical Xpress, MBC News, and Maeil Business News.
Professor Jeong has also developed a wirelessly chargeable soft machine for brain transplants, and the results were published in Nature Communications. He thereby opened a new paradigm for implantable semi-permanent devices for transplants, and is making unprecedented research achievements.
2022.06.13 View 10529
Professor Jae-Woong Jeong Receives Hyonwoo KAIST Academic Award
Professor Jae-Woong Jeong from the School of Electrical Engineering was selected for the Hyonwoo KAIST Academic Award, funded by the HyonWoo Cultural Foundation (Chairman Soo-il Kwak, honorary professor at Seoul National University Business School).
The Hyonwoo KAIST Academic Award, presented for the first time in 2021, is an award newly founded by the donations of Chairman Soo-il Kwak of the HyonWoo Cultural Foundation, who aims to reward excellent KAIST scholars who have made outstanding academic achievements.
Every year, through the strict evaluations of the selection committee of the HyonWoo Cultural Foundation and the faculty reward recommendation board, KAIST will choose one faculty member that may represent the school with their excellent academic achievement, and reward them with a plaque and 100 million won.
Professor Jae-Woong Jeong, the winner of this year’s award, developed the first IoT-based wireless remote brain neural network control system to overcome brain diseases, and has been leading the field. The research was published in 2021 in Nature Biomedical Engineering, one of world’s best scientific journals, and has been recognized as a novel technology that suggested a new vision for the automation of brain research and disease treatment. This study, led by Professor Jeong’s research team, was part of the KAIST College of Engineering Global Initiative Interdisciplinary Research Project, and was jointly studied by Washington University School of Medicine through an international research collaboration. The technology was introduced more than 60 times through both domestic and international media, including Medical Xpress, MBC News, and Maeil Business News.
Professor Jeong has also developed a wirelessly chargeable soft machine for brain transplants, and the results were published in Nature Communications. He thereby opened a new paradigm for implantable semi-permanent devices for transplants, and is making unprecedented research achievements.
2022.06.13 View 10529 -
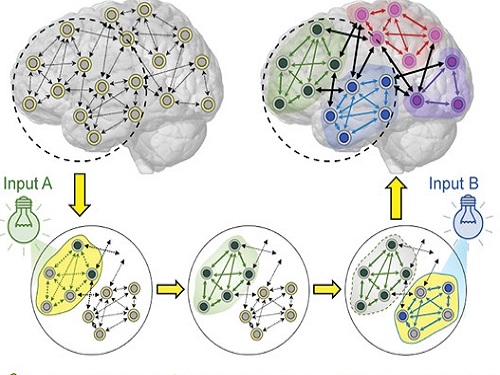 Energy-Efficient AI Hardware Technology Via a Brain-Inspired Stashing System
Researchers demonstrate neuromodulation-inspired stashing system for the energy-efficient learning of a spiking neural network using a self-rectifying memristor array
Researchers have proposed a novel system inspired by the neuromodulation of the brain, referred to as a ‘stashing system,’ that requires less energy consumption. The research group led by Professor Kyung Min Kim from the Department of Materials Science and Engineering has developed a technology that can efficiently handle mathematical operations for artificial intelligence by imitating the continuous changes in the topology of the neural network according to the situation. The human brain changes its neural topology in real time, learning to store or recall memories as needed. The research group presented a new artificial intelligence learning method that directly implements these neural coordination circuit configurations.
Research on artificial intelligence is becoming very active, and the development of artificial intelligence-based electronic devices and product releases are accelerating, especially in the Fourth Industrial Revolution age. To implement artificial intelligence in electronic devices, customized hardware development should also be supported. However most electronic devices for artificial intelligence require high power consumption and highly integrated memory arrays for large-scale tasks. It has been challenging to solve these power consumption and integration limitations, and efforts have been made to find out how the human brain solves problems.
To prove the efficiency of the developed technology, the research group created artificial neural network hardware equipped with a self-rectifying synaptic array and algorithm called a ‘stashing system’ that was developed to conduct artificial intelligence learning. As a result, it was able to reduce energy by 37% within the stashing system without any accuracy degradation. This result proves that emulating the neuromodulation in humans is possible.
Professor Kim said, "In this study, we implemented the learning method of the human brain with only a simple circuit composition and through this we were able to reduce the energy needed by nearly 40 percent.”
This neuromodulation-inspired stashing system that mimics the brain’s neural activity is compatible with existing electronic devices and commercialized semiconductor hardware. It is expected to be used in the design of next-generation semiconductor chips for artificial intelligence.
This study was published in Advanced Functional Materials in March 2022 and supported by KAIST, the National Research Foundation of Korea, the National NanoFab Center, and SK Hynix.
-Publication:
Woon Hyung Cheong, Jae Bum Jeon†, Jae Hyun In, Geunyoung Kim, Hanchan Song, Janho An, Juseong Park, Young Seok Kim, Cheol Seong Hwang, and Kyung Min Kim (2022)
“Demonstration of Neuromodulation-inspired Stashing System for Energy-efficient Learning of Spiking Neural Network using a Self-Rectifying Memristor Array,” Advanced FunctionalMaterials March 31, 2022 (DOI: 10.1002/adfm.202200337)
-Profile:
Professor Kyung Min Kimhttp://semi.kaist.ac.kr https://scholar.google.com/citations?user=BGw8yDYAAAAJ&hl=ko
Department of Materials Science and EngineeringKAIST
2022.05.18 View 13445
Energy-Efficient AI Hardware Technology Via a Brain-Inspired Stashing System
Researchers demonstrate neuromodulation-inspired stashing system for the energy-efficient learning of a spiking neural network using a self-rectifying memristor array
Researchers have proposed a novel system inspired by the neuromodulation of the brain, referred to as a ‘stashing system,’ that requires less energy consumption. The research group led by Professor Kyung Min Kim from the Department of Materials Science and Engineering has developed a technology that can efficiently handle mathematical operations for artificial intelligence by imitating the continuous changes in the topology of the neural network according to the situation. The human brain changes its neural topology in real time, learning to store or recall memories as needed. The research group presented a new artificial intelligence learning method that directly implements these neural coordination circuit configurations.
Research on artificial intelligence is becoming very active, and the development of artificial intelligence-based electronic devices and product releases are accelerating, especially in the Fourth Industrial Revolution age. To implement artificial intelligence in electronic devices, customized hardware development should also be supported. However most electronic devices for artificial intelligence require high power consumption and highly integrated memory arrays for large-scale tasks. It has been challenging to solve these power consumption and integration limitations, and efforts have been made to find out how the human brain solves problems.
To prove the efficiency of the developed technology, the research group created artificial neural network hardware equipped with a self-rectifying synaptic array and algorithm called a ‘stashing system’ that was developed to conduct artificial intelligence learning. As a result, it was able to reduce energy by 37% within the stashing system without any accuracy degradation. This result proves that emulating the neuromodulation in humans is possible.
Professor Kim said, "In this study, we implemented the learning method of the human brain with only a simple circuit composition and through this we were able to reduce the energy needed by nearly 40 percent.”
This neuromodulation-inspired stashing system that mimics the brain’s neural activity is compatible with existing electronic devices and commercialized semiconductor hardware. It is expected to be used in the design of next-generation semiconductor chips for artificial intelligence.
This study was published in Advanced Functional Materials in March 2022 and supported by KAIST, the National Research Foundation of Korea, the National NanoFab Center, and SK Hynix.
-Publication:
Woon Hyung Cheong, Jae Bum Jeon†, Jae Hyun In, Geunyoung Kim, Hanchan Song, Janho An, Juseong Park, Young Seok Kim, Cheol Seong Hwang, and Kyung Min Kim (2022)
“Demonstration of Neuromodulation-inspired Stashing System for Energy-efficient Learning of Spiking Neural Network using a Self-Rectifying Memristor Array,” Advanced FunctionalMaterials March 31, 2022 (DOI: 10.1002/adfm.202200337)
-Profile:
Professor Kyung Min Kimhttp://semi.kaist.ac.kr https://scholar.google.com/citations?user=BGw8yDYAAAAJ&hl=ko
Department of Materials Science and EngineeringKAIST
2022.05.18 View 13445