ACT
-
 Scientists re-writes FDA-recommended equation to improve estimation of drug-drug interaction
Drugs absorbed into the body are metabolized and thus removed by enzymes from several organs like the liver. How fast a drug is cleared out of the system can be affected by other drugs that are taken together because added substance can increase the amount of enzyme secretion in the body. This dramatically decreases the concentration of a drug, reducing its efficacy, often leading to the failure of having any effect at all. Therefore, accurately predicting the clearance rate in the presence of drug-drug interaction* is critical in the process of drug prescription and development of a new drug in order to ensure its efficacy and/or to avoid unwanted side-effects.
*Drug-drug interaction: In terms of metabolism, drug-drug interaction is a phenomenon in which one drug changes the metabolism of another drug to promote or inhibit its excretion from the body when two or more drugs are taken together. As a result, it increases the toxicity of medicines or causes loss of efficacy.
Since it is practically impossible to evaluate all interactions between new drug candidates and all marketed drugs during the development process, the FDA recommends indirect evaluation of drug interactions using a formula suggested in their guidance, first published in 1997, revised in January of 2020, in order to evaluate drug interactions and minimize side effects of having to use more than one type of drugs at once.
The formula relies on the 110-year-old Michaelis-Menten (MM) model, which has a fundamental limit of making a very broad and groundless assumption on the part of the presence of the enzymes that metabolizes the drug. While MM equation has been one of the most widely known equations in biochemistry used in more than 220,000 published papers, the MM equation is accurate only when the concentration of the enzyme that metabolizes the drug is almost non-existent, causing the accuracy of the equation highly unsatisfactory – only 38 percent of the predictions had less than two-fold errors.
“To make up for the gap, researcher resorted to plugging in scientifically unjustified constants into the equation,” Professor Jung-woo Chae of Chungnam National University College of Pharmacy said. “This is comparable to having to have the epicyclic orbits introduced to explain the motion of the planets back in the days in order to explain the now-defunct Ptolemaic theory, because it was 'THE' theory back then.”
< (From left) Ph.D. student Yun Min Song (KAIST, co-first authors), Professor Sang Kyum Kim (Chungnam National University, co-corresponding author), Jae Kyoung Kim, CI (KAIST, co-corresponding author), Professor Jung-woo Chae (Chungnam National University, co-corresponding author), Ph.D. students Quyen Thi Tran and Ngoc-Anh Thi Vu (Chungnam National University, co-first authors) >
A joint research team composed of mathematicians from the Biomedical Mathematics Group within the Institute for Basic Science (IBS) and the Korea Advanced Institute of Science and Technology (KAIST) and pharmacological scientists from the Chungnam National University reported that they identified the major causes of the FDA-recommended equation’s inaccuracies and presented a solution.
When estimating the gut bioavailability (Fg), which is the key parameter of the equation, the fraction absorbed from the gut lumen (Fa) is usually assumed to be 1. However, many experiments have shown that Fa is less than 1, obviously since it can’t be expected that all of the orally taken drugs to be completely absorbed by the intestines. To solve this problem, the research team used an “estimated Fa” value based on factors such as the drug’s transit time, intestine radius, and permeability values and used it to re-calculate Fg.
Also, taking a different approach from the MM equation, the team used an alternative model they derived in a previous study back in 2020, which can more accurately predict the drug metabolism rate regardless of the enzyme concentration. Combining these changes, the modified equation with re-calculated Fg had a dramatically increased accuracy of the resulting estimate. The existing FDA formula predicted drug interactions within a 2-fold margin of error at the rate of 38%, whereas the accuracy rate of the revised formula reached 80%.
“Such drastic improvement in drug-drug interaction prediction accuracy is expected to make great contribution to increasing the success rate of new drug development and drug efficacy in clinical trials. As the results of this study were published in one of the top clinical pharmacology journal, it is expected that the FDA guidance will be revised according to the results of this study.” said Professor Sang Kyum Kim from Chungnam National University College of Pharmacy.
Furthermore, this study highlights the importance of collaborative research between research groups in vastly different disciplines, in a field that is as dynamic as drug interactions.
“Thanks to the collaborative research between mathematics and pharmacy, we were able to recify the formula that we have accepted to be the right answer for so long to finally grasp on the leads toward healthier life for mankind.,” said Professor Jae Kyung Kim. He continued, “I hope seeing a ‘K-formula’ entered into the US FDA guidance one day.”
The results of this study were published in the online edition of Clinical Pharmacology and Therapeutics (IF 7.051), an authoritative journal in the field of clinical pharmacology, on December 15, 2022 (Korean time).
Thesis Title: Beyond the Michaelis-Menten: Accurate Prediction of Drug Interactions through Cytochrome P450 3A4 Induction (doi: 10.1002/cpt.2824)
< Figure 1. The formula proposed by the FDA guidance for predicting drug-drug interactions (top) and the formula newly derived by the researchers (bottom). AUCR (the ratio of substrate area under the plasma concentration-time curve) represents the rate of change in drug concentration due to drug interactions. The research team more than doubled the accuracy of drug interaction prediction compared to the existing formula. >
< Figure 2. Existing FDA formulas tend to underestimate the extent of drug-drug interactions (gray dots) than the actual measured values. On the other hand, the newly derived equation (red dot) has a prediction rate that is within the error range of 2 times (0.5 to 2 times) of the measured value, and is more than twice as high as the existing equation. The solid line in the figure represents the predicted value that matches the measured value. The dotted line represents the predicted value with an error of 0.5 to 2 times. >
For further information or to request media assistance, please contact Jae Kyoung Kim at Biomedical Mathematics Group, Institute for Basic Science (IBS) (jaekkim@ibs.re.kr) or William I. Suh at the IBS Communications Team (willisuh@ibs.re.kr).
- About the Institute for Basic Science (IBS)
IBS was founded in 2011 by the government of the Republic of Korea with the sole purpose of driving forward the development of basic science in South Korea. IBS has 4 research institutes and 33 research centers as of January 2023. There are eleven physics, three mathematics, five chemistry, nine life science, two earth science, and three interdisciplinary research centers.
2023.01.18 View 12755
Scientists re-writes FDA-recommended equation to improve estimation of drug-drug interaction
Drugs absorbed into the body are metabolized and thus removed by enzymes from several organs like the liver. How fast a drug is cleared out of the system can be affected by other drugs that are taken together because added substance can increase the amount of enzyme secretion in the body. This dramatically decreases the concentration of a drug, reducing its efficacy, often leading to the failure of having any effect at all. Therefore, accurately predicting the clearance rate in the presence of drug-drug interaction* is critical in the process of drug prescription and development of a new drug in order to ensure its efficacy and/or to avoid unwanted side-effects.
*Drug-drug interaction: In terms of metabolism, drug-drug interaction is a phenomenon in which one drug changes the metabolism of another drug to promote or inhibit its excretion from the body when two or more drugs are taken together. As a result, it increases the toxicity of medicines or causes loss of efficacy.
Since it is practically impossible to evaluate all interactions between new drug candidates and all marketed drugs during the development process, the FDA recommends indirect evaluation of drug interactions using a formula suggested in their guidance, first published in 1997, revised in January of 2020, in order to evaluate drug interactions and minimize side effects of having to use more than one type of drugs at once.
The formula relies on the 110-year-old Michaelis-Menten (MM) model, which has a fundamental limit of making a very broad and groundless assumption on the part of the presence of the enzymes that metabolizes the drug. While MM equation has been one of the most widely known equations in biochemistry used in more than 220,000 published papers, the MM equation is accurate only when the concentration of the enzyme that metabolizes the drug is almost non-existent, causing the accuracy of the equation highly unsatisfactory – only 38 percent of the predictions had less than two-fold errors.
“To make up for the gap, researcher resorted to plugging in scientifically unjustified constants into the equation,” Professor Jung-woo Chae of Chungnam National University College of Pharmacy said. “This is comparable to having to have the epicyclic orbits introduced to explain the motion of the planets back in the days in order to explain the now-defunct Ptolemaic theory, because it was 'THE' theory back then.”
< (From left) Ph.D. student Yun Min Song (KAIST, co-first authors), Professor Sang Kyum Kim (Chungnam National University, co-corresponding author), Jae Kyoung Kim, CI (KAIST, co-corresponding author), Professor Jung-woo Chae (Chungnam National University, co-corresponding author), Ph.D. students Quyen Thi Tran and Ngoc-Anh Thi Vu (Chungnam National University, co-first authors) >
A joint research team composed of mathematicians from the Biomedical Mathematics Group within the Institute for Basic Science (IBS) and the Korea Advanced Institute of Science and Technology (KAIST) and pharmacological scientists from the Chungnam National University reported that they identified the major causes of the FDA-recommended equation’s inaccuracies and presented a solution.
When estimating the gut bioavailability (Fg), which is the key parameter of the equation, the fraction absorbed from the gut lumen (Fa) is usually assumed to be 1. However, many experiments have shown that Fa is less than 1, obviously since it can’t be expected that all of the orally taken drugs to be completely absorbed by the intestines. To solve this problem, the research team used an “estimated Fa” value based on factors such as the drug’s transit time, intestine radius, and permeability values and used it to re-calculate Fg.
Also, taking a different approach from the MM equation, the team used an alternative model they derived in a previous study back in 2020, which can more accurately predict the drug metabolism rate regardless of the enzyme concentration. Combining these changes, the modified equation with re-calculated Fg had a dramatically increased accuracy of the resulting estimate. The existing FDA formula predicted drug interactions within a 2-fold margin of error at the rate of 38%, whereas the accuracy rate of the revised formula reached 80%.
“Such drastic improvement in drug-drug interaction prediction accuracy is expected to make great contribution to increasing the success rate of new drug development and drug efficacy in clinical trials. As the results of this study were published in one of the top clinical pharmacology journal, it is expected that the FDA guidance will be revised according to the results of this study.” said Professor Sang Kyum Kim from Chungnam National University College of Pharmacy.
Furthermore, this study highlights the importance of collaborative research between research groups in vastly different disciplines, in a field that is as dynamic as drug interactions.
“Thanks to the collaborative research between mathematics and pharmacy, we were able to recify the formula that we have accepted to be the right answer for so long to finally grasp on the leads toward healthier life for mankind.,” said Professor Jae Kyung Kim. He continued, “I hope seeing a ‘K-formula’ entered into the US FDA guidance one day.”
The results of this study were published in the online edition of Clinical Pharmacology and Therapeutics (IF 7.051), an authoritative journal in the field of clinical pharmacology, on December 15, 2022 (Korean time).
Thesis Title: Beyond the Michaelis-Menten: Accurate Prediction of Drug Interactions through Cytochrome P450 3A4 Induction (doi: 10.1002/cpt.2824)
< Figure 1. The formula proposed by the FDA guidance for predicting drug-drug interactions (top) and the formula newly derived by the researchers (bottom). AUCR (the ratio of substrate area under the plasma concentration-time curve) represents the rate of change in drug concentration due to drug interactions. The research team more than doubled the accuracy of drug interaction prediction compared to the existing formula. >
< Figure 2. Existing FDA formulas tend to underestimate the extent of drug-drug interactions (gray dots) than the actual measured values. On the other hand, the newly derived equation (red dot) has a prediction rate that is within the error range of 2 times (0.5 to 2 times) of the measured value, and is more than twice as high as the existing equation. The solid line in the figure represents the predicted value that matches the measured value. The dotted line represents the predicted value with an error of 0.5 to 2 times. >
For further information or to request media assistance, please contact Jae Kyoung Kim at Biomedical Mathematics Group, Institute for Basic Science (IBS) (jaekkim@ibs.re.kr) or William I. Suh at the IBS Communications Team (willisuh@ibs.re.kr).
- About the Institute for Basic Science (IBS)
IBS was founded in 2011 by the government of the Republic of Korea with the sole purpose of driving forward the development of basic science in South Korea. IBS has 4 research institutes and 33 research centers as of January 2023. There are eleven physics, three mathematics, five chemistry, nine life science, two earth science, and three interdisciplinary research centers.
2023.01.18 View 12755 -
 “3D sketch” Your Ideas and Bring Them to Life, Instantly!
Professor Seok-Hyung Bae’s research team at the Department of Industrial Design developed a novel 3D sketching system that rapidly creates animated 3D concepts through simple user interactions like sketching on a piece of paper or playing a toy.
Foldable drones, transforming vehicles, and multi-legged robots from sci-fi movies are now becoming commonplace thanks to technological progress. However, designing them remains a difficult challenge even for skilled experts, because complex design decisions must be made regarding not only their form, but also the structure, poses, and motions, which are interdependent on one another.
Creating a 3D concept comprising of multiple moving parts connected by different types of joints using a traditional 3D CAD tool, which is more suited for processing precise and elaborate modeling, is a painstaking and time-consuming process. This presents a major bottleneck for the workflow during the early stage of design, in which it is preferred that as many ideas are tried and discarded out as quickly as possible in order to explore a wide range of possibilities in the shortest amount of time.
A research team led by Professor Bae has focused on designers’ freehand sketches drew up with a pen on a paper that serve as the starting point for virtually all design projects. This led them to develop their 3D sketching technology to generate desired 3D curves from the rough but expressive 2D strokes drawn with a digital stylus on a digital tablet.
Their latest research helps designers bring their 3D sketches to life almost instantly. Using the intuitive set of multi-touch gestures the team successfully designed and implemented, designers can handle the 3D sketches they are working on with their fingers as if they are playing with toys and put them into animation in no time.
< Figure 1. A novel 3D sketching system for rapidly designing articulated 3D concepts with a small set of coherent pen and multi-touch gestures. (a) Sketching: A 3D sketch curve is created by marking a pen stroke that is projected onto a sketch plane widget. (b) Segmenting: Entire or partial sketch curves are added to separate parts that serve as links in the kinematic chain. (c) Rigging: Repeatedly demonstrating the desired motion of a part leaves behind a trail, from which the system infers a joint. (d) Posing: Desired poses can be achieved through actuating joints via forward or inverse kinematics. (e) Filming: A sequence of keyframes specifying desired poses and viewpoints is connected as a smooth motion. >
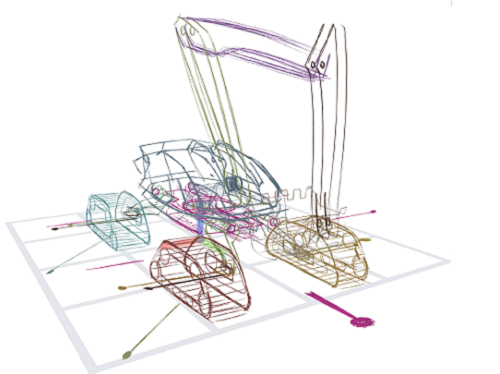
< Figure 2. (a) Concept drawing of an autonomous excavator. It features (b, c) four caterpillars that swivel for high maneuverability, (d) an extendable boom and a bucket connected by multiple links, and (e) a rotating platform. The concept’s designer, who had 8 years of work experience, estimated that it would take 1-2 weeks to express and communicate such a complex articulated object with existing tools. With the proposed system, it took only 2 hours and 52 minutes. >
The major findings of their work were published under the title “Rapid Design of Articulated Objects” in ACM Transactions on Graphics (impact factor: 7.403), the top international journal in the field of computer graphics, and presented at ACM SIGGRAPH 2022 (h5-index: 103), the world’s largest international academic conference in the field, which was held back in August in Vancouver, Canada with Joon Hyub Lee, a Ph.D. student of the Department of Industrial Design as the first author.
The ACM SIGGRAPH 2022 conference was reportedly attended by over 10,000 participants including researchers, artists, and developers from world-renowned universities; film, animation, and game studies, such as Marvel, Pixar, and Blizzard; high-tech manufacturers, such as Lockheed Martin and Boston Dynamics; and metaverse platform companies, such as Meta and Roblox.
< Figure 3. The findings of Professor Bae’s research team were published in ACM Transactions on Graphics, the top international academic journal in the field of computer graphics, and presented at ACM SIGGRAPH 2022, the largest international academic conference held in conjunction early August in Vancouver, Canada. The team’s live demo at the Emerging Technologies program was highly praised by numerous academics and industry officials and received an Honorable Mention. >
The team was also invited to present their technical paper as a demo and a special talk at the Emerging Technologies program at ACM SIGGRAPH 2022 as one of the top-three impactful technologies. The live performance, in which Hanbit Kim, a Ph.D. student of the Department of Industrial Design at KAIST and a co-author, sketched and animated a sophisticated animal-shaped robot from scratch in a matter of a few minutes, wowed the audience and won the Honorable Mention Award from the jury.
Edwin Catmull, the co-founder of Pixar and a keynote speaker at the SIGGRAPH conference, praised the team’s research on 3D sketching as “really excellent work” and “a kind of tool that would be useful to Pixar's creative model designers.”
This technology, which became virally popular in Japan after featuring in an online IT media outlet and attracting more than 600K views, received a special award from the Digital Content Association of Japan (DCAJ) and was invited and exhibited for three days at Tokyo in November, as a part of Inter BEE 2022, the largest broadcasting and media expo in Japan.
“The more we come to understand how designers think and work, the more effective design tools can be built around that understanding,” said Professor Bae, explaining that “the key is to integrate different algorithms into a harmonious system as intuitive interactions.” He added that “this work wouldn’t have been possible if it weren’t for the convergent research environment cultivated by the Department of Industrial Design at KAIST, in which all students see themselves not only as aspiring creative designers, but also as practical engineers.”
By enabling designers to produce highly expressive animated 3D concepts far more quickly and easily in comparison to using existing methods, this new tool is expected to revolutionize design practices and processes in the content creation, manufacturing, and metaverse-related industries.
This research was funded by the Ministry of Science and ICT, and the National Research Foundation of Korea.
More info: https://sketch.kaist.ac.kr/publications/2022_siggraph_rapid_design
Video: https://www.youtube.com/watch?v=rsBl0QvSDqI
< Figure 4. From left to right: Ph.D. students Hanbit Kim, and Joon Hyub Lee and Professor Bae of the Department of Industrial Design, KAIST >
2022.11.23 View 9929
“3D sketch” Your Ideas and Bring Them to Life, Instantly!
Professor Seok-Hyung Bae’s research team at the Department of Industrial Design developed a novel 3D sketching system that rapidly creates animated 3D concepts through simple user interactions like sketching on a piece of paper or playing a toy.
Foldable drones, transforming vehicles, and multi-legged robots from sci-fi movies are now becoming commonplace thanks to technological progress. However, designing them remains a difficult challenge even for skilled experts, because complex design decisions must be made regarding not only their form, but also the structure, poses, and motions, which are interdependent on one another.
Creating a 3D concept comprising of multiple moving parts connected by different types of joints using a traditional 3D CAD tool, which is more suited for processing precise and elaborate modeling, is a painstaking and time-consuming process. This presents a major bottleneck for the workflow during the early stage of design, in which it is preferred that as many ideas are tried and discarded out as quickly as possible in order to explore a wide range of possibilities in the shortest amount of time.
A research team led by Professor Bae has focused on designers’ freehand sketches drew up with a pen on a paper that serve as the starting point for virtually all design projects. This led them to develop their 3D sketching technology to generate desired 3D curves from the rough but expressive 2D strokes drawn with a digital stylus on a digital tablet.
Their latest research helps designers bring their 3D sketches to life almost instantly. Using the intuitive set of multi-touch gestures the team successfully designed and implemented, designers can handle the 3D sketches they are working on with their fingers as if they are playing with toys and put them into animation in no time.
< Figure 1. A novel 3D sketching system for rapidly designing articulated 3D concepts with a small set of coherent pen and multi-touch gestures. (a) Sketching: A 3D sketch curve is created by marking a pen stroke that is projected onto a sketch plane widget. (b) Segmenting: Entire or partial sketch curves are added to separate parts that serve as links in the kinematic chain. (c) Rigging: Repeatedly demonstrating the desired motion of a part leaves behind a trail, from which the system infers a joint. (d) Posing: Desired poses can be achieved through actuating joints via forward or inverse kinematics. (e) Filming: A sequence of keyframes specifying desired poses and viewpoints is connected as a smooth motion. >
< Figure 2. (a) Concept drawing of an autonomous excavator. It features (b, c) four caterpillars that swivel for high maneuverability, (d) an extendable boom and a bucket connected by multiple links, and (e) a rotating platform. The concept’s designer, who had 8 years of work experience, estimated that it would take 1-2 weeks to express and communicate such a complex articulated object with existing tools. With the proposed system, it took only 2 hours and 52 minutes. >
The major findings of their work were published under the title “Rapid Design of Articulated Objects” in ACM Transactions on Graphics (impact factor: 7.403), the top international journal in the field of computer graphics, and presented at ACM SIGGRAPH 2022 (h5-index: 103), the world’s largest international academic conference in the field, which was held back in August in Vancouver, Canada with Joon Hyub Lee, a Ph.D. student of the Department of Industrial Design as the first author.
The ACM SIGGRAPH 2022 conference was reportedly attended by over 10,000 participants including researchers, artists, and developers from world-renowned universities; film, animation, and game studies, such as Marvel, Pixar, and Blizzard; high-tech manufacturers, such as Lockheed Martin and Boston Dynamics; and metaverse platform companies, such as Meta and Roblox.
< Figure 3. The findings of Professor Bae’s research team were published in ACM Transactions on Graphics, the top international academic journal in the field of computer graphics, and presented at ACM SIGGRAPH 2022, the largest international academic conference held in conjunction early August in Vancouver, Canada. The team’s live demo at the Emerging Technologies program was highly praised by numerous academics and industry officials and received an Honorable Mention. >
The team was also invited to present their technical paper as a demo and a special talk at the Emerging Technologies program at ACM SIGGRAPH 2022 as one of the top-three impactful technologies. The live performance, in which Hanbit Kim, a Ph.D. student of the Department of Industrial Design at KAIST and a co-author, sketched and animated a sophisticated animal-shaped robot from scratch in a matter of a few minutes, wowed the audience and won the Honorable Mention Award from the jury.
Edwin Catmull, the co-founder of Pixar and a keynote speaker at the SIGGRAPH conference, praised the team’s research on 3D sketching as “really excellent work” and “a kind of tool that would be useful to Pixar's creative model designers.”
This technology, which became virally popular in Japan after featuring in an online IT media outlet and attracting more than 600K views, received a special award from the Digital Content Association of Japan (DCAJ) and was invited and exhibited for three days at Tokyo in November, as a part of Inter BEE 2022, the largest broadcasting and media expo in Japan.
“The more we come to understand how designers think and work, the more effective design tools can be built around that understanding,” said Professor Bae, explaining that “the key is to integrate different algorithms into a harmonious system as intuitive interactions.” He added that “this work wouldn’t have been possible if it weren’t for the convergent research environment cultivated by the Department of Industrial Design at KAIST, in which all students see themselves not only as aspiring creative designers, but also as practical engineers.”
By enabling designers to produce highly expressive animated 3D concepts far more quickly and easily in comparison to using existing methods, this new tool is expected to revolutionize design practices and processes in the content creation, manufacturing, and metaverse-related industries.
This research was funded by the Ministry of Science and ICT, and the National Research Foundation of Korea.
More info: https://sketch.kaist.ac.kr/publications/2022_siggraph_rapid_design
Video: https://www.youtube.com/watch?v=rsBl0QvSDqI
< Figure 4. From left to right: Ph.D. students Hanbit Kim, and Joon Hyub Lee and Professor Bae of the Department of Industrial Design, KAIST >
2022.11.23 View 9929 -
 KAIST develops biocompatible adhesive applicable to hair transplants
Aside from being used as a new medical adhesive, the new material can be applied to developing a new method of hair transplants, which cannot be repeated multiple times using current method of implanting the wholly intact follicles into the skin.
Medical adhesives are materials that can be applied to various uses such as wound healing, hemostasis, vascular anastomosis, and tissue engineering, and is expected to contribute greatly to the development of minimally invasive surgery and organ transplants. However, adhesives with high adhesion, low toxicity, and capable of decomposing in the body are rare. Adhesives based on natural proteins, such as fibrin and collagen, have high biocompatibility but insufficient adhesive strength. Synthetic polymer adhesives based on urethane or acrylic have greater adhesion but do not decompose well and may cause an inflammatory reaction in the body.
A joint research team led by Professor Myungeun Seo and Professor Haeshin Lee from the KAIST Department of Chemistry developed a bio-friendly adhesive from biocompatible polymers using tannic acid, the source of astringency in wine.
The research team focused on tannic acid, a natural polyphenolic product. Tannic acid is a polyphenol present in large amounts in fruit peels, nuts, and cacao. It has a high affinity and coating ability on other substances, and we sense the astringent taste in wine when tannic acid sticks to the surface of our tongue. When tannic acid is mixed with hydrophilic polymers, they form coacervates, or small droplets of jelly-like fluids that sink. If the polymers used are biocompatible, the mixture can be applied as a medical adhesive with low toxicity. However, coacervates are fundamentally fluid-like and cannot withstand large forces, which limits their adhesive capabilities. Thus, while research to utilize it as an adhesive has been actively discussed, a biodegradable material exhibiting strong adhesion due to its high shear strength has not yet been developed.
The research team figured out a way to enhance adhesion by mixing two biocompatible FDA-approved polymers, polyethylene glycol (PEG) and polylactic acid (PLA). While PEG, which is used widely in eyedrops and cream, is hydrophilic, PLA, a well-known bioplastic derived from lactic acid, is insoluble in water. The team combined the two into a block copolymer, which forms hydrophilic PLA aggregates in water with PEG blocks surrounding them. A coacervate created by mixing the micelles and tannic acid would behave like a solid due to the hard PLA components, and show an elastic modulus improved by a thousand times compared to PEG, enabling it to withstand much greater force as an adhesive.
Figure 1. (Above) Principle of biodegradable adhesive made by mixing poly(ethylene glycol)-poly(lactic acid) diblock copolymer and tannic acid in water. Yellow coacervate is precipitated through hydrogen bonding between the block copolymer micelles and tannic acid, and exhibits adhesion. After heat treatment, hydrogen bonds are rearranged to further improve adhesion. (Bottom) Adhesion comparison. Compared to using poly(ethylene glycol) polymer (d), it can support 10 times more weight when using block copolymer (e) and 60 times more weight after heat treatment (f). The indicated G' values represent the elastic modulus of the material.
Furthermore, the research team observed that the material’s mechanical properties can be improved by over a hundred times through a heating and cooling process that is used to heat-treat metals. They also discovered that this is due to the enforced interactions between micelle and tannic acid arrays.
The research team used the fact that the material shows minimal irritation to the skin and decomposes well in the body to demonstrate its possible application as an adhesive for hair transplantation through an animal experiment. Professor Haeshin Lee, who has pioneered various application fields including medical adhesives, hemostatic agents, and browning shampoo, focused on the adhesive capacities and low toxicity of polyphenols like tannic acid, and now looks forward to it improving the limitations of current hair transplant methods, which still involve follicle transfer and are difficult to be repeated multiple times.
Figure 2. (a) Overview of a hair transplantation method using a biodegradable adhesive (right) compared to a conventional hair transplantation method (left) that transplants hair containing hair follicles. After applying an adhesive to the tip of the hair, it is fixed to the skin by implanting it through a subcutaneous injection, and repeated treatment is possible. (b) Initial animal test results. One day after 15 hair transplantation, 12 strands of hair remain. If you pull the 3 strands of hair, you can see that the whole body is pulled up, indicating that it is firmly implanted into the skin. All strands of hair applied without the new adhesive material fell off, and in the case of adhesive without heat treatment, the efficiency was 1/7.
This research was conducted by first co-authors Dr. Jongmin Park (currently a senior researcher at the Korea Research Institute of Chemical Technology) from Professor Myeongeun Seo’s team and Dr. Eunsook Park from Professor Haeshin Lee’s team in the KAIST Department of Chemistry, and through joint research with the teams led by Professor Hyungjun Kim from the KAIST Department of Chemistry and Professor Siyoung Choi from the Department of Chemical and Biomolecular Engineering. The research was published online on August 22 in the international journal Au (JACS Au) under the title Biodegradable Block Copolymer-Tannic Acid Glue.
This study was funded by the Support Research Under Protection Project of the National Research Foundation (NRF), Leading Research Center Support Project (Research Center for Multiscale Chiral Structure), Biodegradable Plastics Commercialization and Demonstration Project by the Ministry of Trade and Industry, and institutional funding from the Korea Research Institute of Chemical Technology.
2022.10.07 View 10842
KAIST develops biocompatible adhesive applicable to hair transplants
Aside from being used as a new medical adhesive, the new material can be applied to developing a new method of hair transplants, which cannot be repeated multiple times using current method of implanting the wholly intact follicles into the skin.
Medical adhesives are materials that can be applied to various uses such as wound healing, hemostasis, vascular anastomosis, and tissue engineering, and is expected to contribute greatly to the development of minimally invasive surgery and organ transplants. However, adhesives with high adhesion, low toxicity, and capable of decomposing in the body are rare. Adhesives based on natural proteins, such as fibrin and collagen, have high biocompatibility but insufficient adhesive strength. Synthetic polymer adhesives based on urethane or acrylic have greater adhesion but do not decompose well and may cause an inflammatory reaction in the body.
A joint research team led by Professor Myungeun Seo and Professor Haeshin Lee from the KAIST Department of Chemistry developed a bio-friendly adhesive from biocompatible polymers using tannic acid, the source of astringency in wine.
The research team focused on tannic acid, a natural polyphenolic product. Tannic acid is a polyphenol present in large amounts in fruit peels, nuts, and cacao. It has a high affinity and coating ability on other substances, and we sense the astringent taste in wine when tannic acid sticks to the surface of our tongue. When tannic acid is mixed with hydrophilic polymers, they form coacervates, or small droplets of jelly-like fluids that sink. If the polymers used are biocompatible, the mixture can be applied as a medical adhesive with low toxicity. However, coacervates are fundamentally fluid-like and cannot withstand large forces, which limits their adhesive capabilities. Thus, while research to utilize it as an adhesive has been actively discussed, a biodegradable material exhibiting strong adhesion due to its high shear strength has not yet been developed.
The research team figured out a way to enhance adhesion by mixing two biocompatible FDA-approved polymers, polyethylene glycol (PEG) and polylactic acid (PLA). While PEG, which is used widely in eyedrops and cream, is hydrophilic, PLA, a well-known bioplastic derived from lactic acid, is insoluble in water. The team combined the two into a block copolymer, which forms hydrophilic PLA aggregates in water with PEG blocks surrounding them. A coacervate created by mixing the micelles and tannic acid would behave like a solid due to the hard PLA components, and show an elastic modulus improved by a thousand times compared to PEG, enabling it to withstand much greater force as an adhesive.
Figure 1. (Above) Principle of biodegradable adhesive made by mixing poly(ethylene glycol)-poly(lactic acid) diblock copolymer and tannic acid in water. Yellow coacervate is precipitated through hydrogen bonding between the block copolymer micelles and tannic acid, and exhibits adhesion. After heat treatment, hydrogen bonds are rearranged to further improve adhesion. (Bottom) Adhesion comparison. Compared to using poly(ethylene glycol) polymer (d), it can support 10 times more weight when using block copolymer (e) and 60 times more weight after heat treatment (f). The indicated G' values represent the elastic modulus of the material.
Furthermore, the research team observed that the material’s mechanical properties can be improved by over a hundred times through a heating and cooling process that is used to heat-treat metals. They also discovered that this is due to the enforced interactions between micelle and tannic acid arrays.
The research team used the fact that the material shows minimal irritation to the skin and decomposes well in the body to demonstrate its possible application as an adhesive for hair transplantation through an animal experiment. Professor Haeshin Lee, who has pioneered various application fields including medical adhesives, hemostatic agents, and browning shampoo, focused on the adhesive capacities and low toxicity of polyphenols like tannic acid, and now looks forward to it improving the limitations of current hair transplant methods, which still involve follicle transfer and are difficult to be repeated multiple times.
Figure 2. (a) Overview of a hair transplantation method using a biodegradable adhesive (right) compared to a conventional hair transplantation method (left) that transplants hair containing hair follicles. After applying an adhesive to the tip of the hair, it is fixed to the skin by implanting it through a subcutaneous injection, and repeated treatment is possible. (b) Initial animal test results. One day after 15 hair transplantation, 12 strands of hair remain. If you pull the 3 strands of hair, you can see that the whole body is pulled up, indicating that it is firmly implanted into the skin. All strands of hair applied without the new adhesive material fell off, and in the case of adhesive without heat treatment, the efficiency was 1/7.
This research was conducted by first co-authors Dr. Jongmin Park (currently a senior researcher at the Korea Research Institute of Chemical Technology) from Professor Myeongeun Seo’s team and Dr. Eunsook Park from Professor Haeshin Lee’s team in the KAIST Department of Chemistry, and through joint research with the teams led by Professor Hyungjun Kim from the KAIST Department of Chemistry and Professor Siyoung Choi from the Department of Chemical and Biomolecular Engineering. The research was published online on August 22 in the international journal Au (JACS Au) under the title Biodegradable Block Copolymer-Tannic Acid Glue.
This study was funded by the Support Research Under Protection Project of the National Research Foundation (NRF), Leading Research Center Support Project (Research Center for Multiscale Chiral Structure), Biodegradable Plastics Commercialization and Demonstration Project by the Ministry of Trade and Industry, and institutional funding from the Korea Research Institute of Chemical Technology.
2022.10.07 View 10842 -
 Phage resistant Escherichia coli strains developed to reduce fermentation failure
A genome engineering-based systematic strategy for developing phage resistant Escherichia coli strains has been successfully developed through the collaborative efforts of a team led by Professor Sang Yup Lee, Professor Shi Chen, and Professor Lianrong Wang. This study by Xuan Zou et al. was published in Nature Communications in August 2022 and featured in Nature Communications Editors’ Highlights. The collaboration by the School of Pharmaceutical Sciences at Wuhan University, the First Affiliated Hospital of Shenzhen University, and the KAIST Department of Chemical and Biomolecular Engineering has made an important advance in the metabolic engineering and fermentation industry as it solves a big problem of phage infection causing fermentation failure.
Systems metabolic engineering is a highly interdisciplinary field that has made the development of microbial cell factories to produce various bioproducts including chemicals, fuels, and materials possible in a sustainable and environmentally friendly way, mitigating the impact of worldwide resource depletion and climate change. Escherichia coli is one of the most important chassis microbial strains, given its wide applications in the bio-based production of a diverse range of chemicals and materials. With the development of tools and strategies for systems metabolic engineering using E. coli, a highly optimized and well-characterized cell factory will play a crucial role in converting cheap and readily available raw materials into products of great economic and industrial value.
However, the consistent problem of phage contamination in fermentation imposes a devastating impact on host cells and threatens the productivity of bacterial bioprocesses in biotechnology facilities, which can lead to widespread fermentation failure and immeasurable economic loss. Host-controlled defense systems can be developed into effective genetic engineering solutions to address bacteriophage contamination in industrial-scale fermentation; however, most of the resistance mechanisms only narrowly restrict phages and their effect on phage contamination will be limited.
There have been attempts to develop diverse abilities/systems for environmental adaptation or antiviral defense. The team’s collaborative efforts developed a new type II single-stranded DNA phosphorothioation (Ssp) defense system derived from E. coli 3234/A, which can be used in multiple industrial E. coli strains (e.g., E. coli K-12, B and W) to provide broad protection against various types of dsDNA coliphages. Furthermore, they developed a systematic genome engineering strategy involving the simultaneous genomic integration of the Ssp defense module and mutations in components that are essential to the phage life cycle. This strategy can be used to transform E. coli hosts that are highly susceptible to phage attack into strains with powerful restriction effects on the tested bacteriophages. This endows hosts with strong resistance against a wide spectrum of phage infections without affecting bacterial growth and normal physiological function. More importantly, the resulting engineered phage-resistant strains maintained the capabilities of producing the desired chemicals and recombinant proteins even under high levels of phage cocktail challenge, which provides crucial protection against phage attacks.
This is a major step forward, as it provides a systematic solution for engineering phage-resistant bacterial strains, especially industrial bioproduction strains, to protect cells from a wide range of bacteriophages. Considering the functionality of this engineering strategy with diverse E. coli strains, the strategy reported in this study can be widely extended to other bacterial species and industrial applications, which will be of great interest to researchers in academia and industry alike.
Fig. A schematic model of the systematic strategy for engineering phage-sensitive industrial E. coli strains into strains with broad antiphage activities. Through the simultaneous genomic integration of a DNA phosphorothioation-based Ssp defense module and mutations of components essential for the phage life cycle, the engineered E. coli strains show strong resistance against diverse phages tested and maintain the capabilities of producing example recombinant proteins, even under high levels of phage cocktail challenge.
2022.08.23 View 12866
Phage resistant Escherichia coli strains developed to reduce fermentation failure
A genome engineering-based systematic strategy for developing phage resistant Escherichia coli strains has been successfully developed through the collaborative efforts of a team led by Professor Sang Yup Lee, Professor Shi Chen, and Professor Lianrong Wang. This study by Xuan Zou et al. was published in Nature Communications in August 2022 and featured in Nature Communications Editors’ Highlights. The collaboration by the School of Pharmaceutical Sciences at Wuhan University, the First Affiliated Hospital of Shenzhen University, and the KAIST Department of Chemical and Biomolecular Engineering has made an important advance in the metabolic engineering and fermentation industry as it solves a big problem of phage infection causing fermentation failure.
Systems metabolic engineering is a highly interdisciplinary field that has made the development of microbial cell factories to produce various bioproducts including chemicals, fuels, and materials possible in a sustainable and environmentally friendly way, mitigating the impact of worldwide resource depletion and climate change. Escherichia coli is one of the most important chassis microbial strains, given its wide applications in the bio-based production of a diverse range of chemicals and materials. With the development of tools and strategies for systems metabolic engineering using E. coli, a highly optimized and well-characterized cell factory will play a crucial role in converting cheap and readily available raw materials into products of great economic and industrial value.
However, the consistent problem of phage contamination in fermentation imposes a devastating impact on host cells and threatens the productivity of bacterial bioprocesses in biotechnology facilities, which can lead to widespread fermentation failure and immeasurable economic loss. Host-controlled defense systems can be developed into effective genetic engineering solutions to address bacteriophage contamination in industrial-scale fermentation; however, most of the resistance mechanisms only narrowly restrict phages and their effect on phage contamination will be limited.
There have been attempts to develop diverse abilities/systems for environmental adaptation or antiviral defense. The team’s collaborative efforts developed a new type II single-stranded DNA phosphorothioation (Ssp) defense system derived from E. coli 3234/A, which can be used in multiple industrial E. coli strains (e.g., E. coli K-12, B and W) to provide broad protection against various types of dsDNA coliphages. Furthermore, they developed a systematic genome engineering strategy involving the simultaneous genomic integration of the Ssp defense module and mutations in components that are essential to the phage life cycle. This strategy can be used to transform E. coli hosts that are highly susceptible to phage attack into strains with powerful restriction effects on the tested bacteriophages. This endows hosts with strong resistance against a wide spectrum of phage infections without affecting bacterial growth and normal physiological function. More importantly, the resulting engineered phage-resistant strains maintained the capabilities of producing the desired chemicals and recombinant proteins even under high levels of phage cocktail challenge, which provides crucial protection against phage attacks.
This is a major step forward, as it provides a systematic solution for engineering phage-resistant bacterial strains, especially industrial bioproduction strains, to protect cells from a wide range of bacteriophages. Considering the functionality of this engineering strategy with diverse E. coli strains, the strategy reported in this study can be widely extended to other bacterial species and industrial applications, which will be of great interest to researchers in academia and industry alike.
Fig. A schematic model of the systematic strategy for engineering phage-sensitive industrial E. coli strains into strains with broad antiphage activities. Through the simultaneous genomic integration of a DNA phosphorothioation-based Ssp defense module and mutations of components essential for the phage life cycle, the engineered E. coli strains show strong resistance against diverse phages tested and maintain the capabilities of producing example recombinant proteins, even under high levels of phage cocktail challenge.
2022.08.23 View 12866 -
 Interactive Map of Metabolical Synthesis of Chemicals
An interactive map that compiled the chemicals produced by biological, chemical and combined reactions has been distributed on the web
- A team led by Distinguished Professor Sang Yup Lee of the Department of Chemical and Biomolecular Engineering, organized and distributed an all-inclusive listing of chemical substances that can be synthesized using microorganisms
- It is expected to be used by researchers around the world as it enables easy assessment of the synthetic pathway through the web.
A research team comprised of Woo Dae Jang, Gi Bae Kim, and Distinguished Professor Sang Yup Lee of the Department of Chemical and Biomolecular Engineering at KAIST reported an interactive metabolic map of bio-based chemicals. Their research paper “An interactive metabolic map of bio-based chemicals” was published online in Trends in Biotechnology on August 10, 2022.
As a response to rapid climate change and environmental pollution, research on the production of petrochemical products using microorganisms is receiving attention as a sustainable alternative to existing methods of productions. In order to synthesize various chemical substances, materials, and fuel using microorganisms, it is necessary to first construct the biosynthetic pathway toward desired product by exploration and discovery and introduce them into microorganisms. In addition, in order to efficiently synthesize various chemical substances, it is sometimes necessary to employ chemical methods along with bioengineering methods using microorganisms at the same time. For the production of non-native chemicals, novel pathways are designed by recruiting enzymes from heterologous sources or employing enzymes designed though rational engineering, directed evolution, or ab initio design.
The research team had completed a map of chemicals which compiled all available pathways of biological and/or chemical reactions that lead to the production of various bio-based chemicals back in 2019 and published the map in Nature Catalysis. The map was distributed in the form of a poster to industries and academia so that the synthesis paths of bio-based chemicals could be checked at a glance.
The research team has expanded the bio-based chemicals map this time in the form of an interactive map on the web so that anyone with internet access can quickly explore efficient paths to synthesize desired products. The web-based map provides interactive visual tools to allow interactive visualization, exploration, and analysis of complex networks of biological and/or chemical reactions toward the desired products. In addition, the reported paper also discusses the production of natural compounds that are used for diverse purposes such as food and medicine, which will help designing novel pathways through similar approaches or by exploiting the promiscuity of enzymes described in the map. The published bio-based chemicals map is also available at http://systemsbiotech.co.kr.
The co-first authors, Dr. Woo Dae Jang and Ph.D. student Gi Bae Kim, said, “We conducted this study to address the demand for updating the previously distributed chemicals map and enhancing its versatility.” “The map is expected to be utilized in a variety of research and in efforts to set strategies and prospects for chemical production incorporating bio and chemical methods that are detailed in the map.”
Distinguished Professor Sang Yup Lee said, “The interactive bio-based chemicals map is expected to help design and optimization of the metabolic pathways for the biosynthesis of target chemicals together with the strategies of chemical conversions, serving as a blueprint for developing further ideas on the production of desired chemicals through biological and/or chemical reactions.”
The interactive metabolic map of bio-based chemicals.
2022.08.11 View 13996
Interactive Map of Metabolical Synthesis of Chemicals
An interactive map that compiled the chemicals produced by biological, chemical and combined reactions has been distributed on the web
- A team led by Distinguished Professor Sang Yup Lee of the Department of Chemical and Biomolecular Engineering, organized and distributed an all-inclusive listing of chemical substances that can be synthesized using microorganisms
- It is expected to be used by researchers around the world as it enables easy assessment of the synthetic pathway through the web.
A research team comprised of Woo Dae Jang, Gi Bae Kim, and Distinguished Professor Sang Yup Lee of the Department of Chemical and Biomolecular Engineering at KAIST reported an interactive metabolic map of bio-based chemicals. Their research paper “An interactive metabolic map of bio-based chemicals” was published online in Trends in Biotechnology on August 10, 2022.
As a response to rapid climate change and environmental pollution, research on the production of petrochemical products using microorganisms is receiving attention as a sustainable alternative to existing methods of productions. In order to synthesize various chemical substances, materials, and fuel using microorganisms, it is necessary to first construct the biosynthetic pathway toward desired product by exploration and discovery and introduce them into microorganisms. In addition, in order to efficiently synthesize various chemical substances, it is sometimes necessary to employ chemical methods along with bioengineering methods using microorganisms at the same time. For the production of non-native chemicals, novel pathways are designed by recruiting enzymes from heterologous sources or employing enzymes designed though rational engineering, directed evolution, or ab initio design.
The research team had completed a map of chemicals which compiled all available pathways of biological and/or chemical reactions that lead to the production of various bio-based chemicals back in 2019 and published the map in Nature Catalysis. The map was distributed in the form of a poster to industries and academia so that the synthesis paths of bio-based chemicals could be checked at a glance.
The research team has expanded the bio-based chemicals map this time in the form of an interactive map on the web so that anyone with internet access can quickly explore efficient paths to synthesize desired products. The web-based map provides interactive visual tools to allow interactive visualization, exploration, and analysis of complex networks of biological and/or chemical reactions toward the desired products. In addition, the reported paper also discusses the production of natural compounds that are used for diverse purposes such as food and medicine, which will help designing novel pathways through similar approaches or by exploiting the promiscuity of enzymes described in the map. The published bio-based chemicals map is also available at http://systemsbiotech.co.kr.
The co-first authors, Dr. Woo Dae Jang and Ph.D. student Gi Bae Kim, said, “We conducted this study to address the demand for updating the previously distributed chemicals map and enhancing its versatility.” “The map is expected to be utilized in a variety of research and in efforts to set strategies and prospects for chemical production incorporating bio and chemical methods that are detailed in the map.”
Distinguished Professor Sang Yup Lee said, “The interactive bio-based chemicals map is expected to help design and optimization of the metabolic pathways for the biosynthesis of target chemicals together with the strategies of chemical conversions, serving as a blueprint for developing further ideas on the production of desired chemicals through biological and/or chemical reactions.”
The interactive metabolic map of bio-based chemicals.
2022.08.11 View 13996 -
 Metabolically Engineered Bacterium Produces Lutein
A research group at KAIST has engineered a bacterial strain capable of producing lutein. The research team applied systems metabolic engineering strategies, including substrate channeling and electron channeling, to enhance the production of lutein in an engineered Escherichia coli strain. The strategies will be also useful for the efficient production of other industrially important natural products used in the food, pharmaceutical, and cosmetic industries.
Figure: Systems metabolic engineering was employed to construct and optimize the metabolic pathways for lutein production, and substrate channeling and electron channeling strategies were additionally employed to increase the production of the lutein with high productivity.
Lutein is classified as a xanthophyll chemical that is abundant in egg yolk, fruits, and vegetables. It protects the eye from oxidative damage from radiation and reduces the risk of eye diseases including macular degeneration and cataracts. Commercialized products featuring lutein are derived from the extracts of the marigold flower, which is known to harbor abundant amounts of lutein. However, the drawback of lutein production from nature is that it takes a long time to grow and harvest marigold flowers. Furthermore, it requires additional physical and chemical-based extractions with a low yield, which makes it economically unfeasible in terms of productivity. The high cost and low yield of these bioprocesses has made it difficult to readily meet the demand for lutein.
These challenges inspired the metabolic engineers at KAIST, including researchers Dr. Seon Young Park, Ph.D. Candidate Hyunmin Eun, and Distinguished Professor Sang Yup Lee from the Department of Chemical and Biomolecular Engineering. The team’s study entitled “Metabolic engineering of Escherichia coli with electron channeling for the production of natural products” was published in Nature Catalysis on August 5, 2022.
This research details the ability to produce lutein from E. coli with a high yield using a cheap carbon source, glycerol, via systems metabolic engineering. The research group focused on solving the bottlenecks of the biosynthetic pathway for lutein production constructed within an individual cell. First, using systems metabolic engineering, which is an integrated technology to engineer the metabolism of a microorganism, lutein was produced when the lutein biosynthesis pathway was introduced, albeit in very small amounts.
To improve the productivity of lutein production, the bottleneck enzymes within the metabolic pathway were first identified. It turned out that metabolic reactions that involve a promiscuous enzyme, an enzyme that is involved in two or more metabolic reactions, and electron-requiring cytochrome P450 enzymes are the main bottleneck steps of the pathway inhibiting lutein biosynthesis.
To overcome these challenges, substrate channeling, a strategy to artificially recruit enzymes in physical proximity within the cell in order to increase the local concentrations of substrates that can be converted into products, was employed to channel more metabolic flux towards the target chemical while reducing the formation of unwanted byproducts.
Furthermore, electron channeling, a strategy similar to substrate channeling but differing in terms of increasing the local concentrations of electrons required for oxidoreduction reactions mediated by P450 and its reductase partners, was applied to further streamline the metabolic flux towards lutein biosynthesis, which led to the highest titer of lutein production achieved in a bacterial host ever reported. The same electron channeling strategy was successfully applied for the production of other natural products including nootkatone and apigenin in E. coli, showcasing the general applicability of the strategy in the research field.
“It is expected that this microbial cell factory-based production of lutein will be able to replace the current plant extraction-based process,” said Dr. Seon Young Park, the first author of the paper. She explained that another important point of the research is that integrated metabolic engineering strategies developed from this study can be generally applicable for the efficient production of other natural products useful as pharmaceuticals or nutraceuticals.
“As maintaining good health in an aging society is becoming increasingly important, we expect that the technology and strategies developed here will play pivotal roles in producing other valuable natural products of medical or nutritional importance,” explained Distinguished Professor Sang Yup Lee.
This work was supported by the Cooperative Research Program for Agriculture Science & Technology Development funded by the Rural Development Administration of Korea, with further support from the Development of Next-generation Biorefinery Platform Technologies for Leading Bio-based Chemicals Industry Project and by the Development of Platform Technologies of Microbial Cell Factories for the Next-generation Biorefineries Project of the National Research Foundation funded by the Ministry of Science and ICT of Korea.
2022.08.05 View 9866
Metabolically Engineered Bacterium Produces Lutein
A research group at KAIST has engineered a bacterial strain capable of producing lutein. The research team applied systems metabolic engineering strategies, including substrate channeling and electron channeling, to enhance the production of lutein in an engineered Escherichia coli strain. The strategies will be also useful for the efficient production of other industrially important natural products used in the food, pharmaceutical, and cosmetic industries.
Figure: Systems metabolic engineering was employed to construct and optimize the metabolic pathways for lutein production, and substrate channeling and electron channeling strategies were additionally employed to increase the production of the lutein with high productivity.
Lutein is classified as a xanthophyll chemical that is abundant in egg yolk, fruits, and vegetables. It protects the eye from oxidative damage from radiation and reduces the risk of eye diseases including macular degeneration and cataracts. Commercialized products featuring lutein are derived from the extracts of the marigold flower, which is known to harbor abundant amounts of lutein. However, the drawback of lutein production from nature is that it takes a long time to grow and harvest marigold flowers. Furthermore, it requires additional physical and chemical-based extractions with a low yield, which makes it economically unfeasible in terms of productivity. The high cost and low yield of these bioprocesses has made it difficult to readily meet the demand for lutein.
These challenges inspired the metabolic engineers at KAIST, including researchers Dr. Seon Young Park, Ph.D. Candidate Hyunmin Eun, and Distinguished Professor Sang Yup Lee from the Department of Chemical and Biomolecular Engineering. The team’s study entitled “Metabolic engineering of Escherichia coli with electron channeling for the production of natural products” was published in Nature Catalysis on August 5, 2022.
This research details the ability to produce lutein from E. coli with a high yield using a cheap carbon source, glycerol, via systems metabolic engineering. The research group focused on solving the bottlenecks of the biosynthetic pathway for lutein production constructed within an individual cell. First, using systems metabolic engineering, which is an integrated technology to engineer the metabolism of a microorganism, lutein was produced when the lutein biosynthesis pathway was introduced, albeit in very small amounts.
To improve the productivity of lutein production, the bottleneck enzymes within the metabolic pathway were first identified. It turned out that metabolic reactions that involve a promiscuous enzyme, an enzyme that is involved in two or more metabolic reactions, and electron-requiring cytochrome P450 enzymes are the main bottleneck steps of the pathway inhibiting lutein biosynthesis.
To overcome these challenges, substrate channeling, a strategy to artificially recruit enzymes in physical proximity within the cell in order to increase the local concentrations of substrates that can be converted into products, was employed to channel more metabolic flux towards the target chemical while reducing the formation of unwanted byproducts.
Furthermore, electron channeling, a strategy similar to substrate channeling but differing in terms of increasing the local concentrations of electrons required for oxidoreduction reactions mediated by P450 and its reductase partners, was applied to further streamline the metabolic flux towards lutein biosynthesis, which led to the highest titer of lutein production achieved in a bacterial host ever reported. The same electron channeling strategy was successfully applied for the production of other natural products including nootkatone and apigenin in E. coli, showcasing the general applicability of the strategy in the research field.
“It is expected that this microbial cell factory-based production of lutein will be able to replace the current plant extraction-based process,” said Dr. Seon Young Park, the first author of the paper. She explained that another important point of the research is that integrated metabolic engineering strategies developed from this study can be generally applicable for the efficient production of other natural products useful as pharmaceuticals or nutraceuticals.
“As maintaining good health in an aging society is becoming increasingly important, we expect that the technology and strategies developed here will play pivotal roles in producing other valuable natural products of medical or nutritional importance,” explained Distinguished Professor Sang Yup Lee.
This work was supported by the Cooperative Research Program for Agriculture Science & Technology Development funded by the Rural Development Administration of Korea, with further support from the Development of Next-generation Biorefinery Platform Technologies for Leading Bio-based Chemicals Industry Project and by the Development of Platform Technologies of Microbial Cell Factories for the Next-generation Biorefineries Project of the National Research Foundation funded by the Ministry of Science and ICT of Korea.
2022.08.05 View 9866 -
 Professor Juho Kim’s Team Wins Best Paper Award at ACM CHI 2022
The research team led by Professor Juho Kim from the KAIST School of Computing won a Best Paper Award and an Honorable Mention Award at the Association for Computing Machinery Conference on Human Factors in Computing Systems (ACM CHI) held between April 30 and May 6.
ACM CHI is the world’s most recognized conference in the field of human computer interactions (HCI), and is ranked number one out of all HCI-related journals and conferences based on Google Scholar’s h-5 index. Best paper awards are given to works that rank in the top one percent, and honorable mention awards are given to the top five percent of the papers accepted by the conference.
Professor Juho Kim presented a total of seven papers at ACM CHI 2022, and tied for the largest number of papers. A total of 19 papers were affiliated with KAIST, putting it fifth out of all participating institutes and thereby proving KAIST’s competence in research.
One of Professor Kim’s research teams composed of Jeongyeon Kim (first author, MS graduate) from the School of Computing, MS candidate Yubin Choi from the School of Electrical Engineering, and Dr. Meng Xia (post-doctoral associate in the School of Computing, currently a post-doctoral associate at Carnegie Mellon University) received a best paper award for their paper, “Mobile-Friendly Content Design for MOOCs: Challenges, Requirements, and Design Opportunities”.
The study analyzed the difficulties experienced by learners watching video-based educational content in a mobile environment and suggests guidelines for solutions. The research team analyzed 134 survey responses and 21 interviews, and revealed that texts that are too small or overcrowded are what mainly brings down the legibility of video contents. Additionally, lighting, noise, and surrounding environments that change frequently are also important factors that may disturb a learning experience.
Based on these findings, the team analyzed the aptness of 41,722 frames from 101 video lectures for mobile environments, and confirmed that they generally show low levels of adequacy. For instance, in the case of text sizes, only 24.5% of the frames were shown to be adequate for learning in mobile environments. To overcome this issue, the research team suggested a guideline that may improve the legibility of video contents and help overcome the difficulties arising from mobile learning environments.
The importance of and dependency on video-based learning continue to rise, especially in the wake of the pandemic, and it is meaningful that this research suggested a means to analyze and tackle the difficulties of users that learn from the small screens of mobile devices. Furthermore, the paper also suggested technology that can solve problems related to video-based learning through human-AI collaborations, enhancing existing video lectures and improving learning experiences. This technology can be applied to various video-based platforms and content creation.
Meanwhile, a research team composed of Ph.D. candidate Tae Soo Kim (first author), MS candidate DaEun Choi, and Ph.D. candidate Yoonseo Choi from the School of Computing received an honorable mention award for their paper, “Stylette: styling the Web with Natural Language”.
The research team developed a novel interface technology that allows nonexperts who are unfamiliar with technical jargon to edit website features through speech. People often find it difficult to use or find the information they need from various websites due to accessibility issues, device-related constraints, inconvenient design, style preferences, etc. However, it is not easy for laymen to edit website features without expertise in programming or design, and most end up just putting up with the inconveniences. But what if the system could read the intentions of its users from their everyday language like “emphasize this part a little more”, or “I want a more modern design”, and edit the features automatically?
Based on this question, Professor Kim’s research team developed ‘Stylette’, a system in which AI analyses its users’ speech expressed in their natural language and automatically recommends a new style that best fits their intentions. The research team created a new system by putting together language AI, visual AI, and user interface technologies. On the linguistic side, a large-scale language model AI converts the intentions of the users expressed through their everyday language into adequate style elements. On the visual side, computer vision AI compares 1.7 million existing web design features and recommends a style adequate for the current website. In an experiment where 40 nonexperts were asked to edit a website design, the subjects that used this system showed double the success rate in a time span that was 35% shorter compared to the control group.
It is meaningful that this research proposed a practical case in which AI technology constructs intuitive interactions with users. The developed technology can be applied to existing design applications and web browsers in a plug-in format, and can be utilized to improve websites or for advertisements by collecting the natural intention data of users on a large scale.
2022.06.13 View 8522
Professor Juho Kim’s Team Wins Best Paper Award at ACM CHI 2022
The research team led by Professor Juho Kim from the KAIST School of Computing won a Best Paper Award and an Honorable Mention Award at the Association for Computing Machinery Conference on Human Factors in Computing Systems (ACM CHI) held between April 30 and May 6.
ACM CHI is the world’s most recognized conference in the field of human computer interactions (HCI), and is ranked number one out of all HCI-related journals and conferences based on Google Scholar’s h-5 index. Best paper awards are given to works that rank in the top one percent, and honorable mention awards are given to the top five percent of the papers accepted by the conference.
Professor Juho Kim presented a total of seven papers at ACM CHI 2022, and tied for the largest number of papers. A total of 19 papers were affiliated with KAIST, putting it fifth out of all participating institutes and thereby proving KAIST’s competence in research.
One of Professor Kim’s research teams composed of Jeongyeon Kim (first author, MS graduate) from the School of Computing, MS candidate Yubin Choi from the School of Electrical Engineering, and Dr. Meng Xia (post-doctoral associate in the School of Computing, currently a post-doctoral associate at Carnegie Mellon University) received a best paper award for their paper, “Mobile-Friendly Content Design for MOOCs: Challenges, Requirements, and Design Opportunities”.
The study analyzed the difficulties experienced by learners watching video-based educational content in a mobile environment and suggests guidelines for solutions. The research team analyzed 134 survey responses and 21 interviews, and revealed that texts that are too small or overcrowded are what mainly brings down the legibility of video contents. Additionally, lighting, noise, and surrounding environments that change frequently are also important factors that may disturb a learning experience.
Based on these findings, the team analyzed the aptness of 41,722 frames from 101 video lectures for mobile environments, and confirmed that they generally show low levels of adequacy. For instance, in the case of text sizes, only 24.5% of the frames were shown to be adequate for learning in mobile environments. To overcome this issue, the research team suggested a guideline that may improve the legibility of video contents and help overcome the difficulties arising from mobile learning environments.
The importance of and dependency on video-based learning continue to rise, especially in the wake of the pandemic, and it is meaningful that this research suggested a means to analyze and tackle the difficulties of users that learn from the small screens of mobile devices. Furthermore, the paper also suggested technology that can solve problems related to video-based learning through human-AI collaborations, enhancing existing video lectures and improving learning experiences. This technology can be applied to various video-based platforms and content creation.
Meanwhile, a research team composed of Ph.D. candidate Tae Soo Kim (first author), MS candidate DaEun Choi, and Ph.D. candidate Yoonseo Choi from the School of Computing received an honorable mention award for their paper, “Stylette: styling the Web with Natural Language”.
The research team developed a novel interface technology that allows nonexperts who are unfamiliar with technical jargon to edit website features through speech. People often find it difficult to use or find the information they need from various websites due to accessibility issues, device-related constraints, inconvenient design, style preferences, etc. However, it is not easy for laymen to edit website features without expertise in programming or design, and most end up just putting up with the inconveniences. But what if the system could read the intentions of its users from their everyday language like “emphasize this part a little more”, or “I want a more modern design”, and edit the features automatically?
Based on this question, Professor Kim’s research team developed ‘Stylette’, a system in which AI analyses its users’ speech expressed in their natural language and automatically recommends a new style that best fits their intentions. The research team created a new system by putting together language AI, visual AI, and user interface technologies. On the linguistic side, a large-scale language model AI converts the intentions of the users expressed through their everyday language into adequate style elements. On the visual side, computer vision AI compares 1.7 million existing web design features and recommends a style adequate for the current website. In an experiment where 40 nonexperts were asked to edit a website design, the subjects that used this system showed double the success rate in a time span that was 35% shorter compared to the control group.
It is meaningful that this research proposed a practical case in which AI technology constructs intuitive interactions with users. The developed technology can be applied to existing design applications and web browsers in a plug-in format, and can be utilized to improve websites or for advertisements by collecting the natural intention data of users on a large scale.
2022.06.13 View 8522 -
 Now You Can See Floral Scents!
Optical interferometry visualizes how often lilies emit volatile organic compounds
Have you ever thought about when flowers emit their scents?
KAIST mechanical engineers and biological scientists directly visualized how often a lily releases a floral scent using a laser interferometry method. These measurement results can provide new insights for understanding and further exploring the biosynthesis and emission mechanisms of floral volatiles.
Why is it important to know this? It is well known that the fragrance of flowers affects their interactions with pollinators, microorganisms, and florivores. For instance, many flowering plants can tune their scent emission rates when pollinators are active for pollination. Petunias and the wild tobacco Nicotiana attenuata emit floral scents at night to attract night-active pollinators. Thus, visualizing scent emissions can help us understand the ecological evolution of plant-pollinator interactions.
Many groups have been trying to develop methods for scent analysis. Mass spectrometry has been one widely used method for investigating the fragrance of flowers. Although mass spectrometry reveals the quality and quantity of floral scents, it is impossible to directly measure the releasing frequency. A laser-based gas detection system and a smartphone-based detection system using chemo-responsive dyes have also been used to measure volatile organic compounds (VOCs) in real-time, but it is still hard to measure the time-dependent emission rate of floral scents.
However, the KAIST research team co-led by Professor Hyoungsoo Kim from the Department of Mechanical Engineering and Professor Sang-Gyu Kim from the Department of Biological Sciences measured a refractive index difference between the vapor of the VOCs of lilies and the air to measure the emission frequency. The floral scent vapor was detected and the refractive index of air was 1.0 while that of the major floral scent of a linalool lily was 1.46.
Professor Hyoungsoo Kim said, “We expect this technology to be further applicable to various industrial sectors such as developing it to detect hazardous substances in a space.” The research team also plans to identify the DNA mechanism that controls floral scent secretion.
The current work entitled “Real-time visualization of scent accumulation reveals the frequency of floral scent emissions” was published in ‘Frontiers in Plant Science’ on April 18, 2022. (https://doi.org/10.3389/fpls.2022.835305).
This research was supported by the Basic Science Research Program through the National Research Foundation of Korea (NRF-2021R1A2C2007835), the Rural Development Administration (PJ016403), and the KAIST-funded Global Singularity Research PREP-Program.
-Publication:H. Kim, G. Lee, J. Song, and S.-G. Kim, "Real-time visualization of scent accumulation reveals the frequency of floral scent emissions," Frontiers in Plant Science 18, 835305 (2022) (https://doi.org/10.3389/fpls.2022.835305)
-Profile:Professor Hyoungsoo Kimhttp://fil.kaist.ac.kr
@MadeInH on TwitterDepartment of Mechanical EngineeringKAIST
Professor Sang-Gyu Kimhttps://sites.google.com/view/kimlab/home Department of Biological SciencesKAIST
2022.05.25 View 9421
Now You Can See Floral Scents!
Optical interferometry visualizes how often lilies emit volatile organic compounds
Have you ever thought about when flowers emit their scents?
KAIST mechanical engineers and biological scientists directly visualized how often a lily releases a floral scent using a laser interferometry method. These measurement results can provide new insights for understanding and further exploring the biosynthesis and emission mechanisms of floral volatiles.
Why is it important to know this? It is well known that the fragrance of flowers affects their interactions with pollinators, microorganisms, and florivores. For instance, many flowering plants can tune their scent emission rates when pollinators are active for pollination. Petunias and the wild tobacco Nicotiana attenuata emit floral scents at night to attract night-active pollinators. Thus, visualizing scent emissions can help us understand the ecological evolution of plant-pollinator interactions.
Many groups have been trying to develop methods for scent analysis. Mass spectrometry has been one widely used method for investigating the fragrance of flowers. Although mass spectrometry reveals the quality and quantity of floral scents, it is impossible to directly measure the releasing frequency. A laser-based gas detection system and a smartphone-based detection system using chemo-responsive dyes have also been used to measure volatile organic compounds (VOCs) in real-time, but it is still hard to measure the time-dependent emission rate of floral scents.
However, the KAIST research team co-led by Professor Hyoungsoo Kim from the Department of Mechanical Engineering and Professor Sang-Gyu Kim from the Department of Biological Sciences measured a refractive index difference between the vapor of the VOCs of lilies and the air to measure the emission frequency. The floral scent vapor was detected and the refractive index of air was 1.0 while that of the major floral scent of a linalool lily was 1.46.
Professor Hyoungsoo Kim said, “We expect this technology to be further applicable to various industrial sectors such as developing it to detect hazardous substances in a space.” The research team also plans to identify the DNA mechanism that controls floral scent secretion.
The current work entitled “Real-time visualization of scent accumulation reveals the frequency of floral scent emissions” was published in ‘Frontiers in Plant Science’ on April 18, 2022. (https://doi.org/10.3389/fpls.2022.835305).
This research was supported by the Basic Science Research Program through the National Research Foundation of Korea (NRF-2021R1A2C2007835), the Rural Development Administration (PJ016403), and the KAIST-funded Global Singularity Research PREP-Program.
-Publication:H. Kim, G. Lee, J. Song, and S.-G. Kim, "Real-time visualization of scent accumulation reveals the frequency of floral scent emissions," Frontiers in Plant Science 18, 835305 (2022) (https://doi.org/10.3389/fpls.2022.835305)
-Profile:Professor Hyoungsoo Kimhttp://fil.kaist.ac.kr
@MadeInH on TwitterDepartment of Mechanical EngineeringKAIST
Professor Sang-Gyu Kimhttps://sites.google.com/view/kimlab/home Department of Biological SciencesKAIST
2022.05.25 View 9421 -
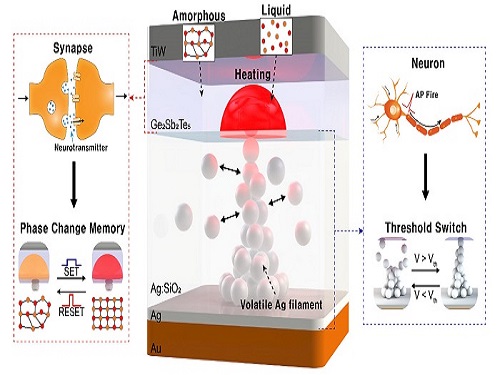 Neuromorphic Memory Device Simulates Neurons and Synapses
Simultaneous emulation of neuronal and synaptic properties promotes the development of brain-like artificial intelligence
Researchers have reported a nano-sized neuromorphic memory device that emulates neurons and synapses simultaneously in a unit cell, another step toward completing the goal of neuromorphic computing designed to rigorously mimic the human brain with semiconductor devices.
Neuromorphic computing aims to realize artificial intelligence (AI) by mimicking the mechanisms of neurons and synapses that make up the human brain. Inspired by the cognitive functions of the human brain that current computers cannot provide, neuromorphic devices have been widely investigated. However, current Complementary Metal-Oxide Semiconductor (CMOS)-based neuromorphic circuits simply connect artificial neurons and synapses without synergistic interactions, and the concomitant implementation of neurons and synapses still remains a challenge. To address these issues, a research team led by Professor Keon Jae Lee from the Department of Materials Science and Engineering implemented the biological working mechanisms of humans by introducing the neuron-synapse interactions in a single memory cell, rather than the conventional approach of electrically connecting artificial neuronal and synaptic devices.
Similar to commercial graphics cards, the artificial synaptic devices previously studied often used to accelerate parallel computations, which shows clear differences from the operational mechanisms of the human brain. The research team implemented the synergistic interactions between neurons and synapses in the neuromorphic memory device, emulating the mechanisms of the biological neural network. In addition, the developed neuromorphic device can replace complex CMOS neuron circuits with a single device, providing high scalability and cost efficiency.
The human brain consists of a complex network of 100 billion neurons and 100 trillion synapses. The functions and structures of neurons and synapses can flexibly change according to the external stimuli, adapting to the surrounding environment. The research team developed a neuromorphic device in which short-term and long-term memories coexist using volatile and non-volatile memory devices that mimic the characteristics of neurons and synapses, respectively. A threshold switch device is used as volatile memory and phase-change memory is used as a non-volatile device. Two thin-film devices are integrated without intermediate electrodes, implementing the functional adaptability of neurons and synapses in the neuromorphic memory.
Professor Keon Jae Lee explained, "Neurons and synapses interact with each other to establish cognitive functions such as memory and learning, so simulating both is an essential element for brain-inspired artificial intelligence. The developed neuromorphic memory device also mimics the retraining effect that allows quick learning of the forgotten information by implementing a positive feedback effect between neurons and synapses.”
This result entitled “Simultaneous emulation of synaptic and intrinsic plasticity using a memristive synapse” was published in the May 19, 2022 issue of Nature Communications.
-Publication:Sang Hyun Sung, Tae Jin Kim, Hyera Shin, Tae Hong Im, and Keon Jae Lee (2022) “Simultaneous emulation of synaptic and intrinsic plasticity using a memristive synapse,” Nature Communications May 19, 2022 (DOI: 10.1038/s41467-022-30432-2)
-Profile:Professor Keon Jae Leehttp://fand.kaist.ac.kr
Department of Materials Science and EngineeringKAIST
2022.05.20 View 13467
Neuromorphic Memory Device Simulates Neurons and Synapses
Simultaneous emulation of neuronal and synaptic properties promotes the development of brain-like artificial intelligence
Researchers have reported a nano-sized neuromorphic memory device that emulates neurons and synapses simultaneously in a unit cell, another step toward completing the goal of neuromorphic computing designed to rigorously mimic the human brain with semiconductor devices.
Neuromorphic computing aims to realize artificial intelligence (AI) by mimicking the mechanisms of neurons and synapses that make up the human brain. Inspired by the cognitive functions of the human brain that current computers cannot provide, neuromorphic devices have been widely investigated. However, current Complementary Metal-Oxide Semiconductor (CMOS)-based neuromorphic circuits simply connect artificial neurons and synapses without synergistic interactions, and the concomitant implementation of neurons and synapses still remains a challenge. To address these issues, a research team led by Professor Keon Jae Lee from the Department of Materials Science and Engineering implemented the biological working mechanisms of humans by introducing the neuron-synapse interactions in a single memory cell, rather than the conventional approach of electrically connecting artificial neuronal and synaptic devices.
Similar to commercial graphics cards, the artificial synaptic devices previously studied often used to accelerate parallel computations, which shows clear differences from the operational mechanisms of the human brain. The research team implemented the synergistic interactions between neurons and synapses in the neuromorphic memory device, emulating the mechanisms of the biological neural network. In addition, the developed neuromorphic device can replace complex CMOS neuron circuits with a single device, providing high scalability and cost efficiency.
The human brain consists of a complex network of 100 billion neurons and 100 trillion synapses. The functions and structures of neurons and synapses can flexibly change according to the external stimuli, adapting to the surrounding environment. The research team developed a neuromorphic device in which short-term and long-term memories coexist using volatile and non-volatile memory devices that mimic the characteristics of neurons and synapses, respectively. A threshold switch device is used as volatile memory and phase-change memory is used as a non-volatile device. Two thin-film devices are integrated without intermediate electrodes, implementing the functional adaptability of neurons and synapses in the neuromorphic memory.
Professor Keon Jae Lee explained, "Neurons and synapses interact with each other to establish cognitive functions such as memory and learning, so simulating both is an essential element for brain-inspired artificial intelligence. The developed neuromorphic memory device also mimics the retraining effect that allows quick learning of the forgotten information by implementing a positive feedback effect between neurons and synapses.”
This result entitled “Simultaneous emulation of synaptic and intrinsic plasticity using a memristive synapse” was published in the May 19, 2022 issue of Nature Communications.
-Publication:Sang Hyun Sung, Tae Jin Kim, Hyera Shin, Tae Hong Im, and Keon Jae Lee (2022) “Simultaneous emulation of synaptic and intrinsic plasticity using a memristive synapse,” Nature Communications May 19, 2022 (DOI: 10.1038/s41467-022-30432-2)
-Profile:Professor Keon Jae Leehttp://fand.kaist.ac.kr
Department of Materials Science and EngineeringKAIST
2022.05.20 View 13467 -
 Anonymous Donor Makes a Gift of Property Valued at 30 Billion KRW
The KAIST Development Foundation announced on May 9 that an anonymous donor in his 50s made a gift of real estate valued at 30 billion KRW. This is the first donation from an anonymous benefactor on such a grand scale. The benefactor expressed his wishes to fund scholarships for students in need and R&D for medical and bio sciences.
According to the Development Foundation official, the benefactor is reported to have said that he felt burdened that he earned much more than he needed and was looking for the right way to share his assets. The benefactor refused to hold an official donation ceremony and meeting with high-level university administrators.
The donor believes that KAIST is filled with young and dynamic energy, saying, “I would like to help KAIST move forward and create breakthroughs that will benefit the nation as well as all humanity.”
Before making up his mind to give his asset to KAIST, he had planned to establish his own social foundation but he changed his mind. “I decided that an investment in education would be the best investment,” he said. He explained that he was inspired by his KAIST graduate friend who is running a company. He was deeply motivated to help KAIST after witnessing the KAIST graduate’s passion for conducting his business.
After receiving the gift, KAIST President Kwang Hyung Lee was thankful for the full support and trust of the benefactor. “We will spare no effort to foster next-generation talents and advance science and technology in the field of biomedicine.”
2022.05.11 View 5558
Anonymous Donor Makes a Gift of Property Valued at 30 Billion KRW
The KAIST Development Foundation announced on May 9 that an anonymous donor in his 50s made a gift of real estate valued at 30 billion KRW. This is the first donation from an anonymous benefactor on such a grand scale. The benefactor expressed his wishes to fund scholarships for students in need and R&D for medical and bio sciences.
According to the Development Foundation official, the benefactor is reported to have said that he felt burdened that he earned much more than he needed and was looking for the right way to share his assets. The benefactor refused to hold an official donation ceremony and meeting with high-level university administrators.
The donor believes that KAIST is filled with young and dynamic energy, saying, “I would like to help KAIST move forward and create breakthroughs that will benefit the nation as well as all humanity.”
Before making up his mind to give his asset to KAIST, he had planned to establish his own social foundation but he changed his mind. “I decided that an investment in education would be the best investment,” he said. He explained that he was inspired by his KAIST graduate friend who is running a company. He was deeply motivated to help KAIST after witnessing the KAIST graduate’s passion for conducting his business.
After receiving the gift, KAIST President Kwang Hyung Lee was thankful for the full support and trust of the benefactor. “We will spare no effort to foster next-generation talents and advance science and technology in the field of biomedicine.”
2022.05.11 View 5558 -
 Mathematicians Identify a Key Source of Cell-to-Cell Variability in Cell Signaling
Systematic inferences identify a major source of heterogeneity in cell signaling dynamics
Why do genetically identical cells respond differently to the same external stimuli, such as antibiotics? This long-standing mystery has been solved by KAIST and IBS mathematicians who have developed a new framework for analyzing cell responses to some stimuli. The team found that the cell-to-cell variability in antibiotic stress response increases as the effective length of the cell signaling pathway (i.e., the number of rate-limiting steps) increases. This finding could identify more effective chemotherapies to overcome the fractional killing of cancer cells caused by cell-to-cell variability.
Cells in the human body contain signal transduction systems that respond to various external stimuli such as antibiotics and changes in osmotic pressure. When an external stimulus is detected, various biochemical reactions occur sequentially. This leads to the expression of relevant genes, allowing the cells to respond to the perturbed external environment. Furthermore, signal transduction leads to a drug response (e.g., antibiotic resistance genes are expressed when antibiotic drugs are given).
However, even when the same external stimuli are detected, the responses of individual cells are greatly heterogeneous. This leads to the emergence of persister cells that are highly resistant to drugs. To identify potential sources of this cell-to cell variability, many studies have been conducted. However, most of the intermediate signal transduction reactions are unobservable with current experimental techniques.
A group of researchers including Dae Wook Kim and Hyukpyo Hong and led by Professor Jae Kyoung Kim from the KAIST Department of Mathematical Sciences and IBS Biomedical Mathematics Group solved the mystery by exploiting queueing theory and Bayesian inference methodology. They proposed a queueing process that describes the signal transduction system in cells. Based on this, they developed Bayesian inference computational software using MBI (the Moment-based Bayesian Inference method). This enables the analysis of the signal transduction system without a direct observation of the intermediate steps. This study was published in Science Advances.
By analyzing experimental data from Escherichia coli using MBI, the research team found that cell-to-cell variability increases as the number of rate-limiting steps in the signaling pathway increases.
The rate-limiting steps denote the slowest steps (i.e., bottlenecks) in sequential biochemical reaction steps composing cell signaling pathways and thus dominates most of the signaling time. As the number of the rate-limiting steps increases, the intensity of the transduced signal becomes greatly heterogeneous even in a population of genetically identical cells. This finding is expected to provide a new paradigm for studying the heterogeneous antibiotic resistance of cells, which is a big challenge in cancer medicine.
Professor Kim said, “As a mathematician, I am excited to help advance the understanding of cell-to-cell variability in response to external stimuli. I hope this finding facilitates the development of more effective chemotherapies.”
This work was supported by the Samsung Science and Technology Foundation, the National Research Foundation of Korea, and the Institute for Basic Science.
-Publication:Dae Wook Kim, Hyukpyo Hong, and Jae Kyoung Kim (2022) “Systematic inference identifies a major source of heterogeneity in cell signaling dynamics: the rate-limiting step number,”Science Advances March 18, 2022 (DOI: 10.1126/sciadv.abl4598)
-Profile:Professor Jae Kyoung Kimhttp://mathsci.kaist.ac.kr/~jaekkim
jaekkim@kaist.ac.kr@umichkim on TwitterDepartment of Mathematical SciencesKAIST
2022.03.29 View 10037
Mathematicians Identify a Key Source of Cell-to-Cell Variability in Cell Signaling
Systematic inferences identify a major source of heterogeneity in cell signaling dynamics
Why do genetically identical cells respond differently to the same external stimuli, such as antibiotics? This long-standing mystery has been solved by KAIST and IBS mathematicians who have developed a new framework for analyzing cell responses to some stimuli. The team found that the cell-to-cell variability in antibiotic stress response increases as the effective length of the cell signaling pathway (i.e., the number of rate-limiting steps) increases. This finding could identify more effective chemotherapies to overcome the fractional killing of cancer cells caused by cell-to-cell variability.
Cells in the human body contain signal transduction systems that respond to various external stimuli such as antibiotics and changes in osmotic pressure. When an external stimulus is detected, various biochemical reactions occur sequentially. This leads to the expression of relevant genes, allowing the cells to respond to the perturbed external environment. Furthermore, signal transduction leads to a drug response (e.g., antibiotic resistance genes are expressed when antibiotic drugs are given).
However, even when the same external stimuli are detected, the responses of individual cells are greatly heterogeneous. This leads to the emergence of persister cells that are highly resistant to drugs. To identify potential sources of this cell-to cell variability, many studies have been conducted. However, most of the intermediate signal transduction reactions are unobservable with current experimental techniques.
A group of researchers including Dae Wook Kim and Hyukpyo Hong and led by Professor Jae Kyoung Kim from the KAIST Department of Mathematical Sciences and IBS Biomedical Mathematics Group solved the mystery by exploiting queueing theory and Bayesian inference methodology. They proposed a queueing process that describes the signal transduction system in cells. Based on this, they developed Bayesian inference computational software using MBI (the Moment-based Bayesian Inference method). This enables the analysis of the signal transduction system without a direct observation of the intermediate steps. This study was published in Science Advances.
By analyzing experimental data from Escherichia coli using MBI, the research team found that cell-to-cell variability increases as the number of rate-limiting steps in the signaling pathway increases.
The rate-limiting steps denote the slowest steps (i.e., bottlenecks) in sequential biochemical reaction steps composing cell signaling pathways and thus dominates most of the signaling time. As the number of the rate-limiting steps increases, the intensity of the transduced signal becomes greatly heterogeneous even in a population of genetically identical cells. This finding is expected to provide a new paradigm for studying the heterogeneous antibiotic resistance of cells, which is a big challenge in cancer medicine.
Professor Kim said, “As a mathematician, I am excited to help advance the understanding of cell-to-cell variability in response to external stimuli. I hope this finding facilitates the development of more effective chemotherapies.”
This work was supported by the Samsung Science and Technology Foundation, the National Research Foundation of Korea, and the Institute for Basic Science.
-Publication:Dae Wook Kim, Hyukpyo Hong, and Jae Kyoung Kim (2022) “Systematic inference identifies a major source of heterogeneity in cell signaling dynamics: the rate-limiting step number,”Science Advances March 18, 2022 (DOI: 10.1126/sciadv.abl4598)
-Profile:Professor Jae Kyoung Kimhttp://mathsci.kaist.ac.kr/~jaekkim
jaekkim@kaist.ac.kr@umichkim on TwitterDepartment of Mathematical SciencesKAIST
2022.03.29 View 10037 -
 'Fingerprint' Machine Learning Technique Identifies Different Bacteria in Seconds
A synergistic combination of surface-enhanced Raman spectroscopy and deep learning serves as an effective platform for separation-free detection of bacteria in arbitrary media
Bacterial identification can take hours and often longer, precious time when diagnosing infections and selecting appropriate treatments. There may be a quicker, more accurate process according to researchers at KAIST. By teaching a deep learning algorithm to identify the “fingerprint” spectra of the molecular components of various bacteria, the researchers could classify various bacteria in different media with accuracies of up to 98%.
Their results were made available online on Jan. 18 in Biosensors and Bioelectronics, ahead of publication in the journal’s April issue.
Bacteria-induced illnesses, those caused by direct bacterial infection or by exposure to bacterial toxins, can induce painful symptoms and even lead to death, so the rapid detection of bacteria is crucial to prevent the intake of contaminated foods and to diagnose infections from clinical samples, such as urine. “By using surface-enhanced Raman spectroscopy (SERS) analysis boosted with a newly proposed deep learning model, we demonstrated a markedly simple, fast, and effective route to classify the signals of two common bacteria and their resident media without any separation procedures,” said Professor Sungho Jo from the School of Computing.
Raman spectroscopy sends light through a sample to see how it scatters. The results reveal structural information about the sample — the spectral fingerprint — allowing researchers to identify its molecules. The surface-enhanced version places sample cells on noble metal nanostructures that help amplify the sample’s signals.
However, it is challenging to obtain consistent and clear spectra of bacteria due to numerous overlapping peak sources, such as proteins in cell walls. “Moreover, strong signals of surrounding media are also enhanced to overwhelm target signals, requiring time-consuming and tedious bacterial separation steps,” said Professor Yeon Sik Jung from the Department of Materials Science and Engineering.
To parse through the noisy signals, the researchers implemented an artificial intelligence method called deep learning that can hierarchically extract certain features of the spectral information to classify data. They specifically designed their model, named the dual-branch wide-kernel network (DualWKNet), to efficiently learn the correlation between spectral features. Such an ability is critical for analyzing one-dimensional spectral data, according to Professor Jo.
“Despite having interfering signals or noise from the media, which make the general shapes of different bacterial spectra and their residing media signals look similar, high classification accuracies of bacterial types and their media were achieved,” Professor Jo said, explaining that DualWKNet allowed the team to identify key peaks in each class that were almost indiscernible in individual spectra, enhancing the classification accuracies. “Ultimately, with the use of DualWKNet replacing the bacteria and media separation steps, our method dramatically reduces analysis time.”
The researchers plan to use their platform to study more bacteria and media types, using the information to build a training data library of various bacterial types in additional media to reduce the collection and detection times for new samples.
“We developed a meaningful universal platform for rapid bacterial detection with the collaboration between SERS and deep learning,” Professor Jo said. “We hope to extend the use of our deep learning-based SERS analysis platform to detect numerous types of bacteria in additional media that are important for food or clinical analysis, such as blood.”
The National R&D Program, through a National Research Foundation of Korea grant funded by the Ministry of Science and ICT, supported this research.
-PublicationEojin Rho, Minjoon Kim, Seunghee H. Cho, Bongjae Choi, Hyungjoon Park, Hanhwi Jang, Yeon Sik Jung, Sungho Jo, “Separation-free bacterial identification in arbitrary media via deepneural network-based SERS analysis,” Biosensors and Bioelectronics online January 18, 2022 (doi.org/10.1016/j.bios.2022.113991)
-ProfileProfessor Yeon Sik JungDepartment of Materials Science and EngineeringKAIST
Professor Sungho JoSchool of ComputingKAIST
2022.03.04 View 21825
'Fingerprint' Machine Learning Technique Identifies Different Bacteria in Seconds
A synergistic combination of surface-enhanced Raman spectroscopy and deep learning serves as an effective platform for separation-free detection of bacteria in arbitrary media
Bacterial identification can take hours and often longer, precious time when diagnosing infections and selecting appropriate treatments. There may be a quicker, more accurate process according to researchers at KAIST. By teaching a deep learning algorithm to identify the “fingerprint” spectra of the molecular components of various bacteria, the researchers could classify various bacteria in different media with accuracies of up to 98%.
Their results were made available online on Jan. 18 in Biosensors and Bioelectronics, ahead of publication in the journal’s April issue.
Bacteria-induced illnesses, those caused by direct bacterial infection or by exposure to bacterial toxins, can induce painful symptoms and even lead to death, so the rapid detection of bacteria is crucial to prevent the intake of contaminated foods and to diagnose infections from clinical samples, such as urine. “By using surface-enhanced Raman spectroscopy (SERS) analysis boosted with a newly proposed deep learning model, we demonstrated a markedly simple, fast, and effective route to classify the signals of two common bacteria and their resident media without any separation procedures,” said Professor Sungho Jo from the School of Computing.
Raman spectroscopy sends light through a sample to see how it scatters. The results reveal structural information about the sample — the spectral fingerprint — allowing researchers to identify its molecules. The surface-enhanced version places sample cells on noble metal nanostructures that help amplify the sample’s signals.
However, it is challenging to obtain consistent and clear spectra of bacteria due to numerous overlapping peak sources, such as proteins in cell walls. “Moreover, strong signals of surrounding media are also enhanced to overwhelm target signals, requiring time-consuming and tedious bacterial separation steps,” said Professor Yeon Sik Jung from the Department of Materials Science and Engineering.
To parse through the noisy signals, the researchers implemented an artificial intelligence method called deep learning that can hierarchically extract certain features of the spectral information to classify data. They specifically designed their model, named the dual-branch wide-kernel network (DualWKNet), to efficiently learn the correlation between spectral features. Such an ability is critical for analyzing one-dimensional spectral data, according to Professor Jo.
“Despite having interfering signals or noise from the media, which make the general shapes of different bacterial spectra and their residing media signals look similar, high classification accuracies of bacterial types and their media were achieved,” Professor Jo said, explaining that DualWKNet allowed the team to identify key peaks in each class that were almost indiscernible in individual spectra, enhancing the classification accuracies. “Ultimately, with the use of DualWKNet replacing the bacteria and media separation steps, our method dramatically reduces analysis time.”
The researchers plan to use their platform to study more bacteria and media types, using the information to build a training data library of various bacterial types in additional media to reduce the collection and detection times for new samples.
“We developed a meaningful universal platform for rapid bacterial detection with the collaboration between SERS and deep learning,” Professor Jo said. “We hope to extend the use of our deep learning-based SERS analysis platform to detect numerous types of bacteria in additional media that are important for food or clinical analysis, such as blood.”
The National R&D Program, through a National Research Foundation of Korea grant funded by the Ministry of Science and ICT, supported this research.
-PublicationEojin Rho, Minjoon Kim, Seunghee H. Cho, Bongjae Choi, Hyungjoon Park, Hanhwi Jang, Yeon Sik Jung, Sungho Jo, “Separation-free bacterial identification in arbitrary media via deepneural network-based SERS analysis,” Biosensors and Bioelectronics online January 18, 2022 (doi.org/10.1016/j.bios.2022.113991)
-ProfileProfessor Yeon Sik JungDepartment of Materials Science and EngineeringKAIST
Professor Sungho JoSchool of ComputingKAIST
2022.03.04 View 21825