NI
-
 New IEEE Fellow, Professor Jong Chul Ye
Professor Jong Chul Ye from the Department of Bio and Brain Engineering was named a new fellow of the Institute of Electrical and Electronics Engineers (IEEE). IEEE announced this on December 1 in recognition of Professor Ye’s contributions to the development of signal processing and artificial intelligence (AI) technology in the field of biomedical imaging.
As the world’s largest society in the electrical and electronics field, IEEE names the top 0.1% of their members as fellows based on their research achievements.Professor Ye has published more than 100 research papers in world-leading journals in the biomedical imaging field, including those affiliated with IEEE.
He also gave a keynote talk at the yearly conference of the International Society for Magnetic Resonance Imaging (ISMRM) on medical AI technology. In addition, Professor Ye has been appointed to serve as the next chair of the Computational Imaging Technical Committee of the IEEE Signal Processing Society, and the chair of the IEEE Symposium on Biomedical Imaging (ISBI) 2020 to be held in April in Iowa, USA.
Professor Ye said, “The importance of AI technology is developing in the biomedical imaging field. I feel proud that my contributions have been internationally recognized and allowed me to be named an IEEE fellow.”
2019.12.18 View 11462
New IEEE Fellow, Professor Jong Chul Ye
Professor Jong Chul Ye from the Department of Bio and Brain Engineering was named a new fellow of the Institute of Electrical and Electronics Engineers (IEEE). IEEE announced this on December 1 in recognition of Professor Ye’s contributions to the development of signal processing and artificial intelligence (AI) technology in the field of biomedical imaging.
As the world’s largest society in the electrical and electronics field, IEEE names the top 0.1% of their members as fellows based on their research achievements.Professor Ye has published more than 100 research papers in world-leading journals in the biomedical imaging field, including those affiliated with IEEE.
He also gave a keynote talk at the yearly conference of the International Society for Magnetic Resonance Imaging (ISMRM) on medical AI technology. In addition, Professor Ye has been appointed to serve as the next chair of the Computational Imaging Technical Committee of the IEEE Signal Processing Society, and the chair of the IEEE Symposium on Biomedical Imaging (ISBI) 2020 to be held in April in Iowa, USA.
Professor Ye said, “The importance of AI technology is developing in the biomedical imaging field. I feel proud that my contributions have been internationally recognized and allowed me to be named an IEEE fellow.”
2019.12.18 View 11462 -
 Professor Il-Doo Kim Named Scientist of the Year by the Journalists
Professor Il-Doo Kim from the Department of Materials Science and Engineering was named the 2019 Scientist of the Year by Korean science journalists. The award was conferred at the 2019 Science Press Night ceremony of the Korea Science Journalists Association (KSJA) on November 29.
Professor Kim focuses on developing nanofiber gas sensors for diagnosing diseases in advance by analyzing exhaled biomarkers with electrospinning technology. His outstanding research was praised and selected as one of the top 10 nanotechnology of 2019 by the Korea Nano Technology Research Society (KoNTRS), the Ministry of Science and ICT (MSIT), and the Ministry of Trade, Industry and Energy (MOTIE).
Professor Kim was honored with the QIAN Baojun Fiber Award, which is awarded every two years by Donghua University in Shanghai, China to recognize outstanding contributions in fiber science and technology. Professor Kim was also elected as an academician of the Asia Pacific Academy of Materials (APAM) on November 21 in Guangzhou, China.
In May, Professor Kim was appointed as an associate editor of ACS Nano, a leading international research journal in the field of nanoscience. In his editorial published in the May issue of ACS Nano, Professor Kim introduced and shared the history of KAIST and its vision for the future with other members of the journal. He hopes this will help with promoting a closer relationship between the members of the journal and KAIST moving forward.
“Above all,” he said in his acceptance speech, “the greatest news for me as an educator is that the first PhD graduate from our lab, Dr. Seonjin Choi, was appointed as the youngest professor in the Division of Materials Science and Engineering at Hanyang University on September 1.”
2019.12.17 View 12753
Professor Il-Doo Kim Named Scientist of the Year by the Journalists
Professor Il-Doo Kim from the Department of Materials Science and Engineering was named the 2019 Scientist of the Year by Korean science journalists. The award was conferred at the 2019 Science Press Night ceremony of the Korea Science Journalists Association (KSJA) on November 29.
Professor Kim focuses on developing nanofiber gas sensors for diagnosing diseases in advance by analyzing exhaled biomarkers with electrospinning technology. His outstanding research was praised and selected as one of the top 10 nanotechnology of 2019 by the Korea Nano Technology Research Society (KoNTRS), the Ministry of Science and ICT (MSIT), and the Ministry of Trade, Industry and Energy (MOTIE).
Professor Kim was honored with the QIAN Baojun Fiber Award, which is awarded every two years by Donghua University in Shanghai, China to recognize outstanding contributions in fiber science and technology. Professor Kim was also elected as an academician of the Asia Pacific Academy of Materials (APAM) on November 21 in Guangzhou, China.
In May, Professor Kim was appointed as an associate editor of ACS Nano, a leading international research journal in the field of nanoscience. In his editorial published in the May issue of ACS Nano, Professor Kim introduced and shared the history of KAIST and its vision for the future with other members of the journal. He hopes this will help with promoting a closer relationship between the members of the journal and KAIST moving forward.
“Above all,” he said in his acceptance speech, “the greatest news for me as an educator is that the first PhD graduate from our lab, Dr. Seonjin Choi, was appointed as the youngest professor in the Division of Materials Science and Engineering at Hanyang University on September 1.”
2019.12.17 View 12753 -
 KAIST Awarded the IPBC R&D Institution Team of the Year
KAIST was awarded the R&D Institution Team of the Year during the annual IPBC (Intellectual Property Business Congress) Asia 2019 held in Tokyo October 28-30. IPBC is a conference dedicated to IP value creation strategies hosted by IAM Media, a world’s leading IP business media platform.
IPBC Asia 2019 recognized the institutions and businesses that employed innovative IP strategies and management to produce the greatest IP value in 11 categories covering automotive, electronics, healthcare and biotechnology, internet and software, R&D institutions, semiconductors, industrials, mobile and telecommunications, Asia IP deals, Asia teams, and Asia individuals. This year, KAIST was recognized as one of the most active patentees in the Asia-Pacific region by significantly increasing its IP value through licensing and tech transfers.
Associate Vice President Kyung Cheol Choi of the Office of University-Industry Cooperation remarked, “We are so delighted to prove the strong research capacity of KAIST. This will help us accomplish our vision of being a leading university that creates global impact.”
2019.12.04 View 9115
KAIST Awarded the IPBC R&D Institution Team of the Year
KAIST was awarded the R&D Institution Team of the Year during the annual IPBC (Intellectual Property Business Congress) Asia 2019 held in Tokyo October 28-30. IPBC is a conference dedicated to IP value creation strategies hosted by IAM Media, a world’s leading IP business media platform.
IPBC Asia 2019 recognized the institutions and businesses that employed innovative IP strategies and management to produce the greatest IP value in 11 categories covering automotive, electronics, healthcare and biotechnology, internet and software, R&D institutions, semiconductors, industrials, mobile and telecommunications, Asia IP deals, Asia teams, and Asia individuals. This year, KAIST was recognized as one of the most active patentees in the Asia-Pacific region by significantly increasing its IP value through licensing and tech transfers.
Associate Vice President Kyung Cheol Choi of the Office of University-Industry Cooperation remarked, “We are so delighted to prove the strong research capacity of KAIST. This will help us accomplish our vision of being a leading university that creates global impact.”
2019.12.04 View 9115 -
 KAIST and Google Jointly Develop AI Curricula
KAIST selected the two professors who will develop AI curriculum under the auspices of the KAIST-Google Partnership for AI Education and Research. The Graduate School of AI announced the two authors among the 20 applicants who will develop the curriculum next year. They will be provided 7,500 USD per subject.
Professor Changho Suh from the School of Electrical Engineering and Professor Yong-Jin Yoon from the Department of Mechanical Engineering will use Google technology such as TensorFlow, Google Cloud, and Android to create the curriculum.
Professor Suh’s “TensorFlow for Information Theory and Convex Optimization “will be used for curriculum in the graduate courses and Professor Yoon’s “AI Convergence Project Based Learning (PBL)” will be used for online courses. Professor Yoon’s course will explore and define problems by utilizing AI and experiencing the process of developing products that use AI through design thinking, which involves product design, production, and verification. Professor Suh’s course will discus“information theory and convergence,” which uses basic sciences and engineering as well as AI, machine learning, and deep learning.
2019.12.04 View 16075
KAIST and Google Jointly Develop AI Curricula
KAIST selected the two professors who will develop AI curriculum under the auspices of the KAIST-Google Partnership for AI Education and Research. The Graduate School of AI announced the two authors among the 20 applicants who will develop the curriculum next year. They will be provided 7,500 USD per subject.
Professor Changho Suh from the School of Electrical Engineering and Professor Yong-Jin Yoon from the Department of Mechanical Engineering will use Google technology such as TensorFlow, Google Cloud, and Android to create the curriculum.
Professor Suh’s “TensorFlow for Information Theory and Convex Optimization “will be used for curriculum in the graduate courses and Professor Yoon’s “AI Convergence Project Based Learning (PBL)” will be used for online courses. Professor Yoon’s course will explore and define problems by utilizing AI and experiencing the process of developing products that use AI through design thinking, which involves product design, production, and verification. Professor Suh’s course will discus“information theory and convergence,” which uses basic sciences and engineering as well as AI, machine learning, and deep learning.
2019.12.04 View 16075 -
 KAIST Alumnus NYU Professor Supports Female AI Researchers
A KAIST alumnus and an associate professor at New York University (NYU), Dr. Kyunghyun Cho donated 3,000 USD to the KAIST Graduate School of AI to support female AI researchers.
Professor Cho spoke as a guest lecturer at the 2019 Samsung AI Forum on November 4 and received 3,000 USD as an honorarium. He donated this honorarium to the KAIST Graduate School of AI with a special request to support the school’s female PhD students attending the 2020 International Conference on Learning Representations (ICLR), where he serves as a program co-chair.
Professor Cho received his BS degree from KAIST’s School of Computing in 2009 and is now serving as an associate professor at NYU’s Computer Science Department and Center for Data Science. His research mainly covers machine learning and natural language processing.
Professor Cho said that he decided to make this donation because “In Korea and even in the US, women in science, technology, engineering, and mathematics (STEM) lack opportunities and environments that allow them to excel.”
Professor Song Chong, the Head of the KAIST Graduate School of AI, responded, “We are so grateful for Professor Kyunghyun Cho’s contribution and we will also use funds from the school in addition to the donation to support our female PhD students who will attend the ICLR.”
(END)
2019.11.15 View 10296
KAIST Alumnus NYU Professor Supports Female AI Researchers
A KAIST alumnus and an associate professor at New York University (NYU), Dr. Kyunghyun Cho donated 3,000 USD to the KAIST Graduate School of AI to support female AI researchers.
Professor Cho spoke as a guest lecturer at the 2019 Samsung AI Forum on November 4 and received 3,000 USD as an honorarium. He donated this honorarium to the KAIST Graduate School of AI with a special request to support the school’s female PhD students attending the 2020 International Conference on Learning Representations (ICLR), where he serves as a program co-chair.
Professor Cho received his BS degree from KAIST’s School of Computing in 2009 and is now serving as an associate professor at NYU’s Computer Science Department and Center for Data Science. His research mainly covers machine learning and natural language processing.
Professor Cho said that he decided to make this donation because “In Korea and even in the US, women in science, technology, engineering, and mathematics (STEM) lack opportunities and environments that allow them to excel.”
Professor Song Chong, the Head of the KAIST Graduate School of AI, responded, “We are so grateful for Professor Kyunghyun Cho’s contribution and we will also use funds from the school in addition to the donation to support our female PhD students who will attend the ICLR.”
(END)
2019.11.15 View 10296 -
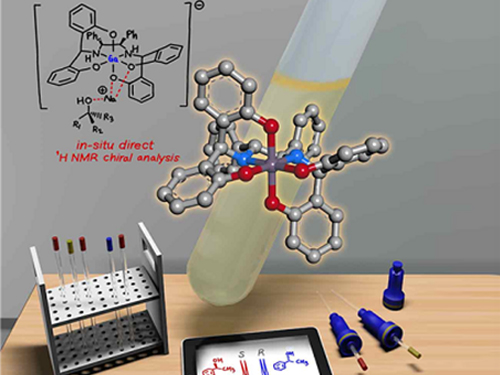 Gallium-Based Solvating Agent Efficiently Analyzes Optically Active Alcohols
A KAIST research team has developed a gallium-based metal complex enabling the rapid chiral analysis of alcohols. A team working under Professor Hyunwoo Kim reported the efficient new alcohol analysis method using nuclear magnetic resonance (NMR) spectroscopy in iScience.
Enantiopure chiral alcohols are ubiquitous in nature and widely utilized as pharmaceuticals. This importance of chirality in synthetic and medicinal chemistry has advanced the search for rapid and facile methods to determine the enantiomeric purities of compounds. To date, chiral analysis has been performed using high-performance liquid chromatography (HPLC) with chiral columns.
Along with the HPLC technique, chiral analysis using NMR spectroscopy has gained tremendous attention as an alternative to traditionally employed chromatographic methods due to its simplicity and rapid detection for real-time measurement. However, this method carries drawbacks such as line-broadening, narrow substrate scope, and poor resolution. Thus, compared with popular methods of chromatographic analysis, NMR spectroscopy is infrequently used for chiral analysis.
In principle, a chiral solvating agent is additionally required for the NMR measurement of chiral alcohols to obtain two distinct signals. However, NMR analysis of chiral alcohols has been challenging due to weak binding interactions with chiral solvating agents. To overcome the intrinsic difficulty of relatively weak molecular interactions that are common for alcohols, many researchers have used multifunctional alcohols to enhance interactions with solvating agents.
Instead, the KAIST team successfully varied the physical properties of metal complexes to induce stronger interactions with alcohols rather than the strategy of using multifunctional analytes, in the hopes of developing a universal chiral solvating agent for alcohols. Compared to the current method of chiral analysis used in the pharmaceutical industry, alcohols that do not possess chromophores can also be directly analyzed with the gallium complexes.
Professor Kim said that this method could be a complementary chiral analysis technique at the industry level in the near future. He added that since the developed gallium complex can determine enantiomeric excess within minutes, it can be further utilized to monitor asymmetric synthesis. This feature will benefit a large number of researchers in the organic chemistry community, as well as the pharmaceutical industry.
(Figure: Schematic view of the in-situ direct 1H NMR chiral analysis.)
-Profile:
Professor Hyunwoo Kim
Department of Chemistry
KAIST
http://mdos.kaist.ac.kr
hwk34@kaist.ac.kr
For more on this article,
please go to https://doi.org/10.1016/j.isci2019.07051
2019.11.14 View 12233
Gallium-Based Solvating Agent Efficiently Analyzes Optically Active Alcohols
A KAIST research team has developed a gallium-based metal complex enabling the rapid chiral analysis of alcohols. A team working under Professor Hyunwoo Kim reported the efficient new alcohol analysis method using nuclear magnetic resonance (NMR) spectroscopy in iScience.
Enantiopure chiral alcohols are ubiquitous in nature and widely utilized as pharmaceuticals. This importance of chirality in synthetic and medicinal chemistry has advanced the search for rapid and facile methods to determine the enantiomeric purities of compounds. To date, chiral analysis has been performed using high-performance liquid chromatography (HPLC) with chiral columns.
Along with the HPLC technique, chiral analysis using NMR spectroscopy has gained tremendous attention as an alternative to traditionally employed chromatographic methods due to its simplicity and rapid detection for real-time measurement. However, this method carries drawbacks such as line-broadening, narrow substrate scope, and poor resolution. Thus, compared with popular methods of chromatographic analysis, NMR spectroscopy is infrequently used for chiral analysis.
In principle, a chiral solvating agent is additionally required for the NMR measurement of chiral alcohols to obtain two distinct signals. However, NMR analysis of chiral alcohols has been challenging due to weak binding interactions with chiral solvating agents. To overcome the intrinsic difficulty of relatively weak molecular interactions that are common for alcohols, many researchers have used multifunctional alcohols to enhance interactions with solvating agents.
Instead, the KAIST team successfully varied the physical properties of metal complexes to induce stronger interactions with alcohols rather than the strategy of using multifunctional analytes, in the hopes of developing a universal chiral solvating agent for alcohols. Compared to the current method of chiral analysis used in the pharmaceutical industry, alcohols that do not possess chromophores can also be directly analyzed with the gallium complexes.
Professor Kim said that this method could be a complementary chiral analysis technique at the industry level in the near future. He added that since the developed gallium complex can determine enantiomeric excess within minutes, it can be further utilized to monitor asymmetric synthesis. This feature will benefit a large number of researchers in the organic chemistry community, as well as the pharmaceutical industry.
(Figure: Schematic view of the in-situ direct 1H NMR chiral analysis.)
-Profile:
Professor Hyunwoo Kim
Department of Chemistry
KAIST
http://mdos.kaist.ac.kr
hwk34@kaist.ac.kr
For more on this article,
please go to https://doi.org/10.1016/j.isci2019.07051
2019.11.14 View 12233 -
 AI to Determine When to Intervene with Your Driving
(Professor Uichin Lee (left) and PhD candidate Auk Kim)
Can your AI agent judge when to talk to you while you are driving? According to a KAIST research team, their in-vehicle conservation service technology will judge when it is appropriate to contact you to ensure your safety.
Professor Uichin Lee from the Department of Industrial and Systems Engineering at KAIST and his research team have developed AI technology that automatically detects safe moments for AI agents to provide conversation services to drivers.
Their research focuses on solving the potential problems of distraction created by in-vehicle conversation services. If an AI agent talks to a driver at an inopportune moment, such as while making a turn, a car accident will be more likely to occur.
In-vehicle conversation services need to be convenient as well as safe. However, the cognitive burden of multitasking negatively influences the quality of the service. Users tend to be more distracted during certain traffic conditions. To address this long-standing challenge of the in-vehicle conversation services, the team introduced a composite cognitive model that considers both safe driving and auditory-verbal service performance and used a machine-learning model for all collected data.
The combination of these individual measures is able to determine the appropriate moments for conversation and most appropriate types of conversational services. For instance, in the case of delivering simple-context information, such as a weather forecast, driver safety alone would be the most appropriate consideration. Meanwhile, when delivering information that requires a driver response, such as a “Yes” or “No,” the combination of driver safety and auditory-verbal performance should be considered.
The research team developed a prototype of an in-vehicle conversation service based on a navigation app that can be used in real driving environments. The app was also connected to the vehicle to collect in-vehicle OBD-II/CAN data, such as the steering wheel angle and brake pedal position, and mobility and environmental data such as the distance between successive cars and traffic flow.
Using pseudo-conversation services, the research team collected a real-world driving dataset consisting of 1,388 interactions and sensor data from 29 drivers who interacted with AI conversational agents. Machine learning analysis based on the dataset demonstrated that the opportune moments for driver interruption could be correctly inferred with 87% accuracy.
The safety enhancement technology developed by the team is expected to minimize driver distractions caused by in-vehicle conversation services. This technology can be directly applied to current in-vehicle systems that provide conversation services. It can also be extended and applied to the real-time detection of driver distraction problems caused by the use of a smartphone while driving.
Professor Lee said, “In the near future, cars will proactively deliver various in-vehicle conversation services. This technology will certainly help vehicles interact with their drivers safely as it can fairly accurately determine when to provide conversation services using only basic sensor data generated by cars.”
The researchers presented their findings at the ACM International Joint Conference on Pervasive and Ubiquitous Computing (Ubicomp’19) in London, UK. This research was supported in part by Hyundai NGV and by the Next-Generation Information Computing Development Program through the National Research Foundation of Korea (NRF) funded by the Ministry of Science and ICT.
(Figure: Visual description of safe enhancement technology for in-vehicle conversation services)
2019.11.13 View 19454
AI to Determine When to Intervene with Your Driving
(Professor Uichin Lee (left) and PhD candidate Auk Kim)
Can your AI agent judge when to talk to you while you are driving? According to a KAIST research team, their in-vehicle conservation service technology will judge when it is appropriate to contact you to ensure your safety.
Professor Uichin Lee from the Department of Industrial and Systems Engineering at KAIST and his research team have developed AI technology that automatically detects safe moments for AI agents to provide conversation services to drivers.
Their research focuses on solving the potential problems of distraction created by in-vehicle conversation services. If an AI agent talks to a driver at an inopportune moment, such as while making a turn, a car accident will be more likely to occur.
In-vehicle conversation services need to be convenient as well as safe. However, the cognitive burden of multitasking negatively influences the quality of the service. Users tend to be more distracted during certain traffic conditions. To address this long-standing challenge of the in-vehicle conversation services, the team introduced a composite cognitive model that considers both safe driving and auditory-verbal service performance and used a machine-learning model for all collected data.
The combination of these individual measures is able to determine the appropriate moments for conversation and most appropriate types of conversational services. For instance, in the case of delivering simple-context information, such as a weather forecast, driver safety alone would be the most appropriate consideration. Meanwhile, when delivering information that requires a driver response, such as a “Yes” or “No,” the combination of driver safety and auditory-verbal performance should be considered.
The research team developed a prototype of an in-vehicle conversation service based on a navigation app that can be used in real driving environments. The app was also connected to the vehicle to collect in-vehicle OBD-II/CAN data, such as the steering wheel angle and brake pedal position, and mobility and environmental data such as the distance between successive cars and traffic flow.
Using pseudo-conversation services, the research team collected a real-world driving dataset consisting of 1,388 interactions and sensor data from 29 drivers who interacted with AI conversational agents. Machine learning analysis based on the dataset demonstrated that the opportune moments for driver interruption could be correctly inferred with 87% accuracy.
The safety enhancement technology developed by the team is expected to minimize driver distractions caused by in-vehicle conversation services. This technology can be directly applied to current in-vehicle systems that provide conversation services. It can also be extended and applied to the real-time detection of driver distraction problems caused by the use of a smartphone while driving.
Professor Lee said, “In the near future, cars will proactively deliver various in-vehicle conversation services. This technology will certainly help vehicles interact with their drivers safely as it can fairly accurately determine when to provide conversation services using only basic sensor data generated by cars.”
The researchers presented their findings at the ACM International Joint Conference on Pervasive and Ubiquitous Computing (Ubicomp’19) in London, UK. This research was supported in part by Hyundai NGV and by the Next-Generation Information Computing Development Program through the National Research Foundation of Korea (NRF) funded by the Ministry of Science and ICT.
(Figure: Visual description of safe enhancement technology for in-vehicle conversation services)
2019.11.13 View 19454 -
 Ultrafast Quantum Motion in a Nanoscale Trap Detected
< Professor Heung-Sun Sim (left) and Co-author Dr. Sungguen Ryu (right) >
KAIST researchers have reported the detection of a picosecond electron motion in a silicon transistor. This study has presented a new protocol for measuring ultrafast electronic dynamics in an effective time-resolved fashion of picosecond resolution. The detection was made in collaboration with Nippon Telegraph and Telephone Corp. (NTT) in Japan and National Physical Laboratory (NPL) in the UK and is the first report to the best of our knowledge.
When an electron is captured in a nanoscale trap in solids, its quantum mechanical wave function can exhibit spatial oscillation at sub-terahertz frequencies. Time-resolved detection of such picosecond dynamics of quantum waves is important, as the detection provides a way of understanding the quantum behavior of electrons in nano-electronics. It also applies to quantum information technologies such as the ultrafast quantum-bit operation of quantum computing and high-sensitivity electromagnetic-field sensing. However, detecting picosecond dynamics has been a challenge since the sub-terahertz scale is far beyond the latest bandwidth measurement tools.
A KAIST team led by Professor Heung-Sun Sim developed a theory of ultrafast electron dynamics in a nanoscale trap, and proposed a scheme for detecting the dynamics, which utilizes a quantum-mechanical resonant state formed beside the trap. The coupling between the electron dynamics and the resonant state is switched on and off at a picosecond so that information on the dynamics is read out on the electric current being generated when the coupling is switched on.
NTT realized, together with NPL, the detection scheme and applied it to electron motions in a nanoscale trap formed in a silicon transistor. A single electron was captured in the trap by controlling electrostatic gates, and a resonant state was formed in the potential barrier of the trap.
The switching on and off of the coupling between the electron and the resonant state was achieved by aligning the resonance energy with the energy of the electron within a picosecond. An electric current from the trap through the resonant state to an electrode was measured at only a few Kelvin degrees, unveiling the spatial quantum-coherent oscillation of the electron with 250 GHz frequency inside the trap.
Professor Sim said, “This work suggests a scheme of detecting picosecond electron motions in submicron scales by utilizing quantum resonance. It will be useful in dynamical control of quantum mechanical electron waves for various purposes in nano-electronics, quantum sensing, and quantum information”.
This work was published online at Nature Nanotechnology on November 4. It was partly supported by the Korea National Research Foundation through the SRC Center for Quantum Coherence in Condensed Matter. For more on the NTT news release this article, please visit https://www.ntt.co.jp/news2019/1911e/191105a.html
-ProfileProfessor Heung-Sun Sim
Department of PhysicsDirector, SRC Center for Quantum Coherence in Condensed Matterhttps://qet.kaist.ac.kr KAIST
-Publication:Gento Yamahata, Sungguen Ryu, Nathan Johnson, H.-S. Sim, Akira Fujiwara, and Masaya Kataoka. 2019. Picosecond coherent electron motion in a silicon single-electron source. Nature Nanotechnology (Online Publication). 6 pages. https://doi.org/10.1038/s41565-019-0563-2
2019.11.05 View 19597
Ultrafast Quantum Motion in a Nanoscale Trap Detected
< Professor Heung-Sun Sim (left) and Co-author Dr. Sungguen Ryu (right) >
KAIST researchers have reported the detection of a picosecond electron motion in a silicon transistor. This study has presented a new protocol for measuring ultrafast electronic dynamics in an effective time-resolved fashion of picosecond resolution. The detection was made in collaboration with Nippon Telegraph and Telephone Corp. (NTT) in Japan and National Physical Laboratory (NPL) in the UK and is the first report to the best of our knowledge.
When an electron is captured in a nanoscale trap in solids, its quantum mechanical wave function can exhibit spatial oscillation at sub-terahertz frequencies. Time-resolved detection of such picosecond dynamics of quantum waves is important, as the detection provides a way of understanding the quantum behavior of electrons in nano-electronics. It also applies to quantum information technologies such as the ultrafast quantum-bit operation of quantum computing and high-sensitivity electromagnetic-field sensing. However, detecting picosecond dynamics has been a challenge since the sub-terahertz scale is far beyond the latest bandwidth measurement tools.
A KAIST team led by Professor Heung-Sun Sim developed a theory of ultrafast electron dynamics in a nanoscale trap, and proposed a scheme for detecting the dynamics, which utilizes a quantum-mechanical resonant state formed beside the trap. The coupling between the electron dynamics and the resonant state is switched on and off at a picosecond so that information on the dynamics is read out on the electric current being generated when the coupling is switched on.
NTT realized, together with NPL, the detection scheme and applied it to electron motions in a nanoscale trap formed in a silicon transistor. A single electron was captured in the trap by controlling electrostatic gates, and a resonant state was formed in the potential barrier of the trap.
The switching on and off of the coupling between the electron and the resonant state was achieved by aligning the resonance energy with the energy of the electron within a picosecond. An electric current from the trap through the resonant state to an electrode was measured at only a few Kelvin degrees, unveiling the spatial quantum-coherent oscillation of the electron with 250 GHz frequency inside the trap.
Professor Sim said, “This work suggests a scheme of detecting picosecond electron motions in submicron scales by utilizing quantum resonance. It will be useful in dynamical control of quantum mechanical electron waves for various purposes in nano-electronics, quantum sensing, and quantum information”.
This work was published online at Nature Nanotechnology on November 4. It was partly supported by the Korea National Research Foundation through the SRC Center for Quantum Coherence in Condensed Matter. For more on the NTT news release this article, please visit https://www.ntt.co.jp/news2019/1911e/191105a.html
-ProfileProfessor Heung-Sun Sim
Department of PhysicsDirector, SRC Center for Quantum Coherence in Condensed Matterhttps://qet.kaist.ac.kr KAIST
-Publication:Gento Yamahata, Sungguen Ryu, Nathan Johnson, H.-S. Sim, Akira Fujiwara, and Masaya Kataoka. 2019. Picosecond coherent electron motion in a silicon single-electron source. Nature Nanotechnology (Online Publication). 6 pages. https://doi.org/10.1038/s41565-019-0563-2
2019.11.05 View 19597 -
 Tungsten Suboxide Improves the Efficiency of Platinum in Hydrogen Production
< PhD Candidate Jinkyu Park and Professor Jinwoo Lee >
Researchers presented a new strategy for enhancing catalytic activity using tungsten suboxide as a single-atom catalyst (SAC). This strategy, which significantly improves hydrogen evolution reaction (HER) in metal platinum (pt) by 16.3 times, sheds light on the development of new electrochemical catalyst technologies.
Hydrogen has been touted as a promising alternative to fossil fuels. However, most of the conventional industrial hydrogen production methods come with environmental issues, releasing significant amounts of carbon dioxide and greenhouse gases.
Electrochemical water splitting is considered a potential approach for clean hydrogen production. Pt is one of the most commonly used catalysts to improve HER performance in electrochemical water splitting, but the high cost and scarcity of Pt remain key obstacles to mass commercial applications.
SACs, where all metal species are individually dispersed on a desired support material, have been identified as one way to reduce the amount of Pt usage, as they offer the maximum number of surface exposed Pt atoms.
Inspired by earlier studies, which mainly focused on SACs supported by carbon-based materials, a KAIST research team led by Professor Jinwoo Lee from the Department of Chemical and Biomolecular Engineering investigated the influence of support materials on the performance of SACs.
Professor Lee and his researchers suggested mesoporous tungsten suboxide as a new support material for atomically dispersed Pt, as this was expected to provide high electronic conductivity and have a synergetic effect with Pt.
They compared the performance of single-atom Pt supported by carbon and tungsten suboxide respectively. The results revealed that the support effect occurred with tungsten suboxide, in which the mass activity of a single-atom Pt supported by tungsten suboxide was 2.1 times greater than that of single-atom Pt supported by carbon, and 16.3 times higher than that of Pt nanoparticles supported by carbon.
The team indicated a change in the electronic structure of Pt via charge transfer from tungsten suboxide to Pt. This phenomenon was reported as a result of strong metal-support interaction between Pt and tungsten suboxide.
HER performance can be improved not only by changing the electronic structure of the supported metal, but also by inducing another support effect, the spillover effect, the research group reported. Hydrogen spillover is a phenomenon where adsorbed hydrogen migrates from one surface to another, and it occurs more easily as the Pt size becomes smaller.
The researchers compared the performance of single-atom Pt and Pt nanoparticles supported by tungsten suboxide. The single-atom Pt supported by tungsten suboxide exhibited a higher degree of hydrogen spillover phenomenon, which enhanced the Pt mass activity for hydrogen evolution up to 10.7 times compared to Pt nanoparticles supported by tungsten suboxide.
Professor Lee said, “Choosing the right support material is important for improving electrocatalysis in hydrogen production. The tungsten suboxide catalyst we used to support Pt in our study implies that interactions between the well-matched metal and support can drastically enhance the efficiency of the process.”
This research was supported by the Ministry of Science and ICT and introduced in the International Edition of the German journal Angewandte Chemie.
Figure. Schematic representation of hydrogen evolution reaction (HER) of pseudo single-atom Pt supported by tungsten suboxide
-Publication
Jinkyu Park, Dr. Seonggyu Lee, Hee-Eun Kim, Ara Cho, Seongbeen Kim, Dr. Youngjin Ye, Prof. Jeong Woo Han, Prof. Hyunjoo Lee, Dr. Jong Hyun Jang, and Prof. Jinwoo Lee. 2019. Investigation of the Support Effect in Atomically Dispersed Pt on WO3−x for Utilization of Pt in the Hydrogen Evolution Reaction. International Edition of Angewandte Chemie. Volume No. 58. Issue No. 45. 6 pages. https://doi.org/10.1002/anie.201908122
-ProfileProfessor Jinwoo LeeConvergence of Energy and Nano Science Laboratoryhttp://cens.kaist.ac.kr
Department of Chemical and Biomolecular EngineeringKAIST
2019.10.28 View 22827
Tungsten Suboxide Improves the Efficiency of Platinum in Hydrogen Production
< PhD Candidate Jinkyu Park and Professor Jinwoo Lee >
Researchers presented a new strategy for enhancing catalytic activity using tungsten suboxide as a single-atom catalyst (SAC). This strategy, which significantly improves hydrogen evolution reaction (HER) in metal platinum (pt) by 16.3 times, sheds light on the development of new electrochemical catalyst technologies.
Hydrogen has been touted as a promising alternative to fossil fuels. However, most of the conventional industrial hydrogen production methods come with environmental issues, releasing significant amounts of carbon dioxide and greenhouse gases.
Electrochemical water splitting is considered a potential approach for clean hydrogen production. Pt is one of the most commonly used catalysts to improve HER performance in electrochemical water splitting, but the high cost and scarcity of Pt remain key obstacles to mass commercial applications.
SACs, where all metal species are individually dispersed on a desired support material, have been identified as one way to reduce the amount of Pt usage, as they offer the maximum number of surface exposed Pt atoms.
Inspired by earlier studies, which mainly focused on SACs supported by carbon-based materials, a KAIST research team led by Professor Jinwoo Lee from the Department of Chemical and Biomolecular Engineering investigated the influence of support materials on the performance of SACs.
Professor Lee and his researchers suggested mesoporous tungsten suboxide as a new support material for atomically dispersed Pt, as this was expected to provide high electronic conductivity and have a synergetic effect with Pt.
They compared the performance of single-atom Pt supported by carbon and tungsten suboxide respectively. The results revealed that the support effect occurred with tungsten suboxide, in which the mass activity of a single-atom Pt supported by tungsten suboxide was 2.1 times greater than that of single-atom Pt supported by carbon, and 16.3 times higher than that of Pt nanoparticles supported by carbon.
The team indicated a change in the electronic structure of Pt via charge transfer from tungsten suboxide to Pt. This phenomenon was reported as a result of strong metal-support interaction between Pt and tungsten suboxide.
HER performance can be improved not only by changing the electronic structure of the supported metal, but also by inducing another support effect, the spillover effect, the research group reported. Hydrogen spillover is a phenomenon where adsorbed hydrogen migrates from one surface to another, and it occurs more easily as the Pt size becomes smaller.
The researchers compared the performance of single-atom Pt and Pt nanoparticles supported by tungsten suboxide. The single-atom Pt supported by tungsten suboxide exhibited a higher degree of hydrogen spillover phenomenon, which enhanced the Pt mass activity for hydrogen evolution up to 10.7 times compared to Pt nanoparticles supported by tungsten suboxide.
Professor Lee said, “Choosing the right support material is important for improving electrocatalysis in hydrogen production. The tungsten suboxide catalyst we used to support Pt in our study implies that interactions between the well-matched metal and support can drastically enhance the efficiency of the process.”
This research was supported by the Ministry of Science and ICT and introduced in the International Edition of the German journal Angewandte Chemie.
Figure. Schematic representation of hydrogen evolution reaction (HER) of pseudo single-atom Pt supported by tungsten suboxide
-Publication
Jinkyu Park, Dr. Seonggyu Lee, Hee-Eun Kim, Ara Cho, Seongbeen Kim, Dr. Youngjin Ye, Prof. Jeong Woo Han, Prof. Hyunjoo Lee, Dr. Jong Hyun Jang, and Prof. Jinwoo Lee. 2019. Investigation of the Support Effect in Atomically Dispersed Pt on WO3−x for Utilization of Pt in the Hydrogen Evolution Reaction. International Edition of Angewandte Chemie. Volume No. 58. Issue No. 45. 6 pages. https://doi.org/10.1002/anie.201908122
-ProfileProfessor Jinwoo LeeConvergence of Energy and Nano Science Laboratoryhttp://cens.kaist.ac.kr
Department of Chemical and Biomolecular EngineeringKAIST
2019.10.28 View 22827 -
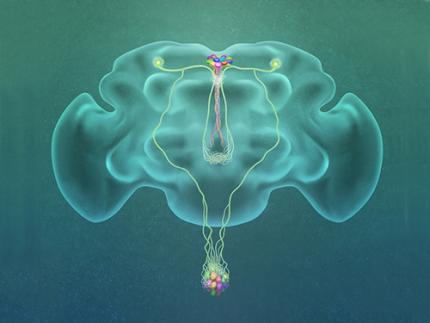 A Single, Master Switch for Sugar Levels?
When a fly eats sugar, a single brain cell sends simultaneous messages to stimulate one hormone and inhibit another to control glucose levels in the body. Further research into this control system with remarkable precision could shed light on the neural mechanisms of diabetes and obesity in humans .
A single neuron appears to monitor and control sugar levels in the fly body, according to research published this week in Nature. This new insight into the mechanisms in the fly brain that maintain a balance of two key hormones controlling glucose levels, insulin and glucagon, can provide a framework for understanding diabetes and obesity in humans.
Neurons that sense and respond to glucose were identified more than 50 years ago, but what they do in our body has remained unclear. Researchers at the Korea Advanced Institute of Science and Technology (KAIST) and New York University School of Medicine have now found a single “glucose-sensing neuron” that appears to be the master controller in Drosophila, the vinegar fly, for maintaining an ideal glucose balance, called homeostasis.
Professor Greg Seong-Bae Suh, Dr. Yangkyun Oh and colleagues identified a key neuron that is excited by glucose, which they called CN neuron. This CN neuron has a unique shape – it has an axon (which is used to transmit information to downstream cells) that is bifurcated. One branch projects to insulin-producing cells, and sends a signal triggering the secretion of the insulin equivalent in flies. The other branch projects to glucagon-producing cells and sends a signal inhibiting the secretion of the glucagon equivalent.
When flies consume food, the levels of glucose in their body increase; this excites the CN neuron, which fires the simultaneous signals to stimulate insulin and inhibit glucagon secretion, thereby maintaining the appropriate balance between the hormones and sugar in the blood. The researchers were able to see this happening in the brain in real time by using a combination of cutting-edge fluorescent calcium imaging technology, as well as measuring hormone and sugar levels and applying highly sophisticated molecular genetic techniques.
When flies were not fed, however, the researchers observed a reduction in the activity of CN neuron, a reduction in insulin secretion and an increase in glucagon secretion. These findings indicate that these key hormones are under the direct control of the glucose-sensing neuron. Furthermore, when they silenced the CN neuron rendering dysfunctional CN neuron in flies, these animals experienced an imbalance, resulting in hyperglycemia – high levels of sugars in the blood, similar to what is observed in diabetes in humans. This further suggests that the CN neuron is critical to maintaining glucose homeostasis in animals.
While further research is required to investigate this process in humans, Suh notes this is a significant step forward in the fields of both neurobiology and endocrinology.
“This work lays the foundation for translational research to better understand how this delicate regulatory process is affected by diabetes, obesity, excessive nutrition and diets high in sugar,” Suh said.
Profile: Greg Seong-Bae Suh
seongbaesuh@kaist.ac.kr
Professor Department of Biological Sciences
KAIST
(Figure: A single glucose-excited CN neuron extends bifurcated axonal branches,
one of which innervates insulin producing cells and stimulates their activity an the other axonal branch projects to glucagon producing cells and inhibits their activity.)
2019.10.24 View 18696
A Single, Master Switch for Sugar Levels?
When a fly eats sugar, a single brain cell sends simultaneous messages to stimulate one hormone and inhibit another to control glucose levels in the body. Further research into this control system with remarkable precision could shed light on the neural mechanisms of diabetes and obesity in humans .
A single neuron appears to monitor and control sugar levels in the fly body, according to research published this week in Nature. This new insight into the mechanisms in the fly brain that maintain a balance of two key hormones controlling glucose levels, insulin and glucagon, can provide a framework for understanding diabetes and obesity in humans.
Neurons that sense and respond to glucose were identified more than 50 years ago, but what they do in our body has remained unclear. Researchers at the Korea Advanced Institute of Science and Technology (KAIST) and New York University School of Medicine have now found a single “glucose-sensing neuron” that appears to be the master controller in Drosophila, the vinegar fly, for maintaining an ideal glucose balance, called homeostasis.
Professor Greg Seong-Bae Suh, Dr. Yangkyun Oh and colleagues identified a key neuron that is excited by glucose, which they called CN neuron. This CN neuron has a unique shape – it has an axon (which is used to transmit information to downstream cells) that is bifurcated. One branch projects to insulin-producing cells, and sends a signal triggering the secretion of the insulin equivalent in flies. The other branch projects to glucagon-producing cells and sends a signal inhibiting the secretion of the glucagon equivalent.
When flies consume food, the levels of glucose in their body increase; this excites the CN neuron, which fires the simultaneous signals to stimulate insulin and inhibit glucagon secretion, thereby maintaining the appropriate balance between the hormones and sugar in the blood. The researchers were able to see this happening in the brain in real time by using a combination of cutting-edge fluorescent calcium imaging technology, as well as measuring hormone and sugar levels and applying highly sophisticated molecular genetic techniques.
When flies were not fed, however, the researchers observed a reduction in the activity of CN neuron, a reduction in insulin secretion and an increase in glucagon secretion. These findings indicate that these key hormones are under the direct control of the glucose-sensing neuron. Furthermore, when they silenced the CN neuron rendering dysfunctional CN neuron in flies, these animals experienced an imbalance, resulting in hyperglycemia – high levels of sugars in the blood, similar to what is observed in diabetes in humans. This further suggests that the CN neuron is critical to maintaining glucose homeostasis in animals.
While further research is required to investigate this process in humans, Suh notes this is a significant step forward in the fields of both neurobiology and endocrinology.
“This work lays the foundation for translational research to better understand how this delicate regulatory process is affected by diabetes, obesity, excessive nutrition and diets high in sugar,” Suh said.
Profile: Greg Seong-Bae Suh
seongbaesuh@kaist.ac.kr
Professor Department of Biological Sciences
KAIST
(Figure: A single glucose-excited CN neuron extends bifurcated axonal branches,
one of which innervates insulin producing cells and stimulates their activity an the other axonal branch projects to glucagon producing cells and inhibits their activity.)
2019.10.24 View 18696 -
 A Mathematical Model Reveals Long-Distance Cell Communication Mechanism
How can tens of thousands of people in a large football stadium all clap together with the same beat even though they can only hear the people near them clapping?
A combination of a partial differential equation and a synthetic circuit in microbes answers this question. An interdisciplinary collaborative team of Professor Jae Kyoung Kim at KAIST, Professor Krešimir Josić at the University of Houston, and Professor Matt Bennett at Rice University has identified how a large community can communicate with each other almost simultaneously even with very short distance signaling. The research was reported at Nature Chemical Biology.
Cells often communicate using signaling molecules, which can travel only a short distance. Nevertheless, the cells can also communicate over large distances to spur collective action. The team revealed a cell communication mechanism that quickly forms a network of local interactions to spur collective action, even in large communities.
The research team used an engineered transcriptional circuit of combined positive and negative feedback loops in E. coli, which can periodically release two types of signaling molecules: activator and repressor. As the signaling molecules travel over a short distance, cells can only talk to their nearest neighbors. However, cell communities synchronize oscillatory gene expression in spatially extended systems as long as the transcriptional circuit contains a positive feedback loop for the activator.
Professor Kim said that analyzing and understanding such high-dimensional dynamics was extremely difficult. He explained, “That’s why we used high-dimensional partial differential equation to describe the system based on the interactions among various types of molecules.” Surprisingly, the mathematical model accurately simulates the synthesis of the signaling molecules in the cell and their spatial diffusion throughout the chamber and their effect on neighboring cells.
The team simplified the high-dimensional system into a one-dimensional orbit, noting that the system repeats periodically. This allowed them to discover that cells can make one voice when they lowered their own voice and listened to the others. “It turns out the positive feedback loop reduces the distance between moving points and finally makes them move all together. That’s why you clap louder when you hear applause from nearby neighbors and everyone eventually claps together at almost the same time,” said Professor Kim.
Professor Kim added, “Math is a powerful as it simplifies complex thing so that we can find an essential underlying property. This finding would not have been possible without the simplification of complex systems using mathematics."
The National Institutes of Health, the National Science Foundation, the Robert A. Welch Foundation, the Hamill Foundation, the National Research Foundation of Korea, and the T.J. Park Science Fellowship of POSCO supported the research.
(Figure: Complex molecular interactions among microbial consortia is simplified as interactions among points on a limit cycle (right).)
2019.10.15 View 29429
A Mathematical Model Reveals Long-Distance Cell Communication Mechanism
How can tens of thousands of people in a large football stadium all clap together with the same beat even though they can only hear the people near them clapping?
A combination of a partial differential equation and a synthetic circuit in microbes answers this question. An interdisciplinary collaborative team of Professor Jae Kyoung Kim at KAIST, Professor Krešimir Josić at the University of Houston, and Professor Matt Bennett at Rice University has identified how a large community can communicate with each other almost simultaneously even with very short distance signaling. The research was reported at Nature Chemical Biology.
Cells often communicate using signaling molecules, which can travel only a short distance. Nevertheless, the cells can also communicate over large distances to spur collective action. The team revealed a cell communication mechanism that quickly forms a network of local interactions to spur collective action, even in large communities.
The research team used an engineered transcriptional circuit of combined positive and negative feedback loops in E. coli, which can periodically release two types of signaling molecules: activator and repressor. As the signaling molecules travel over a short distance, cells can only talk to their nearest neighbors. However, cell communities synchronize oscillatory gene expression in spatially extended systems as long as the transcriptional circuit contains a positive feedback loop for the activator.
Professor Kim said that analyzing and understanding such high-dimensional dynamics was extremely difficult. He explained, “That’s why we used high-dimensional partial differential equation to describe the system based on the interactions among various types of molecules.” Surprisingly, the mathematical model accurately simulates the synthesis of the signaling molecules in the cell and their spatial diffusion throughout the chamber and their effect on neighboring cells.
The team simplified the high-dimensional system into a one-dimensional orbit, noting that the system repeats periodically. This allowed them to discover that cells can make one voice when they lowered their own voice and listened to the others. “It turns out the positive feedback loop reduces the distance between moving points and finally makes them move all together. That’s why you clap louder when you hear applause from nearby neighbors and everyone eventually claps together at almost the same time,” said Professor Kim.
Professor Kim added, “Math is a powerful as it simplifies complex thing so that we can find an essential underlying property. This finding would not have been possible without the simplification of complex systems using mathematics."
The National Institutes of Health, the National Science Foundation, the Robert A. Welch Foundation, the Hamill Foundation, the National Research Foundation of Korea, and the T.J. Park Science Fellowship of POSCO supported the research.
(Figure: Complex molecular interactions among microbial consortia is simplified as interactions among points on a limit cycle (right).)
2019.10.15 View 29429 -
 Two Professors Recognized for the National R&D Excellence 100
< Professor Haeng-Ki Lee (left) and Professor Jeong-Ho Lee (right) >
Two KAIST professors were listed among the 2019 National R&D Excellence 100 announced by the Ministry of Science and ICT and the Korea Institute of S&T Evaluation and Planning.
Professor Haeng-Ki Lee from the Department of Civil and Environmental Engineering was recognized in the field of mechanics and materials for his research on developing new construction materials through the convergence of nano- and biotechnologies.
In the field of life and marine science, Professor Jeong-Ho Lee from the Graduate School of Medical Science and Engineering was lauded for his research of diagnostic tools and therapies for glioblastoma and pediatric brain tumors.
A certificate from the Minister of Ministry of Science and ICT will be conferred to these two professors, and their names will be inscribed on a special 2019 National R&D Excellence 100 plaque to celebrate their achievements. The professors will also be given privileges during the process of new R&D project selection.
(END)
2019.10.15 View 13622
Two Professors Recognized for the National R&D Excellence 100
< Professor Haeng-Ki Lee (left) and Professor Jeong-Ho Lee (right) >
Two KAIST professors were listed among the 2019 National R&D Excellence 100 announced by the Ministry of Science and ICT and the Korea Institute of S&T Evaluation and Planning.
Professor Haeng-Ki Lee from the Department of Civil and Environmental Engineering was recognized in the field of mechanics and materials for his research on developing new construction materials through the convergence of nano- and biotechnologies.
In the field of life and marine science, Professor Jeong-Ho Lee from the Graduate School of Medical Science and Engineering was lauded for his research of diagnostic tools and therapies for glioblastoma and pediatric brain tumors.
A certificate from the Minister of Ministry of Science and ICT will be conferred to these two professors, and their names will be inscribed on a special 2019 National R&D Excellence 100 plaque to celebrate their achievements. The professors will also be given privileges during the process of new R&D project selection.
(END)
2019.10.15 View 13622