renewable+resource
-
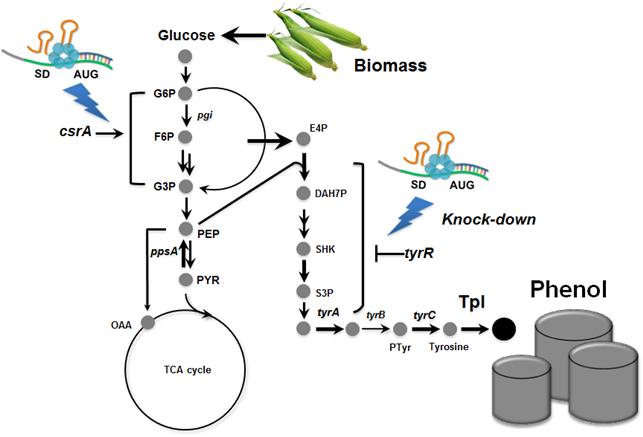 Metabolically engineered E. coli producing phenol
Many chemicals we use in everyday life are derived from fossil resources. Due to the increasing concerns on the use of fossil resources, there has been much interest in producing chemicals from renewable resources through biotechnology.
Phenol is an important commodity chemical, and is a starting material for the production of numerous industrial chemicals and polymers, including bisphenol A and phenolic resins, and others. At present, the production of phenol entirely depends on the chemical synthesis from benzene, and its annual production exceeds 8 million tons worldwide. Microbial production of phenol seems to be a non-viable process considering the high toxicity of phenol to the cell.
In the paper published online in Biotechnology Journal, a Korean research team led by Distinguished Professor Sang Yup Lee at the Department of Chemical and Biomolecular Engineering from the Korea Advanced Institute of Science and Technology (KAIST) reported the successful development of an engineered Escherichia coli (E. coli) strain which can produce phenol from glucose. E. coli has been a workhorse for biological production of various value-added compounds such as succinic acid and 1,4-butanediol in industrial scale. However, due to its low tolerance to phenol, E. coli was not considered a viable host strain for the biological production of phenol.
Professor Lee"s team, a leading research group in metabolic engineering, noted the genetic and physiological differences of various E. coli strains and investigated 18 different E. coli strains with respect to phenol tolerance and engineered all of the 18 strains simultaneously. If the traditional genetic engineering methods were used, this work would have taken years to do. To overcome this challenge, the research team used synthetic small RNA (sRNA) technology they recently developed (Nature Biotechnology, vol 31, pp 170-174, 2013). The sRNA technology allowed the team to screen 18 E. coli strains with respect to the phenol tolerance, and the activities of the metabolic pathway and enzyme involved in the production of phenol. The research team also metabolically engineered the E. coli strains to increase carbon flux toward phenol and finally generated an engineered E. coli strain which can produce phenol from glucose.
Furthermore, the team developed a biphasic extractive fermentation process to minimize the toxicity of phenol to E. coli cells. Glycerol tributyrate was found to have low toxicity to E. coli and allowed efficient extraction of phenol from the culture broth. Through the biphasic fed-batch fermentation using glycerol tributyrate as an in situ extractant, the final engineered E. coli strain produced phenol to the highest titer and productivity reported (3.8 g/L and 0.18 g/L/h, respectively). The strategy used for the strain development and the fermentation process will serve as a framework for metabolic engineering of microorganisms for the production of toxic chemicals from renewable resources.
This work was supported by the Intelligent Synthetic Biology Center through the Global Frontier Project (2011-0031963) of the Ministry of Science, ICT & Future Planning through the National Research Foundation of Korea.
Process of Phenol Production
2013.11.05 View 11458
Metabolically engineered E. coli producing phenol
Many chemicals we use in everyday life are derived from fossil resources. Due to the increasing concerns on the use of fossil resources, there has been much interest in producing chemicals from renewable resources through biotechnology.
Phenol is an important commodity chemical, and is a starting material for the production of numerous industrial chemicals and polymers, including bisphenol A and phenolic resins, and others. At present, the production of phenol entirely depends on the chemical synthesis from benzene, and its annual production exceeds 8 million tons worldwide. Microbial production of phenol seems to be a non-viable process considering the high toxicity of phenol to the cell.
In the paper published online in Biotechnology Journal, a Korean research team led by Distinguished Professor Sang Yup Lee at the Department of Chemical and Biomolecular Engineering from the Korea Advanced Institute of Science and Technology (KAIST) reported the successful development of an engineered Escherichia coli (E. coli) strain which can produce phenol from glucose. E. coli has been a workhorse for biological production of various value-added compounds such as succinic acid and 1,4-butanediol in industrial scale. However, due to its low tolerance to phenol, E. coli was not considered a viable host strain for the biological production of phenol.
Professor Lee"s team, a leading research group in metabolic engineering, noted the genetic and physiological differences of various E. coli strains and investigated 18 different E. coli strains with respect to phenol tolerance and engineered all of the 18 strains simultaneously. If the traditional genetic engineering methods were used, this work would have taken years to do. To overcome this challenge, the research team used synthetic small RNA (sRNA) technology they recently developed (Nature Biotechnology, vol 31, pp 170-174, 2013). The sRNA technology allowed the team to screen 18 E. coli strains with respect to the phenol tolerance, and the activities of the metabolic pathway and enzyme involved in the production of phenol. The research team also metabolically engineered the E. coli strains to increase carbon flux toward phenol and finally generated an engineered E. coli strain which can produce phenol from glucose.
Furthermore, the team developed a biphasic extractive fermentation process to minimize the toxicity of phenol to E. coli cells. Glycerol tributyrate was found to have low toxicity to E. coli and allowed efficient extraction of phenol from the culture broth. Through the biphasic fed-batch fermentation using glycerol tributyrate as an in situ extractant, the final engineered E. coli strain produced phenol to the highest titer and productivity reported (3.8 g/L and 0.18 g/L/h, respectively). The strategy used for the strain development and the fermentation process will serve as a framework for metabolic engineering of microorganisms for the production of toxic chemicals from renewable resources.
This work was supported by the Intelligent Synthetic Biology Center through the Global Frontier Project (2011-0031963) of the Ministry of Science, ICT & Future Planning through the National Research Foundation of Korea.
Process of Phenol Production
2013.11.05 View 11458 -
 A powerful strategy for developing microbial cell factories by employing synthetic small RNAs
The current systems for the production of chemicals, fuels and materials heavily rely on the use of fossil resources. Due to the increasing concerns on climate change and other environmental problems, however, there has been much interest in developing biorefineries for the production of such chemicals, fuels and materials from renewable resources. For the biorefineries to be competitive with the traditional fossil resource-based refineries, development of high performance microorganisms is the most important as it will affect the overall economics of the process most significantly. Metabolic engineering, which can be defined as purposeful modification of cellular metabolic and regulatory networks with an aim to improve the production of a desired product, has been successfully employed to improve the performance of the cell. However, it is not trivial to engineer the cellular metabolism and regulatory circuits in the cell due to their high complexity.
In metabolic engineering, it is important to find the genes that need to be amplified and attenuated in order to increase the product formation rate while minimizing the production of undesirable byproducts. Gene knock-out experiments are often performed to delete those metabolic fluxes that will consequently result in the increase of the desired product formation. However, gene knock-out experiments require much effort and time to perform, and are difficult to do for a large number of genes. Furthermore, the gene knock-out experiments performed in one strain cannot be transferred to another organism and thus the whole experimental process has to be repeated. This is a big problem in developing a high performance microbial cell factory because it is required to find the best platform strain among many different strains. Therefore, researchers have been eager to develop a strategy that allows rapid identification of multiple genes to be attenuated in multiple strains at the same time.
A Korean research team led by Distinguished Professor Sang Yup Lee at the Department of Chemical and Biomolecular Engineering from the Korea Advanced Institute of Science and Technology (KAIST) reported the development of a strategy for efficiently developing microbial cell factories by employing synthetic small RNAs (sRNAs). They first reported the development of such system in Nature Biotechnology last February. This strategy of employing synthetic sRNAs in metabolic engineering has been receiving great interest worldwide as it allows easy, rapid, high-throughput, tunable, and un-doable knock-down of multiple genes in multiple strains at the same time. The research team published a paper online on August 8 as a cover page (September issue) in Nature Protocols, describing the detailed strategy and protocol to employ synthetic sRNAs for metabolic engineering.
In this paper, researchers described the detailed step-by-step protocol for synthetic sRNA-based gene expression control, including the sRNA design principles. Tailor-made synthetic sRNAs can be easily manipulated by using conventional gene cloning method. The use of synthetic sRNAs for gene expression regulation provides several advantages such as portability, conditionality, and tunability in high-throughput experiments. Plasmid-based synthetic sRNA expression system does not leave any scar on the chromosome, and can be easily transferred to many other host strains to be examined. Thus, the construction of libraries and examination of different host strains are much easier than the conventional hard-coded gene manipulation systems. Also, the expression of genes can be conditionally repressed by controlling the production of synthetic sRNAs. Synthetic sRNAs possessing different repression efficiencies make it possible to finely tune the gene expression levels as well. Furthermore, synthetic sRNAs allow knock-down of the expression of essential genes, which was not possible by conventional gene knock-out experiments.
Synthetic sRNAs can be utilized for diverse experiments where gene expression regulation is needed. One of promising applications is high-throughput screening of the target genes to be manipulated and multiple strains simultaneously to enhance the production of chemicals and materials of interest. Such simultaneous optimization of gene targets and strains has been one of the big challenges in metabolic engineering. Another application is to fine tune the expression of the screened genes for flux optimization, which would enhance chemical production further by balancing the flux between biomass formation and target chemical production. Synthetic sRNAs can also be applied to finely regulating genetic interactions in a circuit or network, which is essential in synthetic biology. Once a sRNA scaffold-harboring plasmid is constructed, tailor-made, synthetic sRNAs can be made within 3-4 days, followed by the desired application experiments.
Dr. Eytan Zlotorynski, an editor at Nature Protocols, said "This paper describes the detailed protocol for the design and applications of synthetic sRNA. The method, which has many advantages, is likely to become common practice, and prove useful for metabolic engineering and synthetic biology studies."
This paper published in Nature Protocols will be useful for all researchers in academia and industry who are interested in the use of synthetic sRNAs for fundamental and applied biological and biotechnological studies.
This work was supported by the Technology Development Program to Solve Climate Changes on Systems Metabolic Engineering for Biorefineries (NRF-2012-C1AAA001-2012M1A2A2026556) and the Intelligent Synthetic Biology Center through the Global Frontier Project (2011-0031963) of the Ministry of Science, ICT and Future Planning through the National Research Foundation of Korea.
2013.10.31 View 10971
A powerful strategy for developing microbial cell factories by employing synthetic small RNAs
The current systems for the production of chemicals, fuels and materials heavily rely on the use of fossil resources. Due to the increasing concerns on climate change and other environmental problems, however, there has been much interest in developing biorefineries for the production of such chemicals, fuels and materials from renewable resources. For the biorefineries to be competitive with the traditional fossil resource-based refineries, development of high performance microorganisms is the most important as it will affect the overall economics of the process most significantly. Metabolic engineering, which can be defined as purposeful modification of cellular metabolic and regulatory networks with an aim to improve the production of a desired product, has been successfully employed to improve the performance of the cell. However, it is not trivial to engineer the cellular metabolism and regulatory circuits in the cell due to their high complexity.
In metabolic engineering, it is important to find the genes that need to be amplified and attenuated in order to increase the product formation rate while minimizing the production of undesirable byproducts. Gene knock-out experiments are often performed to delete those metabolic fluxes that will consequently result in the increase of the desired product formation. However, gene knock-out experiments require much effort and time to perform, and are difficult to do for a large number of genes. Furthermore, the gene knock-out experiments performed in one strain cannot be transferred to another organism and thus the whole experimental process has to be repeated. This is a big problem in developing a high performance microbial cell factory because it is required to find the best platform strain among many different strains. Therefore, researchers have been eager to develop a strategy that allows rapid identification of multiple genes to be attenuated in multiple strains at the same time.
A Korean research team led by Distinguished Professor Sang Yup Lee at the Department of Chemical and Biomolecular Engineering from the Korea Advanced Institute of Science and Technology (KAIST) reported the development of a strategy for efficiently developing microbial cell factories by employing synthetic small RNAs (sRNAs). They first reported the development of such system in Nature Biotechnology last February. This strategy of employing synthetic sRNAs in metabolic engineering has been receiving great interest worldwide as it allows easy, rapid, high-throughput, tunable, and un-doable knock-down of multiple genes in multiple strains at the same time. The research team published a paper online on August 8 as a cover page (September issue) in Nature Protocols, describing the detailed strategy and protocol to employ synthetic sRNAs for metabolic engineering.
In this paper, researchers described the detailed step-by-step protocol for synthetic sRNA-based gene expression control, including the sRNA design principles. Tailor-made synthetic sRNAs can be easily manipulated by using conventional gene cloning method. The use of synthetic sRNAs for gene expression regulation provides several advantages such as portability, conditionality, and tunability in high-throughput experiments. Plasmid-based synthetic sRNA expression system does not leave any scar on the chromosome, and can be easily transferred to many other host strains to be examined. Thus, the construction of libraries and examination of different host strains are much easier than the conventional hard-coded gene manipulation systems. Also, the expression of genes can be conditionally repressed by controlling the production of synthetic sRNAs. Synthetic sRNAs possessing different repression efficiencies make it possible to finely tune the gene expression levels as well. Furthermore, synthetic sRNAs allow knock-down of the expression of essential genes, which was not possible by conventional gene knock-out experiments.
Synthetic sRNAs can be utilized for diverse experiments where gene expression regulation is needed. One of promising applications is high-throughput screening of the target genes to be manipulated and multiple strains simultaneously to enhance the production of chemicals and materials of interest. Such simultaneous optimization of gene targets and strains has been one of the big challenges in metabolic engineering. Another application is to fine tune the expression of the screened genes for flux optimization, which would enhance chemical production further by balancing the flux between biomass formation and target chemical production. Synthetic sRNAs can also be applied to finely regulating genetic interactions in a circuit or network, which is essential in synthetic biology. Once a sRNA scaffold-harboring plasmid is constructed, tailor-made, synthetic sRNAs can be made within 3-4 days, followed by the desired application experiments.
Dr. Eytan Zlotorynski, an editor at Nature Protocols, said "This paper describes the detailed protocol for the design and applications of synthetic sRNA. The method, which has many advantages, is likely to become common practice, and prove useful for metabolic engineering and synthetic biology studies."
This paper published in Nature Protocols will be useful for all researchers in academia and industry who are interested in the use of synthetic sRNAs for fundamental and applied biological and biotechnological studies.
This work was supported by the Technology Development Program to Solve Climate Changes on Systems Metabolic Engineering for Biorefineries (NRF-2012-C1AAA001-2012M1A2A2026556) and the Intelligent Synthetic Biology Center through the Global Frontier Project (2011-0031963) of the Ministry of Science, ICT and Future Planning through the National Research Foundation of Korea.
2013.10.31 View 10971