Nature+Chemical+Biology
-
 KAIST develops technology for selective RNA modification in living cells and animals
· A team led by Professor Won Do Heo from the Department of Biological Sciences, KAIST, has developed a pioneering technology that selectively acetylates specific RNA molecules in living cells and tissues.
· The platform uses RNA-targeting CRISPR tools in combination with RNA-modifying enzymes to chemically modify only the intended RNA.
· The method opens new possibilities for gene therapy by enabling precise control of disease-related RNA without affecting the rest of the transcriptome.
< Photo 1. (From left) Professor Won Do Heo and Jihwan Yu, a Ph.D. Candidate of the Department of Biological Sciences >
CRISPR-Cas13, a powerful RNA-targeting technology is gaining increasing attention as a next-generation gene therapy platform due to its precision and reduced side effects. Utilizing this system, researchers at KAIST have now developed the world’s first technology capable of selectively acetylating (chemically modifying) specific RNA molecules among countless transcripts within living cells. This breakthrough enables precise, programmable control of RNA function and is expected to open new avenues in RNA-based therapeutic development.
KAIST (President Kwang Hyung Lee) announced that a research team led by Professor Won Do Heo in the Department of Biological Sciences has recently developed a groundbreaking technology capable of selectively acetylating specific RNA molecules within the human body using the CRISPR-Cas13 system—an RNA-targeting platform gaining increasing attention in the fields of gene regulation and RNA-based therapeutics.
RNA molecules can undergo chemical modifications—the addition of specific chemical groups—which alter their function and behavior without changing the underlying nucleotide sequence. However, some of these modifications, a critical layer of post-transcriptional gene regulation, remain poorly understood. Among them, N4-acetylcytidine (ac4C) has been particularly enigmatic, with ongoing debate about its existence and function in human messenger RNA (mRNA), the RNA that encodes proteins.
To address this gap, the KAIST research team developed a targeted RNA acetylation system, named dCas13-eNAT10. This platform combines a catalytically inactive Cas13 enzyme (dCas13) that guides the system to specific RNA targets, with a hyperactive variant of the NAT10 enzyme (eNAT10), which performs RNA acetylation. This approach enables precise acetylation of only the desired RNA molecules among the vast pool of transcripts within the cell.
< Figure 1. Development of hyperactive variant eNAT10 through NAT10 protein engineering. By engineering the NAT10 protein, which performs RNA acetylation in human cells, based on its domain and structure, eNAT10 was developed, showing approximately a 3-fold increase in RNA acetylation activity compared to the wild-type enzyme. >
Using this system, the researchers demonstrated that guide RNAs could direct the dCas13-eNAT10 complex to acetylate specific RNA targets, and acetylation significantly increased protein expression from the modified mRNA. Moreover, the study revealed, for the first time, that RNA acetylation plays a role in intracellular RNA localization, facilitating the export of RNA from the nucleus to the cytoplasm—a critical step in gene expression regulation.
To validate its therapeutic potential, the team successfully delivered the targeted RNA acetylation system into the livers of live mice using adeno-associated virus (AAV), a commonly used gene therapy vector. This marks the first demonstration of in vivo RNA modification, extending the applicability of RNA chemical modification tools from cell culture models to living organisms.
< Figure 2. Acetylation of various RNA in cells using dCas13-eNAT10 fusion protein. Utilizing the CRISPR-Cas13 system, which can precisely target specific RNA through guide RNA, a dCas13-eNAT10 fusion protein was created, demonstrating its ability to specifically acetylate various endogenous RNA at different locations within cells. >
Professor Won Do Heo, who previously developed COVID-19 treatment technology using RNA gene scissors and technology to activate RNA gene scissors with light, stated, "Existing RNA chemical modification research faced difficulties in controlling specificity, temporality, and spatiality. However, this new technology allows selective acetylation of desired RNA, opening the door for accurate and detailed research into the functions of RNA acetylation." He added, "The RNA chemical modification technology developed in this study can be widely used as an RNA-based therapeutic agent and a tool for regulating RNA functions in living organisms in the future."
< Figure 3. In vivo delivery of targeted RNA acetylation system. The targeted RNA acetylation system was encoded in an AAV vector, commonly used in gene therapy, and delivered intravenously to adult mice, showing that target RNA in liver tissue was specifically acetylated according to the guide RNA. >
This research, with Ph.D. candidate Jihwan Yu from the Department of Biological Sciences at KAIST as the first author, was published in the journal Nature Chemical Biology on June 2, 2025. (Title: Programmable RNA acetylation with CRISPR-Cas13, Impact factor: 12.9, DOI: https://doi.org/10.1038/s41589-025-01922-3)
This research was supported by the Samsung Future Technology Foundation and the Bio & Medical Technology Development Program of the National Research Foundation of Korea.
2025.06.10 View 1187
KAIST develops technology for selective RNA modification in living cells and animals
· A team led by Professor Won Do Heo from the Department of Biological Sciences, KAIST, has developed a pioneering technology that selectively acetylates specific RNA molecules in living cells and tissues.
· The platform uses RNA-targeting CRISPR tools in combination with RNA-modifying enzymes to chemically modify only the intended RNA.
· The method opens new possibilities for gene therapy by enabling precise control of disease-related RNA without affecting the rest of the transcriptome.
< Photo 1. (From left) Professor Won Do Heo and Jihwan Yu, a Ph.D. Candidate of the Department of Biological Sciences >
CRISPR-Cas13, a powerful RNA-targeting technology is gaining increasing attention as a next-generation gene therapy platform due to its precision and reduced side effects. Utilizing this system, researchers at KAIST have now developed the world’s first technology capable of selectively acetylating (chemically modifying) specific RNA molecules among countless transcripts within living cells. This breakthrough enables precise, programmable control of RNA function and is expected to open new avenues in RNA-based therapeutic development.
KAIST (President Kwang Hyung Lee) announced that a research team led by Professor Won Do Heo in the Department of Biological Sciences has recently developed a groundbreaking technology capable of selectively acetylating specific RNA molecules within the human body using the CRISPR-Cas13 system—an RNA-targeting platform gaining increasing attention in the fields of gene regulation and RNA-based therapeutics.
RNA molecules can undergo chemical modifications—the addition of specific chemical groups—which alter their function and behavior without changing the underlying nucleotide sequence. However, some of these modifications, a critical layer of post-transcriptional gene regulation, remain poorly understood. Among them, N4-acetylcytidine (ac4C) has been particularly enigmatic, with ongoing debate about its existence and function in human messenger RNA (mRNA), the RNA that encodes proteins.
To address this gap, the KAIST research team developed a targeted RNA acetylation system, named dCas13-eNAT10. This platform combines a catalytically inactive Cas13 enzyme (dCas13) that guides the system to specific RNA targets, with a hyperactive variant of the NAT10 enzyme (eNAT10), which performs RNA acetylation. This approach enables precise acetylation of only the desired RNA molecules among the vast pool of transcripts within the cell.
< Figure 1. Development of hyperactive variant eNAT10 through NAT10 protein engineering. By engineering the NAT10 protein, which performs RNA acetylation in human cells, based on its domain and structure, eNAT10 was developed, showing approximately a 3-fold increase in RNA acetylation activity compared to the wild-type enzyme. >
Using this system, the researchers demonstrated that guide RNAs could direct the dCas13-eNAT10 complex to acetylate specific RNA targets, and acetylation significantly increased protein expression from the modified mRNA. Moreover, the study revealed, for the first time, that RNA acetylation plays a role in intracellular RNA localization, facilitating the export of RNA from the nucleus to the cytoplasm—a critical step in gene expression regulation.
To validate its therapeutic potential, the team successfully delivered the targeted RNA acetylation system into the livers of live mice using adeno-associated virus (AAV), a commonly used gene therapy vector. This marks the first demonstration of in vivo RNA modification, extending the applicability of RNA chemical modification tools from cell culture models to living organisms.
< Figure 2. Acetylation of various RNA in cells using dCas13-eNAT10 fusion protein. Utilizing the CRISPR-Cas13 system, which can precisely target specific RNA through guide RNA, a dCas13-eNAT10 fusion protein was created, demonstrating its ability to specifically acetylate various endogenous RNA at different locations within cells. >
Professor Won Do Heo, who previously developed COVID-19 treatment technology using RNA gene scissors and technology to activate RNA gene scissors with light, stated, "Existing RNA chemical modification research faced difficulties in controlling specificity, temporality, and spatiality. However, this new technology allows selective acetylation of desired RNA, opening the door for accurate and detailed research into the functions of RNA acetylation." He added, "The RNA chemical modification technology developed in this study can be widely used as an RNA-based therapeutic agent and a tool for regulating RNA functions in living organisms in the future."
< Figure 3. In vivo delivery of targeted RNA acetylation system. The targeted RNA acetylation system was encoded in an AAV vector, commonly used in gene therapy, and delivered intravenously to adult mice, showing that target RNA in liver tissue was specifically acetylated according to the guide RNA. >
This research, with Ph.D. candidate Jihwan Yu from the Department of Biological Sciences at KAIST as the first author, was published in the journal Nature Chemical Biology on June 2, 2025. (Title: Programmable RNA acetylation with CRISPR-Cas13, Impact factor: 12.9, DOI: https://doi.org/10.1038/s41589-025-01922-3)
This research was supported by the Samsung Future Technology Foundation and the Bio & Medical Technology Development Program of the National Research Foundation of Korea.
2025.06.10 View 1187 -
 KAIST Develops Eco-Friendly, Nylon-Like Plastic Using Microorganisms
Poly(ester amide) amide is a next-generation material that combines the advantages of PET (polyester) and nylon (polyamide), two widely used plastics. However, it could only be produced from fossil fuels, which posed environmental concerns. Using microorganisms, KAIST researchers have successfully developed a new bio-based plastic to replace conventional plastic.
KAIST (represented by President Kwang Hyung Lee) announced on the 20th of March that a research team led by Distinguished Professor Sang Yup Lee from the Department of Chemical and Biomolecular Engineering has developed microbial strains through systems metabolic engineering to produce various eco-friendly, bio-based poly(ester amide)s. The team collaborated with researchers from the Korea Research Institute of Chemical Technology (KRICT, President Young-Kook Lee) to analyze and confirm the properties of the resulting plastic.
Professor Sang Yup Lee’s research team designed new metabolic pathways that do not naturally exist in microorganisms, and developed a platform microbial strain capable of producing nine different types of poly(ester amide)s, including poly(3-hydroxybutyrate-ran-3-aminopropionate) and poly(3-hydroxybutyrate-ran-4-aminobutyrate).
Using glucose derived from abundant biomass sources such as waste wood and weeds, the team successfully produced poly(ester amide)s in an eco-friendly manner. The researchers also confirmed the potential for industrial-scale production by demonstrating high production efficiency (54.57 g/L) using fed-batch fermentation of the engineered strain.
In collaboration with researchers Haemin Jeong and Jihoon Shin from KRICT, the KAIST team analyzed the properties of the bio-based plastic and found that it exhibited characteristics similar to high-density polyethylene (HDPE). This means the new plastic is not only eco-friendly but also strong and durable enough to replace conventional plastics.
The engineered strains and strategies developed in this study are expected to be useful not only for producing various poly(ester amide)s but also for constructing metabolic pathways for the biosynthesis of other types of polymers.
Professor Sang Yup Lee stated, “This study is the first to demonstrate the possibility of producing poly(ester amide)s (plastics) through a renewable bio-based chemical process rather than relying on the petroleum-based chemical industry. We plan to further enhance the production yield and efficiency through continued research.”
The study was published online on March 17 in the international journal Nature Chemical Biology.
·Title: Biosynthesis of poly(ester amide)s in engineered Escherichia coli
·DOI: 10.1038/s41589-025-01842-2
·Authors: A total of seven authors including Tong Un Chae (KAIST, first author), So Young Choi (KAIST, second author), Da-Hee Ahn (KAIST, third author), Woo Dae Jang (KAIST, fourth author), Haemin Jeong (KRICT, fifth author), Jihoon Shin (KRICT, sixth author), and Sang Yup Lee (KAIST, corresponding author).
This research was supported by the Ministry of Science and ICT (MSIT) under the Eco-Friendly Chemical Technology Development Project as part of the "Next-Generation Biorefinery Technology Development to Lead the Bio-Chemical Industry" initiative (project led by Distinguished Professor Sang Yup Lee at KAIST).
2025.03.24 View 4940
KAIST Develops Eco-Friendly, Nylon-Like Plastic Using Microorganisms
Poly(ester amide) amide is a next-generation material that combines the advantages of PET (polyester) and nylon (polyamide), two widely used plastics. However, it could only be produced from fossil fuels, which posed environmental concerns. Using microorganisms, KAIST researchers have successfully developed a new bio-based plastic to replace conventional plastic.
KAIST (represented by President Kwang Hyung Lee) announced on the 20th of March that a research team led by Distinguished Professor Sang Yup Lee from the Department of Chemical and Biomolecular Engineering has developed microbial strains through systems metabolic engineering to produce various eco-friendly, bio-based poly(ester amide)s. The team collaborated with researchers from the Korea Research Institute of Chemical Technology (KRICT, President Young-Kook Lee) to analyze and confirm the properties of the resulting plastic.
Professor Sang Yup Lee’s research team designed new metabolic pathways that do not naturally exist in microorganisms, and developed a platform microbial strain capable of producing nine different types of poly(ester amide)s, including poly(3-hydroxybutyrate-ran-3-aminopropionate) and poly(3-hydroxybutyrate-ran-4-aminobutyrate).
Using glucose derived from abundant biomass sources such as waste wood and weeds, the team successfully produced poly(ester amide)s in an eco-friendly manner. The researchers also confirmed the potential for industrial-scale production by demonstrating high production efficiency (54.57 g/L) using fed-batch fermentation of the engineered strain.
In collaboration with researchers Haemin Jeong and Jihoon Shin from KRICT, the KAIST team analyzed the properties of the bio-based plastic and found that it exhibited characteristics similar to high-density polyethylene (HDPE). This means the new plastic is not only eco-friendly but also strong and durable enough to replace conventional plastics.
The engineered strains and strategies developed in this study are expected to be useful not only for producing various poly(ester amide)s but also for constructing metabolic pathways for the biosynthesis of other types of polymers.
Professor Sang Yup Lee stated, “This study is the first to demonstrate the possibility of producing poly(ester amide)s (plastics) through a renewable bio-based chemical process rather than relying on the petroleum-based chemical industry. We plan to further enhance the production yield and efficiency through continued research.”
The study was published online on March 17 in the international journal Nature Chemical Biology.
·Title: Biosynthesis of poly(ester amide)s in engineered Escherichia coli
·DOI: 10.1038/s41589-025-01842-2
·Authors: A total of seven authors including Tong Un Chae (KAIST, first author), So Young Choi (KAIST, second author), Da-Hee Ahn (KAIST, third author), Woo Dae Jang (KAIST, fourth author), Haemin Jeong (KRICT, fifth author), Jihoon Shin (KRICT, sixth author), and Sang Yup Lee (KAIST, corresponding author).
This research was supported by the Ministry of Science and ICT (MSIT) under the Eco-Friendly Chemical Technology Development Project as part of the "Next-Generation Biorefinery Technology Development to Lead the Bio-Chemical Industry" initiative (project led by Distinguished Professor Sang Yup Lee at KAIST).
2025.03.24 View 4940 -
 A Mathematical Model Reveals Long-Distance Cell Communication Mechanism
How can tens of thousands of people in a large football stadium all clap together with the same beat even though they can only hear the people near them clapping?
A combination of a partial differential equation and a synthetic circuit in microbes answers this question. An interdisciplinary collaborative team of Professor Jae Kyoung Kim at KAIST, Professor Krešimir Josić at the University of Houston, and Professor Matt Bennett at Rice University has identified how a large community can communicate with each other almost simultaneously even with very short distance signaling. The research was reported at Nature Chemical Biology.
Cells often communicate using signaling molecules, which can travel only a short distance. Nevertheless, the cells can also communicate over large distances to spur collective action. The team revealed a cell communication mechanism that quickly forms a network of local interactions to spur collective action, even in large communities.
The research team used an engineered transcriptional circuit of combined positive and negative feedback loops in E. coli, which can periodically release two types of signaling molecules: activator and repressor. As the signaling molecules travel over a short distance, cells can only talk to their nearest neighbors. However, cell communities synchronize oscillatory gene expression in spatially extended systems as long as the transcriptional circuit contains a positive feedback loop for the activator.
Professor Kim said that analyzing and understanding such high-dimensional dynamics was extremely difficult. He explained, “That’s why we used high-dimensional partial differential equation to describe the system based on the interactions among various types of molecules.” Surprisingly, the mathematical model accurately simulates the synthesis of the signaling molecules in the cell and their spatial diffusion throughout the chamber and their effect on neighboring cells.
The team simplified the high-dimensional system into a one-dimensional orbit, noting that the system repeats periodically. This allowed them to discover that cells can make one voice when they lowered their own voice and listened to the others. “It turns out the positive feedback loop reduces the distance between moving points and finally makes them move all together. That’s why you clap louder when you hear applause from nearby neighbors and everyone eventually claps together at almost the same time,” said Professor Kim.
Professor Kim added, “Math is a powerful as it simplifies complex thing so that we can find an essential underlying property. This finding would not have been possible without the simplification of complex systems using mathematics."
The National Institutes of Health, the National Science Foundation, the Robert A. Welch Foundation, the Hamill Foundation, the National Research Foundation of Korea, and the T.J. Park Science Fellowship of POSCO supported the research.
(Figure: Complex molecular interactions among microbial consortia is simplified as interactions among points on a limit cycle (right).)
2019.10.15 View 29711
A Mathematical Model Reveals Long-Distance Cell Communication Mechanism
How can tens of thousands of people in a large football stadium all clap together with the same beat even though they can only hear the people near them clapping?
A combination of a partial differential equation and a synthetic circuit in microbes answers this question. An interdisciplinary collaborative team of Professor Jae Kyoung Kim at KAIST, Professor Krešimir Josić at the University of Houston, and Professor Matt Bennett at Rice University has identified how a large community can communicate with each other almost simultaneously even with very short distance signaling. The research was reported at Nature Chemical Biology.
Cells often communicate using signaling molecules, which can travel only a short distance. Nevertheless, the cells can also communicate over large distances to spur collective action. The team revealed a cell communication mechanism that quickly forms a network of local interactions to spur collective action, even in large communities.
The research team used an engineered transcriptional circuit of combined positive and negative feedback loops in E. coli, which can periodically release two types of signaling molecules: activator and repressor. As the signaling molecules travel over a short distance, cells can only talk to their nearest neighbors. However, cell communities synchronize oscillatory gene expression in spatially extended systems as long as the transcriptional circuit contains a positive feedback loop for the activator.
Professor Kim said that analyzing and understanding such high-dimensional dynamics was extremely difficult. He explained, “That’s why we used high-dimensional partial differential equation to describe the system based on the interactions among various types of molecules.” Surprisingly, the mathematical model accurately simulates the synthesis of the signaling molecules in the cell and their spatial diffusion throughout the chamber and their effect on neighboring cells.
The team simplified the high-dimensional system into a one-dimensional orbit, noting that the system repeats periodically. This allowed them to discover that cells can make one voice when they lowered their own voice and listened to the others. “It turns out the positive feedback loop reduces the distance between moving points and finally makes them move all together. That’s why you clap louder when you hear applause from nearby neighbors and everyone eventually claps together at almost the same time,” said Professor Kim.
Professor Kim added, “Math is a powerful as it simplifies complex thing so that we can find an essential underlying property. This finding would not have been possible without the simplification of complex systems using mathematics."
The National Institutes of Health, the National Science Foundation, the Robert A. Welch Foundation, the Hamill Foundation, the National Research Foundation of Korea, and the T.J. Park Science Fellowship of POSCO supported the research.
(Figure: Complex molecular interactions among microbial consortia is simplified as interactions among points on a limit cycle (right).)
2019.10.15 View 29711 -
 Efficiently Producing Fatty Acids and Biofuels from Glucose
Researchers have presented a new strategy for efficiently producing fatty acids and biofuels that can transform glucose and oleaginous microorganisms into microbial diesel fuel, with one-step direct fermentative production.
The newly developed strain, created by Distinguished Professor Sang Yup Lee and his team, showed the highest efficiency in producing fatty acids and biodiesels ever reported. It will be expected to serve as a new platform to sustainably produce a wide array of fatty acid-based products from glucose and other carbon substrates.
Fossil fuels, which have long been energy resources for our daily lives, are now facing serious challenges: depletion of their reserves and their role in global warming. The production of sustainable bio-based renewable energy has emerged as an essential alternative and many studies to replace fossil fuels are underway. One of the representative examples is biodiesel. Currently, it is mainly being produced through the transesterification of vegetable oils or animal fats.
The research team engineered oleaginous microorganisms, Rhodococcus opacus, to produce fatty acids and their derivatives that can be used as biodiesel from glucose, one of the most abundant and cheap sugars derived from non-edible biomass.
Professor Lee’s team has already engineered Escherichia coli to produce short-chain hydrocarbons, which can be used as gasoline (published in Nature as the cover paper in 2013). However, the production efficiency of the short-chain hydrocarbons using E. coli (0.58 g/L) fell short of the levels required for commercialization.
To overcome these issues, the team employed oil-accumulating Rhodococcus opacus as a host strain in this study. First, the team optimized the cultivation conditions of Rhodococcus opacus to maximize the accumulation of oil (triacylglycerol), which serves as a precursor for the biosynthesis of fatty acids and their derivatives. Then, they systematically analyzed the metabolism of the strain and redesigned it to enable higher levels of fatty acids and two kinds of fatty acid-derived biodiesels (fatty acid ethyl esters and long-chain hydrocarbons) to be produced.
They found that the resulting strains produced 50.2, 21.3, and 5.2 g/L of fatty acids, fatty acid ethyl esters, and long-chain hydrocarbons, respectively. These are all the highest concentrations ever reported by microbial fermentations. It is expected that these strains can contribute to the future industrialization of microbial-based biodiesel production.
“This technology creates fatty acids and biodiesel with high efficiency by utilizing lignocellulose, one of the most abundant resources on the Earth, without depending on fossil fuels and vegetable or animal oils. This will provide new opportunities for oil and petroleum industries, which have long relied on fossil fuels, to turn to sustainable and eco-friendly biotechnologies,” said Professor Lee.
This paper titled “Engineering of an oleaginous bacterium for the production of fatty acids and fuels” was published in Nature Chemical Biology on June 17.
This work was supported by the Technology Development Program to Solve Climate Changes on Systems Metabolic Engineering for Biorefineries from the Ministry of Science and ICT through the National Research Foundation (NRF) of Korea (NRF-2012M1A2A2026556 and NRF-2012M1A2A2026557).
(Figure: Metabolic engineering for the production of free fatty acids (FFAs), fatty acid ethyl esters (FAEEs), and long-chain hydrocarbons (LCHCs) in Rhodococcus opacus PD630. Researchers have presented a new strategy for efficiently producing fatty acids and biofuels that can transform glucose and oleaginous microorganisms into microbial diesel fuel, with one-step direct fermentative production.)
# # #
Source:
Hye Mi Kim, Tong Un Chae, So Young Choi, Won Jun Kim and Sang Yup Lee. Engineering of an oleaginous bacterium for the production of fatty acids and fuels. Nature Chemical Biology ( https://www.nature.com/nchembio/ ) DOI: 10.1038/s41589-019-0295-5
Profile
Dr. Sang Yup Lee
leesy@kaist.ac.kr
Distinguished Professor at the Department of Chemical and Biomolecular Engineering
KAIST
2019.06.19 View 51434
Efficiently Producing Fatty Acids and Biofuels from Glucose
Researchers have presented a new strategy for efficiently producing fatty acids and biofuels that can transform glucose and oleaginous microorganisms into microbial diesel fuel, with one-step direct fermentative production.
The newly developed strain, created by Distinguished Professor Sang Yup Lee and his team, showed the highest efficiency in producing fatty acids and biodiesels ever reported. It will be expected to serve as a new platform to sustainably produce a wide array of fatty acid-based products from glucose and other carbon substrates.
Fossil fuels, which have long been energy resources for our daily lives, are now facing serious challenges: depletion of their reserves and their role in global warming. The production of sustainable bio-based renewable energy has emerged as an essential alternative and many studies to replace fossil fuels are underway. One of the representative examples is biodiesel. Currently, it is mainly being produced through the transesterification of vegetable oils or animal fats.
The research team engineered oleaginous microorganisms, Rhodococcus opacus, to produce fatty acids and their derivatives that can be used as biodiesel from glucose, one of the most abundant and cheap sugars derived from non-edible biomass.
Professor Lee’s team has already engineered Escherichia coli to produce short-chain hydrocarbons, which can be used as gasoline (published in Nature as the cover paper in 2013). However, the production efficiency of the short-chain hydrocarbons using E. coli (0.58 g/L) fell short of the levels required for commercialization.
To overcome these issues, the team employed oil-accumulating Rhodococcus opacus as a host strain in this study. First, the team optimized the cultivation conditions of Rhodococcus opacus to maximize the accumulation of oil (triacylglycerol), which serves as a precursor for the biosynthesis of fatty acids and their derivatives. Then, they systematically analyzed the metabolism of the strain and redesigned it to enable higher levels of fatty acids and two kinds of fatty acid-derived biodiesels (fatty acid ethyl esters and long-chain hydrocarbons) to be produced.
They found that the resulting strains produced 50.2, 21.3, and 5.2 g/L of fatty acids, fatty acid ethyl esters, and long-chain hydrocarbons, respectively. These are all the highest concentrations ever reported by microbial fermentations. It is expected that these strains can contribute to the future industrialization of microbial-based biodiesel production.
“This technology creates fatty acids and biodiesel with high efficiency by utilizing lignocellulose, one of the most abundant resources on the Earth, without depending on fossil fuels and vegetable or animal oils. This will provide new opportunities for oil and petroleum industries, which have long relied on fossil fuels, to turn to sustainable and eco-friendly biotechnologies,” said Professor Lee.
This paper titled “Engineering of an oleaginous bacterium for the production of fatty acids and fuels” was published in Nature Chemical Biology on June 17.
This work was supported by the Technology Development Program to Solve Climate Changes on Systems Metabolic Engineering for Biorefineries from the Ministry of Science and ICT through the National Research Foundation (NRF) of Korea (NRF-2012M1A2A2026556 and NRF-2012M1A2A2026557).
(Figure: Metabolic engineering for the production of free fatty acids (FFAs), fatty acid ethyl esters (FAEEs), and long-chain hydrocarbons (LCHCs) in Rhodococcus opacus PD630. Researchers have presented a new strategy for efficiently producing fatty acids and biofuels that can transform glucose and oleaginous microorganisms into microbial diesel fuel, with one-step direct fermentative production.)
# # #
Source:
Hye Mi Kim, Tong Un Chae, So Young Choi, Won Jun Kim and Sang Yup Lee. Engineering of an oleaginous bacterium for the production of fatty acids and fuels. Nature Chemical Biology ( https://www.nature.com/nchembio/ ) DOI: 10.1038/s41589-019-0295-5
Profile
Dr. Sang Yup Lee
leesy@kaist.ac.kr
Distinguished Professor at the Department of Chemical and Biomolecular Engineering
KAIST
2019.06.19 View 51434 -
 Ligand Recognition Mechanism of Protein Identified
Professor Hak-Sung Kim
-“Solved the 50 year old mystery of how protein recognises and binds to ligands”
- Exciting potential for understanding life phenomena and the further development of highly effective therapeutic agent development
KAIST’s Biological Science Department’s Professor Hak-Sung Kim, working in collaboration with Professor Sung-Chul Hong of Department of Physics, Seoul National University, has identified the mechanism of how the protein recognizes and binds to ligands within the human body.
The research findings were published in the online edition of Nature Chemical Biology (March 18), which is the most prestigious journal in the field of life science.
Since the research identified the mechanism, of which protein recognises and binds to ligands, it will take an essential role in understanding complex life phenomenon by understanding regulatory function of protein.
Also, ligand recognition of proteins is closely related to the cause of various diseases. Therefore the research team hopes to contribute to the development of highly effective treatments.
Ligands, well-known examples include nucleic acid and proteins, form the structure of an organism or are essential constituents with special functions such as information signalling.
In particular, the most important role of protein is recognising and binding to a particular ligand and hence regulating and maintaining life phenomena. The abnormal occurrence of an error in recognition of ligands may lead to various diseases.
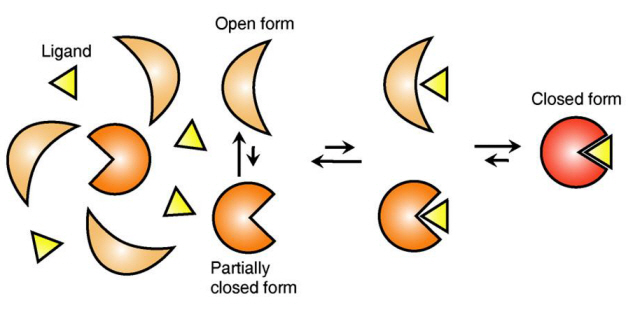
The research team focused on the repetition of change in protein structure from the most stable “open form” to a relatively unstable “partially closed form”.
Professor Kim’s team analysed the change in protein structure when binding to a ligand on a molecular level in real time to explain the ligand recognition mechanism.
The research findings showed that ligands prefer the most stable protein structure. The team was the first in the world to identify that ligands alter protein structure to the most stable, the lowest energy level, when it binds to the protein.
In addition, the team found that ligands bind to unstable partially-closed forms to change protein structure.
The existing models to explain ligand recognition mechanism of protein are “Induced Custom Model”, which involves change in protein structure in binding to ligands, and the “Structure Selection Model”, which argues that ligands select and recognise only the best protein structure out of many. The academic world considers that the team’s research findings have perfectly proved the models through experiments for the first time in the world.
Professor Kim explained, “In the presence of ligands, there exists a phenomenon where the speed of altering protein structure is changed. This phenomenon is analysed on a molecular level to prove ligand recognition mechanism of protein for the first time”. He also said, “The 50-year old mystery, that existed only as a hypothesis on biology textbooks and was thought never to be solved, has been confirmed through experiments for the first time.”
Figure 1: Proteins, with open and partially open form, recognising and binding to ligands.
Figure 2: Ligands temporarily bind to a stable protein structure, open form, which changes into the most stable structure, closed form. In addition, binding to partially closed form also changes protein structure to closed form.
2013.04.01 View 13367
Ligand Recognition Mechanism of Protein Identified
Professor Hak-Sung Kim
-“Solved the 50 year old mystery of how protein recognises and binds to ligands”
- Exciting potential for understanding life phenomena and the further development of highly effective therapeutic agent development
KAIST’s Biological Science Department’s Professor Hak-Sung Kim, working in collaboration with Professor Sung-Chul Hong of Department of Physics, Seoul National University, has identified the mechanism of how the protein recognizes and binds to ligands within the human body.
The research findings were published in the online edition of Nature Chemical Biology (March 18), which is the most prestigious journal in the field of life science.
Since the research identified the mechanism, of which protein recognises and binds to ligands, it will take an essential role in understanding complex life phenomenon by understanding regulatory function of protein.
Also, ligand recognition of proteins is closely related to the cause of various diseases. Therefore the research team hopes to contribute to the development of highly effective treatments.
Ligands, well-known examples include nucleic acid and proteins, form the structure of an organism or are essential constituents with special functions such as information signalling.
In particular, the most important role of protein is recognising and binding to a particular ligand and hence regulating and maintaining life phenomena. The abnormal occurrence of an error in recognition of ligands may lead to various diseases.
The research team focused on the repetition of change in protein structure from the most stable “open form” to a relatively unstable “partially closed form”.
Professor Kim’s team analysed the change in protein structure when binding to a ligand on a molecular level in real time to explain the ligand recognition mechanism.
The research findings showed that ligands prefer the most stable protein structure. The team was the first in the world to identify that ligands alter protein structure to the most stable, the lowest energy level, when it binds to the protein.
In addition, the team found that ligands bind to unstable partially-closed forms to change protein structure.
The existing models to explain ligand recognition mechanism of protein are “Induced Custom Model”, which involves change in protein structure in binding to ligands, and the “Structure Selection Model”, which argues that ligands select and recognise only the best protein structure out of many. The academic world considers that the team’s research findings have perfectly proved the models through experiments for the first time in the world.
Professor Kim explained, “In the presence of ligands, there exists a phenomenon where the speed of altering protein structure is changed. This phenomenon is analysed on a molecular level to prove ligand recognition mechanism of protein for the first time”. He also said, “The 50-year old mystery, that existed only as a hypothesis on biology textbooks and was thought never to be solved, has been confirmed through experiments for the first time.”
Figure 1: Proteins, with open and partially open form, recognising and binding to ligands.
Figure 2: Ligands temporarily bind to a stable protein structure, open form, which changes into the most stable structure, closed form. In addition, binding to partially closed form also changes protein structure to closed form.
2013.04.01 View 13367