Nature
-
 Finding Human Thermal Comfort with a Watch-type Sweat Rate Sensor
(from left: Professor Young-Ho Cho and Researcher SungHyun Yoon)
KAIST developed a watch-type sweat rate sensor. This subminiature device can detect human thermal comfort accurately and steadily by measuring an individual’s sweat rate.
It is natural to sweat more in the summer and less in the winter; however, an individual’s sweat rate may vary in a given environment. Therefore, sweat can be an excellent proxy for sensing core body temperature.
Conventional sweat rate sensors using natural ventilation require bulky external devices, such as pumps and ice condensers. They are usually for physiological experiments, hence they need a manual ventilation process or high power, bulky thermos-pneumatic actuators to lift sweat rate detection chambers above skin for continuous measurement. There is also a small sweat rate sensor, but it needs a long recovery period.
To overcome these problems, Professor Young-Ho Cho and his team from the Department of Bio and Brain Engineering developed a lightweight, watch-type sweat sensor. The team integrated miniaturized thermos-pneumatic actuators for automatic natural ventilation, which allows sweat to be measured continuously.
This watch-type sensor measures sweat rate with the humidity rising rate when the chamber is closed during skin contact. Since the team integrated thermos-pneumatic actuators, the chamber no longer needs to be separated manually from skin after each measurement in order for the chamber to ventilate the collected humidity.
Moreover, this sensor is wind-resistant enough to be used for portable and wearable devices. The team identified that the sensor operates steadily with air velocity ranging up to 1.5m/s, equivalent to the average human walking speed.
Although this subminiature sensor (35mm x 25mm) only weighs 30 grams, it operates continuously for more than four hours using the conventional wrist watch batteries.
The team plans to utilize this technology for developing a new concept of cognitive air-conditioning systems recognizing Human thermal status directly; while the conventional air-conditioning systems measuring air temperature and humidity.
Professor Cho said, “Our sensor for human thermal comfort monitoring can be applied to customized or smart air conditioners. Furthermore, there will be more demands for both physical and mental healthcare, hence this technology will serve as a new platform for personalized emotional communion between humans and devices.”
This research, led by researchers Jai Kyoung Sim and SungHyun Yoon, was published in Scientific Reports on January 19, 2018.
Figure1. The fabricated watch-type sweat rate sensor for human thermal comfort monitoring
Figure 2. Views of the watch-type sweat rate sensor
Figure 3. Operation of the watch-type sweat rate sensor
2018.02.08 View 9296
Finding Human Thermal Comfort with a Watch-type Sweat Rate Sensor
(from left: Professor Young-Ho Cho and Researcher SungHyun Yoon)
KAIST developed a watch-type sweat rate sensor. This subminiature device can detect human thermal comfort accurately and steadily by measuring an individual’s sweat rate.
It is natural to sweat more in the summer and less in the winter; however, an individual’s sweat rate may vary in a given environment. Therefore, sweat can be an excellent proxy for sensing core body temperature.
Conventional sweat rate sensors using natural ventilation require bulky external devices, such as pumps and ice condensers. They are usually for physiological experiments, hence they need a manual ventilation process or high power, bulky thermos-pneumatic actuators to lift sweat rate detection chambers above skin for continuous measurement. There is also a small sweat rate sensor, but it needs a long recovery period.
To overcome these problems, Professor Young-Ho Cho and his team from the Department of Bio and Brain Engineering developed a lightweight, watch-type sweat sensor. The team integrated miniaturized thermos-pneumatic actuators for automatic natural ventilation, which allows sweat to be measured continuously.
This watch-type sensor measures sweat rate with the humidity rising rate when the chamber is closed during skin contact. Since the team integrated thermos-pneumatic actuators, the chamber no longer needs to be separated manually from skin after each measurement in order for the chamber to ventilate the collected humidity.
Moreover, this sensor is wind-resistant enough to be used for portable and wearable devices. The team identified that the sensor operates steadily with air velocity ranging up to 1.5m/s, equivalent to the average human walking speed.
Although this subminiature sensor (35mm x 25mm) only weighs 30 grams, it operates continuously for more than four hours using the conventional wrist watch batteries.
The team plans to utilize this technology for developing a new concept of cognitive air-conditioning systems recognizing Human thermal status directly; while the conventional air-conditioning systems measuring air temperature and humidity.
Professor Cho said, “Our sensor for human thermal comfort monitoring can be applied to customized or smart air conditioners. Furthermore, there will be more demands for both physical and mental healthcare, hence this technology will serve as a new platform for personalized emotional communion between humans and devices.”
This research, led by researchers Jai Kyoung Sim and SungHyun Yoon, was published in Scientific Reports on January 19, 2018.
Figure1. The fabricated watch-type sweat rate sensor for human thermal comfort monitoring
Figure 2. Views of the watch-type sweat rate sensor
Figure 3. Operation of the watch-type sweat rate sensor
2018.02.08 View 9296 -
 New Arylation Inducing Reaction Developed
(Professor Chang(left) and Professor Baik)
KAIST researchers have identified a reaction mechanism that selectively introduces aryl groups at the desired position of a molecule at room temperature. A team, co-led by Professor Sukbok Chang and Mu-Hyun Baik of the Department of Chemistry, used an iridium catalyst for the reaction. The team also proved that the reaction proceeds by an unusual mechanism by employing computer simulations that were substantiated with targeted experimental probes.
Hydrocarbon is an omnipresent material in nature. But its low reactivity makes it difficult to process to value-added products at the room temperature. Thus, designing catalysts that can accelerate the reaction remains an important challenge in chemistry.
In particular, since most chemicals used in medicine, pharmacy, or material chemistry contain aryl groups, an effective reaction to selectively introduce the aryl group has been an area of intensive research in organic chemistry.
In order to introduce an aryl group into stable carbon-hydrogen (C-H) bond, activation of the C-H bond with a halogen atom or organic metal is required prior to the introduction of the aryl group, or C-H functionalization directly on C-H bond is needed. Direct functionalization is more effective and economical, but most reactions require harsh reaction conditions such as high temperature or excess additives. And adding the aryl fragment selectively to only one among the many possible sites in the molecule is difficult. The new catalyst developed by these KAIST researchers is highly selective.
This work is the latest example of a successful teamwork between experimental and theoretical research groups: Computer simulations revealed that traditional approaches to arylation required high energies because the intermediates produced during the reaction are too low in energy. Based on this insight, the researchers thought of changing the character of the intermediate by oxidizing it, which was predicted to be a great way of increasing the reactivity of the catalyst. Subsequent experimental work showed that this design strategy is highly effective resulting in unprecedented chemical transformations.
Professor Chang said, “We have been able to carry out location-selective arylation at room temperature, as well as identifying a new reaction pathway, different from the conventionally suggested mechanism.” He continued, “This research is significant for identifying the reaction pathway and developing a novel selective reaction method that does not require high temperature or additives based on the mechanistic understanding. This work is a triumph of rational design, rather than fortuitous discovery.” The research findings were published online in Nature Chemistry on December 11, 2017.
(Figure 1: X-ray crystal structure transmetallation intermediate)
(Figure 2: Correlation between oxidation state of intermediate and energy barrier required for reductive elimination of intermediate as calculated using density function from computational chemistry )
(Figure 3: Arylation mechanism using iridium catalyst as suggested by the research team)
2018.01.11 View 6382
New Arylation Inducing Reaction Developed
(Professor Chang(left) and Professor Baik)
KAIST researchers have identified a reaction mechanism that selectively introduces aryl groups at the desired position of a molecule at room temperature. A team, co-led by Professor Sukbok Chang and Mu-Hyun Baik of the Department of Chemistry, used an iridium catalyst for the reaction. The team also proved that the reaction proceeds by an unusual mechanism by employing computer simulations that were substantiated with targeted experimental probes.
Hydrocarbon is an omnipresent material in nature. But its low reactivity makes it difficult to process to value-added products at the room temperature. Thus, designing catalysts that can accelerate the reaction remains an important challenge in chemistry.
In particular, since most chemicals used in medicine, pharmacy, or material chemistry contain aryl groups, an effective reaction to selectively introduce the aryl group has been an area of intensive research in organic chemistry.
In order to introduce an aryl group into stable carbon-hydrogen (C-H) bond, activation of the C-H bond with a halogen atom or organic metal is required prior to the introduction of the aryl group, or C-H functionalization directly on C-H bond is needed. Direct functionalization is more effective and economical, but most reactions require harsh reaction conditions such as high temperature or excess additives. And adding the aryl fragment selectively to only one among the many possible sites in the molecule is difficult. The new catalyst developed by these KAIST researchers is highly selective.
This work is the latest example of a successful teamwork between experimental and theoretical research groups: Computer simulations revealed that traditional approaches to arylation required high energies because the intermediates produced during the reaction are too low in energy. Based on this insight, the researchers thought of changing the character of the intermediate by oxidizing it, which was predicted to be a great way of increasing the reactivity of the catalyst. Subsequent experimental work showed that this design strategy is highly effective resulting in unprecedented chemical transformations.
Professor Chang said, “We have been able to carry out location-selective arylation at room temperature, as well as identifying a new reaction pathway, different from the conventionally suggested mechanism.” He continued, “This research is significant for identifying the reaction pathway and developing a novel selective reaction method that does not require high temperature or additives based on the mechanistic understanding. This work is a triumph of rational design, rather than fortuitous discovery.” The research findings were published online in Nature Chemistry on December 11, 2017.
(Figure 1: X-ray crystal structure transmetallation intermediate)
(Figure 2: Correlation between oxidation state of intermediate and energy barrier required for reductive elimination of intermediate as calculated using density function from computational chemistry )
(Figure 3: Arylation mechanism using iridium catalyst as suggested by the research team)
2018.01.11 View 6382 -
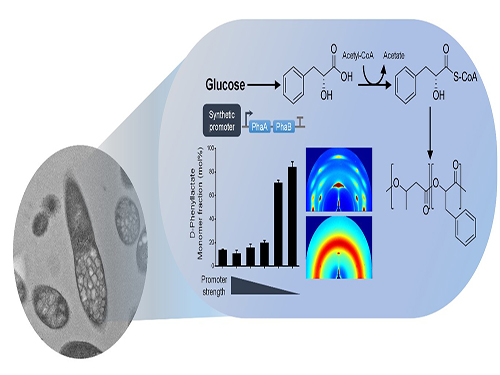 One-Step Production of Aromatic Polyesters by E. coli Strains
KAIST systems metabolic engineers defined a novel strategy for microbial aromatic polyesters production fused with synthetic biology from renewable biomass. The team of Distinguished Professor Sang Yup Lee of the Department of Chemical and Biomolecular Engineering produced aromatic polyesters from Escherichia coli (E. coli) strains by applying microbial fermentation, employing direct microbial fermentation from renewable feedstock carbohydrates.
This is the first report to determine a platform strain of engineered E. coli capable of producing environmentally friendly aromatic polyesters. This engineered E. coli strain, if desired, has the potential to be used as a platform strain capable of producing various high-valued aromatic polyesters from renewable biomass. This research was published in Nature Communications on January 8.
Conventionally, aromatic polyesters boast solid strength and heat stability so that there has been a great deal of interest in fermentative production of aromatic polyesters from renewable non-food biomass, but without success.
However, aromatic polyesters are only made by feeding the cells with corresponding aromatic monomers as substrates, and have not been produced by direct fermentation from renewable feedstock carbohydrates such as glucose.
To address this issue, the team prescribed the detailed procedure for aromatic polyester production through identifying CoA-transferase that activates phenylalkanoates into their corresponding CoA derivatives. In this process, researchers employed metabolic engineering of E. coli to produce phenylalkanoates from glucose based on genome-scale metabolic flux analysis. In particular, the KAIST team made a modulation of gene expression to produce various aromatic polyesters having different monomer fractions.
The research team successfully produced aromatic polyesters, a non-natural polymer using the strategy that combines systems metabolic engineering and synthetic biology. They succeeded in biosynthesis of various kinds of aromatic polyesters through the system, thus proving the technical excellence of the environmentally friendly biosynthetic system of this research. Furthermore, his team also proved the potential of expanding the range of aromatic polyesters from renewable resources, which is expected to play an important role in the bio-plastic industry.
Professor Lee said, “An eco-friendly and sustainable chemical industry is the key global agenda every nation faces. We are making a research focus to a biochemical industry free from petroleum dependence, and conducting diverse research activities to address the issue. This novel technology we are presenting will serve as an opportunity to advance the biochemical industry moving forward.”
This work was supported by the Intelligent Synthetic Biology Center through the Global Frontier Project (2011-0031963) and also by the Technology Development Program to Solve Climate Changes on Systems Metabolic Engineering for Biorefineries (NRF-2012M1A2A2026556 and NRF-2012M1A2A2026557) from the Ministry of Science and ICT through the National Research Foundation of Korea.
Figure: Biosynthesis of aromatic polyesters by metabolically engineered E. coli.This schematic diagram shows the overall conceptualization of how metabolically engineered E. coli produced aromatic polyesters from glucose.
2018.01.09 View 7997
One-Step Production of Aromatic Polyesters by E. coli Strains
KAIST systems metabolic engineers defined a novel strategy for microbial aromatic polyesters production fused with synthetic biology from renewable biomass. The team of Distinguished Professor Sang Yup Lee of the Department of Chemical and Biomolecular Engineering produced aromatic polyesters from Escherichia coli (E. coli) strains by applying microbial fermentation, employing direct microbial fermentation from renewable feedstock carbohydrates.
This is the first report to determine a platform strain of engineered E. coli capable of producing environmentally friendly aromatic polyesters. This engineered E. coli strain, if desired, has the potential to be used as a platform strain capable of producing various high-valued aromatic polyesters from renewable biomass. This research was published in Nature Communications on January 8.
Conventionally, aromatic polyesters boast solid strength and heat stability so that there has been a great deal of interest in fermentative production of aromatic polyesters from renewable non-food biomass, but without success.
However, aromatic polyesters are only made by feeding the cells with corresponding aromatic monomers as substrates, and have not been produced by direct fermentation from renewable feedstock carbohydrates such as glucose.
To address this issue, the team prescribed the detailed procedure for aromatic polyester production through identifying CoA-transferase that activates phenylalkanoates into their corresponding CoA derivatives. In this process, researchers employed metabolic engineering of E. coli to produce phenylalkanoates from glucose based on genome-scale metabolic flux analysis. In particular, the KAIST team made a modulation of gene expression to produce various aromatic polyesters having different monomer fractions.
The research team successfully produced aromatic polyesters, a non-natural polymer using the strategy that combines systems metabolic engineering and synthetic biology. They succeeded in biosynthesis of various kinds of aromatic polyesters through the system, thus proving the technical excellence of the environmentally friendly biosynthetic system of this research. Furthermore, his team also proved the potential of expanding the range of aromatic polyesters from renewable resources, which is expected to play an important role in the bio-plastic industry.
Professor Lee said, “An eco-friendly and sustainable chemical industry is the key global agenda every nation faces. We are making a research focus to a biochemical industry free from petroleum dependence, and conducting diverse research activities to address the issue. This novel technology we are presenting will serve as an opportunity to advance the biochemical industry moving forward.”
This work was supported by the Intelligent Synthetic Biology Center through the Global Frontier Project (2011-0031963) and also by the Technology Development Program to Solve Climate Changes on Systems Metabolic Engineering for Biorefineries (NRF-2012M1A2A2026556 and NRF-2012M1A2A2026557) from the Ministry of Science and ICT through the National Research Foundation of Korea.
Figure: Biosynthesis of aromatic polyesters by metabolically engineered E. coli.This schematic diagram shows the overall conceptualization of how metabolically engineered E. coli produced aromatic polyesters from glucose.
2018.01.09 View 7997 -
 Technology to Find Optimum Drug Target for Cancer Developed
(Professor Kwang-Hyun Cho (right) and lead author Dr. Minsoo Choi)
A KAIST research team led by Professor Kwang-Hyun Cho of the Department of Bio and Brain Engineering developed technology to find the optimum drug target according to the type of cancer cell. The team used systems biology to analyze molecular network dynamics that reflect genetic mutations in cancer cells and to predict drug response. The technology could contribute greatly to future anti-cancer drug development.
There are many types of genetic variations found in cancer cells, including gene mutations and copy number variations. These variations differ in cancer cells even within the same type of cancer, and thus the drug response varies cell by cell. Cancer researchers worked towards identifying frequently occurring genetic variations in cancer patients and, in particular, the mutations that can be used as an index for specific drugs. Previous studies focused on identifying a single genetic mutation or creating an analysis of the structural characteristics of a gene network. However, this approach was limited in its inability to explain the biological properties of cancer which are induced by various gene and protein interactions in cancer cells, which result in differences in drug response.
Gene mutations in cancer cells not only affect the function of the affected gene, but also other genes that interact with the mutated gene and proteins. As a consequence, one mutation could lead to changes in the dynamical properties of the molecular network. Therefore, the responses to anti-cancer drugs by cancer cells differ. The current treatment approach that ignores molecular network dynamics and targets a few cancer-related genes is only effective on a fraction of patients, while many other patients exhibit resistance to the drug.
Professor Cho’s team integrated a large-scale computer simulation using super-computing and cellular experiments to analyze changes in molecular network dynamics in cancer cells.
This led to development of technology to find the optimum drug target according to the type of cancer cells by predicting drug response. This technology was applied to the molecular network of known tumor suppressor p53. The team used large-scale cancer cell genomic data available from The Cancer Cell Line Encyclopedia (CCLE) to construct different molecular networks specific to the characteristics of genetic variations.
Perturbation analysis on drug response in each molecular network was used to quantify changes in cancer cells from drug response and similar networks were clustered. Then, computer simulations were used to analyze the synergetic effects in terms of efficacy and combination to predict the level of drug response. Based on the simulation results from various cancer cell lines including lung, breast, bone, skin, kidney, and ovary cancers were used in drug response experiments for compare analysis.
This technique can be applied in any molecular network to identify the optimum drug target for personalized medicine.
The research team suggests that the technology can analyze varying drug response due to the heterogeneity of cancer cells by considering the overall modulatory interactions rather than focusing only on a specific gene or protein. Further, the technology aids the prediction of causes of drug resistance and thus the identification of the optimum drug target to inhibit the resistance. This could be core source technology that can be used in drug repositioning, a process of applying existing drugs to new disease targets.
Professor Cho said, “Genetic variations in cancer cells are the cause of diverse drug response, but a complete analysis had not yet been made.” He continued, “Systems biology allowed the simulation of drug responses by cancer cell molecular networks to identify fundamental principles of drug response and optimum drug targets using a new conceptual approach.”
This research was published in Nature Communications on December 5 and was funded by Ministry of Science and ICT and National Research Foundation of Korea.
(Figure 1. Drug response prediction for each cancer cell type from computer simulation and cellular experiment verification for comparison)
(Figure 2. Drug response prediction based on cancer cell molecular network dynamics and clustering of cancer cells by their molecular networks)
(Figure 3. Identification of drug target for each cancer cell type by cellular molecular network analysis and establishment for personalized medicine strategy for each cancer patient)
2017.12.15 View 7578
Technology to Find Optimum Drug Target for Cancer Developed
(Professor Kwang-Hyun Cho (right) and lead author Dr. Minsoo Choi)
A KAIST research team led by Professor Kwang-Hyun Cho of the Department of Bio and Brain Engineering developed technology to find the optimum drug target according to the type of cancer cell. The team used systems biology to analyze molecular network dynamics that reflect genetic mutations in cancer cells and to predict drug response. The technology could contribute greatly to future anti-cancer drug development.
There are many types of genetic variations found in cancer cells, including gene mutations and copy number variations. These variations differ in cancer cells even within the same type of cancer, and thus the drug response varies cell by cell. Cancer researchers worked towards identifying frequently occurring genetic variations in cancer patients and, in particular, the mutations that can be used as an index for specific drugs. Previous studies focused on identifying a single genetic mutation or creating an analysis of the structural characteristics of a gene network. However, this approach was limited in its inability to explain the biological properties of cancer which are induced by various gene and protein interactions in cancer cells, which result in differences in drug response.
Gene mutations in cancer cells not only affect the function of the affected gene, but also other genes that interact with the mutated gene and proteins. As a consequence, one mutation could lead to changes in the dynamical properties of the molecular network. Therefore, the responses to anti-cancer drugs by cancer cells differ. The current treatment approach that ignores molecular network dynamics and targets a few cancer-related genes is only effective on a fraction of patients, while many other patients exhibit resistance to the drug.
Professor Cho’s team integrated a large-scale computer simulation using super-computing and cellular experiments to analyze changes in molecular network dynamics in cancer cells.
This led to development of technology to find the optimum drug target according to the type of cancer cells by predicting drug response. This technology was applied to the molecular network of known tumor suppressor p53. The team used large-scale cancer cell genomic data available from The Cancer Cell Line Encyclopedia (CCLE) to construct different molecular networks specific to the characteristics of genetic variations.
Perturbation analysis on drug response in each molecular network was used to quantify changes in cancer cells from drug response and similar networks were clustered. Then, computer simulations were used to analyze the synergetic effects in terms of efficacy and combination to predict the level of drug response. Based on the simulation results from various cancer cell lines including lung, breast, bone, skin, kidney, and ovary cancers were used in drug response experiments for compare analysis.
This technique can be applied in any molecular network to identify the optimum drug target for personalized medicine.
The research team suggests that the technology can analyze varying drug response due to the heterogeneity of cancer cells by considering the overall modulatory interactions rather than focusing only on a specific gene or protein. Further, the technology aids the prediction of causes of drug resistance and thus the identification of the optimum drug target to inhibit the resistance. This could be core source technology that can be used in drug repositioning, a process of applying existing drugs to new disease targets.
Professor Cho said, “Genetic variations in cancer cells are the cause of diverse drug response, but a complete analysis had not yet been made.” He continued, “Systems biology allowed the simulation of drug responses by cancer cell molecular networks to identify fundamental principles of drug response and optimum drug targets using a new conceptual approach.”
This research was published in Nature Communications on December 5 and was funded by Ministry of Science and ICT and National Research Foundation of Korea.
(Figure 1. Drug response prediction for each cancer cell type from computer simulation and cellular experiment verification for comparison)
(Figure 2. Drug response prediction based on cancer cell molecular network dynamics and clustering of cancer cells by their molecular networks)
(Figure 3. Identification of drug target for each cancer cell type by cellular molecular network analysis and establishment for personalized medicine strategy for each cancer patient)
2017.12.15 View 7578 -
 A New Spin Current Generating Material Developed
(Professor Park(left) and Ph.D. candidate Kim)
Magnetic random-access memory (MRAM) is a non-volatile device made of thin magnetic film that can maintain information without an external power supply, in contrast to conventional silicon-based semiconductor memory. It also has the potential for high-density integration and high-speed operation.
The operation of MRAM involves the control of the magnetization direction by exerting spin current-induced torque on a magnetic material. Spin current is generated using electricity in conventional MRAM, but this study developed materials technology that generates spin current using heat.
A KAIST research team led by Professor Byong-Guk Park of the Department of Materials Science and Engineering developed a material that generates spin current from heat, which can be utilized for a new operation principle for MRAM.
There have been theoretical reports on the spin Nernst effect, the phenomenon of the thermal generation of spin current, but is yet to have been experimentally proven due to technological limitations. However, the research team introduced a spin Nernst magnetoresistance measurement method using tungsten (W) and platinum (Pt) with high spin orbit coupling which allows for the experimental identification of the spin Nernst effect. They also demonstrated that the efficiency of spin current generation from heat is similar to that of spin current generated from electricity.
Professor Park said, “This research has great significance in experimentally proving spin current generation from heat, a new physical phenomenon. We aim to develop the technology as a new operational method for MRAM through further research. This can lower power consumption, and is expected to contribute to the advancement of electronics requiring low power requirement such as wearable, mobile, and IOT devices”.
This research was conducted as a joint research project with Professor Kyung-Jin Lee at Korea University and Professor Jong-Ryul Jeong at Chungnam National University. It was published in Nature Communications online on November 9 titled “Observation of transverse spin Nernst magnetoresistance induced by thermal spin current in ferromagnet/non-magnet bilayers.” Ph.D. candidate Dong-Jun Kim at KAIST is the first author. This research was funded by the Ministry of Science and ICT.
(Schematic diagram of spin Nernst magnetoresistance)
(Research result of new spin current generating materials)
2017.12.08 View 9095
A New Spin Current Generating Material Developed
(Professor Park(left) and Ph.D. candidate Kim)
Magnetic random-access memory (MRAM) is a non-volatile device made of thin magnetic film that can maintain information without an external power supply, in contrast to conventional silicon-based semiconductor memory. It also has the potential for high-density integration and high-speed operation.
The operation of MRAM involves the control of the magnetization direction by exerting spin current-induced torque on a magnetic material. Spin current is generated using electricity in conventional MRAM, but this study developed materials technology that generates spin current using heat.
A KAIST research team led by Professor Byong-Guk Park of the Department of Materials Science and Engineering developed a material that generates spin current from heat, which can be utilized for a new operation principle for MRAM.
There have been theoretical reports on the spin Nernst effect, the phenomenon of the thermal generation of spin current, but is yet to have been experimentally proven due to technological limitations. However, the research team introduced a spin Nernst magnetoresistance measurement method using tungsten (W) and platinum (Pt) with high spin orbit coupling which allows for the experimental identification of the spin Nernst effect. They also demonstrated that the efficiency of spin current generation from heat is similar to that of spin current generated from electricity.
Professor Park said, “This research has great significance in experimentally proving spin current generation from heat, a new physical phenomenon. We aim to develop the technology as a new operational method for MRAM through further research. This can lower power consumption, and is expected to contribute to the advancement of electronics requiring low power requirement such as wearable, mobile, and IOT devices”.
This research was conducted as a joint research project with Professor Kyung-Jin Lee at Korea University and Professor Jong-Ryul Jeong at Chungnam National University. It was published in Nature Communications online on November 9 titled “Observation of transverse spin Nernst magnetoresistance induced by thermal spin current in ferromagnet/non-magnet bilayers.” Ph.D. candidate Dong-Jun Kim at KAIST is the first author. This research was funded by the Ministry of Science and ICT.
(Schematic diagram of spin Nernst magnetoresistance)
(Research result of new spin current generating materials)
2017.12.08 View 9095 -
 Expanding Gas Storage Capacity of Nanoporous Materials
A KAIST research team led by Professor Jihan Kim of the Department of Chemical and Biomolecular Engineering has successfully proposed a rational defect engineering methodology that can greatly enhance the gas storage capacity of nanoporous materials. The team conducted a high-throughput computational screening of a large experimental metal-organic framework database to identify 13 candidate materials that could experience significant methane uptake enhancement with only a small proportion of linker vacancy defects.
This research was published online on November 16 in Nature Communications, with M.S. candidate Sanggyu Chong from KAIST as the first author and post-doctorate researcher Günther Thiele from the Department of Chemistry at UC Berkeley as a contributing author.
Metal-organic frameworks, hereinafter MOF, are crystalline nanoporous materials that are comprised of metal clusters and organic linkers continuously bound together by coordination bonds. Due to their ultrahigh surface areas and pore volumes, they have been widely studied for various energy and environment applications.
Similar to other crystalline materials, MOFs are never perfectly crystalline and are likely to contain several different types of defects within their crystalline structures. Among these defects, linker vacancy defects, or the random absence of linker vacancies in their designated bonding positions, are known to be controllable by practicing careful control over the synthesis conditions.
The research team combined the concepts of rational defect engineering over the linker vacancy defects and the potential presence of inaccessible pores within MOFs to propose a methodology where controlled the introduction of linker vacancy defects could lead to a dramatic enhancement in gas adsorption and storage capacities.
The study utilized a Graphic Processing Unit (GPU) code developed by Professor Kim in a high-throughput computational screening of 12,000 experimentally synthesized MOFs to identify the structures with significant amounts of pores that were inaccessible for methane. In determining the presence of inaccessible pores, a flood-fill algorithm was performed over the energy-low regions of the structure, which is the same algorithm used for filling an area with color in Microsoft Paint.
For the MOFs with significant amounts of inaccessible pores, as determined from the screening, the research team emulated linker vacancy defects in their crystalline structures so that the previously inaccessible pores would be newly merged into the main adsorption channel with the introduction of defects for additional surface area and pore volume available for adsorption. The research team successfully identified 13 structures that would experience up to a 55.56% increase in their methane uptake with less than 8.33% of the linker vacancy defects.
The research team believes that this rational defect engineering scheme can be further utilized for many other applications in areas such as selective adsorption of an adsorbate from a gas mixture and the semi-permanent capture of gas molecules.
This research was conducted with the support of the Mid-career Research Program of the National Research Foundation of Korea.
Figure1. A diagram for flood fill algorithm and example of identification of inaccessible regions within the MOFs, using the flood fill algorithm
Figure2. Methane energy contours before and after detect introduction
2017.12.04 View 8720
Expanding Gas Storage Capacity of Nanoporous Materials
A KAIST research team led by Professor Jihan Kim of the Department of Chemical and Biomolecular Engineering has successfully proposed a rational defect engineering methodology that can greatly enhance the gas storage capacity of nanoporous materials. The team conducted a high-throughput computational screening of a large experimental metal-organic framework database to identify 13 candidate materials that could experience significant methane uptake enhancement with only a small proportion of linker vacancy defects.
This research was published online on November 16 in Nature Communications, with M.S. candidate Sanggyu Chong from KAIST as the first author and post-doctorate researcher Günther Thiele from the Department of Chemistry at UC Berkeley as a contributing author.
Metal-organic frameworks, hereinafter MOF, are crystalline nanoporous materials that are comprised of metal clusters and organic linkers continuously bound together by coordination bonds. Due to their ultrahigh surface areas and pore volumes, they have been widely studied for various energy and environment applications.
Similar to other crystalline materials, MOFs are never perfectly crystalline and are likely to contain several different types of defects within their crystalline structures. Among these defects, linker vacancy defects, or the random absence of linker vacancies in their designated bonding positions, are known to be controllable by practicing careful control over the synthesis conditions.
The research team combined the concepts of rational defect engineering over the linker vacancy defects and the potential presence of inaccessible pores within MOFs to propose a methodology where controlled the introduction of linker vacancy defects could lead to a dramatic enhancement in gas adsorption and storage capacities.
The study utilized a Graphic Processing Unit (GPU) code developed by Professor Kim in a high-throughput computational screening of 12,000 experimentally synthesized MOFs to identify the structures with significant amounts of pores that were inaccessible for methane. In determining the presence of inaccessible pores, a flood-fill algorithm was performed over the energy-low regions of the structure, which is the same algorithm used for filling an area with color in Microsoft Paint.
For the MOFs with significant amounts of inaccessible pores, as determined from the screening, the research team emulated linker vacancy defects in their crystalline structures so that the previously inaccessible pores would be newly merged into the main adsorption channel with the introduction of defects for additional surface area and pore volume available for adsorption. The research team successfully identified 13 structures that would experience up to a 55.56% increase in their methane uptake with less than 8.33% of the linker vacancy defects.
The research team believes that this rational defect engineering scheme can be further utilized for many other applications in areas such as selective adsorption of an adsorbate from a gas mixture and the semi-permanent capture of gas molecules.
This research was conducted with the support of the Mid-career Research Program of the National Research Foundation of Korea.
Figure1. A diagram for flood fill algorithm and example of identification of inaccessible regions within the MOFs, using the flood fill algorithm
Figure2. Methane energy contours before and after detect introduction
2017.12.04 View 8720 -
 New Photocatalyst Converts Carbon Dioxide to 99% Pure Fuel
(Professor Song, Ph.D. candidates Kim, and Lim (from left))
A KAIST research team led by Professor Hyunjoon Song of the Department of Chemistry developed a metal oxide nanocatalyst that converts carbon dioxide to 99% pure methane. This technology directly uses sunlight to convert carbon dioxide into methane, which is more efficient in terms of energy storage capacity, compared to the conventional way of storing the electricity produced by solar cells in batteries. The research team used cheap catalytic materials to significantly enhance the reaction efficiency and selectivity of the chemical energy storage method.
This research was conducted as a joint research project with Professor Ki Min Nam at Mokpo National University with co-first authors Dr. Kyung-Lyul Bae and Ph.D. candidates Jinmo Kim and Chan Kyu Lim. The study was published in Nature Communications on November 7.
Although there is growing interest in sunlight as an energy resource, its usage has been limited to daytime and the power output varies with the weather. If sunlight could be directly converted to chemical energy, such as fuel, the limitations of energy storage and its usage could be overcome. In particular, the usage of sunlight to convert carbon dioxide, a main cause of the greenhouse effect in our atmosphere, is of great interest since both energy and environmental issues can be addressed. However, the stability of carbon dioxide made it difficult to convert it to other molecules. Thus, there was a need for a catalyst with enhanced efficiency and selectivity.
Professor Song’s team synthesized zinc oxide nanoparticles, often used in sun cream. The nanoparticles were then bound to copper oxide as single particles, forming a colloidal form of zinc oxide-copper oxide nanoparticles. Zinc oxides produce high energy electrons using light, and this energy is used to convert carbon dioxide into methane. Further, zinc oxide can also produce electrons with light and transfer the electrons to copper oxide. Similar to the principles of photosynthesis in leaves, the electron transfer reaction could be maintained for a long time. As a consequence, although the reaction was conducted in aqueous solution, methane of 99% purity could be obtained from carbon dioxide.
Conventional heterogeneous photocatalysts were in solid powder form with irregular structures and were not dispersed in water. Professor Song’s team used a nanochemical synthesis method to control the structure of the catalyst particles to be regular and maintained over a large surface area. This led to increasing carbon dioxide conversion activity by hundreds of fold in solution compared to existing catalysts.
Professor Song said, “A long time will be needed for the commercialization of the direct conversion reaction of carbon dioxide using sunlight. However, the precise control of catalyst structures at nanoscale would enhance the efficiency of photocatalyst reactions.” He continued, “Applying this method to various phtocatalysts will maximize the catalysts performance.”
(Figure 1. Scheme for carbon dioxide conversion reaction using nano photocatalyst in aqueous solution)
(Figure 2. Structure, photocatalytic CO2 conversion, and stability of ZnO-Cu2O nanocatalyst )
2017.11.13 View 9217
New Photocatalyst Converts Carbon Dioxide to 99% Pure Fuel
(Professor Song, Ph.D. candidates Kim, and Lim (from left))
A KAIST research team led by Professor Hyunjoon Song of the Department of Chemistry developed a metal oxide nanocatalyst that converts carbon dioxide to 99% pure methane. This technology directly uses sunlight to convert carbon dioxide into methane, which is more efficient in terms of energy storage capacity, compared to the conventional way of storing the electricity produced by solar cells in batteries. The research team used cheap catalytic materials to significantly enhance the reaction efficiency and selectivity of the chemical energy storage method.
This research was conducted as a joint research project with Professor Ki Min Nam at Mokpo National University with co-first authors Dr. Kyung-Lyul Bae and Ph.D. candidates Jinmo Kim and Chan Kyu Lim. The study was published in Nature Communications on November 7.
Although there is growing interest in sunlight as an energy resource, its usage has been limited to daytime and the power output varies with the weather. If sunlight could be directly converted to chemical energy, such as fuel, the limitations of energy storage and its usage could be overcome. In particular, the usage of sunlight to convert carbon dioxide, a main cause of the greenhouse effect in our atmosphere, is of great interest since both energy and environmental issues can be addressed. However, the stability of carbon dioxide made it difficult to convert it to other molecules. Thus, there was a need for a catalyst with enhanced efficiency and selectivity.
Professor Song’s team synthesized zinc oxide nanoparticles, often used in sun cream. The nanoparticles were then bound to copper oxide as single particles, forming a colloidal form of zinc oxide-copper oxide nanoparticles. Zinc oxides produce high energy electrons using light, and this energy is used to convert carbon dioxide into methane. Further, zinc oxide can also produce electrons with light and transfer the electrons to copper oxide. Similar to the principles of photosynthesis in leaves, the electron transfer reaction could be maintained for a long time. As a consequence, although the reaction was conducted in aqueous solution, methane of 99% purity could be obtained from carbon dioxide.
Conventional heterogeneous photocatalysts were in solid powder form with irregular structures and were not dispersed in water. Professor Song’s team used a nanochemical synthesis method to control the structure of the catalyst particles to be regular and maintained over a large surface area. This led to increasing carbon dioxide conversion activity by hundreds of fold in solution compared to existing catalysts.
Professor Song said, “A long time will be needed for the commercialization of the direct conversion reaction of carbon dioxide using sunlight. However, the precise control of catalyst structures at nanoscale would enhance the efficiency of photocatalyst reactions.” He continued, “Applying this method to various phtocatalysts will maximize the catalysts performance.”
(Figure 1. Scheme for carbon dioxide conversion reaction using nano photocatalyst in aqueous solution)
(Figure 2. Structure, photocatalytic CO2 conversion, and stability of ZnO-Cu2O nanocatalyst )
2017.11.13 View 9217 -
 Mutant Gene Network in Colon Cancer Identified
The principles of the gene network for colon tumorigenesis have been identified by a KAIST research team. The principles will be used to find the molecular target for effective anti-cancer drugs in the future. Further, this research gained attention for using a systems biology approach, which is an integrated research area of IT and BT.
The KAIST research team led by Professor Kwang-Hyun Cho for the Department of Bio and Brain Engineering succeeded in the identification. Conducted by Dr. Dongkwan Shin and student researchers Jonghoon Lee and Jeong-Ryeol Gong, the research was published in Nature Communications online on November 2.
Human cancer is caused by genetic mutations. The frequency of the mutations differs by the type of cancer; for example, only around 10 mutations are found in leukemia and childhood cancer, but an average of 50 mutations are found in adult solid cancers and even hundreds of mutations are found in cancers due to external factors, such as with lung cancer.
Cancer researchers around the world are working to identify frequently found genetic mutations in patients, and in turn identify important cancer-inducing genes (called ‘driver genes’) to develop targets for anti-cancer drugs. However, gene mutations not only affect their own functions but also affect other genes through interactions. Therefore, there are limitations in current treatments targeting a few cancer-inducing genes without further knowledge on gene networks, hence current drugs are only effective in a few patients and often induce drug resistance.
Professor Cho’s team used large-scale genomic data from cancer patients to construct a mathematical model on the cooperative effects of multiple genetic mutations found in gene interaction networks. The basis of the model construction was The Cancer Genome Atlas (TCGA) presented at the International Cancer Genome Consortium. The team successfully quantified the effects of mutations in gene networks to group colon cancer patients by clinical characteristics.
Further, the critical transition phenomenon that occurs in tumorigenesis was identified using large-scale computer simulation analysis, which was the first hidden gene network principle to be identified. Critical transition is the phenomenon in which the state of matter is suddenly changed through phase transition. It was not possible to identify the presence of transition phenomenon in the past, as it was difficult to track the sequence of gene mutations during tumorigenesis.
The research team used a systems biology-based research method to find that colon cancer tumorigenesis shows a critical transition phenomenon if the known driver gene mutations follow sequentially. Using the developed mathematical model, it can be possible to develop a new anti-cancer targeting drug that most effectively inhibits the effects of many gene mutations found in cancer patients. In particular, not only driver genes, but also other passenger genes affected by the gene mutations, could be evaluated to find the most effective drug targets.
Professor Cho said, “Little was known about the contribution of many gene mutations during tumorigenesis.” He continued, “In this research, a systems biology approach identified the principle of gene networks for the first time to suggest the possibility of anti-cancer drug target identification from a new perspective.”
This research was funded by the Ministry of Science and ICT and the National Research Foundation of Korea.
Figure1. Formation of giant clusters via mutation propagation
Figure2. Critical transition phenomenon by cooperative effect of mutations in tumorigenesis
2017.11.10 View 8652
Mutant Gene Network in Colon Cancer Identified
The principles of the gene network for colon tumorigenesis have been identified by a KAIST research team. The principles will be used to find the molecular target for effective anti-cancer drugs in the future. Further, this research gained attention for using a systems biology approach, which is an integrated research area of IT and BT.
The KAIST research team led by Professor Kwang-Hyun Cho for the Department of Bio and Brain Engineering succeeded in the identification. Conducted by Dr. Dongkwan Shin and student researchers Jonghoon Lee and Jeong-Ryeol Gong, the research was published in Nature Communications online on November 2.
Human cancer is caused by genetic mutations. The frequency of the mutations differs by the type of cancer; for example, only around 10 mutations are found in leukemia and childhood cancer, but an average of 50 mutations are found in adult solid cancers and even hundreds of mutations are found in cancers due to external factors, such as with lung cancer.
Cancer researchers around the world are working to identify frequently found genetic mutations in patients, and in turn identify important cancer-inducing genes (called ‘driver genes’) to develop targets for anti-cancer drugs. However, gene mutations not only affect their own functions but also affect other genes through interactions. Therefore, there are limitations in current treatments targeting a few cancer-inducing genes without further knowledge on gene networks, hence current drugs are only effective in a few patients and often induce drug resistance.
Professor Cho’s team used large-scale genomic data from cancer patients to construct a mathematical model on the cooperative effects of multiple genetic mutations found in gene interaction networks. The basis of the model construction was The Cancer Genome Atlas (TCGA) presented at the International Cancer Genome Consortium. The team successfully quantified the effects of mutations in gene networks to group colon cancer patients by clinical characteristics.
Further, the critical transition phenomenon that occurs in tumorigenesis was identified using large-scale computer simulation analysis, which was the first hidden gene network principle to be identified. Critical transition is the phenomenon in which the state of matter is suddenly changed through phase transition. It was not possible to identify the presence of transition phenomenon in the past, as it was difficult to track the sequence of gene mutations during tumorigenesis.
The research team used a systems biology-based research method to find that colon cancer tumorigenesis shows a critical transition phenomenon if the known driver gene mutations follow sequentially. Using the developed mathematical model, it can be possible to develop a new anti-cancer targeting drug that most effectively inhibits the effects of many gene mutations found in cancer patients. In particular, not only driver genes, but also other passenger genes affected by the gene mutations, could be evaluated to find the most effective drug targets.
Professor Cho said, “Little was known about the contribution of many gene mutations during tumorigenesis.” He continued, “In this research, a systems biology approach identified the principle of gene networks for the first time to suggest the possibility of anti-cancer drug target identification from a new perspective.”
This research was funded by the Ministry of Science and ICT and the National Research Foundation of Korea.
Figure1. Formation of giant clusters via mutation propagation
Figure2. Critical transition phenomenon by cooperative effect of mutations in tumorigenesis
2017.11.10 View 8652 -
 Highly Flexible Organic Flash Memory for Foldable and Disposable Electronics
A KAIST team reported ultra-flexible organic flash memory that is bendable down to a radius of 300μm. The memory exhibits a significantly-long projected retention rate with a programming voltage on par with the present industrial standards.
A joint research team led by Professor Seunghyup Yoo of the School of Electrical Engineering and Professor Sung Gap Im of the Department of Chemical and Biomolecular Engineering said that their memory technology can be applied to non-conventional substrates, such as plastics and papers, to demonstrate its feasibility over a wide range of applications.
With Dr. Seungwon Lee and Dr. Hanul Moon playing the role of leading authors, the research was published in Nature Communications on September 28.
Flash memory is a non-volatile, transistor-based data-storage device that has become essential in most electronic systems in daily life. With straightforward operation mechanisms and easy integration into NAND or NOR array architecture, flash memory has been established as the most successful and dominant non-volatile memory technology by far.
Despite promising demonstrations in the early stages of organic electronics, the overall progress in this field has been far slower than that of thin-film transistors (TFTs) or other devices based on flexible materials. It has been challenging, in particular, to develop flash memory that simultaneously exhibits a significant level of flexibility and performance. This is mainly due to the scarcity of flexible dielectric layers, which are responsible for the tunneling and blocking of charges.
The solution processing used for the preparation of most of the polymeric dielectric layers also makes it difficult to use them in flash memory due to the complexity involved in the formation of the bilayer dielectric structure, which is the key to flash memory operations.
The research team tried to overcome these hurdles and realize highly flexible flash memory by employing thin polymeric insulators grown with initiated chemical vapor deposition (iCVD), a vapor-phase growth technique for polymers that was previously shown to be promising for the fabrication of flexible TFTs. It was further shown that these iCVD-based polymeric insulators, when coupled with rational device design and material choice, can make a significant contribution to flash memory as well.
Memory using conventional polymer insulating films has often required a voltage as high as 100 V (volt) in order to attain long memory retention. If the device is made to operate at a low voltage, the short retention period of less than a month was problematic.
The KAIST team produced flash memory with programming voltages around 10 V and a projected data retention time of over 10 years, while maintaining its memory performance even at a mechanical strain of 2.8%. This is a significant improvement over the existing inorganic insulation layer-based flash memory that allowed only a 1% strain.
The team demonstrated the virtually foldable memory devices by fabricating the proposed flash memory on a 6-micrometer-thick ultrathin plastic film. In addition, it succeeded in producing them on printing paper, opening a way for disposable smart electronic products such as electronic paper and electronic business card.
Professor Yoo said, " This study well illustrates that even highly flexible flash memory can be made to have a practically viable level of performance, so that it contributes to full-fledged wearable electronic devices and smart electronic paper."
(Figure 1. Structure of flexible flash memory )
(Figure 2. Foldable flash memory)
2017.11.06 View 9606
Highly Flexible Organic Flash Memory for Foldable and Disposable Electronics
A KAIST team reported ultra-flexible organic flash memory that is bendable down to a radius of 300μm. The memory exhibits a significantly-long projected retention rate with a programming voltage on par with the present industrial standards.
A joint research team led by Professor Seunghyup Yoo of the School of Electrical Engineering and Professor Sung Gap Im of the Department of Chemical and Biomolecular Engineering said that their memory technology can be applied to non-conventional substrates, such as plastics and papers, to demonstrate its feasibility over a wide range of applications.
With Dr. Seungwon Lee and Dr. Hanul Moon playing the role of leading authors, the research was published in Nature Communications on September 28.
Flash memory is a non-volatile, transistor-based data-storage device that has become essential in most electronic systems in daily life. With straightforward operation mechanisms and easy integration into NAND or NOR array architecture, flash memory has been established as the most successful and dominant non-volatile memory technology by far.
Despite promising demonstrations in the early stages of organic electronics, the overall progress in this field has been far slower than that of thin-film transistors (TFTs) or other devices based on flexible materials. It has been challenging, in particular, to develop flash memory that simultaneously exhibits a significant level of flexibility and performance. This is mainly due to the scarcity of flexible dielectric layers, which are responsible for the tunneling and blocking of charges.
The solution processing used for the preparation of most of the polymeric dielectric layers also makes it difficult to use them in flash memory due to the complexity involved in the formation of the bilayer dielectric structure, which is the key to flash memory operations.
The research team tried to overcome these hurdles and realize highly flexible flash memory by employing thin polymeric insulators grown with initiated chemical vapor deposition (iCVD), a vapor-phase growth technique for polymers that was previously shown to be promising for the fabrication of flexible TFTs. It was further shown that these iCVD-based polymeric insulators, when coupled with rational device design and material choice, can make a significant contribution to flash memory as well.
Memory using conventional polymer insulating films has often required a voltage as high as 100 V (volt) in order to attain long memory retention. If the device is made to operate at a low voltage, the short retention period of less than a month was problematic.
The KAIST team produced flash memory with programming voltages around 10 V and a projected data retention time of over 10 years, while maintaining its memory performance even at a mechanical strain of 2.8%. This is a significant improvement over the existing inorganic insulation layer-based flash memory that allowed only a 1% strain.
The team demonstrated the virtually foldable memory devices by fabricating the proposed flash memory on a 6-micrometer-thick ultrathin plastic film. In addition, it succeeded in producing them on printing paper, opening a way for disposable smart electronic products such as electronic paper and electronic business card.
Professor Yoo said, " This study well illustrates that even highly flexible flash memory can be made to have a practically viable level of performance, so that it contributes to full-fledged wearable electronic devices and smart electronic paper."
(Figure 1. Structure of flexible flash memory )
(Figure 2. Foldable flash memory)
2017.11.06 View 9606 -
 High-Speed Motion Core Technology for Magnetic Memory
(Professor Kab-Jin Kim of the Department of Physics)
A joint research team led by Professor Kab-Jin Kim of the Department of Physics, KAIST and Professor Kyung-Jin Lee at Korea University developed technology to dramatically enhance the speed of next generation domain wall-based magnetic memory. This research was published online in Nature Materials on September 25.
Currently-used memory materials, D-RAM and S-RAM, are fast but volatile, leading to memory loss when the power is switched off. Flash memory is non-volatile but slow, while hard disk drives (HDD) have greater storage but are high in energy usage and weak in physical shock tolerance.
To overcome the limitations of existing memory materials, ‘domain wall-based, magnetic memory’ is being researched. The core mechanism of domain wall magnetic memory is the movement of a domain wall by the current. Non-volatility is secured by using magnetic nanowires and the lack of mechanical rotation reduced power usage. This is a new form of high density, low power next-generation memory.
However, previous studies showed the speed limit of domain wall memory to be hundreds m/s at maximum due to the ‘Walker breakdown phenomenon’, which refers to velocity breakdown from the angular precession of a domain wall. Therefore, there was a need to develop core technology to remove the Walker breakdown phenomenon and increase the speed for the commercialization of domain wall memory.
Most domain wall memory studies used ferromagnetic bodies, which cannot overcome the Walker breakdown phenomenon. The team discovered that the use of ‘ferrimagnetic‘ GdFeCo at certain conditions could overcome the Walker breakdown phenomenon and using this mechanism they could increase domain wall speed to over 2Km/s at room temperature.
Domain wall memory is high-density, low-power, and non-volatile memory. The memory could be the leading next-generation memory with the addition of the high speed property discovered in this research.
Professor Kim said, “This research is significant in discovering a new physical phenomenon at the point at which the angular momentum of a ferrimagnetic body is 0 and it is expected to advance the implementation of next-generation memory in the future.”
This research was funded by the National Research Foundation of Korea (NRF) grant funded by the Korea Government (MSIP) (No. 2017R1C1B2009686, NRF-2016R1A5A1008184) and by the DGIST R&D Program of the Ministry of Science, ICT and Future Planning (17-BT-02).
(Figure 1. Concept Map of Domain Wall Memory Material using Ferrimagnetic Body)
(Figure 2. Scheme and Experimental Results of Domain Wall Speed Measurements)
2017.10.30 View 9500
High-Speed Motion Core Technology for Magnetic Memory
(Professor Kab-Jin Kim of the Department of Physics)
A joint research team led by Professor Kab-Jin Kim of the Department of Physics, KAIST and Professor Kyung-Jin Lee at Korea University developed technology to dramatically enhance the speed of next generation domain wall-based magnetic memory. This research was published online in Nature Materials on September 25.
Currently-used memory materials, D-RAM and S-RAM, are fast but volatile, leading to memory loss when the power is switched off. Flash memory is non-volatile but slow, while hard disk drives (HDD) have greater storage but are high in energy usage and weak in physical shock tolerance.
To overcome the limitations of existing memory materials, ‘domain wall-based, magnetic memory’ is being researched. The core mechanism of domain wall magnetic memory is the movement of a domain wall by the current. Non-volatility is secured by using magnetic nanowires and the lack of mechanical rotation reduced power usage. This is a new form of high density, low power next-generation memory.
However, previous studies showed the speed limit of domain wall memory to be hundreds m/s at maximum due to the ‘Walker breakdown phenomenon’, which refers to velocity breakdown from the angular precession of a domain wall. Therefore, there was a need to develop core technology to remove the Walker breakdown phenomenon and increase the speed for the commercialization of domain wall memory.
Most domain wall memory studies used ferromagnetic bodies, which cannot overcome the Walker breakdown phenomenon. The team discovered that the use of ‘ferrimagnetic‘ GdFeCo at certain conditions could overcome the Walker breakdown phenomenon and using this mechanism they could increase domain wall speed to over 2Km/s at room temperature.
Domain wall memory is high-density, low-power, and non-volatile memory. The memory could be the leading next-generation memory with the addition of the high speed property discovered in this research.
Professor Kim said, “This research is significant in discovering a new physical phenomenon at the point at which the angular momentum of a ferrimagnetic body is 0 and it is expected to advance the implementation of next-generation memory in the future.”
This research was funded by the National Research Foundation of Korea (NRF) grant funded by the Korea Government (MSIP) (No. 2017R1C1B2009686, NRF-2016R1A5A1008184) and by the DGIST R&D Program of the Ministry of Science, ICT and Future Planning (17-BT-02).
(Figure 1. Concept Map of Domain Wall Memory Material using Ferrimagnetic Body)
(Figure 2. Scheme and Experimental Results of Domain Wall Speed Measurements)
2017.10.30 View 9500 -
 The Medici Effect: Highly Flexible, Wearable Displays Born in KAIST
(Ph.D. candidate Seungyeop Choi)
How do you feel when technology you saw in a movie is made into reality? Collaboration between the electrical engineering and textile industries has made TVs or smartphone screens displaying on clothing a reality.
A research team led by Professor Kyung Cheol Choi at the School of Electrical Engineering presented wearable displays for various applications including fashion, IT, and healthcare. Integrating OLED (organic light-emitting diode) into fabrics, the team developed the most highly flexible and reliable technology for wearable displays in the world.
Recently, information displays have become increasingly important as they construct the external part of smart devices for the next generation. As world trends are focusing on the Internet of Things (IoTs) and wearable technology, the team drew a lot of attention by making great progress towards commercializing clothing-shaped ‘wearable displays’.
The research for realizing displays on clothing gained considerable attention from academia as well as industry when research on luminescence formed in fabrics was introduced in 2011; however, there was no technology for commercializing it due to its surface roughness and flexibility.
Because of this technical limitation, clothing-shaped wearable displays were thought to be unreachable technology. However, the KAIST team recently succeeded in developing the world’s most highly efficient, light-emitting clothes that can be commercialized.
The research team used two different approaches, fabric-type and fiber-type, in order to realize clothing-shaped wearable displays. In 2015, the team successfully laminated a thin planarization sheet thermally onto fabric to form a surface that is compatible with the OLEDs approximately 200 hundred nanometers thick. Also, the team reported their research outcomes on enhancing the reliability of operating fiber-based OLEDs. In 2016, the team introduced a dip-coating method, capable of uniformly depositing layers, to develop polymer light-emitting diodes, which show high luminance even on thin fabric.
Based on the research performance in 2015 and 2016, Ph.D. candidate Seungyeop Choi took the lead in the research team and succeeded in realizing fabric-based OLEDs, showing high luminance and efficiency while maintaining the flexibility of the fabric.
The long-term reliability of this wearable device that has the world’s best electrical and optical characteristics was verified through their self-developed, organic and inorganic encapsulation technology. According to the team, their wearable device facilitates the operation of OLEDs even at a bending radius of 2mm.
According to Choi, “Having wavy structures and empty spaces, fiber plays a significant role in lowering the mechanical stress on the OLEDs.”
“Screen displayed on our daily clothing is no longer a future technology,” said Professor Choi. “Light-emitting clothes will have considerable influence on not only the e-textile industry but also the automobile and healthcare industries.”
Moreover, the research team remarked, “It means a lot to realize clothing-shaped OLEDs that have the world’s best luminance and efficiency. It is the most flexible fabric-based light-emitting device among those reported. Moreover, noting that this research carried out an in-depth analysis of the mechanical characteristics of the clothing-spared, light-emitting device, the research performance will become a guideline for developing the fabric-based electronics industry.”
This research was funded by the Ministry of Trade, Industry and Energy and collaborated with KOLON Glotech, INC. The research performance was published in Scientific Reports in July.
(OLEDs operating in fabrics)
(Current-voltage-luminance and efficiency of the highly flexible, fabric-based OLEDs;Image of OLEDs after repetitive bending tests;Verification of flexibility through mechanical simulation)
2017.08.24 View 16560
The Medici Effect: Highly Flexible, Wearable Displays Born in KAIST
(Ph.D. candidate Seungyeop Choi)
How do you feel when technology you saw in a movie is made into reality? Collaboration between the electrical engineering and textile industries has made TVs or smartphone screens displaying on clothing a reality.
A research team led by Professor Kyung Cheol Choi at the School of Electrical Engineering presented wearable displays for various applications including fashion, IT, and healthcare. Integrating OLED (organic light-emitting diode) into fabrics, the team developed the most highly flexible and reliable technology for wearable displays in the world.
Recently, information displays have become increasingly important as they construct the external part of smart devices for the next generation. As world trends are focusing on the Internet of Things (IoTs) and wearable technology, the team drew a lot of attention by making great progress towards commercializing clothing-shaped ‘wearable displays’.
The research for realizing displays on clothing gained considerable attention from academia as well as industry when research on luminescence formed in fabrics was introduced in 2011; however, there was no technology for commercializing it due to its surface roughness and flexibility.
Because of this technical limitation, clothing-shaped wearable displays were thought to be unreachable technology. However, the KAIST team recently succeeded in developing the world’s most highly efficient, light-emitting clothes that can be commercialized.
The research team used two different approaches, fabric-type and fiber-type, in order to realize clothing-shaped wearable displays. In 2015, the team successfully laminated a thin planarization sheet thermally onto fabric to form a surface that is compatible with the OLEDs approximately 200 hundred nanometers thick. Also, the team reported their research outcomes on enhancing the reliability of operating fiber-based OLEDs. In 2016, the team introduced a dip-coating method, capable of uniformly depositing layers, to develop polymer light-emitting diodes, which show high luminance even on thin fabric.
Based on the research performance in 2015 and 2016, Ph.D. candidate Seungyeop Choi took the lead in the research team and succeeded in realizing fabric-based OLEDs, showing high luminance and efficiency while maintaining the flexibility of the fabric.
The long-term reliability of this wearable device that has the world’s best electrical and optical characteristics was verified through their self-developed, organic and inorganic encapsulation technology. According to the team, their wearable device facilitates the operation of OLEDs even at a bending radius of 2mm.
According to Choi, “Having wavy structures and empty spaces, fiber plays a significant role in lowering the mechanical stress on the OLEDs.”
“Screen displayed on our daily clothing is no longer a future technology,” said Professor Choi. “Light-emitting clothes will have considerable influence on not only the e-textile industry but also the automobile and healthcare industries.”
Moreover, the research team remarked, “It means a lot to realize clothing-shaped OLEDs that have the world’s best luminance and efficiency. It is the most flexible fabric-based light-emitting device among those reported. Moreover, noting that this research carried out an in-depth analysis of the mechanical characteristics of the clothing-spared, light-emitting device, the research performance will become a guideline for developing the fabric-based electronics industry.”
This research was funded by the Ministry of Trade, Industry and Energy and collaborated with KOLON Glotech, INC. The research performance was published in Scientific Reports in July.
(OLEDs operating in fabrics)
(Current-voltage-luminance and efficiency of the highly flexible, fabric-based OLEDs;Image of OLEDs after repetitive bending tests;Verification of flexibility through mechanical simulation)
2017.08.24 View 16560 -
 Solutal Marangoni Flows of Miscible Liquid Drive Transport without Surface Contamination
(Professor Hyoungsoo Kim, Department of Mechanical Engineering, KAIST)
A research team led by Hyoungsoo Kim, a professor of Mechanical Engineering at KAIST, succeeded in quantifying the phenomenon called, the Marangoni effect, which occurs at the interface between alcohol and water. It is expected that this finding will be a valuable resource used for effectively removing impurities from a surface fluid without any contamination, and developing materials that can replace surfactants.
This research, co-conducted with a research team led by Professor Howard A. Stone at Princeton University, was published online in Nature Physics on July 31.
The Marangoni effect, also known as tears of wine, is generated when two fluids having a different surface tension meet, causing finite mixing, spreading time and length scale. Typically, people believe that infinitely miscible liquids immediately mix together; however, it is not always true according to this paper.
The typical surface tension of alcohol is three times lower than that of water, and this different surface tension generates the Marangoni-driven convection flow at the interface of the two liquids. In addition, there is a certain amount of time required for them to mix.
This phenomenon has been discussed many times since it was discovered in early the 20th century, yet there was a limit to quantifying and explaining it.
Professor Kim, considering the mixing and spreading mechanism, used various flow visualization techniques and equipment for capturing high speed images in his experiment.
Through the flow visualization methods, the team succeeded in quantifying and explaining the complex, physicochemical phenomenon generated between water and alcohol. Moreover, they developed a theoretical model to predict the physicochemical hydrodynamic phenomena.
The theoretical model can predict the speed of Marangoni-driven convection flow, the area of a drop of alcohol and the time required to develop the flow field. Hence, this model can map out types of materials (e.g., alcohol) and the volume of a drop of liquid as applicable to target a specific situation.
Moreover, the research team believes that the interfacial flow enables the driving of bulk flows and that it can be a source of technology for effectively delivering drugs and removing impurities from a surface of substance without causing secondary contamination.
Above all, the results show a possibility for replacing surfactant with alcohol as a material used for delivering drugs. In the case of the drug delivery, some drugs are encapsulated with a surfactant in order to be effectively transported in vivo; however, the surfactant accumulates in the body, which can cause various side effects, such as heart disease. Therefore, using new materials like alcohol for drug delivery will contribute to preventing the side effects caused by the surfactant.
“The surfactant is used for delivering drugs, but it is difficult to be expelled from the body. This will cause various side effects, such as heart diseases in asthmatic patients,” said Professor Kim. “I hope that using new materials, like alcohol, will free people from these side effects.”
(Marangoni-driven convection flow generated at the interface between water and alcohol, and the flow visualization results)
- A drop of alcohol on a water surface
- Comparison of mixing structures on the surface
- Marangoni mixing flow under the free surface
2017.08.18 View 9185
Solutal Marangoni Flows of Miscible Liquid Drive Transport without Surface Contamination
(Professor Hyoungsoo Kim, Department of Mechanical Engineering, KAIST)
A research team led by Hyoungsoo Kim, a professor of Mechanical Engineering at KAIST, succeeded in quantifying the phenomenon called, the Marangoni effect, which occurs at the interface between alcohol and water. It is expected that this finding will be a valuable resource used for effectively removing impurities from a surface fluid without any contamination, and developing materials that can replace surfactants.
This research, co-conducted with a research team led by Professor Howard A. Stone at Princeton University, was published online in Nature Physics on July 31.
The Marangoni effect, also known as tears of wine, is generated when two fluids having a different surface tension meet, causing finite mixing, spreading time and length scale. Typically, people believe that infinitely miscible liquids immediately mix together; however, it is not always true according to this paper.
The typical surface tension of alcohol is three times lower than that of water, and this different surface tension generates the Marangoni-driven convection flow at the interface of the two liquids. In addition, there is a certain amount of time required for them to mix.
This phenomenon has been discussed many times since it was discovered in early the 20th century, yet there was a limit to quantifying and explaining it.
Professor Kim, considering the mixing and spreading mechanism, used various flow visualization techniques and equipment for capturing high speed images in his experiment.
Through the flow visualization methods, the team succeeded in quantifying and explaining the complex, physicochemical phenomenon generated between water and alcohol. Moreover, they developed a theoretical model to predict the physicochemical hydrodynamic phenomena.
The theoretical model can predict the speed of Marangoni-driven convection flow, the area of a drop of alcohol and the time required to develop the flow field. Hence, this model can map out types of materials (e.g., alcohol) and the volume of a drop of liquid as applicable to target a specific situation.
Moreover, the research team believes that the interfacial flow enables the driving of bulk flows and that it can be a source of technology for effectively delivering drugs and removing impurities from a surface of substance without causing secondary contamination.
Above all, the results show a possibility for replacing surfactant with alcohol as a material used for delivering drugs. In the case of the drug delivery, some drugs are encapsulated with a surfactant in order to be effectively transported in vivo; however, the surfactant accumulates in the body, which can cause various side effects, such as heart disease. Therefore, using new materials like alcohol for drug delivery will contribute to preventing the side effects caused by the surfactant.
“The surfactant is used for delivering drugs, but it is difficult to be expelled from the body. This will cause various side effects, such as heart diseases in asthmatic patients,” said Professor Kim. “I hope that using new materials, like alcohol, will free people from these side effects.”
(Marangoni-driven convection flow generated at the interface between water and alcohol, and the flow visualization results)
- A drop of alcohol on a water surface
- Comparison of mixing structures on the surface
- Marangoni mixing flow under the free surface
2017.08.18 View 9185