graduate+school+of+medical+science+and+engineering
-
 First Mutations in Human Life Discovered
The earliest mutations of human life have been observed by research team led by the Wellcome Trust Sanger Institute and their collaborators. Analyzing genomes from adult cells, the scientists could look back in time to reveal how each embryo developed.
Research team of the Sanger Institute including Professor Young Seok Ju of the Graduate School of Medical Science and Engineering at KAIST published an article of “Somatic Mutations Reveal Asymmetric Cellular Dynamics in the Early Human Embryo” in Nature on March 22.
The study shows that from the two-cell stage of the human embryo, one of these cells becomes more dominant than the other and leads to a higher proportion of the adult body.
A longstanding question for researchers has been what happens in the very early human development as this has proved impossible to study directly. Now, researchers have analyzed the whole genome sequences of blood samples (collected from 279 individuals with breast cancer) and discovered 163 mutations that occurred very early in the embryonic development of those people.
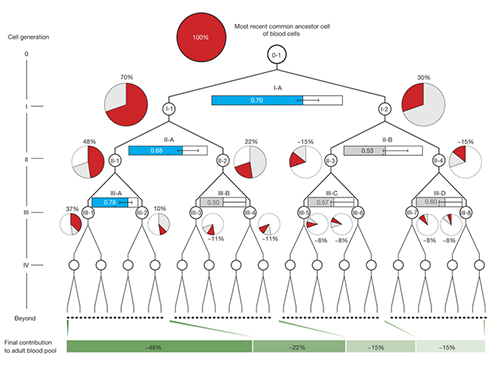
Once identified, the researchers used mutations from the first, second and third divisions of the fertilized egg to calculate which proportion of adult cells resulted from each of the first two cells in the embryo. They found that these first two cells contribute differently to the whole body. One cell gives rise to about 70 percent of the adult body tissues, whereas the other cell has a more minor contribution, leading to about 30percent of the tissues. This skewed contribution continues for some cells in the second and third generation too.
Originally pinpointed in normal blood cells from cancer patients, the researchers then looked for these mutations in cancer samples that had been surgically removed from the patients during treatment. Unlike normal tissues composed of multiple somatic cell clones, a cancer develops from one mutant cell. Therefore, each proposed embryonic mutation should either be present in all of the cancer cells in a tumor, or none of them. This proved to be the case, and by using these cancer samples, the researchers were able to validate that the mutations had originated during early development.
Dr. Young Seok Ju, first author from the Wellcome Trust Sanger Institute and KAIST, said: "This is the first time that anyone has seen where mutations arise in the very early human development. It is like finding a needle in a haystack. There are just a handful of these mutations, compared with millions of inherited genetic variations, and finding them allowed us to track what happened during embryogenesis."
Dr. Inigo Martincorena, from the Sanger Institute, said: "Having identified the mutations, we were able to use statistical analysis to better understand cell dynamics during embryo development. We determined the relative contribution of the first embryonic cells to the adult blood cell pool and found one dominant cell - that led to 70 percent of the blood cells - and one minor cell. We also sequenced normal lymph and breast cells, and the results suggested that the dominant cell also contributes to these other tissues at a similar level. This opens an unprecedented window into the earliest stages of human development."
During this study, the researchers were also able to measure the rate of mutation in early human development for the first time, up to three generations of cell division. Previous researchers had estimated one mutation per cell division, but this study measured three mutations for each cell doubling, in every daughter cell.
Mutations during the development of the embryo occur by two processes - known as mutational signatures 1 and 5. These mutations are fairly randomly distributed through the genome, and the vast majority of them will not affect the developing embryo. However, a mutation that occurs in an important gene can lead to disease such as developmental disorders.
Professor Sir Mike Stratton, lead author on the paper and Director of the Sanger Institute, said: "This is a significant step forward in widening the range of biological insights that can be extracted using genome sequences and mutations. Essentially, the mutations are archaeological traces of embryonic development left in our adult tissues, so if we can find and interpret them, we can understand human embryology better. This is just one early insight into human development, with hopefully many more to come in the future."
(Figure 1. Detection of somatic mutations acquired in early human embryogenesis )
(Figure 2. Unequal contributions of early embryonic cells to adult somatic tissues )
2017.03.23 View 6559
First Mutations in Human Life Discovered
The earliest mutations of human life have been observed by research team led by the Wellcome Trust Sanger Institute and their collaborators. Analyzing genomes from adult cells, the scientists could look back in time to reveal how each embryo developed.
Research team of the Sanger Institute including Professor Young Seok Ju of the Graduate School of Medical Science and Engineering at KAIST published an article of “Somatic Mutations Reveal Asymmetric Cellular Dynamics in the Early Human Embryo” in Nature on March 22.
The study shows that from the two-cell stage of the human embryo, one of these cells becomes more dominant than the other and leads to a higher proportion of the adult body.
A longstanding question for researchers has been what happens in the very early human development as this has proved impossible to study directly. Now, researchers have analyzed the whole genome sequences of blood samples (collected from 279 individuals with breast cancer) and discovered 163 mutations that occurred very early in the embryonic development of those people.
Once identified, the researchers used mutations from the first, second and third divisions of the fertilized egg to calculate which proportion of adult cells resulted from each of the first two cells in the embryo. They found that these first two cells contribute differently to the whole body. One cell gives rise to about 70 percent of the adult body tissues, whereas the other cell has a more minor contribution, leading to about 30percent of the tissues. This skewed contribution continues for some cells in the second and third generation too.
Originally pinpointed in normal blood cells from cancer patients, the researchers then looked for these mutations in cancer samples that had been surgically removed from the patients during treatment. Unlike normal tissues composed of multiple somatic cell clones, a cancer develops from one mutant cell. Therefore, each proposed embryonic mutation should either be present in all of the cancer cells in a tumor, or none of them. This proved to be the case, and by using these cancer samples, the researchers were able to validate that the mutations had originated during early development.
Dr. Young Seok Ju, first author from the Wellcome Trust Sanger Institute and KAIST, said: "This is the first time that anyone has seen where mutations arise in the very early human development. It is like finding a needle in a haystack. There are just a handful of these mutations, compared with millions of inherited genetic variations, and finding them allowed us to track what happened during embryogenesis."
Dr. Inigo Martincorena, from the Sanger Institute, said: "Having identified the mutations, we were able to use statistical analysis to better understand cell dynamics during embryo development. We determined the relative contribution of the first embryonic cells to the adult blood cell pool and found one dominant cell - that led to 70 percent of the blood cells - and one minor cell. We also sequenced normal lymph and breast cells, and the results suggested that the dominant cell also contributes to these other tissues at a similar level. This opens an unprecedented window into the earliest stages of human development."
During this study, the researchers were also able to measure the rate of mutation in early human development for the first time, up to three generations of cell division. Previous researchers had estimated one mutation per cell division, but this study measured three mutations for each cell doubling, in every daughter cell.
Mutations during the development of the embryo occur by two processes - known as mutational signatures 1 and 5. These mutations are fairly randomly distributed through the genome, and the vast majority of them will not affect the developing embryo. However, a mutation that occurs in an important gene can lead to disease such as developmental disorders.
Professor Sir Mike Stratton, lead author on the paper and Director of the Sanger Institute, said: "This is a significant step forward in widening the range of biological insights that can be extracted using genome sequences and mutations. Essentially, the mutations are archaeological traces of embryonic development left in our adult tissues, so if we can find and interpret them, we can understand human embryology better. This is just one early insight into human development, with hopefully many more to come in the future."
(Figure 1. Detection of somatic mutations acquired in early human embryogenesis )
(Figure 2. Unequal contributions of early embryonic cells to adult somatic tissues )
2017.03.23 View 6559 -
 The Antibody That Normalizes Tumor Vessels
Researchers also discover that their antisepsis antibody reduces glioma, lung and breast cancer progression in mice.
A research team at the Center for Vascular Research within the Institute for Basic Science (IBS) discovered that the antisepsis antibody ABTAA (Ang2-Binding and Tie2-Activating Antibody) reduces tumor volume and improves the delivery of anti-cancer drugs. Published in Cancer Cell, this study demonstrates that ABTAA restores the structural and functional integrity of tumor blood vessels in three different tumor models: breast, lungs, and brain.
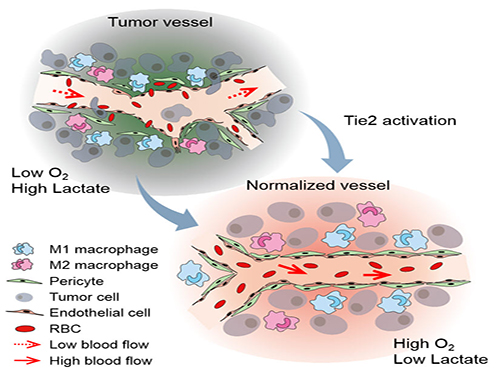
Blood vessels inside and around an established tumor can be described as a chaotic and dysfunctional labyrinth. While the inner walls of healthy blood vessels are surrounded and supported by endothelial cells and other cells called pericytes, in the established tumor, the endothelial junctions are broken apart and pericytes are also detached. Blood flow into and from the tumor is severely retarded and tumor vessels lacking an intact vessel wall become leaky. This microenvironment causes limited drug delivery to the tumor and leads to inadequate oxygen supply (hypoxia) and even metastasis.
The research team led by Professor Gou-Young Koh at KAIST’s Graduate School of Medical Science and Engineering found that the antibody ABTAA normalizes the tumor vessels and hence, change the whole tumor microenvironment. “We call it normalization of tumor vessels, because it resembles closely the wall architecture of healthy, normal vessels,” explains PARK Jin-Sung, first author of the study. And continues: “Tumor can adapt to hypoxia and get more aggressive, so we tried to prevent this transition by normalizing tumor vessels. ABTAA changes the whole tumor environment, oxygenation status and level of lactate, so that the immune cells and drugs can reach the core regions of the tumor more easily. In this way, we create a favorable ground for tumor treatment.”
In an attempt to generate antibodies targeting the protein Ang2, which is specifically expressed by endothelial cells in stressful conditions like in tumor, the team unexpectedly discovered that ABTAA has a peculiar way of working and a dual function. ABTAA indeed not only blocks Ang2, but also activates Tie2 at the same time. Tie2 is a receptor present on the cell membrane of endothelial cells. ABTAA causes Ang2 to cluster together and to strongly activate Tie2 receptors. “If we activate Tie2, we can efficiently normalize tumor vessels, enhance drug delivery and change the whole microenvironment,” explains KOH Gou Young, Director of the Center for Vascular Research.
Several pharmaceutical companies are developing Ang2-blocking antibodies to cure cancer. However, even if these antibodies significantly inhibit tumor progression, they do not stop tumor hypoxia. Moreover, most of the anti-cancer drugs target the tumor at its early stage, when tumors are still hard to diagnose. ABTAA, instead, works with tumors that are already rooted: “When the tumor is established, hypoxia is the main driver of tumor progression. So, if we eliminate hypoxia, we make the tumor milder, by reducing its progression and metastasis,” comments Koh.
Figure: Schematic drawing of a blood vessel around tumors before and after treatment with ABTAA. The picture above shows a typical tumor vasculature characterized by damaged walls, red blood cells leakage and detached pericytes. Activating Tie2 on endothelial cells with the antibody ABTAA restores the normal vessel architecture: endothelial and pericytes on the vessel walls are stabilized, the delivery of blood is improved, and the anticancer drugs are more likely to reach the tumor core.
The researchers tested ABTAA in mice with three different types of tumors that show high levels of Ang2: glioma (a type of a brain tumor), lung carcinoma, and breast cancer. They also compared the effect of ABTAA with ABA, another antibody that blocks Ang2 but misses the Tie2 activating properties. In all three cases, ABTAA was superior to ABA in inducing tumor vessel normalization, which led to a better delivery of the anti-cancer drugs into the tumor core region.
Glioma is one of the so-called intractable diseases, because of its poor prognosis and treatment. Professor Koh’s team found that the glioma volume was reduced 39% by ABTAA and 17% by ABA. ABTAA profoundly reduced vascular leakage and edema formation in glioma through promoting vascular tightening. Moreover, when ABTAA was administered together with the chemotherapeutic drug temozolomide (TMZ), the tumor volume reduces further (76% by ABTAA+TMZ, 51% by ABA+TMZ, and 36% by TMZ).
In the Lewis Lung Carcinoma (LLC) tumor model, the team administered ABTAA together with a chemotherapeutic drug called cisplatin (Cpt) and observed a greater suppression of tumor growth (52%) compared with the controls and increased overall survival. Moreover, ABTAA+Cpt led to a marked increase in necrotic area within tumors.
Finally, in a spontaneous breast cancer model, ABTAA delayed tumor growth and enhanced the anti-tumor effect of Cpt.
Courtesy of the Institute for Basic Sciences (IBS)
Figure: The antibody ABTAA alone and in combination with other anti-cancer drugs have a beneficial effect in reducing tumor volume. ABTAA was tested in mice with brain tumor (glioma), lung or breast cancer. The image shows the improvements: reduction in glioma tumor size, reduction in metastatic colonies in lung tumor and decrease in necrotic regions in breast tumor.
In the future, the team would like to further understand the underlying relationship between faulty blood vessels and diseases. “We would like to apply this antibody to an organ that is rich in blood vessels, that is the eye, and see if this antibody can be useful to treat eye diseases such as age-related macular degeneration and diabetic retinopathy,” concludes Koh.
Professor Gou-Young Koh (left) and Jin-Sung Park (right)
2016.12.16 View 7611
The Antibody That Normalizes Tumor Vessels
Researchers also discover that their antisepsis antibody reduces glioma, lung and breast cancer progression in mice.
A research team at the Center for Vascular Research within the Institute for Basic Science (IBS) discovered that the antisepsis antibody ABTAA (Ang2-Binding and Tie2-Activating Antibody) reduces tumor volume and improves the delivery of anti-cancer drugs. Published in Cancer Cell, this study demonstrates that ABTAA restores the structural and functional integrity of tumor blood vessels in three different tumor models: breast, lungs, and brain.
Blood vessels inside and around an established tumor can be described as a chaotic and dysfunctional labyrinth. While the inner walls of healthy blood vessels are surrounded and supported by endothelial cells and other cells called pericytes, in the established tumor, the endothelial junctions are broken apart and pericytes are also detached. Blood flow into and from the tumor is severely retarded and tumor vessels lacking an intact vessel wall become leaky. This microenvironment causes limited drug delivery to the tumor and leads to inadequate oxygen supply (hypoxia) and even metastasis.
The research team led by Professor Gou-Young Koh at KAIST’s Graduate School of Medical Science and Engineering found that the antibody ABTAA normalizes the tumor vessels and hence, change the whole tumor microenvironment. “We call it normalization of tumor vessels, because it resembles closely the wall architecture of healthy, normal vessels,” explains PARK Jin-Sung, first author of the study. And continues: “Tumor can adapt to hypoxia and get more aggressive, so we tried to prevent this transition by normalizing tumor vessels. ABTAA changes the whole tumor environment, oxygenation status and level of lactate, so that the immune cells and drugs can reach the core regions of the tumor more easily. In this way, we create a favorable ground for tumor treatment.”
In an attempt to generate antibodies targeting the protein Ang2, which is specifically expressed by endothelial cells in stressful conditions like in tumor, the team unexpectedly discovered that ABTAA has a peculiar way of working and a dual function. ABTAA indeed not only blocks Ang2, but also activates Tie2 at the same time. Tie2 is a receptor present on the cell membrane of endothelial cells. ABTAA causes Ang2 to cluster together and to strongly activate Tie2 receptors. “If we activate Tie2, we can efficiently normalize tumor vessels, enhance drug delivery and change the whole microenvironment,” explains KOH Gou Young, Director of the Center for Vascular Research.
Several pharmaceutical companies are developing Ang2-blocking antibodies to cure cancer. However, even if these antibodies significantly inhibit tumor progression, they do not stop tumor hypoxia. Moreover, most of the anti-cancer drugs target the tumor at its early stage, when tumors are still hard to diagnose. ABTAA, instead, works with tumors that are already rooted: “When the tumor is established, hypoxia is the main driver of tumor progression. So, if we eliminate hypoxia, we make the tumor milder, by reducing its progression and metastasis,” comments Koh.
Figure: Schematic drawing of a blood vessel around tumors before and after treatment with ABTAA. The picture above shows a typical tumor vasculature characterized by damaged walls, red blood cells leakage and detached pericytes. Activating Tie2 on endothelial cells with the antibody ABTAA restores the normal vessel architecture: endothelial and pericytes on the vessel walls are stabilized, the delivery of blood is improved, and the anticancer drugs are more likely to reach the tumor core.
The researchers tested ABTAA in mice with three different types of tumors that show high levels of Ang2: glioma (a type of a brain tumor), lung carcinoma, and breast cancer. They also compared the effect of ABTAA with ABA, another antibody that blocks Ang2 but misses the Tie2 activating properties. In all three cases, ABTAA was superior to ABA in inducing tumor vessel normalization, which led to a better delivery of the anti-cancer drugs into the tumor core region.
Glioma is one of the so-called intractable diseases, because of its poor prognosis and treatment. Professor Koh’s team found that the glioma volume was reduced 39% by ABTAA and 17% by ABA. ABTAA profoundly reduced vascular leakage and edema formation in glioma through promoting vascular tightening. Moreover, when ABTAA was administered together with the chemotherapeutic drug temozolomide (TMZ), the tumor volume reduces further (76% by ABTAA+TMZ, 51% by ABA+TMZ, and 36% by TMZ).
In the Lewis Lung Carcinoma (LLC) tumor model, the team administered ABTAA together with a chemotherapeutic drug called cisplatin (Cpt) and observed a greater suppression of tumor growth (52%) compared with the controls and increased overall survival. Moreover, ABTAA+Cpt led to a marked increase in necrotic area within tumors.
Finally, in a spontaneous breast cancer model, ABTAA delayed tumor growth and enhanced the anti-tumor effect of Cpt.
Courtesy of the Institute for Basic Sciences (IBS)
Figure: The antibody ABTAA alone and in combination with other anti-cancer drugs have a beneficial effect in reducing tumor volume. ABTAA was tested in mice with brain tumor (glioma), lung or breast cancer. The image shows the improvements: reduction in glioma tumor size, reduction in metastatic colonies in lung tumor and decrease in necrotic regions in breast tumor.
In the future, the team would like to further understand the underlying relationship between faulty blood vessels and diseases. “We would like to apply this antibody to an organ that is rich in blood vessels, that is the eye, and see if this antibody can be useful to treat eye diseases such as age-related macular degeneration and diabetic retinopathy,” concludes Koh.
Professor Gou-Young Koh (left) and Jin-Sung Park (right)
2016.12.16 View 7611 -
 KAIST and Sejong City Goverment Agree to Establish a Graduate School of Medical Science and Engineering
KAIST and the government of Sejong City will cooperate to establish a graduate school of medical science and engineering. On June 11, 2015, President Steve Kang of KAIST and Mayor Choon-Hee Lee of Sejong City signed a memorandum of understanding at the city hall of Sejong to establish the school.
Under the agreement, the two organizations will work out details to establish the graduate school in Sejong on such issues as administrative assistance, financial support, curriculum development, and the creation of an environment conducive to the growth of medical science.
President Kang said, “Once this graduate school is established, KAIST will be able to offer Korea and the world top-notch researchers in the field of medical science. I have high hopes that the school will produce high-impact research breakthroughs and lead in the advancement of interdisciplinary studies in biotechnology.”
In the picture below, President Steve Kang of KAIST (third from the left) holds the signed memorandum of understanding with Mayor Choon-Hee Lee of Sejong (fourth from the left).
2015.06.16 View 5481
KAIST and Sejong City Goverment Agree to Establish a Graduate School of Medical Science and Engineering
KAIST and the government of Sejong City will cooperate to establish a graduate school of medical science and engineering. On June 11, 2015, President Steve Kang of KAIST and Mayor Choon-Hee Lee of Sejong City signed a memorandum of understanding at the city hall of Sejong to establish the school.
Under the agreement, the two organizations will work out details to establish the graduate school in Sejong on such issues as administrative assistance, financial support, curriculum development, and the creation of an environment conducive to the growth of medical science.
President Kang said, “Once this graduate school is established, KAIST will be able to offer Korea and the world top-notch researchers in the field of medical science. I have high hopes that the school will produce high-impact research breakthroughs and lead in the advancement of interdisciplinary studies in biotechnology.”
In the picture below, President Steve Kang of KAIST (third from the left) holds the signed memorandum of understanding with Mayor Choon-Hee Lee of Sejong (fourth from the left).
2015.06.16 View 5481 -
 Mutations Occurring Only in Brain Responsible for Intractable Epilepsy Identified
KAIST researchers have discovered that brain somatic mutations in MTOR gene induce intractable epilepsy and suggest a precision medicine to treat epileptic seizures.
Epilepsy is a brain disorder which afflicts more than 50 million people worldwide. Many epilepsy patients can control their symptoms through medication, but about 30% suffer from intractable epilepsy and are unable to manage the disease with drugs. Intractable epilepsy causes multiple seizures, permanent mental, physical, and developmental disabilities, and even death. Therefore, surgical removal of the affected area from the brain has been practiced as a treatment for patients with medically refractory seizures, but this too fails to provide a complete solution because only 60% of the patients who undergo surgery are rendered free of seizures.
A Korean research team led by Professor Jeong Ho Lee of the Graduate School of Medical Science and Engineering at the Korea Advanced Institute of Science and Technology (KAIST) and Professor Dong-Seok Kim of Epilepsy Research Center at Yonsei University College of Medicine has recently identified brain somatic mutations in the gene of mechanistic target of rapamycin (MTOR) as the cause of focal cortical dysplasia type II (FCDII), one of the most important and common inducers to intractable epilepsy, particularly in children. They propose a targeted therapy to lessen epileptic seizures by suppressing the activation of mTOR kinase, a signaling protein in the brain. Their research results were published online in Nature Medicine on March 23, 2015.
FCDII contributes to the abnormal developments of the cerebral cortex, ranging from cortical disruption to severe forms of cortical dyslamination, balloon cells, and dysplastic neurons. The research team studied 77 FCDII patients with intractable epilepsy who had received a surgery to remove the affected regions from the brain. The researchers used various deep sequencing technologies to conduct comparative DNA analysis of the samples obtained from the patients’ brain and blood, or saliva. They reported that about 16% of the studied patients had somatic mutations in their brain. Such mutations, however, did not take place in their blood or saliva DNA.
Professor Jeong Ho Lee of KAIST said, “This is an important finding. Unlike our previous belief that genetic mutations causing intractable epilepsy exist anywhere in the human body including blood, specific gene mutations incurred only in the brain can lead to intractable epilepsy. From our animal models, we could see how a small fraction of mutations carrying neurons in the brain could affect its entire function.”
The research team recapitulated the pathogenesis of intractable epilepsy by inducing the focal cortical expression of mutated mTOR in the mouse brain via electroporation method and observed as the mouse develop epileptic symptoms. They then treated these mice with the drug called “rapamycin” to inhibit the activity of mTOR protein and observed that it suppressed the development of epileptic seizures with cytomegalic neurons.
“Our study offers the first evidence that brain-somatic activating mutations in MTOR cause FCDII and identifies mTOR as a treatment target for intractable epilepsy,” said co-author Dr. Dong-Seok Kim, a neurosurgeon at Yonsei Medical Center with the country’s largest surgical experiences in treating patients with this condition.
The research paper is titled “Brain somatic mutations in MTOR cause focal cortical dysplasia type II leading to intractable epilepsy.” (Digital Object Identifier #: 10.1038/nm.3824)
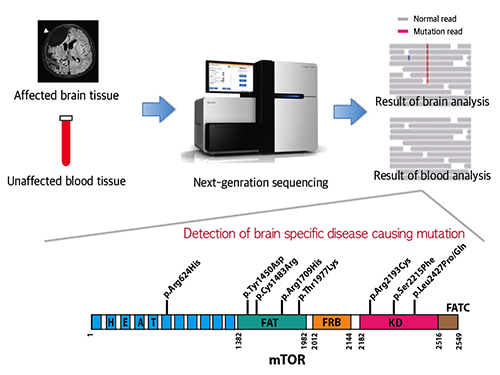
Picture 1: A schematic image to show how to detect brain specific mutation using next-generation sequencing technology with blood-brain paired sample. Simple comparison of non-overlapping mutations between affected and unaffected tissues is able to detect brain specific mutations.
Picture 2: A schematic image to show how to generate focal cortical dysplasia mouse model. This mouse model open the new window of drug screening for seizure patients.
Picture 3: Targeted medicine can rescue the focal cortical dysplasia symptoms including cytomegalic neuron & intractable epilepsy.
2015.03.25 View 12540
Mutations Occurring Only in Brain Responsible for Intractable Epilepsy Identified
KAIST researchers have discovered that brain somatic mutations in MTOR gene induce intractable epilepsy and suggest a precision medicine to treat epileptic seizures.
Epilepsy is a brain disorder which afflicts more than 50 million people worldwide. Many epilepsy patients can control their symptoms through medication, but about 30% suffer from intractable epilepsy and are unable to manage the disease with drugs. Intractable epilepsy causes multiple seizures, permanent mental, physical, and developmental disabilities, and even death. Therefore, surgical removal of the affected area from the brain has been practiced as a treatment for patients with medically refractory seizures, but this too fails to provide a complete solution because only 60% of the patients who undergo surgery are rendered free of seizures.
A Korean research team led by Professor Jeong Ho Lee of the Graduate School of Medical Science and Engineering at the Korea Advanced Institute of Science and Technology (KAIST) and Professor Dong-Seok Kim of Epilepsy Research Center at Yonsei University College of Medicine has recently identified brain somatic mutations in the gene of mechanistic target of rapamycin (MTOR) as the cause of focal cortical dysplasia type II (FCDII), one of the most important and common inducers to intractable epilepsy, particularly in children. They propose a targeted therapy to lessen epileptic seizures by suppressing the activation of mTOR kinase, a signaling protein in the brain. Their research results were published online in Nature Medicine on March 23, 2015.
FCDII contributes to the abnormal developments of the cerebral cortex, ranging from cortical disruption to severe forms of cortical dyslamination, balloon cells, and dysplastic neurons. The research team studied 77 FCDII patients with intractable epilepsy who had received a surgery to remove the affected regions from the brain. The researchers used various deep sequencing technologies to conduct comparative DNA analysis of the samples obtained from the patients’ brain and blood, or saliva. They reported that about 16% of the studied patients had somatic mutations in their brain. Such mutations, however, did not take place in their blood or saliva DNA.
Professor Jeong Ho Lee of KAIST said, “This is an important finding. Unlike our previous belief that genetic mutations causing intractable epilepsy exist anywhere in the human body including blood, specific gene mutations incurred only in the brain can lead to intractable epilepsy. From our animal models, we could see how a small fraction of mutations carrying neurons in the brain could affect its entire function.”
The research team recapitulated the pathogenesis of intractable epilepsy by inducing the focal cortical expression of mutated mTOR in the mouse brain via electroporation method and observed as the mouse develop epileptic symptoms. They then treated these mice with the drug called “rapamycin” to inhibit the activity of mTOR protein and observed that it suppressed the development of epileptic seizures with cytomegalic neurons.
“Our study offers the first evidence that brain-somatic activating mutations in MTOR cause FCDII and identifies mTOR as a treatment target for intractable epilepsy,” said co-author Dr. Dong-Seok Kim, a neurosurgeon at Yonsei Medical Center with the country’s largest surgical experiences in treating patients with this condition.
The research paper is titled “Brain somatic mutations in MTOR cause focal cortical dysplasia type II leading to intractable epilepsy.” (Digital Object Identifier #: 10.1038/nm.3824)
Picture 1: A schematic image to show how to detect brain specific mutation using next-generation sequencing technology with blood-brain paired sample. Simple comparison of non-overlapping mutations between affected and unaffected tissues is able to detect brain specific mutations.
Picture 2: A schematic image to show how to generate focal cortical dysplasia mouse model. This mouse model open the new window of drug screening for seizure patients.
Picture 3: Targeted medicine can rescue the focal cortical dysplasia symptoms including cytomegalic neuron & intractable epilepsy.
2015.03.25 View 12540 -
 Structure of Neuron-Connecting Synaptic Adhesion Molecules Discovered
A research team has found the three-dimensional structure of synaptic adhesion molecules, which orchestrate synaptogenesis. The research findings also propose the mechanism of synapses in its initial formation. Some brain diseases such as obsessive compulsive disorder (OCD) or bipolar disorders arise from a malfunction of synapses. The team expects the findings to be applied in investigating pathogenesis and developing medicines for such diseases.
The research was conducted by a Master’s candidate Kee Hun Kim, Professor Ji Won Um from Yonsei University, and Professor Beom Seok Park from Eulji University under the guidance of Professor Homin Kim from the Graduate School of Medical Science and Engineering, KAIST, and Professor Jaewon Ko from Yonsei University. Sponsored by the Ministry of Science, ICT and Future Planning and the National Research Foundation of Korea, the research findings were published online in the November 14th issue of Nature Communications.
A protein that exists in the neuronal transmembrane, Slitrk, interacts with the presynaptic leukocyte common antigen-related receptor protein tyrosine phosphatases (LAR-RPTPs) and forms a protein complex. It is involved in the development of synapses in the initial stage, and balances excitatory and inhibitory signals of neurons.
It is known that a disorder in those two proteins cause a malfunction of synapses, resulting in neuropsychosis such as autism, epilepsy, OCD, and bipolar disorders. However, because the structure as well as synaptogenic function of these proteins were not understood, the development of cures could not progress.
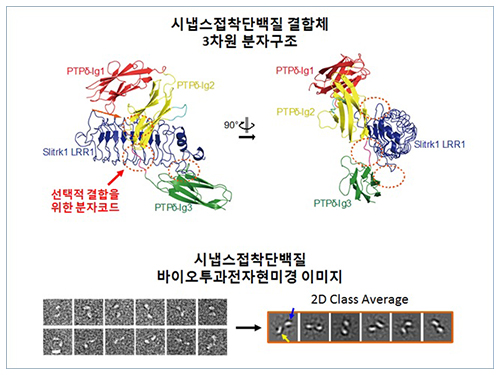
The research team discovered the three-dimensional structure of two synaptic adhesion molecules like Slitrk and LAR-RPTPs and identified the regions of interaction through protein crystallography and transmission electron microscopy (TEM). Furthermore, they found that the formation of the synapse is induced after the combination of two synaptic adhesion molecules develops a cluster.
Professor Kim said, “The research findings will serve as a basis of understanding the pathogenesis of brain diseases which arises from a malfunction of synaptic adhesion molecules. In particular, this is a good example in which collaboration between structural biology and neurobiology has led to a fruitful result.” Professor Ko commented that “this will give new directions to synaptic formation-related-researches by revealing the molecular mechanism of synaptic adhesion molecules.”
Figure 1: Overview of the PTPd Ig1–3/Slitrk1 LRR1 complex.
Figure 2: Representative negative-stained electron microscopy images of Slitrk1 Full ectodomain (yellow arrows indicate the horseshoe-shaped LRR domains). The typical horseshoe-shaped structures and the randomness of the relative positions of each LRR domain can be observed from the two-dimensional class averages displayed in the orange box.
Figure 3: Model of the two-step presynaptic differentiation process mediated by the biding of Slitrks to LAR-RPTPs and subsequent lateral assembly of trans-synaptic LAR-RPTPs/Slitrik complexes.
2014.11.28 View 10102
Structure of Neuron-Connecting Synaptic Adhesion Molecules Discovered
A research team has found the three-dimensional structure of synaptic adhesion molecules, which orchestrate synaptogenesis. The research findings also propose the mechanism of synapses in its initial formation. Some brain diseases such as obsessive compulsive disorder (OCD) or bipolar disorders arise from a malfunction of synapses. The team expects the findings to be applied in investigating pathogenesis and developing medicines for such diseases.
The research was conducted by a Master’s candidate Kee Hun Kim, Professor Ji Won Um from Yonsei University, and Professor Beom Seok Park from Eulji University under the guidance of Professor Homin Kim from the Graduate School of Medical Science and Engineering, KAIST, and Professor Jaewon Ko from Yonsei University. Sponsored by the Ministry of Science, ICT and Future Planning and the National Research Foundation of Korea, the research findings were published online in the November 14th issue of Nature Communications.
A protein that exists in the neuronal transmembrane, Slitrk, interacts with the presynaptic leukocyte common antigen-related receptor protein tyrosine phosphatases (LAR-RPTPs) and forms a protein complex. It is involved in the development of synapses in the initial stage, and balances excitatory and inhibitory signals of neurons.
It is known that a disorder in those two proteins cause a malfunction of synapses, resulting in neuropsychosis such as autism, epilepsy, OCD, and bipolar disorders. However, because the structure as well as synaptogenic function of these proteins were not understood, the development of cures could not progress.
The research team discovered the three-dimensional structure of two synaptic adhesion molecules like Slitrk and LAR-RPTPs and identified the regions of interaction through protein crystallography and transmission electron microscopy (TEM). Furthermore, they found that the formation of the synapse is induced after the combination of two synaptic adhesion molecules develops a cluster.
Professor Kim said, “The research findings will serve as a basis of understanding the pathogenesis of brain diseases which arises from a malfunction of synaptic adhesion molecules. In particular, this is a good example in which collaboration between structural biology and neurobiology has led to a fruitful result.” Professor Ko commented that “this will give new directions to synaptic formation-related-researches by revealing the molecular mechanism of synaptic adhesion molecules.”
Figure 1: Overview of the PTPd Ig1–3/Slitrk1 LRR1 complex.
Figure 2: Representative negative-stained electron microscopy images of Slitrk1 Full ectodomain (yellow arrows indicate the horseshoe-shaped LRR domains). The typical horseshoe-shaped structures and the randomness of the relative positions of each LRR domain can be observed from the two-dimensional class averages displayed in the orange box.
Figure 3: Model of the two-step presynaptic differentiation process mediated by the biding of Slitrks to LAR-RPTPs and subsequent lateral assembly of trans-synaptic LAR-RPTPs/Slitrik complexes.
2014.11.28 View 10102 -
 Regulatory T Cells Influence Liver Damage of Hepatitis A Patients
Liver damage becomes more severe with the decrease of regulatory T cells
“This research will aid the development of hepatitis A targeted drug,” said a KAIST researcher.
The KAIST Graduate School of Medical Science and Engineering’s Professor Eui-Cheol Shin and his research team have identified the mechanism, explaining how the regulatory T cells are responsible for the body’s immune system and how they have induced liver damage of hepatitis A patients.
The research results were published online in the July 9th edition of ‘Gut,’ the world’s most prominent journal in the field of gastroenterology.
Hepatitis A is an acute form of hepatitis caused by hepatitis A virus. The virus spreads through oral contact and enters the body via digestive organs.
Regulatory T cells play an important role in maintaining the homeostasis of the body’s immune system by inhibiting the activation of other immune cells. In the case of chronic viral infections, regulatory T cells are known to contribute to the duration of the infection, weakening the immune response to virus infections. However, there has been no information on what roles the regulatory T cells perform in the case of acute viral infections.
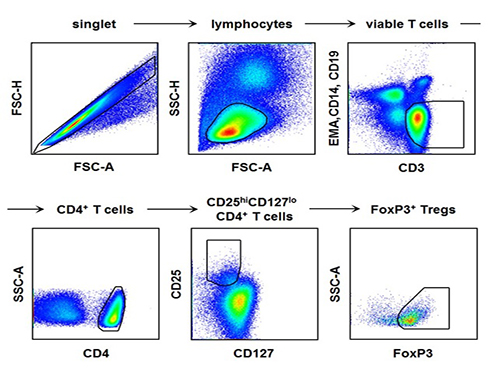
The research team used the fluorescence flow cytometry technique to determine the number and characteristics of a variety of immune cells, including regulatory T cells, in the blood of hepatitis A patients.
Consequently, the researchers confirmed that the decrease in the regulatory T cells immune inhibitory ability was consistent with a significant reduction in the number of regulatory T cells in the blood of hepatitis A patients. Furthermore, it was identified that the more noticeable decrease of regulatory T cells led to the occurrence of a more severe liver injury.
The analysis of hepatitis A patient’s blood proved that the cause of the decrease in the number and function of regulatory T cells was the increased expression of cell surface protein ‘Fas,’ which induces cell death.
Professor Shin said, “This study is the first case which proposes the mechanism for clinical aspects in not only hepatitis A, but also acute virus infection.” He added on the future prospect of the research that: “In the future, we can prevent tissue damage by inhibiting cell death of regulatory T cells for severe acute viral infections that do not have an effective treatment for the virus itself.”
[Picture]
The picture shows the process of fluorescence flow cytometry technique to study regulatory T cell in the blood of hepatitis A patients.
2014.08.11 View 9049
Regulatory T Cells Influence Liver Damage of Hepatitis A Patients
Liver damage becomes more severe with the decrease of regulatory T cells
“This research will aid the development of hepatitis A targeted drug,” said a KAIST researcher.
The KAIST Graduate School of Medical Science and Engineering’s Professor Eui-Cheol Shin and his research team have identified the mechanism, explaining how the regulatory T cells are responsible for the body’s immune system and how they have induced liver damage of hepatitis A patients.
The research results were published online in the July 9th edition of ‘Gut,’ the world’s most prominent journal in the field of gastroenterology.
Hepatitis A is an acute form of hepatitis caused by hepatitis A virus. The virus spreads through oral contact and enters the body via digestive organs.
Regulatory T cells play an important role in maintaining the homeostasis of the body’s immune system by inhibiting the activation of other immune cells. In the case of chronic viral infections, regulatory T cells are known to contribute to the duration of the infection, weakening the immune response to virus infections. However, there has been no information on what roles the regulatory T cells perform in the case of acute viral infections.
The research team used the fluorescence flow cytometry technique to determine the number and characteristics of a variety of immune cells, including regulatory T cells, in the blood of hepatitis A patients.
Consequently, the researchers confirmed that the decrease in the regulatory T cells immune inhibitory ability was consistent with a significant reduction in the number of regulatory T cells in the blood of hepatitis A patients. Furthermore, it was identified that the more noticeable decrease of regulatory T cells led to the occurrence of a more severe liver injury.
The analysis of hepatitis A patient’s blood proved that the cause of the decrease in the number and function of regulatory T cells was the increased expression of cell surface protein ‘Fas,’ which induces cell death.
Professor Shin said, “This study is the first case which proposes the mechanism for clinical aspects in not only hepatitis A, but also acute virus infection.” He added on the future prospect of the research that: “In the future, we can prevent tissue damage by inhibiting cell death of regulatory T cells for severe acute viral infections that do not have an effective treatment for the virus itself.”
[Picture]
The picture shows the process of fluorescence flow cytometry technique to study regulatory T cell in the blood of hepatitis A patients.
2014.08.11 View 9049 -
 Immune Evasion Mechanism of Hepatitis C Virus Revealed
Professor Ui-Cheol Shin
Inhibiting major histocompatibility complex [MHC] class I protein expression, T cell immune response is evaded.
The research will be a great help to the development of C hepatitis vaccine.
Roughly 1-2% of the population in Korea is known to be infected with Hepatitis C. Most Hepatitis C Virus (HCV) infections progress to a chronic disease and can cause liver cirrhosis or liver cancer, which may lead to death.
Unlike Hepatitis type A or B, there is no vaccine for Hepatitis C Virus and therefore avoiding exposure to the virus is the best known method of prevention. However, a team of researchers at KAIST has produced research results, which may contribute significantly to the vaccine development.
KAIST Graduate School of Medical Sciences & Engineering’ Professor Ui-Cheol Shin and his team have successfully identified why Hepatitis C Virus does not cause an immune response within the human body. The research results were published in the May edition of The Journal of Gastroenterology, a world-renowned journal in the field of gastroenterology.
The immune response occurs to eliminate the virus that has invaded our body. During this process, a major histocompatibility complex [MHC] class I plays a key role in inducing T cell response, which is needed for the elimination of virus-infected cells.
When a cell is infected by a virus, a substance called interferon causes the increased expression of major histocompatibility complex class I. T cell recognizes the increased MHC class I and therefore finds the virus-infected cells.
However, the effect that Hepatitis C Virus has on major histocompatibility complex class I has not been clearly identified until now.
The research team has revealed, using a cell culture for infection systems, that the Hepatitis C Virus suppresses the expression of major histocompatibility complex class I. Also, the mechanism to prove that HCV activates a protein called PKR within the cell to inhibit MHC class I protein expression was identified at a molecular level.
In this study, researchers established the hypothesis that regulating PKR protein in the cell can enhance the T cell immune response, which was then proved through experiments.
Professor Ui-Cheol Shin said, “There are a lot of new drugs to treat Hepatitis C Virus, while its vaccine has not been developed yet. Revealing the HCV immune evasion mechanism will help stimulate momentum for the HCV vaccine development.”
The first author of the journal, Dr. Won-Seok Kang is a graduate from Yonsei College of Medicine. After earning his medical degree, he has continued his training as a ‘doctor-scientist’ at KAIST Graduate School of Medical Sciences & Engineering to study Hepatitis C Virus immune evasion mechanism in this research.
Hepatitis C Virus activates PKR-eIF2a pathway, which inhibits the major histocompatibility complex class I, and therefore weakens the T cell activation to the viral activity.
2014.05.19 View 9849
Immune Evasion Mechanism of Hepatitis C Virus Revealed
Professor Ui-Cheol Shin
Inhibiting major histocompatibility complex [MHC] class I protein expression, T cell immune response is evaded.
The research will be a great help to the development of C hepatitis vaccine.
Roughly 1-2% of the population in Korea is known to be infected with Hepatitis C. Most Hepatitis C Virus (HCV) infections progress to a chronic disease and can cause liver cirrhosis or liver cancer, which may lead to death.
Unlike Hepatitis type A or B, there is no vaccine for Hepatitis C Virus and therefore avoiding exposure to the virus is the best known method of prevention. However, a team of researchers at KAIST has produced research results, which may contribute significantly to the vaccine development.
KAIST Graduate School of Medical Sciences & Engineering’ Professor Ui-Cheol Shin and his team have successfully identified why Hepatitis C Virus does not cause an immune response within the human body. The research results were published in the May edition of The Journal of Gastroenterology, a world-renowned journal in the field of gastroenterology.
The immune response occurs to eliminate the virus that has invaded our body. During this process, a major histocompatibility complex [MHC] class I plays a key role in inducing T cell response, which is needed for the elimination of virus-infected cells.
When a cell is infected by a virus, a substance called interferon causes the increased expression of major histocompatibility complex class I. T cell recognizes the increased MHC class I and therefore finds the virus-infected cells.
However, the effect that Hepatitis C Virus has on major histocompatibility complex class I has not been clearly identified until now.
The research team has revealed, using a cell culture for infection systems, that the Hepatitis C Virus suppresses the expression of major histocompatibility complex class I. Also, the mechanism to prove that HCV activates a protein called PKR within the cell to inhibit MHC class I protein expression was identified at a molecular level.
In this study, researchers established the hypothesis that regulating PKR protein in the cell can enhance the T cell immune response, which was then proved through experiments.
Professor Ui-Cheol Shin said, “There are a lot of new drugs to treat Hepatitis C Virus, while its vaccine has not been developed yet. Revealing the HCV immune evasion mechanism will help stimulate momentum for the HCV vaccine development.”
The first author of the journal, Dr. Won-Seok Kang is a graduate from Yonsei College of Medicine. After earning his medical degree, he has continued his training as a ‘doctor-scientist’ at KAIST Graduate School of Medical Sciences & Engineering to study Hepatitis C Virus immune evasion mechanism in this research.
Hepatitis C Virus activates PKR-eIF2a pathway, which inhibits the major histocompatibility complex class I, and therefore weakens the T cell activation to the viral activity.
2014.05.19 View 9849 -
 Complex responsible for protein breakdown in cells identified using Bio TEM
Professor Ho-Min Kim
- High resolution 3D structure analysis success using Bio Transmission Electron Microscopy (TEM), a giant step towards new anticancer treatment development
- Published in Nature on May 5th
Using TEM to observe protein molecules and analysing its high resolution 3D structure is now possible. KAIST Biomedical Science and Engineering Department’s Professor Ho-Min Kim has identified the high resolution structure of proteasome complexes, which is responsible for protein breakdown in cells, using Bio TEM.
This research has been published on the world"s most prestigious journal, Nature, online on May 5th. Our body controls many cellular processes through production and degradation of proteins to maintain homeostasis. A proteasome complex acts as a garbage disposal system and degrades cellular proteins when needed for regulation, which is one of the central roles of the body.
However, a mutation in proteasome complex leads to diseases such as cancer, degenerative brain diseases, and autoimmune diseases.
Currently, the anticancer drug Velcade is used to decrease proteasome function to treat Multiple Myeloma, a form of blood cancer. Research concerning proteasome complexes for more effective anticancer drugs and treatments with fewer side effects has been taking place for more than 20 years. There have been many difficulties in understanding proteasome function through 3D structure analysis since a proteasome complex, consisting of around 30 different proteins, has a great size and complexity.
The research team used Bio TEM instead of conventionally used protein crystallography technique. The protein sample was inserted into Bio TEM, hundreds of photographs were taken from various angles, and then a high–performance computer was used to analyse its structure. Bio TEM requires a smaller sample and can analyse the complexes of great size of proteins.
Professor Ho-Min Kim said, “Identifying proteasome complex assembly process and 3D structure will increase our understanding of cellular protein degradation process and hence assist in new drug development using this knowledge.” He added, “High resolution protein structure analysis using Bio TEM, used for the first time in Korea, will enable us to observe structure analysis of large protein complexes that were difficult to approach using protein crystallography.” Professor Kim continued, “If protein crystallography technology and Bio TEM could be used together to complement one another, it would bring a great synergetic effect to protein complex 3D structure analysis research in the future.”
Professor Ho-Min Kim has conducted this research since his post-doctorate at the University of California, San Francisco, under the advice of Professor Yifan Cheng; in co-operation with Harvard University and Colorado University.
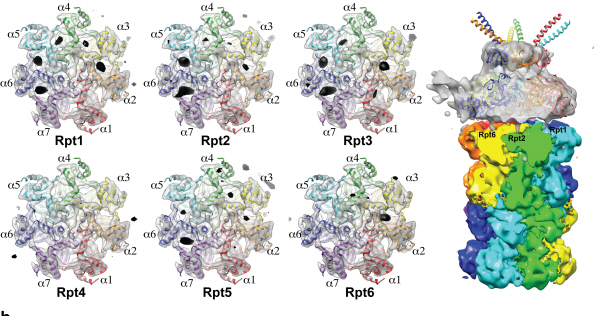
Figure 1: A picture taken by Bio TEM of open state protein sample (proteasome complex)
Figure 2: Bio TEM image analysis showing protein 3D structure
2013.05.25 View 9352
Complex responsible for protein breakdown in cells identified using Bio TEM
Professor Ho-Min Kim
- High resolution 3D structure analysis success using Bio Transmission Electron Microscopy (TEM), a giant step towards new anticancer treatment development
- Published in Nature on May 5th
Using TEM to observe protein molecules and analysing its high resolution 3D structure is now possible. KAIST Biomedical Science and Engineering Department’s Professor Ho-Min Kim has identified the high resolution structure of proteasome complexes, which is responsible for protein breakdown in cells, using Bio TEM.
This research has been published on the world"s most prestigious journal, Nature, online on May 5th. Our body controls many cellular processes through production and degradation of proteins to maintain homeostasis. A proteasome complex acts as a garbage disposal system and degrades cellular proteins when needed for regulation, which is one of the central roles of the body.
However, a mutation in proteasome complex leads to diseases such as cancer, degenerative brain diseases, and autoimmune diseases.
Currently, the anticancer drug Velcade is used to decrease proteasome function to treat Multiple Myeloma, a form of blood cancer. Research concerning proteasome complexes for more effective anticancer drugs and treatments with fewer side effects has been taking place for more than 20 years. There have been many difficulties in understanding proteasome function through 3D structure analysis since a proteasome complex, consisting of around 30 different proteins, has a great size and complexity.
The research team used Bio TEM instead of conventionally used protein crystallography technique. The protein sample was inserted into Bio TEM, hundreds of photographs were taken from various angles, and then a high–performance computer was used to analyse its structure. Bio TEM requires a smaller sample and can analyse the complexes of great size of proteins.
Professor Ho-Min Kim said, “Identifying proteasome complex assembly process and 3D structure will increase our understanding of cellular protein degradation process and hence assist in new drug development using this knowledge.” He added, “High resolution protein structure analysis using Bio TEM, used for the first time in Korea, will enable us to observe structure analysis of large protein complexes that were difficult to approach using protein crystallography.” Professor Kim continued, “If protein crystallography technology and Bio TEM could be used together to complement one another, it would bring a great synergetic effect to protein complex 3D structure analysis research in the future.”
Professor Ho-Min Kim has conducted this research since his post-doctorate at the University of California, San Francisco, under the advice of Professor Yifan Cheng; in co-operation with Harvard University and Colorado University.
Figure 1: A picture taken by Bio TEM of open state protein sample (proteasome complex)
Figure 2: Bio TEM image analysis showing protein 3D structure
2013.05.25 View 9352