Science
-
 Mystery Solved with Math: Cytoplasmic Traffic Jam Disrupts Sleep-Wake Cycles
KAIST mathematicians and their collaborators at Florida State University have identified the principle of how aging and diseases like dementia and obesity cause sleep disorders. A combination of mathematical modelling and experiments demonstrated that the cytoplasmic congestion caused by aging, dementia, and/or obesity disrupts the circadian rhythms in the human body and leads to irregular sleep-wake cycles. This finding suggests new treatment strategies for addressing unstable sleep-wake cycles.
Human bodies adjust sleep schedules in accordance with the ‘circadian rhythms’, which are regulated by our time keeping system, the ‘circadian clock’. This clock tells our body when to rest by generating the 24-hour rhythms of a protein called PERIOD (PER) (See Figure 1).
The amount of the PER protein increases for half of the day and then decreases for the remaining half. The principle is that the PER protein accumulating in the cytoplasm for several hours enters the cell nucleus all at once, hindering the transcription of PER genes and thereby reducing the amount of PER.
However, it has remained a mystery how thousands of PER molecules can simultaneously enter into the nucleus in a complex cell environment where a variety of materials co-exist and can interfere with the motion of PER. This would be like finding a way for thousands of employees from all over New York City to enter an office building at the same time every day.
A group of researchers led by Professor Jae Kyoung Kim from the KAIST Department of Mathematical Sciences solved the mystery by developing a spatiotemporal and probabilistic model that describes the motion of PER molecules in a cell environment.
This study was conducted in collaboration with Professor Choogon Lee’s group from Florida State University, where the experiments were carried out, and the results were published in the Proceedings of the National Academy of Sciences (PNAS) last month.
The joint research team’s spatial stochastic model (See Figure 2) described the motion of PER molecules in cells and demonstrated that the PER molecule should be sufficiently condensed around the cell nucleus to be phosphorylated simultaneously and enter the nucleus together (See Figure 3 Left). Thanks to this phosphorylation synchronization switch, thousands of PER molecules can enter the nucleus at the same time every day and maintain stable circadian rhythms.
However, when aging and/or diseases including dementia and obesity cause the cytoplasm to become congested with increased cytoplasmic obstacles such as protein aggregates and fat vacuoles, it hinders the timely condensation of PER molecules around the cell nucleus (See Figure 3 Right). As a result, the phosphorylation synchronization switch does not work and PER proteins enter into the nucleus at irregular times, making the circadian rhythms and sleep-wake cycles unstable, the study revealed.
Professor Kim said, “As a mathematician, I am excited to help enable the advancement of new treatment strategies that can improve the lives of so many patients who suffer from irregular sleep-wake cycles. Taking these findings as an opportunity, I hope to see more active interchanges of ideas and collaboration between mathematical and biological sciences.”
This work was supported by the National Institutes of Health and the National Science Foundation in the US, and the International Human Frontiers Science Program Organization and the National Research Foundation of Korea.
Publication:
Beesley, S. and Kim, D. W, et al. (2020) Wake-sleep cycles are severely disrupted by diseases affecting cytoplasmic homeostasis. Proceedings of the National Academy of Sciences (PNAS), Vol. 117, No. 45, 28402-28411. Available online at https://doi.org/10.1073/pnas.2003524117
Profile:
Jae Kyoung Kim, Ph.D.
Associate Professor
jaekkim@kaist.ac.kr
http://mathsci.kaist.ac.kr/~jaekkim
@umichkim on Twitter
Department of Mathematical Sciences
Korea Advanced Institute of Science and Technology (KAIST)
Daejeon, Republic of Korea
Profile:
Choogon Lee, Ph.D.
Associate Professor
clee@neuro.fsu.edu
https://med.fsu.edu/biosci/lee-lab
Department of Biomedical Sciences
Florida State University
Florida, USA
(END)
2020.12.11 View 6079
Mystery Solved with Math: Cytoplasmic Traffic Jam Disrupts Sleep-Wake Cycles
KAIST mathematicians and their collaborators at Florida State University have identified the principle of how aging and diseases like dementia and obesity cause sleep disorders. A combination of mathematical modelling and experiments demonstrated that the cytoplasmic congestion caused by aging, dementia, and/or obesity disrupts the circadian rhythms in the human body and leads to irregular sleep-wake cycles. This finding suggests new treatment strategies for addressing unstable sleep-wake cycles.
Human bodies adjust sleep schedules in accordance with the ‘circadian rhythms’, which are regulated by our time keeping system, the ‘circadian clock’. This clock tells our body when to rest by generating the 24-hour rhythms of a protein called PERIOD (PER) (See Figure 1).
The amount of the PER protein increases for half of the day and then decreases for the remaining half. The principle is that the PER protein accumulating in the cytoplasm for several hours enters the cell nucleus all at once, hindering the transcription of PER genes and thereby reducing the amount of PER.
However, it has remained a mystery how thousands of PER molecules can simultaneously enter into the nucleus in a complex cell environment where a variety of materials co-exist and can interfere with the motion of PER. This would be like finding a way for thousands of employees from all over New York City to enter an office building at the same time every day.
A group of researchers led by Professor Jae Kyoung Kim from the KAIST Department of Mathematical Sciences solved the mystery by developing a spatiotemporal and probabilistic model that describes the motion of PER molecules in a cell environment.
This study was conducted in collaboration with Professor Choogon Lee’s group from Florida State University, where the experiments were carried out, and the results were published in the Proceedings of the National Academy of Sciences (PNAS) last month.
The joint research team’s spatial stochastic model (See Figure 2) described the motion of PER molecules in cells and demonstrated that the PER molecule should be sufficiently condensed around the cell nucleus to be phosphorylated simultaneously and enter the nucleus together (See Figure 3 Left). Thanks to this phosphorylation synchronization switch, thousands of PER molecules can enter the nucleus at the same time every day and maintain stable circadian rhythms.
However, when aging and/or diseases including dementia and obesity cause the cytoplasm to become congested with increased cytoplasmic obstacles such as protein aggregates and fat vacuoles, it hinders the timely condensation of PER molecules around the cell nucleus (See Figure 3 Right). As a result, the phosphorylation synchronization switch does not work and PER proteins enter into the nucleus at irregular times, making the circadian rhythms and sleep-wake cycles unstable, the study revealed.
Professor Kim said, “As a mathematician, I am excited to help enable the advancement of new treatment strategies that can improve the lives of so many patients who suffer from irregular sleep-wake cycles. Taking these findings as an opportunity, I hope to see more active interchanges of ideas and collaboration between mathematical and biological sciences.”
This work was supported by the National Institutes of Health and the National Science Foundation in the US, and the International Human Frontiers Science Program Organization and the National Research Foundation of Korea.
Publication:
Beesley, S. and Kim, D. W, et al. (2020) Wake-sleep cycles are severely disrupted by diseases affecting cytoplasmic homeostasis. Proceedings of the National Academy of Sciences (PNAS), Vol. 117, No. 45, 28402-28411. Available online at https://doi.org/10.1073/pnas.2003524117
Profile:
Jae Kyoung Kim, Ph.D.
Associate Professor
jaekkim@kaist.ac.kr
http://mathsci.kaist.ac.kr/~jaekkim
@umichkim on Twitter
Department of Mathematical Sciences
Korea Advanced Institute of Science and Technology (KAIST)
Daejeon, Republic of Korea
Profile:
Choogon Lee, Ph.D.
Associate Professor
clee@neuro.fsu.edu
https://med.fsu.edu/biosci/lee-lab
Department of Biomedical Sciences
Florida State University
Florida, USA
(END)
2020.12.11 View 6079 -
 Drawing the Line to Answer Art’s Big Questions
- KAIST scientists show how statistical physics can reveal art trends across time and culture. -
Algorithms have shown that the compositional structure of Western landscape paintings changed “suspiciously” smoothly between 1500 and 2000 AD, potentially indicating a selection bias by art curators or in art historical literature, physicists from the Korea Advanced Institute of Science and Technology (KAIST) and colleagues report in the Proceedings of the National Academy of Sciences (PNAS).
KAIST statistical physicist Hawoong Jeong worked with statisticians, digital analysts and art historians in Korea, Estonia and the US to clarify whether computer algorithms could help resolve long-standing questions about design principles used in landscape paintings, such as the placement of the horizon and other primary features.
“A foundational question among art historians is whether artwork contains organizing principles that transcend culture and time and, if yes, how these principles evolved over time,” explains Jeong. “We developed an information-theoretic approach that can capture compositional proportion in landscape paintings and found that the preferred compositional proportion systematically evolved over time.”
Digital versions of almost 15,000 canonical landscape paintings from the Western renaissance in the 1500s to the more recent contemporary art period were run through a computer algorithm. The algorithm progressively divides artwork into horizontal and vertical lines depending on the amount of information in each subsequent partition. It allows scientists to evaluate how artists and various art styles compose landscape artwork, in terms of placement of a piece’s most important components, in addition to how high or low the landscape’s horizon is placed.
The scientists started by analysing the first two partitioning lines identified by the algorithm in the paintings and found they could be categorized into four groups: an initial horizontal line followed by a second horizontal line (H-H); an initial horizontal line followed by a second vertical line (H-V); a vertical followed by horizontal line (V-H); or a vertical followed by a vertical line (V-V) (see image 1 and 2). They then looked at the categorizations over time.
They found that before the mid-nineteenth century, H-V was the dominant composition type, followed by H-H, V-H, and V-V. The mid-nineteenth century then brought change, with the H-V composition style decreasing in popularity with a rise in the H-H composition style. The other two styles remained relatively stable.
The scientists also looked at how the horizon line, which separates sky from land, changed over time. In the 16th century, the dominant horizon line of the painting was above the middle of the canvas, but it gradually descended to the lower middle of the canvas by the 17th century, where it remained until the mid-nineteenth century. After that, the horizon line began gradually rising again.
Interestingly, the algorithm showed that these findings were similar across cultures and artistic periods, even through periods dominated by a diversity in art styles. This similarity may well be a function, then, of a bias in the dataset.
“In recent decades, art historians have prioritized the argument that there is great diversity in the evolution of artistic expression rather than offering a relatively smoother consensus story in Western art,” Jeong says. “This study serves as a reminder that the available large-scale datasets might be perpetuating severe biases.”
The scientists next aim to broaden their analyses to include more diverse artwork, as this particular dataset was ultimately Western and male biased. Future analyses should also consider diagonal compositions in paintings, they say.
This work was supported by the National Research Foundation (NRF) of Korea.
Publication:
Lee, B, et al. (2020) Dissecting landscape art history with information theory. Proceedings of the National Academy of Sciences (PNAS), Vol. 117, No. 43, 26580-26590. Available online at https://doi.org/10.1073/pnas.2011927117
Profile:
Hawoong Jeong, Ph.D.
Professor
hjeong@kaist.ac.kr
https://www.kaist.ac.kr
Department of Physics
Korea Advanced Institute of Science and Technology (KAIST)
Daejeon, Republic of Korea
(END)
2020.11.13 View 7292
Drawing the Line to Answer Art’s Big Questions
- KAIST scientists show how statistical physics can reveal art trends across time and culture. -
Algorithms have shown that the compositional structure of Western landscape paintings changed “suspiciously” smoothly between 1500 and 2000 AD, potentially indicating a selection bias by art curators or in art historical literature, physicists from the Korea Advanced Institute of Science and Technology (KAIST) and colleagues report in the Proceedings of the National Academy of Sciences (PNAS).
KAIST statistical physicist Hawoong Jeong worked with statisticians, digital analysts and art historians in Korea, Estonia and the US to clarify whether computer algorithms could help resolve long-standing questions about design principles used in landscape paintings, such as the placement of the horizon and other primary features.
“A foundational question among art historians is whether artwork contains organizing principles that transcend culture and time and, if yes, how these principles evolved over time,” explains Jeong. “We developed an information-theoretic approach that can capture compositional proportion in landscape paintings and found that the preferred compositional proportion systematically evolved over time.”
Digital versions of almost 15,000 canonical landscape paintings from the Western renaissance in the 1500s to the more recent contemporary art period were run through a computer algorithm. The algorithm progressively divides artwork into horizontal and vertical lines depending on the amount of information in each subsequent partition. It allows scientists to evaluate how artists and various art styles compose landscape artwork, in terms of placement of a piece’s most important components, in addition to how high or low the landscape’s horizon is placed.
The scientists started by analysing the first two partitioning lines identified by the algorithm in the paintings and found they could be categorized into four groups: an initial horizontal line followed by a second horizontal line (H-H); an initial horizontal line followed by a second vertical line (H-V); a vertical followed by horizontal line (V-H); or a vertical followed by a vertical line (V-V) (see image 1 and 2). They then looked at the categorizations over time.
They found that before the mid-nineteenth century, H-V was the dominant composition type, followed by H-H, V-H, and V-V. The mid-nineteenth century then brought change, with the H-V composition style decreasing in popularity with a rise in the H-H composition style. The other two styles remained relatively stable.
The scientists also looked at how the horizon line, which separates sky from land, changed over time. In the 16th century, the dominant horizon line of the painting was above the middle of the canvas, but it gradually descended to the lower middle of the canvas by the 17th century, where it remained until the mid-nineteenth century. After that, the horizon line began gradually rising again.
Interestingly, the algorithm showed that these findings were similar across cultures and artistic periods, even through periods dominated by a diversity in art styles. This similarity may well be a function, then, of a bias in the dataset.
“In recent decades, art historians have prioritized the argument that there is great diversity in the evolution of artistic expression rather than offering a relatively smoother consensus story in Western art,” Jeong says. “This study serves as a reminder that the available large-scale datasets might be perpetuating severe biases.”
The scientists next aim to broaden their analyses to include more diverse artwork, as this particular dataset was ultimately Western and male biased. Future analyses should also consider diagonal compositions in paintings, they say.
This work was supported by the National Research Foundation (NRF) of Korea.
Publication:
Lee, B, et al. (2020) Dissecting landscape art history with information theory. Proceedings of the National Academy of Sciences (PNAS), Vol. 117, No. 43, 26580-26590. Available online at https://doi.org/10.1073/pnas.2011927117
Profile:
Hawoong Jeong, Ph.D.
Professor
hjeong@kaist.ac.kr
https://www.kaist.ac.kr
Department of Physics
Korea Advanced Institute of Science and Technology (KAIST)
Daejeon, Republic of Korea
(END)
2020.11.13 View 7292 -
 'Mini-Lungs' Reveal Early Stages of SARS-CoV-2 Infection
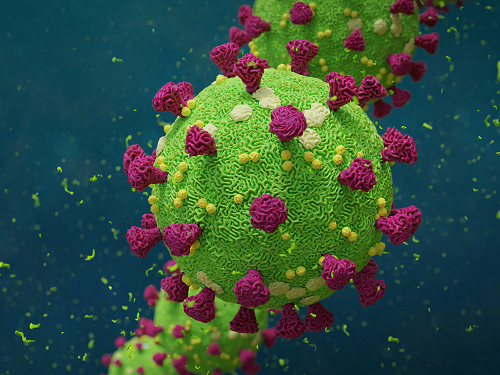
Researchers in Korea and the UK have successfully grown miniature models of critical lung structures called alveoli, and used them to study how the coronavirus that causes COVID-19 infects the lungs.
To date, there have been more than 40 million cases of COVID-19 and almost 1.13 million deaths worldwide. The main target tissues of SARS-CoV-2, the virus that causes COVID-19, especially in patients that develop pneumonia, appear to be alveoli – tiny air sacs in the lungs that take up the oxygen we breathe and exchange it with carbon dioxide to exhale.
To better understand how SARS-CoV-2 infects the lungs and causes disease, a team of Professor Young Seok Ju from the Graduate School of Medical Science and Engineering at KAIST in collaboration with the Wellcome-MRC Cambridge Stem Cell Institute at the University of Cambridge turned to organoids – ‘mini-organs’ grown in three dimensions to mimic the behaviour of tissue and organs.
The team used tissue donated to tissue banks at the Royal Papworth Hospital NHS Foundation Trust and Addenbrooke’s Hospital, Cambridge University NHS Foundations Trust, UK, and Seoul National University Hospital to extract a type of lung cell known as human lung alveolar type 2 cells. By reprogramming these cells back to their earlier ‘stem cell’ stage, they were able to grow self-organizing alveolar-like 3D structures that mimic the behaviour of key lung tissue.
“The research community now has a powerful new platform to study precisely how the virus infects the lungs, as well as explore possible treatments,” said Professor Ju, co-senior author of the research.
Dr. Joo-Hyeon Lee, another co-senior author at the Wellcome-MRC Cambridge Stem Cell Institute, said: “We still know surprisingly little about how SARS-CoV-2 infects the lungs and causes disease. Our approach has allowed us to grow 3D models of key lung tissue – in a sense, ‘mini-lungs’ – in the lab and study what happens when they become infected.”
The team infected the organoids with a strain of SARS-CoV-2 taken from a patient in Korea who was diagnosed with COVID-19 on January 26 after traveling to Wuhan, China. Using a combination of fluorescence imaging and single cell genetic analysis, they were able to study how the cells responded to the virus.
When the 3D models were exposed to SARS-CoV-2, the virus began to replicate rapidly, reaching full cellular infection just six hours after infection. Replication enables the virus to spread throughout the body, infecting other cells and tissue.
Around the same time, the cells began to produce interferons – proteins that act as warning signals to neighbouring cells, telling them to activate their antiviral defences. After 48 hours, the interferons triggered the innate immune response – its first line of defence – and the cells started fighting back against infection.
Sixty hours after infection, a subset of alveolar cells began to disintegrate, leading to cell death and damage to the lung tissue.
Although the researchers observed changes to the lung cells within three days of infection, clinical symptoms of COVID-19 rarely occur so quickly and can sometimes take more than ten days after exposure to appear. The team say there are several possible reasons for this. It may take several days from the virus first infiltrating the upper respiratory tract to it reaching the alveoli. It may also require a substantial proportion of alveolar cells to be infected or for further interactions with immune cells resulting in inflammation before a patient displays symptoms.
“Based on our model we can tackle many unanswered key questions, such as understanding genetic susceptibility to SARS-CoV-2, assessing relative infectivity of viral mutants, and revealing the damage processes of the virus in human alveolar cells,” said Professor Ju. “Most importantly, it provides the opportunity to develop and screen potential therapeutic agents against SARS-CoV-2 infection.”
“We hope to use our technique to grow these 3D models from cells of patients who are particularly vulnerable to infection, such as the elderly or people with diseased lungs, and find out what happens to their tissue,” added Dr. Lee.
The research was a collaboration involving scientists from KAIST, the University of Cambridge, Korea National Institute of Health, Institute for Basic Science (IBS), Seoul National University Hospital and Genome Insight in Korea.
- ProfileProfessor Young Seok JuLaboratory of Cancer Genomics https://julab.kaist.ac.kr the Graduate School of Medical Science and EngineeringKAIST
2020.10.26 View 7343
'Mini-Lungs' Reveal Early Stages of SARS-CoV-2 Infection
Researchers in Korea and the UK have successfully grown miniature models of critical lung structures called alveoli, and used them to study how the coronavirus that causes COVID-19 infects the lungs.
To date, there have been more than 40 million cases of COVID-19 and almost 1.13 million deaths worldwide. The main target tissues of SARS-CoV-2, the virus that causes COVID-19, especially in patients that develop pneumonia, appear to be alveoli – tiny air sacs in the lungs that take up the oxygen we breathe and exchange it with carbon dioxide to exhale.
To better understand how SARS-CoV-2 infects the lungs and causes disease, a team of Professor Young Seok Ju from the Graduate School of Medical Science and Engineering at KAIST in collaboration with the Wellcome-MRC Cambridge Stem Cell Institute at the University of Cambridge turned to organoids – ‘mini-organs’ grown in three dimensions to mimic the behaviour of tissue and organs.
The team used tissue donated to tissue banks at the Royal Papworth Hospital NHS Foundation Trust and Addenbrooke’s Hospital, Cambridge University NHS Foundations Trust, UK, and Seoul National University Hospital to extract a type of lung cell known as human lung alveolar type 2 cells. By reprogramming these cells back to their earlier ‘stem cell’ stage, they were able to grow self-organizing alveolar-like 3D structures that mimic the behaviour of key lung tissue.
“The research community now has a powerful new platform to study precisely how the virus infects the lungs, as well as explore possible treatments,” said Professor Ju, co-senior author of the research.
Dr. Joo-Hyeon Lee, another co-senior author at the Wellcome-MRC Cambridge Stem Cell Institute, said: “We still know surprisingly little about how SARS-CoV-2 infects the lungs and causes disease. Our approach has allowed us to grow 3D models of key lung tissue – in a sense, ‘mini-lungs’ – in the lab and study what happens when they become infected.”
The team infected the organoids with a strain of SARS-CoV-2 taken from a patient in Korea who was diagnosed with COVID-19 on January 26 after traveling to Wuhan, China. Using a combination of fluorescence imaging and single cell genetic analysis, they were able to study how the cells responded to the virus.
When the 3D models were exposed to SARS-CoV-2, the virus began to replicate rapidly, reaching full cellular infection just six hours after infection. Replication enables the virus to spread throughout the body, infecting other cells and tissue.
Around the same time, the cells began to produce interferons – proteins that act as warning signals to neighbouring cells, telling them to activate their antiviral defences. After 48 hours, the interferons triggered the innate immune response – its first line of defence – and the cells started fighting back against infection.
Sixty hours after infection, a subset of alveolar cells began to disintegrate, leading to cell death and damage to the lung tissue.
Although the researchers observed changes to the lung cells within three days of infection, clinical symptoms of COVID-19 rarely occur so quickly and can sometimes take more than ten days after exposure to appear. The team say there are several possible reasons for this. It may take several days from the virus first infiltrating the upper respiratory tract to it reaching the alveoli. It may also require a substantial proportion of alveolar cells to be infected or for further interactions with immune cells resulting in inflammation before a patient displays symptoms.
“Based on our model we can tackle many unanswered key questions, such as understanding genetic susceptibility to SARS-CoV-2, assessing relative infectivity of viral mutants, and revealing the damage processes of the virus in human alveolar cells,” said Professor Ju. “Most importantly, it provides the opportunity to develop and screen potential therapeutic agents against SARS-CoV-2 infection.”
“We hope to use our technique to grow these 3D models from cells of patients who are particularly vulnerable to infection, such as the elderly or people with diseased lungs, and find out what happens to their tissue,” added Dr. Lee.
The research was a collaboration involving scientists from KAIST, the University of Cambridge, Korea National Institute of Health, Institute for Basic Science (IBS), Seoul National University Hospital and Genome Insight in Korea.
- ProfileProfessor Young Seok JuLaboratory of Cancer Genomics https://julab.kaist.ac.kr the Graduate School of Medical Science and EngineeringKAIST
2020.10.26 View 7343 -
 Professor Won-Ki Cho Selected as the 2020 SUHF Young Investigator
Professor Won-Ki Cho from the Department of Biological Sciences was named one of three recipients of the 2020 Suh Kyung-Bae Science Foundation (SUHF) Young Investigator Award.
The SUHF is a non-profit organization established in 2016 and funded by a personal donation of 300 billion KRW in shares from Chairman and CEO Kyung-Bae Suh of the Amorepacific Group. The primary purpose of the foundation is to serve as a platform to nurture and provide comprehensive long-term support for creative and passionate young Korean scientists committed to pursuing research in the field of life sciences. The SUHF selects three to five scientists through an open recruiting process every year and grants each scientist a maximum of 2.5 billion KRW over a period of up to five years.
Since January this year, the foundation received 67 research proposals from scientists across the nation, especially from those who had less than five years of experience as professors, and selected the three recipients.
Professor Cho proposed research on how to observe the interactions between nuclear structures and constantly-changing chromatin monomers in four dimensions through ultra-high-resolution imaging of single living cells. This proposal was recognized as one that could help us better understand the process of transcription regulation, which remains a long-standing question in biology.
The other awards were given to Professor Soung-hun Roh of Seoul National University and Professor Joo-Hyeon Lee of the University of Cambridge.
With these three new awardees, a total of 17 scientists have been named SUHF Young Investigators to date, and the funding to support these scientists now totals 42.5 billion KRW.
Professor Inkyung Jung and Professor Ki-Jun Yoon from the Department of Biological Sciences, and Professor Young Seok Ju and Professor Jeong Ho Lee from the Graduate School of Medical Science and Engineering are the four previous winners from KAIST in the years 2017 through 2019.
(END)
2020.10.15 View 8460
Professor Won-Ki Cho Selected as the 2020 SUHF Young Investigator
Professor Won-Ki Cho from the Department of Biological Sciences was named one of three recipients of the 2020 Suh Kyung-Bae Science Foundation (SUHF) Young Investigator Award.
The SUHF is a non-profit organization established in 2016 and funded by a personal donation of 300 billion KRW in shares from Chairman and CEO Kyung-Bae Suh of the Amorepacific Group. The primary purpose of the foundation is to serve as a platform to nurture and provide comprehensive long-term support for creative and passionate young Korean scientists committed to pursuing research in the field of life sciences. The SUHF selects three to five scientists through an open recruiting process every year and grants each scientist a maximum of 2.5 billion KRW over a period of up to five years.
Since January this year, the foundation received 67 research proposals from scientists across the nation, especially from those who had less than five years of experience as professors, and selected the three recipients.
Professor Cho proposed research on how to observe the interactions between nuclear structures and constantly-changing chromatin monomers in four dimensions through ultra-high-resolution imaging of single living cells. This proposal was recognized as one that could help us better understand the process of transcription regulation, which remains a long-standing question in biology.
The other awards were given to Professor Soung-hun Roh of Seoul National University and Professor Joo-Hyeon Lee of the University of Cambridge.
With these three new awardees, a total of 17 scientists have been named SUHF Young Investigators to date, and the funding to support these scientists now totals 42.5 billion KRW.
Professor Inkyung Jung and Professor Ki-Jun Yoon from the Department of Biological Sciences, and Professor Young Seok Ju and Professor Jeong Ho Lee from the Graduate School of Medical Science and Engineering are the four previous winners from KAIST in the years 2017 through 2019.
(END)
2020.10.15 View 8460 -
 Biomarker Predicts Who Will Have Severe COVID-19
- Airway cell analyses showing an activated immune axis could pinpoint the COVID-19 patients who will most benefit from targeted therapies.-
KAIST researchers have identified key markers that could help pinpoint patients who are bound to get a severe reaction to COVID-19 infection. This would help doctors provide the right treatments at the right time, potentially saving lives. The findings were published in the journal Frontiers in Immunology on August 28.
People’s immune systems react differently to infection with SARS-CoV-2, the virus that causes COVID-19, ranging from mild to severe, life-threatening responses.
To understand the differences in responses, Professor Heung Kyu Lee and PhD candidate Jang Hyun Park from the Graduate School of Medical Science and Engineering at KAIST analysed ribonucleic acid (RNA) sequencing data extracted from individual airway cells of healthy controls and of mildly and severely ill patients with COVID-19. The data was available in a public database previously published by a group of Chinese researchers.
“Our analyses identified an association between immune cells called neutrophils and special cell receptors that bind to the steroid hormone glucocorticoid,” Professor Lee explained. “This finding could be used as a biomarker for predicting disease severity in patients and thus selecting a targeted therapy that can help treat them at an appropriate time,” he added.
Severe illness in COVID-19 is associated with an exaggerated immune response that leads to excessive airway-damaging inflammation. This condition, known as acute respiratory distress syndrome (ARDS), accounts for 70% of deaths in fatal COVID-19 infections.
Scientists already know that this excessive inflammation involves heightened neutrophil recruitment to the airways, but the detailed mechanisms of this reaction are still unclear.
Lee and Park’s analyses found that a group of immune cells called myeloid cells produced excess amounts of neutrophil-recruiting chemicals in severely ill patients, including a cytokine called tumour necrosis factor (TNF) and a chemokine called CXCL8.
Further RNA analyses of neutrophils in severely ill patients showed they were less able to recruit very important T cells needed for attacking the virus. At the same time, the neutrophils produced too many extracellular molecules that normally trap pathogens, but damage airway cells when produced in excess.
The researchers additionally found that the airway cells in severely ill patients were not expressing enough glucocorticoid receptors. This was correlated with increased CXCL8 expression and neutrophil recruitment.
Glucocorticoids, like the well-known drug dexamethasone, are anti-inflammatory agents that could play a role in treating COVID-19. However, using them in early or mild forms of the infection could suppress the necessary immune reactions to combat the virus. But if airway damage has already happened in more severe cases, glucocorticoid treatment would be ineffective.
Knowing who to give this treatment to and when is really important. COVID-19 patients showing reduced glucocorticoid receptor expression, increased CXCL8 expression, and excess neutrophil recruitment to the airways could benefit from treatment with glucocorticoids to prevent airway damage. Further research is needed, however, to confirm the relationship between glucocorticoids and neutrophil inflammation at the protein level.
“Our study could serve as a springboard towards more accurate and reliable COVID-19 treatments,” Professor Lee said.
This work was supported by the National Research Foundation of Korea, and Mobile Clinic Module Project funded by KAIST.
Figure. Low glucocorticoid receptor (GR) expression led to excessive inflammation and lung damage by neutrophils through enhancing the expression of CXCL8 and other cytokines.
Image credit: Professor Heung Kyu Lee, KAIST. Created with Biorender.com.
Image usage restrictions: News organizations may use or redistribute these figures and image, with proper attribution, as part of news coverage of this paper only.
-Publication:
Jang Hyun Park, and Heung Kyu Lee. (2020). Re-analysis of Single Cell Transcriptome Reveals That the NR3C1-CXCL8-Neutrophil Axis Determines the Severity of COVID-19. Frontiers in Immunology, Available online at https://doi.org/10.3389/fimmu.2020.02145
-Profile: Heung Kyu Lee
Associate Professor
heungkyu.lee@kaist.ac.kr
https://www.heungkyulee.kaist.ac.kr/
Laboratory of Host Defenses
Graduate School of Medical Science and Engineering (GSMSE)
The Center for Epidemic Preparedness at KAIST Institute
http://kaist.ac.kr
Korea Advanced Institute of Science and Technology (KAIST)
Daejeon, Republic of Korea
Profile: Jang Hyun Park
PhD Candidate
janghyun.park@kaist.ac.kr
GSMSE, KAIST
2020.09.17 View 10425
Biomarker Predicts Who Will Have Severe COVID-19
- Airway cell analyses showing an activated immune axis could pinpoint the COVID-19 patients who will most benefit from targeted therapies.-
KAIST researchers have identified key markers that could help pinpoint patients who are bound to get a severe reaction to COVID-19 infection. This would help doctors provide the right treatments at the right time, potentially saving lives. The findings were published in the journal Frontiers in Immunology on August 28.
People’s immune systems react differently to infection with SARS-CoV-2, the virus that causes COVID-19, ranging from mild to severe, life-threatening responses.
To understand the differences in responses, Professor Heung Kyu Lee and PhD candidate Jang Hyun Park from the Graduate School of Medical Science and Engineering at KAIST analysed ribonucleic acid (RNA) sequencing data extracted from individual airway cells of healthy controls and of mildly and severely ill patients with COVID-19. The data was available in a public database previously published by a group of Chinese researchers.
“Our analyses identified an association between immune cells called neutrophils and special cell receptors that bind to the steroid hormone glucocorticoid,” Professor Lee explained. “This finding could be used as a biomarker for predicting disease severity in patients and thus selecting a targeted therapy that can help treat them at an appropriate time,” he added.
Severe illness in COVID-19 is associated with an exaggerated immune response that leads to excessive airway-damaging inflammation. This condition, known as acute respiratory distress syndrome (ARDS), accounts for 70% of deaths in fatal COVID-19 infections.
Scientists already know that this excessive inflammation involves heightened neutrophil recruitment to the airways, but the detailed mechanisms of this reaction are still unclear.
Lee and Park’s analyses found that a group of immune cells called myeloid cells produced excess amounts of neutrophil-recruiting chemicals in severely ill patients, including a cytokine called tumour necrosis factor (TNF) and a chemokine called CXCL8.
Further RNA analyses of neutrophils in severely ill patients showed they were less able to recruit very important T cells needed for attacking the virus. At the same time, the neutrophils produced too many extracellular molecules that normally trap pathogens, but damage airway cells when produced in excess.
The researchers additionally found that the airway cells in severely ill patients were not expressing enough glucocorticoid receptors. This was correlated with increased CXCL8 expression and neutrophil recruitment.
Glucocorticoids, like the well-known drug dexamethasone, are anti-inflammatory agents that could play a role in treating COVID-19. However, using them in early or mild forms of the infection could suppress the necessary immune reactions to combat the virus. But if airway damage has already happened in more severe cases, glucocorticoid treatment would be ineffective.
Knowing who to give this treatment to and when is really important. COVID-19 patients showing reduced glucocorticoid receptor expression, increased CXCL8 expression, and excess neutrophil recruitment to the airways could benefit from treatment with glucocorticoids to prevent airway damage. Further research is needed, however, to confirm the relationship between glucocorticoids and neutrophil inflammation at the protein level.
“Our study could serve as a springboard towards more accurate and reliable COVID-19 treatments,” Professor Lee said.
This work was supported by the National Research Foundation of Korea, and Mobile Clinic Module Project funded by KAIST.
Figure. Low glucocorticoid receptor (GR) expression led to excessive inflammation and lung damage by neutrophils through enhancing the expression of CXCL8 and other cytokines.
Image credit: Professor Heung Kyu Lee, KAIST. Created with Biorender.com.
Image usage restrictions: News organizations may use or redistribute these figures and image, with proper attribution, as part of news coverage of this paper only.
-Publication:
Jang Hyun Park, and Heung Kyu Lee. (2020). Re-analysis of Single Cell Transcriptome Reveals That the NR3C1-CXCL8-Neutrophil Axis Determines the Severity of COVID-19. Frontiers in Immunology, Available online at https://doi.org/10.3389/fimmu.2020.02145
-Profile: Heung Kyu Lee
Associate Professor
heungkyu.lee@kaist.ac.kr
https://www.heungkyulee.kaist.ac.kr/
Laboratory of Host Defenses
Graduate School of Medical Science and Engineering (GSMSE)
The Center for Epidemic Preparedness at KAIST Institute
http://kaist.ac.kr
Korea Advanced Institute of Science and Technology (KAIST)
Daejeon, Republic of Korea
Profile: Jang Hyun Park
PhD Candidate
janghyun.park@kaist.ac.kr
GSMSE, KAIST
2020.09.17 View 10425 -
 Microscopy Approach Poised to Offer New Insights into Liver Diseases
Researchers have developed a new way to visualize the progression of nonalcoholic fatty liver disease (NAFLD) in mouse models of the disease. The new microscopy method provides a high-resolution 3D view that could lead to important new insights into NAFLD, a condition in which too much fat is stored in the liver.
“It is estimated that a quarter of the adult global population has NAFLD, yet an effective treatment strategy has not been found,” said professor Pilhan Kim from the Graduate School of Medical Science and Engineering at KAIST. “NAFLD is associated with obesity and type 2 diabetes and can sometimes progress to liver failure in serious case.”
In the Optical Society (OSA) journal Biomedical Optics Express, Professor Kim and colleagues reported their new imaging technique and showed that it can be used to observe how tiny droplets of fat, or lipids, accumulate in the liver cells of living mice over time.
“It has been challenging to find a treatment strategy for NAFLD because most studies examine excised liver tissue that represents just one timepoint in disease progression,” said Professor Kim. “Our technique can capture details of lipid accumulation over time, providing a highly useful research tool for identifying the multiple parameters that likely contribute to the disease and could be targeted with treatment.”
Capturing the dynamics of NAFLD in living mouse models of the disease requires the ability to observe quickly changing interactions of biological components in intact tissue in real-time. To accomplish this, the researchers developed a custom intravital confocal and two-photon microscopy system that acquires images of multiple fluorescent labels at video-rate with cellular resolution.
“With video-rate imaging capability, the continuous movement of liver tissue in live mice due to breathing and heart beating could be tracked in real time and precisely compensated,” said Professor Kim. “This provided motion-artifact free high-resolution images of cellular and sub-cellular sized individual lipid droplets.”
The key to fast imaging was a polygonal mirror that rotated at more than 240 miles per hour to provide extremely fast laser scanning. The researchers also incorporated four different lasers and four high-sensitivity optical detectors into the setup so that they could acquire multi-color images to capture different color fluorescent probes used to label the lipid droplets and microvasculature in the livers of live mice.
“Our approach can capture real-time changes in cell behavior and morphology, vascular structure and function, and the spatiotemporal localization of biological components while directly visualizing of lipid droplet development in NAFLD progression,” said Professor Kim. “It also allows the analysis of the highly complex behaviors of various immune cells as NAFLD progresses.”
The researchers demonstrated their approach by using it to observe the development and spatial distribution of lipid droplets in individual mice with NAFLD induced by a methionine and choline-deficient diet. Next, they plan to use it to study how the liver microenvironment changes during NAFLD progression by imaging the same mouse over time. They also want to use their microscope technique to visualize various immune cells and lipid droplets to better understand the complex liver microenvironment in NAFLD progression.
2020.08.21 View 5684
Microscopy Approach Poised to Offer New Insights into Liver Diseases
Researchers have developed a new way to visualize the progression of nonalcoholic fatty liver disease (NAFLD) in mouse models of the disease. The new microscopy method provides a high-resolution 3D view that could lead to important new insights into NAFLD, a condition in which too much fat is stored in the liver.
“It is estimated that a quarter of the adult global population has NAFLD, yet an effective treatment strategy has not been found,” said professor Pilhan Kim from the Graduate School of Medical Science and Engineering at KAIST. “NAFLD is associated with obesity and type 2 diabetes and can sometimes progress to liver failure in serious case.”
In the Optical Society (OSA) journal Biomedical Optics Express, Professor Kim and colleagues reported their new imaging technique and showed that it can be used to observe how tiny droplets of fat, or lipids, accumulate in the liver cells of living mice over time.
“It has been challenging to find a treatment strategy for NAFLD because most studies examine excised liver tissue that represents just one timepoint in disease progression,” said Professor Kim. “Our technique can capture details of lipid accumulation over time, providing a highly useful research tool for identifying the multiple parameters that likely contribute to the disease and could be targeted with treatment.”
Capturing the dynamics of NAFLD in living mouse models of the disease requires the ability to observe quickly changing interactions of biological components in intact tissue in real-time. To accomplish this, the researchers developed a custom intravital confocal and two-photon microscopy system that acquires images of multiple fluorescent labels at video-rate with cellular resolution.
“With video-rate imaging capability, the continuous movement of liver tissue in live mice due to breathing and heart beating could be tracked in real time and precisely compensated,” said Professor Kim. “This provided motion-artifact free high-resolution images of cellular and sub-cellular sized individual lipid droplets.”
The key to fast imaging was a polygonal mirror that rotated at more than 240 miles per hour to provide extremely fast laser scanning. The researchers also incorporated four different lasers and four high-sensitivity optical detectors into the setup so that they could acquire multi-color images to capture different color fluorescent probes used to label the lipid droplets and microvasculature in the livers of live mice.
“Our approach can capture real-time changes in cell behavior and morphology, vascular structure and function, and the spatiotemporal localization of biological components while directly visualizing of lipid droplet development in NAFLD progression,” said Professor Kim. “It also allows the analysis of the highly complex behaviors of various immune cells as NAFLD progresses.”
The researchers demonstrated their approach by using it to observe the development and spatial distribution of lipid droplets in individual mice with NAFLD induced by a methionine and choline-deficient diet. Next, they plan to use it to study how the liver microenvironment changes during NAFLD progression by imaging the same mouse over time. They also want to use their microscope technique to visualize various immune cells and lipid droplets to better understand the complex liver microenvironment in NAFLD progression.
2020.08.21 View 5684 -
 Tinkering with Roundworm Proteins Offers Hope for Anti-aging Drugs
- The somatic nuclear protein kinase VRK-1 increases the worm’s lifespan through AMPK activation, and this mechanism can be applied to promoting human longevity, the study reveals. -
KAIST researchers have been able to dial up and down creatures’ lifespans by altering the activity of proteins found in roundworm cells that tell them to convert sugar into energy when their cellular energy is running low. Humans also have these proteins, offering up the intriguing possibilities for developing longevity-promoting drugs. These new findings were published on July 1 in Science Advances.
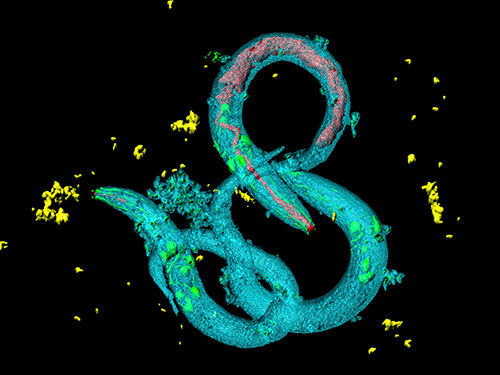
The roundworm Caenorhabditis elegans (C. elegans), a millimeter-long nematode commonly used in lab testing, enjoyed a boost in its lifespan when researchers tinkered with a couple of proteins involved in monitoring the energy use by its cells.
The proteins VRK-1 and AMPK work in tandem in roundworm cells, with the former telling the latter to get to work by sticking a phosphate molecule, composed of one phosphorus and four oxygen atoms, on it. In turn, AMPK’s role is to monitor energy levels in cells, when cellular energy is running low. In essence, VRK-1 regulates AMPK, and AMPK regulates the cellular energy status.
Using a range of different biological research tools, including introducing foreign genes into the worm, a group of researchers led by Professor Seung-Jae V. Lee from the Department of Biological Sciences at KAIST were able to dial up and down the activity of the gene that tells cells to produce the VRK-1 protein. This gene has remained pretty much unchanged throughout evolution. Most complex organisms have this same gene, including humans.
Lead author of the study Sangsoon Park and his colleagues confirmed that the overexpression, or increased production, of the VRK-1 protein boosted the lifespan of the C. elegans, which normally lives just two to three weeks, and the inhibition of VRK-1 production reduced its lifespan.
The research team found that the activity of the VRK-1-to-AMPK cellular-energy monitoring process is increased in low cellular energy status by reduced mitochondrial respiration, the set of metabolic chemical reactions that make use of the oxygen the worm breathes to convert macronutrients from food into the energy “currency” that cells spend to do everything they need to do.
It is already known that mitochondria, the energy-producing engine rooms in cells, play a crucial role in aging, and declines in the functioning of mitochondria are associated with age-related diseases. At the same time, the mild inhibition of mitochondrial respiration has been shown to promote longevity in a range of species, including flies and mammals.
When the research team performed similar tinkering with cultured human cells, they found they could also replicate this ramping up and down of the VRK-1-to-AMPK process that occurs in roundworms.
“This raises the intriguing possibility that VRK-1 also functions as a factor in governing human longevity, and so perhaps we can start developing longevity-promoting drugs that alter the activity of VRK-1,” explained Professor Lee.
At the very least, the research points us in an interesting direction for investigating new therapeutic strategies to combat metabolic disorders by targeting the modulation of VRK-1. Metabolic disorders involve the disruption of chemical reactions in the body, including diseases of the mitochondria.
But before metabolic disorder therapeutics or longevity drugs can be contemplated by scientists, further research still needs to be carried out to better understand how VRK-1 works to activate AMPK, as well as figure out the precise mechanics of how AMPK controls cellular energy.
This work was supported by the National Research Foundation (NRF), and the Ministry of Science and ICT (MSIT) of Korea.
Image credit: Seung-Jae V. LEE, KAIST.
Image usage restrictions: News organizations may use or redistribute this image, with proper attribution, as part of news coverage of this paper only.
Publication:
Park, S., et al. (2020) ‘VRK-1 extends life span by activation of AMPK via phosphorylation’. Science Advances, Volume 6. No. 27, eaaw7824. Available online at https://doi.org/10.1126/sciadv.aaw7824
Profile: Seung-Jae V. Lee, Ph.D.
Professor
seungjaevlee@kaist.ac.kr
https://sites.google.com/view/mgakaist
Molecular Genetics of Aging Laboratory
Department of Biological Sciences
Korea Advanced Institute of Science and Technology (KAIST)
https://www.kaist.ac.krDaejeon 34141, Korea
(END)
2020.07.31 View 8450
Tinkering with Roundworm Proteins Offers Hope for Anti-aging Drugs
- The somatic nuclear protein kinase VRK-1 increases the worm’s lifespan through AMPK activation, and this mechanism can be applied to promoting human longevity, the study reveals. -
KAIST researchers have been able to dial up and down creatures’ lifespans by altering the activity of proteins found in roundworm cells that tell them to convert sugar into energy when their cellular energy is running low. Humans also have these proteins, offering up the intriguing possibilities for developing longevity-promoting drugs. These new findings were published on July 1 in Science Advances.
The roundworm Caenorhabditis elegans (C. elegans), a millimeter-long nematode commonly used in lab testing, enjoyed a boost in its lifespan when researchers tinkered with a couple of proteins involved in monitoring the energy use by its cells.
The proteins VRK-1 and AMPK work in tandem in roundworm cells, with the former telling the latter to get to work by sticking a phosphate molecule, composed of one phosphorus and four oxygen atoms, on it. In turn, AMPK’s role is to monitor energy levels in cells, when cellular energy is running low. In essence, VRK-1 regulates AMPK, and AMPK regulates the cellular energy status.
Using a range of different biological research tools, including introducing foreign genes into the worm, a group of researchers led by Professor Seung-Jae V. Lee from the Department of Biological Sciences at KAIST were able to dial up and down the activity of the gene that tells cells to produce the VRK-1 protein. This gene has remained pretty much unchanged throughout evolution. Most complex organisms have this same gene, including humans.
Lead author of the study Sangsoon Park and his colleagues confirmed that the overexpression, or increased production, of the VRK-1 protein boosted the lifespan of the C. elegans, which normally lives just two to three weeks, and the inhibition of VRK-1 production reduced its lifespan.
The research team found that the activity of the VRK-1-to-AMPK cellular-energy monitoring process is increased in low cellular energy status by reduced mitochondrial respiration, the set of metabolic chemical reactions that make use of the oxygen the worm breathes to convert macronutrients from food into the energy “currency” that cells spend to do everything they need to do.
It is already known that mitochondria, the energy-producing engine rooms in cells, play a crucial role in aging, and declines in the functioning of mitochondria are associated with age-related diseases. At the same time, the mild inhibition of mitochondrial respiration has been shown to promote longevity in a range of species, including flies and mammals.
When the research team performed similar tinkering with cultured human cells, they found they could also replicate this ramping up and down of the VRK-1-to-AMPK process that occurs in roundworms.
“This raises the intriguing possibility that VRK-1 also functions as a factor in governing human longevity, and so perhaps we can start developing longevity-promoting drugs that alter the activity of VRK-1,” explained Professor Lee.
At the very least, the research points us in an interesting direction for investigating new therapeutic strategies to combat metabolic disorders by targeting the modulation of VRK-1. Metabolic disorders involve the disruption of chemical reactions in the body, including diseases of the mitochondria.
But before metabolic disorder therapeutics or longevity drugs can be contemplated by scientists, further research still needs to be carried out to better understand how VRK-1 works to activate AMPK, as well as figure out the precise mechanics of how AMPK controls cellular energy.
This work was supported by the National Research Foundation (NRF), and the Ministry of Science and ICT (MSIT) of Korea.
Image credit: Seung-Jae V. LEE, KAIST.
Image usage restrictions: News organizations may use or redistribute this image, with proper attribution, as part of news coverage of this paper only.
Publication:
Park, S., et al. (2020) ‘VRK-1 extends life span by activation of AMPK via phosphorylation’. Science Advances, Volume 6. No. 27, eaaw7824. Available online at https://doi.org/10.1126/sciadv.aaw7824
Profile: Seung-Jae V. Lee, Ph.D.
Professor
seungjaevlee@kaist.ac.kr
https://sites.google.com/view/mgakaist
Molecular Genetics of Aging Laboratory
Department of Biological Sciences
Korea Advanced Institute of Science and Technology (KAIST)
https://www.kaist.ac.krDaejeon 34141, Korea
(END)
2020.07.31 View 8450 -
 Study Finds Interferon Triggers Inflammation in Severe COVID-19
KAIST medical scientists and their colleagues confirmed that the type I interferon response plays a pivotal role in exacerbating inflammation in severe COVID-19 cases. Severe COVID-19 has been shown to be caused by a hyper-inflammatory response. Particularly, inflammatory cytokines secreted by classical monocytes and macrophages are believed to play a crucial role in the severe progression of COVID-19.
A new single-cell RNA sequencing analysis of more than 59,000 cells from three different patient cohorts provided a detailed look at patients’ immune responses in severe cases of COVID-19. The results suggest that patients with severe cases of COVID-19 experience increased regulation of the type I interferon (IFN-I) inflammation-triggering pathway, a signature that the researchers also observed in patients hospitalized with severe cases of influenza.
Their findings suggest that anti-inflammatory treatment strategies for COVID-19 should also be aimed toward the IFN-I signaling pathway, in addition to targeting inflammatory molecules such as TNF, IL-1, and IL-6, which have been associated with COVID-19.
The research team under Professor Eui-Cheol Shin from the Graduate School of Medical Science and Engineering sequenced the RNA from a total of 59,572 blood cells obtained from four healthy donors, eight patients with mild or severe COVID-19, and five patients with severe influenza.
By comparison, patients with severe cases of influenza showed increased expression of various IFN-stimulated genes, but did not experience TNF/IL-1 responses as seen in COVID-19 patients. Unlike the flu cohort, patients in the severe COVID-19 cohort exhibited the IFN-I signature concurrently with TNF/IL-1-driven inflammation – a combination also not seen in patients with milder cases of COVID-19.
Their result, along with past mouse studies that highlight how the timing of IFN-I expression is critical to determining the outcome of SARS, support targeting IFN-I as a potential treatment strategy for severe COVID-19.
Professor Shin said, “This research provides insights for designing therapeutic options for COVID-19 by investigating very closely how the immune cells of COVDI-19 patients develop. We will continue to conduct research on novel therapeutic immune mechanisms and target therapeutic anti-inflammatory medication to improve the survival of severe COVID-19 patients.”
This study, conducted in collaboration with Severance Hospital at Yonsei University, Asan Medical Center, and Chungbuk National University, was featured in Science Immunology on July 10. This work was funded by Samsung Science and Technology Foundation and SUHF Fellowship.
-PublicationScience Immunology 10 Jul 2020:Vol. 5, Issue 49, eabd1554DOI: 10.1126/sciimmunol.abd1554
-ProfileProfessorEui-Cheol ShinGraduate School of Medical Science and EngineeringLaboratory of Immunology & Infectious Diseases (http://liid.kaist.ac.kr/)euicheols@kaist.ac.krKAIST
2020.07.14 View 6150
Study Finds Interferon Triggers Inflammation in Severe COVID-19
KAIST medical scientists and their colleagues confirmed that the type I interferon response plays a pivotal role in exacerbating inflammation in severe COVID-19 cases. Severe COVID-19 has been shown to be caused by a hyper-inflammatory response. Particularly, inflammatory cytokines secreted by classical monocytes and macrophages are believed to play a crucial role in the severe progression of COVID-19.
A new single-cell RNA sequencing analysis of more than 59,000 cells from three different patient cohorts provided a detailed look at patients’ immune responses in severe cases of COVID-19. The results suggest that patients with severe cases of COVID-19 experience increased regulation of the type I interferon (IFN-I) inflammation-triggering pathway, a signature that the researchers also observed in patients hospitalized with severe cases of influenza.
Their findings suggest that anti-inflammatory treatment strategies for COVID-19 should also be aimed toward the IFN-I signaling pathway, in addition to targeting inflammatory molecules such as TNF, IL-1, and IL-6, which have been associated with COVID-19.
The research team under Professor Eui-Cheol Shin from the Graduate School of Medical Science and Engineering sequenced the RNA from a total of 59,572 blood cells obtained from four healthy donors, eight patients with mild or severe COVID-19, and five patients with severe influenza.
By comparison, patients with severe cases of influenza showed increased expression of various IFN-stimulated genes, but did not experience TNF/IL-1 responses as seen in COVID-19 patients. Unlike the flu cohort, patients in the severe COVID-19 cohort exhibited the IFN-I signature concurrently with TNF/IL-1-driven inflammation – a combination also not seen in patients with milder cases of COVID-19.
Their result, along with past mouse studies that highlight how the timing of IFN-I expression is critical to determining the outcome of SARS, support targeting IFN-I as a potential treatment strategy for severe COVID-19.
Professor Shin said, “This research provides insights for designing therapeutic options for COVID-19 by investigating very closely how the immune cells of COVDI-19 patients develop. We will continue to conduct research on novel therapeutic immune mechanisms and target therapeutic anti-inflammatory medication to improve the survival of severe COVID-19 patients.”
This study, conducted in collaboration with Severance Hospital at Yonsei University, Asan Medical Center, and Chungbuk National University, was featured in Science Immunology on July 10. This work was funded by Samsung Science and Technology Foundation and SUHF Fellowship.
-PublicationScience Immunology 10 Jul 2020:Vol. 5, Issue 49, eabd1554DOI: 10.1126/sciimmunol.abd1554
-ProfileProfessorEui-Cheol ShinGraduate School of Medical Science and EngineeringLaboratory of Immunology & Infectious Diseases (http://liid.kaist.ac.kr/)euicheols@kaist.ac.krKAIST
2020.07.14 View 6150 -
 X-ray Scattering Shines Light on Protein Folding
- Multiple forms of a non-functional, unfolded protein follow different pathways and timelines to reach its folded, functional state, a study reveals. -
KAIST researchers have used an X-ray method to track how proteins fold, which could improve computer simulations of this process, with implications for understanding diseases and improving drug discovery. Their findings were reported in the Proceedings of the National Academy of Sciences of the United States of America (PNAS) on June 30.
When proteins are translated from their DNA codes, they quickly transform from a non-functional, unfolded state into their folded, functional state. Problems in folding can lead to diseases like Alzheimer’s and Parkinson’s.
“Protein folding is one of the most important biological processes, as it forms the functioning 3D protein structure,” explained the physical chemist Hyotcherl Ihee of the Department of Chemistry at KAIST. Dr. Tae Wu Kim, the lead author of this research from Ihee’s group, added, “Understanding the mechanisms of protein folding is important, and could pave the way for disease study and drug development.”
Ihee’s team developed an approach using an X-ray scattering technique to uncover how the protein cytochrome c folds from its initial unfolded state. This protein is composed of a chain of 104 amino acids with an iron-containing heme molecule. It is often used for protein folding studies.
The researchers placed the protein in a solution and shined ultraviolet light on it. This process provides electrons to cytochrome c, reducing the iron within it from the ferric to the ferrous form, which initiates folding. As this was happening, the researchers beamed X-rays at very short intervals onto the sample. The X-rays scattered off all the atomic pairs in the sample and a detector continuously recorded the X-ray scattering patterns. The X-ray scattering patterns provided direct information regarding the 3D protein structure and the changes made in these patterns over time showed real-time motion of the protein during the folding process.
The team found cytochrome c proteins initially exist in a wide variety of unfolded states. Once the folding process is triggered, they stop by a group of intermediates within 31.6 microseconds, and then those intermediates follow different pathways with different folding times to reach an energetically stable folded state.
“We don’t know if this diversity in folding paths can be generalized to other proteins,” Ihee confessed. He continued, “However, we believe that our approach can be used to study other protein folding systems.”
Ihee hopes this approach can improve the accuracy of models that simulate protein interactions by including information on their unstructured states. These simulations are important as they can help identify barriers to proper folding and predict a protein’s folded state given its amino acid sequence. Ultimately, the models could help clarify how some diseases develop and how drugs interact with various protein structures.
Ihee’s group collaborated with Professor Young Min Rhee at the KAIST Department of Chemistry, and this work was supported by the National Research Foundation of Korea (NRF) and the Institute for Basic Science (IBS).
Figure. The scientists found that non-functional unfolded forms of the protein cytochrome c follow different pathways and timelines to reach a stable functional folded state.
Publications:
Kim, T. W., et al. (2020) ‘Protein folding from heterogeneous unfolded state revealed by time-resolved X-ray solution scattering’. PNAS. Volume 117. Issue 26. Page 14996-15005. Available online at https://doi.org/10.1073/pnas.1913442117
Profile: Hyotcherl Ihee, Ph.D.
Professor
hyotcherl.ihee@kaist.ac.kr
http://time.kaist.ac.kr/
Ihee Laboratory
Department of Chemistry
KAIST
https://www.kaist.ac.kr
Daejeon 34141, Korea
Profile: Young Min Rhee, Ph.D.
Professor
ymrhee@kaist.ac.kr
http://singlet.kaist.ac.kr
Rhee Research Group
Department of Chemistry
KAIST
https://www.kaist.ac.kr
Daejeon 34141, Korea
(END)
2020.07.09 View 9795
X-ray Scattering Shines Light on Protein Folding
- Multiple forms of a non-functional, unfolded protein follow different pathways and timelines to reach its folded, functional state, a study reveals. -
KAIST researchers have used an X-ray method to track how proteins fold, which could improve computer simulations of this process, with implications for understanding diseases and improving drug discovery. Their findings were reported in the Proceedings of the National Academy of Sciences of the United States of America (PNAS) on June 30.
When proteins are translated from their DNA codes, they quickly transform from a non-functional, unfolded state into their folded, functional state. Problems in folding can lead to diseases like Alzheimer’s and Parkinson’s.
“Protein folding is one of the most important biological processes, as it forms the functioning 3D protein structure,” explained the physical chemist Hyotcherl Ihee of the Department of Chemistry at KAIST. Dr. Tae Wu Kim, the lead author of this research from Ihee’s group, added, “Understanding the mechanisms of protein folding is important, and could pave the way for disease study and drug development.”
Ihee’s team developed an approach using an X-ray scattering technique to uncover how the protein cytochrome c folds from its initial unfolded state. This protein is composed of a chain of 104 amino acids with an iron-containing heme molecule. It is often used for protein folding studies.
The researchers placed the protein in a solution and shined ultraviolet light on it. This process provides electrons to cytochrome c, reducing the iron within it from the ferric to the ferrous form, which initiates folding. As this was happening, the researchers beamed X-rays at very short intervals onto the sample. The X-rays scattered off all the atomic pairs in the sample and a detector continuously recorded the X-ray scattering patterns. The X-ray scattering patterns provided direct information regarding the 3D protein structure and the changes made in these patterns over time showed real-time motion of the protein during the folding process.
The team found cytochrome c proteins initially exist in a wide variety of unfolded states. Once the folding process is triggered, they stop by a group of intermediates within 31.6 microseconds, and then those intermediates follow different pathways with different folding times to reach an energetically stable folded state.
“We don’t know if this diversity in folding paths can be generalized to other proteins,” Ihee confessed. He continued, “However, we believe that our approach can be used to study other protein folding systems.”
Ihee hopes this approach can improve the accuracy of models that simulate protein interactions by including information on their unstructured states. These simulations are important as they can help identify barriers to proper folding and predict a protein’s folded state given its amino acid sequence. Ultimately, the models could help clarify how some diseases develop and how drugs interact with various protein structures.
Ihee’s group collaborated with Professor Young Min Rhee at the KAIST Department of Chemistry, and this work was supported by the National Research Foundation of Korea (NRF) and the Institute for Basic Science (IBS).
Figure. The scientists found that non-functional unfolded forms of the protein cytochrome c follow different pathways and timelines to reach a stable functional folded state.
Publications:
Kim, T. W., et al. (2020) ‘Protein folding from heterogeneous unfolded state revealed by time-resolved X-ray solution scattering’. PNAS. Volume 117. Issue 26. Page 14996-15005. Available online at https://doi.org/10.1073/pnas.1913442117
Profile: Hyotcherl Ihee, Ph.D.
Professor
hyotcherl.ihee@kaist.ac.kr
http://time.kaist.ac.kr/
Ihee Laboratory
Department of Chemistry
KAIST
https://www.kaist.ac.kr
Daejeon 34141, Korea
Profile: Young Min Rhee, Ph.D.
Professor
ymrhee@kaist.ac.kr
http://singlet.kaist.ac.kr
Rhee Research Group
Department of Chemistry
KAIST
https://www.kaist.ac.kr
Daejeon 34141, Korea
(END)
2020.07.09 View 9795 -
 Professor J.H. Lee Wins the Innovators in Science Award
Professor Jeong Ho Lee from the Graduate School of Medical Science and Engineering won the Early-Career Scientist Award of the 2020 Innovators in Science Award. The New York Academy of Sciences administers the award in partnership with Takeda Pharmaceutical Company.
The Innovators in Science Award grants two prizes of US $200,000 each year: one to an Early-Career Scientist and the other to a well-established Senior Scientist who have distinguished themselves for the creative thinking and impact of their rare disease research. The Senior Scientist Awardee is Dr. Adrian R. Krainer, at Cold Spring Harbor Laboratory whose research focused on the mechanisms and control of RNA splicing.
Prof. Lee is recognized for his research investigating genetic mutations in stem cells in the brain that result in rare developmental brain disorders. He was the first to identify the causes of intractable epilepsies and has identified the genes responsible for several developmental brain disorders, including focal cortical dysplasia, Joubert syndrome—a disorder characterized by an underdevelopment of the brainstem—and hemimegaloencephaly, which is the abnormal enlargement of one side of the brain.
“It is a great honor to be recognized by a jury of such globally respected scientists whom I greatly admire,” said Prof. Lee. “More importantly, this award validates research into brain somatic mutations as an important area of exploration to help patients suffering from devastating and untreatable neurological disorders.”
Prof. Lee also is the Director of the National Creative Research Initiative Center for Brain Somatic Mutations, and Co-founder and Chief Technology Officer of SoVarGen, a biopharmaceutical company aiming to discover novel therapeutics and diagnosis for intractable central nervous system (CNS) diseases caused by low-level somatic mutation.
The Innovators in Science Award is a limited submission competition in which research universities, academic institutions, government or non-profit institutions, or equivalent from around the globe with a well-established record of scientific excellence are invited to nominate their most promising Early-Career Scientists and their most outstanding Senior Scientists working in one of four selected therapeutic fields of neuroscience, gastroenterology, oncology, and regenerative medicine. The 2020 Winners will be honored at the virtual Innovators in Science Award Ceremony and Symposium in October 2020.
2020.07.09 View 6214
Professor J.H. Lee Wins the Innovators in Science Award
Professor Jeong Ho Lee from the Graduate School of Medical Science and Engineering won the Early-Career Scientist Award of the 2020 Innovators in Science Award. The New York Academy of Sciences administers the award in partnership with Takeda Pharmaceutical Company.
The Innovators in Science Award grants two prizes of US $200,000 each year: one to an Early-Career Scientist and the other to a well-established Senior Scientist who have distinguished themselves for the creative thinking and impact of their rare disease research. The Senior Scientist Awardee is Dr. Adrian R. Krainer, at Cold Spring Harbor Laboratory whose research focused on the mechanisms and control of RNA splicing.
Prof. Lee is recognized for his research investigating genetic mutations in stem cells in the brain that result in rare developmental brain disorders. He was the first to identify the causes of intractable epilepsies and has identified the genes responsible for several developmental brain disorders, including focal cortical dysplasia, Joubert syndrome—a disorder characterized by an underdevelopment of the brainstem—and hemimegaloencephaly, which is the abnormal enlargement of one side of the brain.
“It is a great honor to be recognized by a jury of such globally respected scientists whom I greatly admire,” said Prof. Lee. “More importantly, this award validates research into brain somatic mutations as an important area of exploration to help patients suffering from devastating and untreatable neurological disorders.”
Prof. Lee also is the Director of the National Creative Research Initiative Center for Brain Somatic Mutations, and Co-founder and Chief Technology Officer of SoVarGen, a biopharmaceutical company aiming to discover novel therapeutics and diagnosis for intractable central nervous system (CNS) diseases caused by low-level somatic mutation.
The Innovators in Science Award is a limited submission competition in which research universities, academic institutions, government or non-profit institutions, or equivalent from around the globe with a well-established record of scientific excellence are invited to nominate their most promising Early-Career Scientists and their most outstanding Senior Scientists working in one of four selected therapeutic fields of neuroscience, gastroenterology, oncology, and regenerative medicine. The 2020 Winners will be honored at the virtual Innovators in Science Award Ceremony and Symposium in October 2020.
2020.07.09 View 6214 -
 Every Moment of Ultrafast Chemical Bonding Now Captured on Film
- The emerging moment of bond formation, two separate bonding steps, and subsequent vibrational motions were visualized. -
< Emergence of molecular vibrations and the evolution to covalent bonds observed in the research. Video Credit: KEK IMSS >
A team of South Korean researchers led by Professor Hyotcherl Ihee from the Department of Chemistry at KAIST reported the direct observation of the birthing moment of chemical bonds by tracking real-time atomic positions in the molecule. Professor Ihee, who also serves as Associate Director of the Center for Nanomaterials and Chemical Reactions at the Institute for Basic Science (IBS), conducted this study in collaboration with scientists at the Institute of Materials Structure Science of High Energy Accelerator Research Organization (KEK IMSS, Japan), RIKEN (Japan), and Pohang Accelerator Laboratory (PAL, South Korea). This work was published in Nature on June 24.
Targeted cancer drugs work by striking a tight bond between cancer cell and specific molecular targets that are involved in the growth and spread of cancer. Detailed images of such chemical bonding sites or pathways can provide key information necessary for maximizing the efficacy of oncogene treatments. However, atomic movements in a molecule have never been captured in the middle of the action, not even for an extremely simple molecule such as a triatomic molecule, made of only three atoms.
Professor Ihee's group and their international collaborators finally succeeded in capturing the ongoing reaction process of the chemical bond formation in the gold trimer. "The femtosecond-resolution images revealed that such molecular events took place in two separate stages, not simultaneously as previously assumed," says Professor Ihee, the corresponding author of the study. "The atoms in the gold trimer complex atoms remain in motion even after the chemical bonding is complete. The distance between the atoms increased and decreased periodically, exhibiting the molecular vibration. These visualized molecular vibrations allowed us to name the characteristic motion of each observed vibrational mode." adds Professor Ihee.
Atoms move extremely fast at a scale of femtosecond (fs) ― quadrillionths (or millionths of a billionth) of a second. Its movement is minute in the level of angstrom equal to one ten-billionth of a meter. They are especially elusive during the transition state where reaction intermediates are transitioning from reactants to products in a flash. The KAIST-IBS research team made this experimentally challenging task possible by using femtosecond x-ray liquidography (solution scattering). This experimental technique combines laser photolysis and x-ray scattering techniques. When a laser pulse strikes the sample, X-rays scatter and initiate the chemical bond formation reaction in the gold trimer complex. Femtosecond x-ray pulses obtained from a special light source called an x-ray free-electron laser (XFEL) were used to interrogate the bond-forming process. The experiments were performed at two XFEL facilities (4th generation linear accelerator) that are PAL-XFEL in South Korea and SACLA in Japan, and this study was conducted in collaboration with researchers from KEK IMSS, PAL, RIKEN, and the Japan Synchrotron Radiation Research Institute (JASRI).
Scattered waves from each atom interfere with each other and thus their x-ray scattering images are characterized by specific travel directions. The KAIST-IBS research team traced real-time positions of the three gold atoms over time by analyzing x-ray scattering images, which are determined by a three-dimensional structure of a molecule. Structural changes in the molecule complex resulted in multiple characteristic scattering images over time. When a molecule is excited by a laser pulse, multiple vibrational quantum states are simultaneously excited. The superposition of several excited vibrational quantum states is called a wave packet. The researchers tracked the wave packet in three-dimensional nuclear coordinates and found that the first half round of chemical bonding was formed within 35 fs after photoexcitation. The second half of the reaction followed within 360 fs to complete the entire reaction dynamics.
They also accurately illustrated molecular vibration motions in both temporal- and spatial-wise. This is quite a remarkable feat considering that such an ultrafast speed and a minute length of motion are quite challenging conditions for acquiring precise experimental data.
In this study, the KAIST-IBS research team improved upon their 2015 study published by Nature. In the previous study in 2015, the speed of the x-ray camera (time resolution) was limited to 500 fs, and the molecular structure had already changed to be linear with two chemical bonds within 500 fs. In this study, the progress of the bond formation and bent-to-linear structural transformation could be observed in real time, thanks to the improvement time resolution down to 100 fs. Thereby, the asynchronous bond formation mechanism in which two chemical bonds are formed in 35 fs and 360 fs, respectively, and the bent-to-linear transformation completed in 335 fs were visualized. In short, in addition to observing the beginning and end of chemical reactions, they reported every moment of the intermediate, ongoing rearrangement of nuclear configurations with dramatically improved experimental and analytical methods.
They will push this method of 'real-time tracking of atomic positions in a molecule and molecular vibration using femtosecond x-ray scattering' to reveal the mechanisms of organic and inorganic catalytic reactions and reactions involving proteins in the human body. "By directly tracking the molecular vibrations and real-time positions of all atoms in a molecule in the middle of reaction, we will be able to uncover mechanisms of various unknown organic and inorganic catalytic reactions and biochemical reactions," notes Dr. Jong Goo Kim, the lead author of the study.
Publications:
Kim, J. G., et al. (2020) ‘Mapping the emergence of molecular vibrations mediating bond formation’. Nature. Volume 582. Page 520-524. Available online at https://doi.org/10.1038/s41586-020-2417-3
Profile: Hyotcherl Ihee, Ph.D.
Professor
hyotcherl.ihee@kaist.ac.kr
http://time.kaist.ac.kr/
Ihee Laboratory
Department of Chemistry
KAIST
https://www.kaist.ac.kr
Daejeon 34141, Korea
(END)
2020.06.24 View 11279
Every Moment of Ultrafast Chemical Bonding Now Captured on Film
- The emerging moment of bond formation, two separate bonding steps, and subsequent vibrational motions were visualized. -
< Emergence of molecular vibrations and the evolution to covalent bonds observed in the research. Video Credit: KEK IMSS >
A team of South Korean researchers led by Professor Hyotcherl Ihee from the Department of Chemistry at KAIST reported the direct observation of the birthing moment of chemical bonds by tracking real-time atomic positions in the molecule. Professor Ihee, who also serves as Associate Director of the Center for Nanomaterials and Chemical Reactions at the Institute for Basic Science (IBS), conducted this study in collaboration with scientists at the Institute of Materials Structure Science of High Energy Accelerator Research Organization (KEK IMSS, Japan), RIKEN (Japan), and Pohang Accelerator Laboratory (PAL, South Korea). This work was published in Nature on June 24.
Targeted cancer drugs work by striking a tight bond between cancer cell and specific molecular targets that are involved in the growth and spread of cancer. Detailed images of such chemical bonding sites or pathways can provide key information necessary for maximizing the efficacy of oncogene treatments. However, atomic movements in a molecule have never been captured in the middle of the action, not even for an extremely simple molecule such as a triatomic molecule, made of only three atoms.
Professor Ihee's group and their international collaborators finally succeeded in capturing the ongoing reaction process of the chemical bond formation in the gold trimer. "The femtosecond-resolution images revealed that such molecular events took place in two separate stages, not simultaneously as previously assumed," says Professor Ihee, the corresponding author of the study. "The atoms in the gold trimer complex atoms remain in motion even after the chemical bonding is complete. The distance between the atoms increased and decreased periodically, exhibiting the molecular vibration. These visualized molecular vibrations allowed us to name the characteristic motion of each observed vibrational mode." adds Professor Ihee.
Atoms move extremely fast at a scale of femtosecond (fs) ― quadrillionths (or millionths of a billionth) of a second. Its movement is minute in the level of angstrom equal to one ten-billionth of a meter. They are especially elusive during the transition state where reaction intermediates are transitioning from reactants to products in a flash. The KAIST-IBS research team made this experimentally challenging task possible by using femtosecond x-ray liquidography (solution scattering). This experimental technique combines laser photolysis and x-ray scattering techniques. When a laser pulse strikes the sample, X-rays scatter and initiate the chemical bond formation reaction in the gold trimer complex. Femtosecond x-ray pulses obtained from a special light source called an x-ray free-electron laser (XFEL) were used to interrogate the bond-forming process. The experiments were performed at two XFEL facilities (4th generation linear accelerator) that are PAL-XFEL in South Korea and SACLA in Japan, and this study was conducted in collaboration with researchers from KEK IMSS, PAL, RIKEN, and the Japan Synchrotron Radiation Research Institute (JASRI).
Scattered waves from each atom interfere with each other and thus their x-ray scattering images are characterized by specific travel directions. The KAIST-IBS research team traced real-time positions of the three gold atoms over time by analyzing x-ray scattering images, which are determined by a three-dimensional structure of a molecule. Structural changes in the molecule complex resulted in multiple characteristic scattering images over time. When a molecule is excited by a laser pulse, multiple vibrational quantum states are simultaneously excited. The superposition of several excited vibrational quantum states is called a wave packet. The researchers tracked the wave packet in three-dimensional nuclear coordinates and found that the first half round of chemical bonding was formed within 35 fs after photoexcitation. The second half of the reaction followed within 360 fs to complete the entire reaction dynamics.
They also accurately illustrated molecular vibration motions in both temporal- and spatial-wise. This is quite a remarkable feat considering that such an ultrafast speed and a minute length of motion are quite challenging conditions for acquiring precise experimental data.
In this study, the KAIST-IBS research team improved upon their 2015 study published by Nature. In the previous study in 2015, the speed of the x-ray camera (time resolution) was limited to 500 fs, and the molecular structure had already changed to be linear with two chemical bonds within 500 fs. In this study, the progress of the bond formation and bent-to-linear structural transformation could be observed in real time, thanks to the improvement time resolution down to 100 fs. Thereby, the asynchronous bond formation mechanism in which two chemical bonds are formed in 35 fs and 360 fs, respectively, and the bent-to-linear transformation completed in 335 fs were visualized. In short, in addition to observing the beginning and end of chemical reactions, they reported every moment of the intermediate, ongoing rearrangement of nuclear configurations with dramatically improved experimental and analytical methods.
They will push this method of 'real-time tracking of atomic positions in a molecule and molecular vibration using femtosecond x-ray scattering' to reveal the mechanisms of organic and inorganic catalytic reactions and reactions involving proteins in the human body. "By directly tracking the molecular vibrations and real-time positions of all atoms in a molecule in the middle of reaction, we will be able to uncover mechanisms of various unknown organic and inorganic catalytic reactions and biochemical reactions," notes Dr. Jong Goo Kim, the lead author of the study.
Publications:
Kim, J. G., et al. (2020) ‘Mapping the emergence of molecular vibrations mediating bond formation’. Nature. Volume 582. Page 520-524. Available online at https://doi.org/10.1038/s41586-020-2417-3
Profile: Hyotcherl Ihee, Ph.D.
Professor
hyotcherl.ihee@kaist.ac.kr
http://time.kaist.ac.kr/
Ihee Laboratory
Department of Chemistry
KAIST
https://www.kaist.ac.kr
Daejeon 34141, Korea
(END)
2020.06.24 View 11279 -
 Highly Efficient Charge-to-Spin Interconversion in Graphene Heterostructures
Researchers present a new route for designing a graphene-based active spintronic component
KAIST physicists described a route to design the energy-efficient generation, manipulation and detection of spin currents using nonmagnetic two-dimensional materials. The research team, led by Professor Sungjae Cho, observed highly efficient charge-to-spin interconversion via the gate-tunable Rashba-Edelstien effect (REE) in graphene heterostructures.
This research paves the way for the application of graphene as an active spintronic component for generating, controlling, and detecting spin current without ferromagnetic electrodes or magnetic fields.
Graphene is a promising spintronic component owing to its long spin diffusion length. However, its small spin-orbit coupling limits the potential of graphene in spintronic applications since graphene cannot be used to generate, control, or detect spin current.
“We successfully increased the spin-orbit coupling of graphene by stacking graphene on top of 2H-TaS2, which is one of the transition metal dichalcogenide materials with the largest spin-orbit coupling. Graphene now can be used to generate, control, and detect spin current,” Professor Cho said.
The Rashba-Edelstein effect is a physical mechanism that enables charge current-to-spin current interconversion by spin-dependent band structure induced by the Rashba effect, a momentum-dependent splitting of spin bands in low-dimensional condensed matter systems.
Professor Cho’s group demonstrated the gate-tunable Rashba-Edelstein effect in a multilayer graphene for the first time. The Rahsba-Edelstein effect allows the two-dimensional conduction electrons of graphene to be magnetized by an applied charge current and form a spin current. Furthermore, as the Fermi level of graphene, tuned by gate voltage, moves from the valence to conduction band, the spin current generated by graphene reversed its spin direction.
This spin reversal is useful in the design of low-power-consumption transistors utilizing spins in that it provides the carrier “On” state with spin up holes (or spin down electrons) and the "Off" state with zero net spin polarization at so called “charge neutrality point” where numbers of electrons and holes are equal.
“Our work is the first demonstration of charge-to-spin interconversion in a metallic TMD (transition-metal dichalcogenides) and graphene heterostructure with a spin polarization state controlled by a gate. We expect that the all-electrical spin-switching effect and the reversal of non-equilibrium spin polarization by the application of gate voltage is applicable for the energy-efficient generation and manipulation of spin currents using nonmagnetic van der Waals materials,” explained Professor Cho.
This study (https://pubs.acs.org/doi/10.1021/acsnano.0c01037) was supported by the National Research Foundation of Korea.
Publication:
Lijun Li, Jin Zhang, Gyuho Myeong, Wongil Shin, Hongsik Lim, Boram Kim, Seungho Kim, Taehyeok Jin, Stuart Cavill, Beom Seo Kim, Changyoung Kim, Johannes Lischner, Aires Ferreira, and Sungjae Cho, Gate-Tunable Reversible Rashba−Edelstein Effect in a Few-Layer Graphene/2H-TaS2 Heterostructure at Room Temperature. ACS Nano 2020. Link to download the paper: https://pubs.acs.org/doi/10.1021/acsnano.0c01037
Profile:
Professor Sungjae Cho, PhD
sungjae.cho@kaist.ac.kr
http://qtak.kaist.ac.kr
Department of Physics
Korea Advanced Institute of Science and Technology (KAIST)
https://www.kaist.ac.kr
Daejeon 34141, Korea
2020.05.18 View 6448
Highly Efficient Charge-to-Spin Interconversion in Graphene Heterostructures
Researchers present a new route for designing a graphene-based active spintronic component
KAIST physicists described a route to design the energy-efficient generation, manipulation and detection of spin currents using nonmagnetic two-dimensional materials. The research team, led by Professor Sungjae Cho, observed highly efficient charge-to-spin interconversion via the gate-tunable Rashba-Edelstien effect (REE) in graphene heterostructures.
This research paves the way for the application of graphene as an active spintronic component for generating, controlling, and detecting spin current without ferromagnetic electrodes or magnetic fields.
Graphene is a promising spintronic component owing to its long spin diffusion length. However, its small spin-orbit coupling limits the potential of graphene in spintronic applications since graphene cannot be used to generate, control, or detect spin current.
“We successfully increased the spin-orbit coupling of graphene by stacking graphene on top of 2H-TaS2, which is one of the transition metal dichalcogenide materials with the largest spin-orbit coupling. Graphene now can be used to generate, control, and detect spin current,” Professor Cho said.
The Rashba-Edelstein effect is a physical mechanism that enables charge current-to-spin current interconversion by spin-dependent band structure induced by the Rashba effect, a momentum-dependent splitting of spin bands in low-dimensional condensed matter systems.
Professor Cho’s group demonstrated the gate-tunable Rashba-Edelstein effect in a multilayer graphene for the first time. The Rahsba-Edelstein effect allows the two-dimensional conduction electrons of graphene to be magnetized by an applied charge current and form a spin current. Furthermore, as the Fermi level of graphene, tuned by gate voltage, moves from the valence to conduction band, the spin current generated by graphene reversed its spin direction.
This spin reversal is useful in the design of low-power-consumption transistors utilizing spins in that it provides the carrier “On” state with spin up holes (or spin down electrons) and the "Off" state with zero net spin polarization at so called “charge neutrality point” where numbers of electrons and holes are equal.
“Our work is the first demonstration of charge-to-spin interconversion in a metallic TMD (transition-metal dichalcogenides) and graphene heterostructure with a spin polarization state controlled by a gate. We expect that the all-electrical spin-switching effect and the reversal of non-equilibrium spin polarization by the application of gate voltage is applicable for the energy-efficient generation and manipulation of spin currents using nonmagnetic van der Waals materials,” explained Professor Cho.
This study (https://pubs.acs.org/doi/10.1021/acsnano.0c01037) was supported by the National Research Foundation of Korea.
Publication:
Lijun Li, Jin Zhang, Gyuho Myeong, Wongil Shin, Hongsik Lim, Boram Kim, Seungho Kim, Taehyeok Jin, Stuart Cavill, Beom Seo Kim, Changyoung Kim, Johannes Lischner, Aires Ferreira, and Sungjae Cho, Gate-Tunable Reversible Rashba−Edelstein Effect in a Few-Layer Graphene/2H-TaS2 Heterostructure at Room Temperature. ACS Nano 2020. Link to download the paper: https://pubs.acs.org/doi/10.1021/acsnano.0c01037
Profile:
Professor Sungjae Cho, PhD
sungjae.cho@kaist.ac.kr
http://qtak.kaist.ac.kr
Department of Physics
Korea Advanced Institute of Science and Technology (KAIST)
https://www.kaist.ac.kr
Daejeon 34141, Korea
2020.05.18 View 6448