research
-
 KAIST Develops a Fire-risk Free Self-Powered Hydrogen Production System
KAIST researchers have developed a new hydrogen production system that overcomes the current limitations of green hydrogen production. By using a water-splitting system with an aqueous electrolyte, this system is expected to block fire risks and enable stable hydrogen production.
KAIST (represented by President Kwang Hyung Lee) announced on the 22nd of October that a research team led by Professor Jeung Ku Kang from the Department of Materials Science and Engineering developed a self-powered hydrogen production system based on a high-performance zinc-air battery*.
*Zinc-air battery: A primary battery that absorbs oxygen from the air and uses it as an oxidant. Its advantage is long life, but its low electromotive force is a disadvantage.
Hydrogen (H₂) is a key raw material for synthesizing high-value-added substances, and it is gaining attention as a clean fuel with an energy density (142 MJ/kg) more than three times higher than traditional fossil fuels (gasoline, diesel, etc.). However, most current hydrogen production methods impose environmental burden as they emit carbon dioxide (CO₂).
While green hydrogen can be produced by splitting water using renewable energy sources such as solar cells and wind power, these sources are subject to irregular power generation due to weather and temperature fluctuations, leading to low water-splitting efficiency.
To overcome this, air batteries that can emit sufficient voltage (greater than 1.23V) for water splitting have been gaining attention. However, achieving sufficient capacity requires expensive precious metal catalysts and the performance of the catalyst materials becomes significantly degraded during prolonged charge and discharge cycles. Thus, it is essential to develop catalysts that are effective for the water-splitting reactions (oxygen and hydrogen evolution) and materials that can stabilize the repeated charge and discharge reactions (oxygen reduction and evolution) in zinc-air battery electrodes.
In response, Professor Kang's research team proposed a method to synthesize a non-precious metal catalyst material (G-SHELL) that is effective for three different catalytic reactions (oxygen evolution, hydrogen evolution, and oxygen reduction) by growing nano-sized, metal-organic frameworks on graphene oxide.
The team incorporated the developed catalyst material into the air cathode of a zinc-air battery, confirming that it achieved approximately five times higher energy density (797Wh/kg), high power characteristics (275.8mW/cm²), and long-term stability even under repeated charge and discharge conditions compared to conventional batteries.
Additionally, the zinc-air battery, which operates using an aqueous electrolyte, is safe from fire risks. It is expected that this system can be applied as a next-generation energy storage device when linked with water electrolysis systems, offering an environmentally friendly method for hydrogen production.
< Figure 1. Illustrations of a trifunctional graphene-sandwiched heterojunction-embedded layered lattice (G-SHELL) structure. Schematic representation of a) synthesis procedures of G-SHELL from a zeolitic imidazole framework, b) hollow core-layered shell structure with trifunctional sites for oxygen reduction evolution (ORR), oxygen evolution reaction (OER), and hydrogen evolution reaction (HER), and c) heterojunctions, eterojunction-induced internal electric fields, and the corresponding band structure. >
Professor Kang explained, "By developing a catalyst material with high activity and durability for three different electrochemical catalytic reactions at low temperatures using simple methods, the self-powered hydrogen production system we implemented based on zinc-air batteries presents a new breakthrough to overcome the current limitations of green hydrogen production."
<Figure 2. Electrochemical performance of a ZAB-driven water-splitting cell with G-SHELL. Diagram of a self-driven water-splitting cell integrated by combining a ZAB with an alkaline water electrolyzer.>
PhD candidate Dong Won Kim and Jihoon Kim, a master's student in the Department of Materials Science and Engineering at KAIST, were co-first authors of this research, which was published in the international journal Advanced Science on September 17th in the multidisciplinary field of materials science. (Paper Title: “Trifunctional Graphene-Sandwiched Heterojunction-Embedded Layered Lattice Electrocatalyst for High Performance in Zn-Air Battery-Driven Water Splitting”)
This research was supported by the Nano and Material Technology Development Program of the Ministry of Science and ICT and the National Research Foundation of Korea’s Future Technology Research Laboratory.
2024.10.22 View 5930
KAIST Develops a Fire-risk Free Self-Powered Hydrogen Production System
KAIST researchers have developed a new hydrogen production system that overcomes the current limitations of green hydrogen production. By using a water-splitting system with an aqueous electrolyte, this system is expected to block fire risks and enable stable hydrogen production.
KAIST (represented by President Kwang Hyung Lee) announced on the 22nd of October that a research team led by Professor Jeung Ku Kang from the Department of Materials Science and Engineering developed a self-powered hydrogen production system based on a high-performance zinc-air battery*.
*Zinc-air battery: A primary battery that absorbs oxygen from the air and uses it as an oxidant. Its advantage is long life, but its low electromotive force is a disadvantage.
Hydrogen (H₂) is a key raw material for synthesizing high-value-added substances, and it is gaining attention as a clean fuel with an energy density (142 MJ/kg) more than three times higher than traditional fossil fuels (gasoline, diesel, etc.). However, most current hydrogen production methods impose environmental burden as they emit carbon dioxide (CO₂).
While green hydrogen can be produced by splitting water using renewable energy sources such as solar cells and wind power, these sources are subject to irregular power generation due to weather and temperature fluctuations, leading to low water-splitting efficiency.
To overcome this, air batteries that can emit sufficient voltage (greater than 1.23V) for water splitting have been gaining attention. However, achieving sufficient capacity requires expensive precious metal catalysts and the performance of the catalyst materials becomes significantly degraded during prolonged charge and discharge cycles. Thus, it is essential to develop catalysts that are effective for the water-splitting reactions (oxygen and hydrogen evolution) and materials that can stabilize the repeated charge and discharge reactions (oxygen reduction and evolution) in zinc-air battery electrodes.
In response, Professor Kang's research team proposed a method to synthesize a non-precious metal catalyst material (G-SHELL) that is effective for three different catalytic reactions (oxygen evolution, hydrogen evolution, and oxygen reduction) by growing nano-sized, metal-organic frameworks on graphene oxide.
The team incorporated the developed catalyst material into the air cathode of a zinc-air battery, confirming that it achieved approximately five times higher energy density (797Wh/kg), high power characteristics (275.8mW/cm²), and long-term stability even under repeated charge and discharge conditions compared to conventional batteries.
Additionally, the zinc-air battery, which operates using an aqueous electrolyte, is safe from fire risks. It is expected that this system can be applied as a next-generation energy storage device when linked with water electrolysis systems, offering an environmentally friendly method for hydrogen production.
< Figure 1. Illustrations of a trifunctional graphene-sandwiched heterojunction-embedded layered lattice (G-SHELL) structure. Schematic representation of a) synthesis procedures of G-SHELL from a zeolitic imidazole framework, b) hollow core-layered shell structure with trifunctional sites for oxygen reduction evolution (ORR), oxygen evolution reaction (OER), and hydrogen evolution reaction (HER), and c) heterojunctions, eterojunction-induced internal electric fields, and the corresponding band structure. >
Professor Kang explained, "By developing a catalyst material with high activity and durability for three different electrochemical catalytic reactions at low temperatures using simple methods, the self-powered hydrogen production system we implemented based on zinc-air batteries presents a new breakthrough to overcome the current limitations of green hydrogen production."
<Figure 2. Electrochemical performance of a ZAB-driven water-splitting cell with G-SHELL. Diagram of a self-driven water-splitting cell integrated by combining a ZAB with an alkaline water electrolyzer.>
PhD candidate Dong Won Kim and Jihoon Kim, a master's student in the Department of Materials Science and Engineering at KAIST, were co-first authors of this research, which was published in the international journal Advanced Science on September 17th in the multidisciplinary field of materials science. (Paper Title: “Trifunctional Graphene-Sandwiched Heterojunction-Embedded Layered Lattice Electrocatalyst for High Performance in Zn-Air Battery-Driven Water Splitting”)
This research was supported by the Nano and Material Technology Development Program of the Ministry of Science and ICT and the National Research Foundation of Korea’s Future Technology Research Laboratory.
2024.10.22 View 5930 -
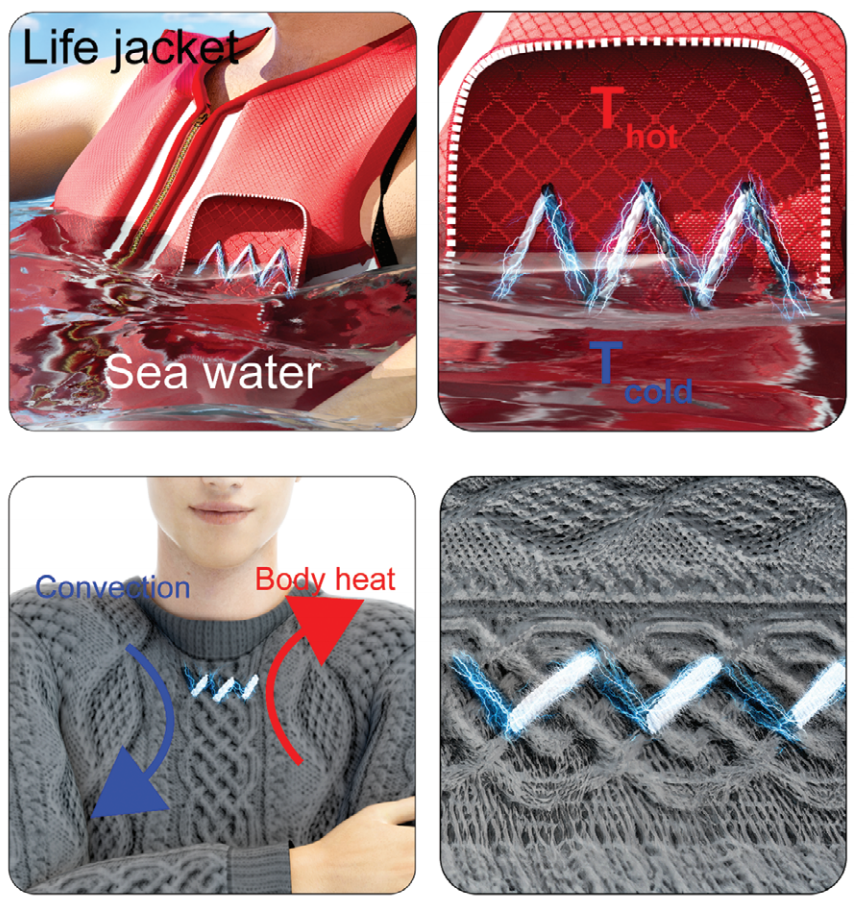 KAIST Develops Thread-like, Flexible Thermoelectric Materials Applicable in Extreme Environments
A team of Korean researchers developed a thermoelectric material that can be used in wearable devices, such as smart clothing, and while maintaining stable thermal energy performance even in extreme environments. It has dramatically resolved the dilemma of striking the balance between achieving good performance and the mechanical flexibility of thermoelectric materials, which has been a long-standing challenge in the field of thermoelectric materials, and has also proven the possibility of commercialization.
KAIST (President Kwang-Hyung Lee) announced on the 21st that a joint research team of Professor Yeon Sik Jung of the Department of Materials Science and Engineering and Professor Inkyu Park of the Department of Mechanical Engineering, in collaboration with the research teams of Professor Min-Wook Oh of Hanbat National University (President Yong Jun Oh) and Dr. Jun-Ho Jeong of the Korea Institute of Machinery and Materials (President Seoghyun Ryu), have successfully developed ‘bismuth telluride (Bi2Te3) thermoelectric fibers,’ an innovative energy harvesting solution for next-generation flexible electronic devices.
Thermoelectric materials are materials that generate voltage when there is a temperature difference and convert thermal energy into electrical energy. Currently, about 70% of energy being lost as wasted heat, so due attention is being given to research on these as sustainable energy materials that can recover and harvesting energy from this waste heat.
Most of the heat sources around us are curved, such as the human body, vehicle exhaust pipes, and cooling fins. Inorganic thermoelectric materials based on ceramic materials boast high thermoelectric performance, but they are fragile and difficult to produce in curved shapes. On the other hand, flexible thermoelectric materials using existing polymer binders can be applied to surfaces of various shapes, but their performance was limited due to the low electrical conductivity and high thermal resistance of the polymer.
Existing flexible thermoelectric materials contain polymer additives, but the inorganic thermoelectric material developed by the research team is not flexible, so they overcame these limitations by twisting nano ribbons instead of additives to produce a thread-shaped thermoelectric material. Inspired by the flexibility of inorganic nano ribbons, the research team used a nanomold-based electron beam deposition technique to continuously deposit nano ribbons and then twisted them into a thread shape to create bismuth telluride (Bi2Te3) inorganic thermoelectric fibers.
These inorganic thermoelectric fibers have higher bending strength than existing thermoelectric materials, and showed almost no change in electrical properties even after repeated bending and tensile tests of more than 1,000 times. The thermoelectric device created by the research team generates electricity using temperature differences, and if clothes are made with fiber-type thermoelectric devices, electricity can be generated from body temperature to operate other electronic devices.
< Figure 1. Schematic diagram and actual image of the all-inorganic flexible thermoelectric yarn made without polymer additives >
In fact, the possibility of commercialization was proven through a demonstration of collecting energy by embedding thermoelectric fibers in life jackets or clothing. In addition, it opened up the possibility of building a high-efficiency energy harvesting system that recycles waste heat by utilizing the temperature difference between the hot fluid inside a pipe and the cold air outside in industrial settings.
Professor Yeon Sik Jung said, "The inorganic flexible thermoelectric material developed in this study can be used in wearable devices such as smart clothing, and it can maintain stable performance even in extreme environments, so it has a high possibility of being commercialized through additional research in the future." Professor Inkyu Park also emphasized, "This technology will become the core of next-generation energy harvesting technology, and it is expected to play an important role in various fields from waste heat utilization in industrial sites to personal wearable self-power generation devices."
This study, in which Hanhwi Jang, a Ph.D. student at KAIST's Department of Materials Science and Engineering, Professor Junseong Ahn of Korea University, Sejong Campus, and Dr. Yongrok Jeong of Korea Atomic Energy Research Institute contributed equally as joint first authors, was published in the online edition of the international academic journal Advanced Materials on September 17, and was selected as the back-cover paper in recognition of its excellence. (Paper title: Flexible All-Inorganic Thermoelectric Yarns)
Meanwhile, this study was conducted through the Mid-career Researcher Support Program and the Future Materials Discovery Program of the National Research Foundation of Korea, and the support from the Global Bio-Integrated Materials Center, the Ministry of Trade, Industry and Energy, and the Korea Institute of Industrial Technology Evaluation and Planning (KEIT) upon the support by the Ministry of Science and ICT.
2024.10.21 View 4710
KAIST Develops Thread-like, Flexible Thermoelectric Materials Applicable in Extreme Environments
A team of Korean researchers developed a thermoelectric material that can be used in wearable devices, such as smart clothing, and while maintaining stable thermal energy performance even in extreme environments. It has dramatically resolved the dilemma of striking the balance between achieving good performance and the mechanical flexibility of thermoelectric materials, which has been a long-standing challenge in the field of thermoelectric materials, and has also proven the possibility of commercialization.
KAIST (President Kwang-Hyung Lee) announced on the 21st that a joint research team of Professor Yeon Sik Jung of the Department of Materials Science and Engineering and Professor Inkyu Park of the Department of Mechanical Engineering, in collaboration with the research teams of Professor Min-Wook Oh of Hanbat National University (President Yong Jun Oh) and Dr. Jun-Ho Jeong of the Korea Institute of Machinery and Materials (President Seoghyun Ryu), have successfully developed ‘bismuth telluride (Bi2Te3) thermoelectric fibers,’ an innovative energy harvesting solution for next-generation flexible electronic devices.
Thermoelectric materials are materials that generate voltage when there is a temperature difference and convert thermal energy into electrical energy. Currently, about 70% of energy being lost as wasted heat, so due attention is being given to research on these as sustainable energy materials that can recover and harvesting energy from this waste heat.
Most of the heat sources around us are curved, such as the human body, vehicle exhaust pipes, and cooling fins. Inorganic thermoelectric materials based on ceramic materials boast high thermoelectric performance, but they are fragile and difficult to produce in curved shapes. On the other hand, flexible thermoelectric materials using existing polymer binders can be applied to surfaces of various shapes, but their performance was limited due to the low electrical conductivity and high thermal resistance of the polymer.
Existing flexible thermoelectric materials contain polymer additives, but the inorganic thermoelectric material developed by the research team is not flexible, so they overcame these limitations by twisting nano ribbons instead of additives to produce a thread-shaped thermoelectric material. Inspired by the flexibility of inorganic nano ribbons, the research team used a nanomold-based electron beam deposition technique to continuously deposit nano ribbons and then twisted them into a thread shape to create bismuth telluride (Bi2Te3) inorganic thermoelectric fibers.
These inorganic thermoelectric fibers have higher bending strength than existing thermoelectric materials, and showed almost no change in electrical properties even after repeated bending and tensile tests of more than 1,000 times. The thermoelectric device created by the research team generates electricity using temperature differences, and if clothes are made with fiber-type thermoelectric devices, electricity can be generated from body temperature to operate other electronic devices.
< Figure 1. Schematic diagram and actual image of the all-inorganic flexible thermoelectric yarn made without polymer additives >
In fact, the possibility of commercialization was proven through a demonstration of collecting energy by embedding thermoelectric fibers in life jackets or clothing. In addition, it opened up the possibility of building a high-efficiency energy harvesting system that recycles waste heat by utilizing the temperature difference between the hot fluid inside a pipe and the cold air outside in industrial settings.
Professor Yeon Sik Jung said, "The inorganic flexible thermoelectric material developed in this study can be used in wearable devices such as smart clothing, and it can maintain stable performance even in extreme environments, so it has a high possibility of being commercialized through additional research in the future." Professor Inkyu Park also emphasized, "This technology will become the core of next-generation energy harvesting technology, and it is expected to play an important role in various fields from waste heat utilization in industrial sites to personal wearable self-power generation devices."
This study, in which Hanhwi Jang, a Ph.D. student at KAIST's Department of Materials Science and Engineering, Professor Junseong Ahn of Korea University, Sejong Campus, and Dr. Yongrok Jeong of Korea Atomic Energy Research Institute contributed equally as joint first authors, was published in the online edition of the international academic journal Advanced Materials on September 17, and was selected as the back-cover paper in recognition of its excellence. (Paper title: Flexible All-Inorganic Thermoelectric Yarns)
Meanwhile, this study was conducted through the Mid-career Researcher Support Program and the Future Materials Discovery Program of the National Research Foundation of Korea, and the support from the Global Bio-Integrated Materials Center, the Ministry of Trade, Industry and Energy, and the Korea Institute of Industrial Technology Evaluation and Planning (KEIT) upon the support by the Ministry of Science and ICT.
2024.10.21 View 4710 -
 KAIST Develops Technology for the Precise Diagnosis of Electric Vehicle Batteries Using Small Currents
Accurately diagnosing the state of electric vehicle (EV) batteries is essential for their efficient management and safe use. KAIST researchers have developed a new technology that can diagnose and monitor the state of batteries with high precision using only small amounts of current, which is expected to maximize the batteries’ long-term stability and efficiency.
KAIST (represented by President Kwang Hyung Lee) announced on the 17th of October that a research team led by Professors Kyeongha Kwon and Sang-Gug Lee from the School of Electrical Engineering had developed electrochemical impedance spectroscopy (EIS) technology that can be used to improve the stability and performance of high-capacity batteries in electric vehicles.
EIS is a powerful tool that measures the impedance* magnitude and changes in a battery, allowing the evaluation of battery efficiency and loss. It is considered an important tool for assessing the state of charge (SOC) and state of health (SOH) of batteries. Additionally, it can be used to identify thermal characteristics, chemical/physical changes, predict battery life, and determine the causes of failures. *Battery Impedance: A measure of the resistance to current flow within the battery that is used to assess battery performance and condition.
However, traditional EIS equipment is expensive and complex, making it difficult to install, operate, and maintain. Moreover, due to sensitivity and precision limitations, applying current disturbances of several amperes (A) to a battery can cause significant electrical stress, increasing the risk of battery failure or fire and making it difficult to use in practice.
< Figure 1. Flow chart for diagnosis and prevention of unexpected combustion via the use of the electrochemical impedance spectroscopy (EIS) for the batteries for electric vehicles. >
To address this, the KAIST research team developed and validated a low-current EIS system for diagnosing the condition and health of high-capacity EV batteries. This EIS system can precisely measure battery impedance with low current disturbances (10mA), minimizing thermal effects and safety issues during the measurement process.
In addition, the system minimizes bulky and costly components, making it easy to integrate into vehicles. The system was proven effective in identifying the electrochemical properties of batteries under various operating conditions, including different temperatures and SOC levels.
Professor Kyeongha Kwon (the corresponding author) explained, “This system can be easily integrated into the battery management system (BMS) of electric vehicles and has demonstrated high measurement accuracy while significantly reducing the cost and complexity compared to traditional high-current EIS methods. It can contribute to battery diagnosis and performance improvements not only for electric vehicles but also for energy storage systems (ESS).”
This research, in which Young-Nam Lee, a doctoral student in the School of Electrical Engineering at KAIST participated as the first author, was published in the prestigious international journal IEEE Transactions on Industrial Electronics (top 2% in the field; IF 7.5) on September 5th. (Paper Title: Small-Perturbation Electrochemical Impedance Spectroscopy System With High Accuracy for High-Capacity Batteries in Electric Vehicles, Link: https://ieeexplore.ieee.org/document/10666864)
< Figure 2. Impedance measurement results of large-capacity batteries for electric vehicles. ZEW (commercial EW; MP10, Wonatech) versus ZMEAS (proposed system) >
This research was supported by the Basic Research Program of the National Research Foundation of Korea, the Next-Generation Intelligent Semiconductor Technology Development Program of the Korea Evaluation Institute of Industrial Technology, and the AI Semiconductor Graduate Program of the Institute of Information & Communications Technology Planning & Evaluation.
2024.10.17 View 6428
KAIST Develops Technology for the Precise Diagnosis of Electric Vehicle Batteries Using Small Currents
Accurately diagnosing the state of electric vehicle (EV) batteries is essential for their efficient management and safe use. KAIST researchers have developed a new technology that can diagnose and monitor the state of batteries with high precision using only small amounts of current, which is expected to maximize the batteries’ long-term stability and efficiency.
KAIST (represented by President Kwang Hyung Lee) announced on the 17th of October that a research team led by Professors Kyeongha Kwon and Sang-Gug Lee from the School of Electrical Engineering had developed electrochemical impedance spectroscopy (EIS) technology that can be used to improve the stability and performance of high-capacity batteries in electric vehicles.
EIS is a powerful tool that measures the impedance* magnitude and changes in a battery, allowing the evaluation of battery efficiency and loss. It is considered an important tool for assessing the state of charge (SOC) and state of health (SOH) of batteries. Additionally, it can be used to identify thermal characteristics, chemical/physical changes, predict battery life, and determine the causes of failures. *Battery Impedance: A measure of the resistance to current flow within the battery that is used to assess battery performance and condition.
However, traditional EIS equipment is expensive and complex, making it difficult to install, operate, and maintain. Moreover, due to sensitivity and precision limitations, applying current disturbances of several amperes (A) to a battery can cause significant electrical stress, increasing the risk of battery failure or fire and making it difficult to use in practice.
< Figure 1. Flow chart for diagnosis and prevention of unexpected combustion via the use of the electrochemical impedance spectroscopy (EIS) for the batteries for electric vehicles. >
To address this, the KAIST research team developed and validated a low-current EIS system for diagnosing the condition and health of high-capacity EV batteries. This EIS system can precisely measure battery impedance with low current disturbances (10mA), minimizing thermal effects and safety issues during the measurement process.
In addition, the system minimizes bulky and costly components, making it easy to integrate into vehicles. The system was proven effective in identifying the electrochemical properties of batteries under various operating conditions, including different temperatures and SOC levels.
Professor Kyeongha Kwon (the corresponding author) explained, “This system can be easily integrated into the battery management system (BMS) of electric vehicles and has demonstrated high measurement accuracy while significantly reducing the cost and complexity compared to traditional high-current EIS methods. It can contribute to battery diagnosis and performance improvements not only for electric vehicles but also for energy storage systems (ESS).”
This research, in which Young-Nam Lee, a doctoral student in the School of Electrical Engineering at KAIST participated as the first author, was published in the prestigious international journal IEEE Transactions on Industrial Electronics (top 2% in the field; IF 7.5) on September 5th. (Paper Title: Small-Perturbation Electrochemical Impedance Spectroscopy System With High Accuracy for High-Capacity Batteries in Electric Vehicles, Link: https://ieeexplore.ieee.org/document/10666864)
< Figure 2. Impedance measurement results of large-capacity batteries for electric vehicles. ZEW (commercial EW; MP10, Wonatech) versus ZMEAS (proposed system) >
This research was supported by the Basic Research Program of the National Research Foundation of Korea, the Next-Generation Intelligent Semiconductor Technology Development Program of the Korea Evaluation Institute of Industrial Technology, and the AI Semiconductor Graduate Program of the Institute of Information & Communications Technology Planning & Evaluation.
2024.10.17 View 6428 -
 KAIST Develops Janus-like Metasurface Technology that Acts According to the Direction of Light
Metasurface technology is an advanced optical technology that is thinner, lighter, and capable of precisely controlling light through nanometer-sized artificial structures compared to conventional technologies. KAIST researchers have overcome the limitations of existing metasurface technologies and successfully designed a Janus metasurface capable of perfectly controlling asymmetric light transmission. By applying this technology, they also proposed an innovative method to significantly enhance security by only decoding information under specific conditions.
KAIST (represented by President Kwang Hyung Lee) announced on the 15th of October that a research team led by Professor Jonghwa Shin from the Department of Materials Science and Engineering had developed a Janus metasurface capable of perfectly controlling asymmetric light transmission.
Asymmetric properties, which react differently depending on the direction, play a crucial role in various fields of science and engineering. The Janus metasurface developed by the research team implements an optical system capable of performing different functions in both directions.
Like the Roman god Janus with two faces, this metasurface shows entirely different optical responses depending on the direction of incoming light, effectively operating two independent optical systems with a single device (for example, a metasurface that acts as a magnifying lens in one direction and as a polarized camera in the other). In other words, by using this technology, it's possible to operate two different optical systems (e.g., a lens and a hologram) depending on the direction of the light.
This achievement addresses a challenge that existing metasurface technologies had not resolved. Conventional metasurface technology had limitations in selectively controlling the three properties of light—intensity, phase, and polarization—based on the direction of incidence.
The research team proposed a solution based on mathematical and physical principles, and succeeded in experimentally implementing different vector holograms in both directions. Through this achievement, they showcased a complete asymmetric light transmission control technology.
< Figure 1. Schematics of a device featuring asymmetric transmission. a) Device operating as a magnifying lens for back-side illumination. b) Device operating as a polarization camera for front-side illumination. >
Additionally, the research team developed a new optical encryption technology based on this metasurface technology. By using the Janus metasurface, they implemented a vector hologram that generates different images depending on the direction and polarization state of incoming light, showcasing an optical encryption system that significantly enhances security by allowing information to be decoded only under specific conditions.
This technology is expected to serve as a next-generation security solution, applicable in various fields such as quantum communication and secure data transmission.
Furthermore, the ultra-thin structure of the metasurface is expected to significantly reduce the volume and weight of traditional optical devices, contributing greatly to the miniaturization and lightweight design of next-generation devices.
< Figure 2. Experimental demonstration of Janus vectorial holograms. With front illuminations, vector images of the butterfly and the grasshopper are created, and with the back-side illuminations, vector images of the ladybug and the beetle are created. >
Professor Jonghwa Shin from the Department of Materials Science and Engineering at KAIST stated, "This research has enabled the complete asymmetric transmission control of light’s intensity, phase, and polarization, which has been a long-standing challenge in optics. It has opened up the possibility of developing various applied optical devices." He added, "We plan to continue developing optical devices that can be applied to various fields such as augmented reality (AR), holographic displays, and LiDAR systems for autonomous vehicles, utilizing the full potential of metasurface technology."
This research, in which Hyeonhee Kim (a doctoral student in the Department of Materials Science and Engineering at KAIST) and Joonkyo Jung participated as co-first authors, was published online in the international journal Advanced Materials and is scheduled to be published in the October 31 issue. (Title of the paper: "Bidirectional Vectorial Holography Using Bi-Layer Metasurfaces and Its Application to Optical Encryption")
The research was supported by the Nano Materials Technology Development Program and the Mid-Career Researcher Program of the National Research Foundation of Korea.
2024.10.15 View 4101
KAIST Develops Janus-like Metasurface Technology that Acts According to the Direction of Light
Metasurface technology is an advanced optical technology that is thinner, lighter, and capable of precisely controlling light through nanometer-sized artificial structures compared to conventional technologies. KAIST researchers have overcome the limitations of existing metasurface technologies and successfully designed a Janus metasurface capable of perfectly controlling asymmetric light transmission. By applying this technology, they also proposed an innovative method to significantly enhance security by only decoding information under specific conditions.
KAIST (represented by President Kwang Hyung Lee) announced on the 15th of October that a research team led by Professor Jonghwa Shin from the Department of Materials Science and Engineering had developed a Janus metasurface capable of perfectly controlling asymmetric light transmission.
Asymmetric properties, which react differently depending on the direction, play a crucial role in various fields of science and engineering. The Janus metasurface developed by the research team implements an optical system capable of performing different functions in both directions.
Like the Roman god Janus with two faces, this metasurface shows entirely different optical responses depending on the direction of incoming light, effectively operating two independent optical systems with a single device (for example, a metasurface that acts as a magnifying lens in one direction and as a polarized camera in the other). In other words, by using this technology, it's possible to operate two different optical systems (e.g., a lens and a hologram) depending on the direction of the light.
This achievement addresses a challenge that existing metasurface technologies had not resolved. Conventional metasurface technology had limitations in selectively controlling the three properties of light—intensity, phase, and polarization—based on the direction of incidence.
The research team proposed a solution based on mathematical and physical principles, and succeeded in experimentally implementing different vector holograms in both directions. Through this achievement, they showcased a complete asymmetric light transmission control technology.
< Figure 1. Schematics of a device featuring asymmetric transmission. a) Device operating as a magnifying lens for back-side illumination. b) Device operating as a polarization camera for front-side illumination. >
Additionally, the research team developed a new optical encryption technology based on this metasurface technology. By using the Janus metasurface, they implemented a vector hologram that generates different images depending on the direction and polarization state of incoming light, showcasing an optical encryption system that significantly enhances security by allowing information to be decoded only under specific conditions.
This technology is expected to serve as a next-generation security solution, applicable in various fields such as quantum communication and secure data transmission.
Furthermore, the ultra-thin structure of the metasurface is expected to significantly reduce the volume and weight of traditional optical devices, contributing greatly to the miniaturization and lightweight design of next-generation devices.
< Figure 2. Experimental demonstration of Janus vectorial holograms. With front illuminations, vector images of the butterfly and the grasshopper are created, and with the back-side illuminations, vector images of the ladybug and the beetle are created. >
Professor Jonghwa Shin from the Department of Materials Science and Engineering at KAIST stated, "This research has enabled the complete asymmetric transmission control of light’s intensity, phase, and polarization, which has been a long-standing challenge in optics. It has opened up the possibility of developing various applied optical devices." He added, "We plan to continue developing optical devices that can be applied to various fields such as augmented reality (AR), holographic displays, and LiDAR systems for autonomous vehicles, utilizing the full potential of metasurface technology."
This research, in which Hyeonhee Kim (a doctoral student in the Department of Materials Science and Engineering at KAIST) and Joonkyo Jung participated as co-first authors, was published online in the international journal Advanced Materials and is scheduled to be published in the October 31 issue. (Title of the paper: "Bidirectional Vectorial Holography Using Bi-Layer Metasurfaces and Its Application to Optical Encryption")
The research was supported by the Nano Materials Technology Development Program and the Mid-Career Researcher Program of the National Research Foundation of Korea.
2024.10.15 View 4101 -
 KAIST Succeeds in the Real-time Observation of Organoids using Holotomography
Organoids, which are 3D miniature organs that mimic the structure and function of human organs, play an essential role in disease research and drug development. A Korean research team has overcome the limitations of existing imaging technologies, succeeding in the real-time, high-resolution observation of living organoids.
KAIST (represented by President Kwang Hyung Lee) announced on the 14th of October that Professor YongKeun Park’s research team from the Department of Physics, in collaboration with the Genome Editing Research Center (Director Bon-Kyoung Koo) of the Institute for Basic Science (IBS President Do-Young Noh) and Tomocube Inc., has developed an imaging technology using holotomography to observe live, small intestinal organoids in real time at a high resolution.
Existing imaging techniques have struggled to observe living organoids in high resolution over extended periods and often required additional treatments like fluorescent staining.
< Figure 1. Overview of the low-coherence HT workflow. Using holotomography, 3D morphological restoration and quantitative analysis of organoids can be performed. In order to improve the limited field of view, which is a limitation of the microscope, our research team utilized a large-area field of view combination algorithm and made a 3D restoration by acquiring multi-focus holographic images for 3D measurements. After that, the organoids were compartmentalized to divide the parts necessary for analysis and quantitatively evaluated the protein concentration measurable from the refractive index and the survival rate of the organoids. >
The research team introduced holotomography technology to address these issues, which provides high-resolution images without the need for fluorescent staining and allows for the long-term observation of dynamic changes in real time without causing cell damage.
The team validated this technology using small intestinal organoids from experimental mice and were able to observe various cell structures inside the organoids in detail. They also captured dynamic changes such as growth processes, cell division, and cell death in real time using holotomography.
Additionally, the technology allowed for the precise analysis of the organoids' responses to drug treatments, verifying the survival of the cells.
The researchers believe that this breakthrough will open new horizons in organoid research, enabling the greater utilization of organoids in drug development, personalized medicine, and regenerative medicine.
Future research is expected to more accurately replicate the in vivo environment of organoids, contributing significantly to a more detailed understanding of various life phenomena at the cellular level through more precise 3D imaging.
< Figure 2. Real-time organoid morphology analysis. Using holotomography, it is possible to observe the lumen and villus development process of intestinal organoids in real time, which was difficult to observe with a conventional microscope. In addition, various information about intestinal organoids can be obtained by quantifying the size and protein amount of intestinal organoids through image analysis. >
Dr. Mahn Jae Lee, a graduate of KAIST's Graduate School of Medical Science and Engineering, currently at Chungnam National University Hospital and the first author of the paper, commented, "This research represents a new imaging technology that surpasses previous limitations and is expected to make a major contribution to disease modeling, personalized treatments, and drug development research using organoids."
The research results were published online in the international journal Experimental & Molecular Medicine on October 1, 2024, and the technology has been recognized for its applicability in various fields of life sciences. (Paper title: “Long-term three-dimensional high-resolution imaging of live unlabeled small intestinal organoids via low-coherence holotomography”)
This research was supported by the National Research Foundation of Korea, KAIST Institutes, and the Institute for Basic Science.
2024.10.14 View 5211
KAIST Succeeds in the Real-time Observation of Organoids using Holotomography
Organoids, which are 3D miniature organs that mimic the structure and function of human organs, play an essential role in disease research and drug development. A Korean research team has overcome the limitations of existing imaging technologies, succeeding in the real-time, high-resolution observation of living organoids.
KAIST (represented by President Kwang Hyung Lee) announced on the 14th of October that Professor YongKeun Park’s research team from the Department of Physics, in collaboration with the Genome Editing Research Center (Director Bon-Kyoung Koo) of the Institute for Basic Science (IBS President Do-Young Noh) and Tomocube Inc., has developed an imaging technology using holotomography to observe live, small intestinal organoids in real time at a high resolution.
Existing imaging techniques have struggled to observe living organoids in high resolution over extended periods and often required additional treatments like fluorescent staining.
< Figure 1. Overview of the low-coherence HT workflow. Using holotomography, 3D morphological restoration and quantitative analysis of organoids can be performed. In order to improve the limited field of view, which is a limitation of the microscope, our research team utilized a large-area field of view combination algorithm and made a 3D restoration by acquiring multi-focus holographic images for 3D measurements. After that, the organoids were compartmentalized to divide the parts necessary for analysis and quantitatively evaluated the protein concentration measurable from the refractive index and the survival rate of the organoids. >
The research team introduced holotomography technology to address these issues, which provides high-resolution images without the need for fluorescent staining and allows for the long-term observation of dynamic changes in real time without causing cell damage.
The team validated this technology using small intestinal organoids from experimental mice and were able to observe various cell structures inside the organoids in detail. They also captured dynamic changes such as growth processes, cell division, and cell death in real time using holotomography.
Additionally, the technology allowed for the precise analysis of the organoids' responses to drug treatments, verifying the survival of the cells.
The researchers believe that this breakthrough will open new horizons in organoid research, enabling the greater utilization of organoids in drug development, personalized medicine, and regenerative medicine.
Future research is expected to more accurately replicate the in vivo environment of organoids, contributing significantly to a more detailed understanding of various life phenomena at the cellular level through more precise 3D imaging.
< Figure 2. Real-time organoid morphology analysis. Using holotomography, it is possible to observe the lumen and villus development process of intestinal organoids in real time, which was difficult to observe with a conventional microscope. In addition, various information about intestinal organoids can be obtained by quantifying the size and protein amount of intestinal organoids through image analysis. >
Dr. Mahn Jae Lee, a graduate of KAIST's Graduate School of Medical Science and Engineering, currently at Chungnam National University Hospital and the first author of the paper, commented, "This research represents a new imaging technology that surpasses previous limitations and is expected to make a major contribution to disease modeling, personalized treatments, and drug development research using organoids."
The research results were published online in the international journal Experimental & Molecular Medicine on October 1, 2024, and the technology has been recognized for its applicability in various fields of life sciences. (Paper title: “Long-term three-dimensional high-resolution imaging of live unlabeled small intestinal organoids via low-coherence holotomography”)
This research was supported by the National Research Foundation of Korea, KAIST Institutes, and the Institute for Basic Science.
2024.10.14 View 5211 -
 KAIST Changes the Paradigm of Drug Discovery with World's First Atomic Editing
In pioneering drug development, the new technology that enables the easy and rapid editing of key atoms responsible for drug efficacy has been regarded as a fundamental and "dream" technology, revolutionizing the process of discovering potential drug candidates. KAIST researchers have become the first in the world to successfully develop single-atom editing technology that maximizes drug efficacy.
On October 8th, KAIST (represented by President Kwang-Hyung Lee) announced that Professor Yoonsu Park’s research team from the Department of Chemistry successfully developed technology that enables the easy editing and correction of oxygen atoms in furan compounds into nitrogen atoms, directly converting them into pyrrole frameworks, which are widely used in pharmaceuticals.
< Image. Conceptual image illustrating the main idea of the research >
This research was published in the prestigious scientific journal Science on October 3rd under the title "Photocatalytic Furan-to-Pyrrole Conversion."
Many drugs have complex chemical structures, but their efficacy is often determined by a single critical atom. Atoms like oxygen and nitrogen play a central role in enhancing the pharmacological effects of these drugs, particularly against viruses.
This phenomenon, where the introduction of specific atoms into a drug molecule dramatically affects its efficacy, is known as the "Single Atom Effect." In leading-edge drug development, discovering atoms that maximize drug efficacy is key.
However, evaluating the Single Atom Effect has traditionally required multi-step, costly synthesis processes, as it has been difficult to selectively edit single atoms within stable ring structures containing oxygen or nitrogen.
Professor Park’s team overcame this challenge by introducing a photocatalyst that uses light energy. They developed a photocatalyst that acts as a “molecular scissor,” freely cutting and attaching five-membered rings, enabling single-atom editing at room temperature and atmospheric pressure—a world first.
The team discovered a new reaction mechanism in which the excited molecular scissor removes oxygen from furan via single-electron oxidation and then sequentially adds a nitrogen atom.
Donghyeon Kim and Jaehyun You, the study's first authors and candidates in KAIST’s integrated master's and doctoral program in the Department of Chemistry, explained that this technique offers high versatility by utilizing light energy to replace harsh conditions. They further noted that the technology enables selective editing, even when applied to complex natural products or pharmaceuticals. Professor Yoonsu Park, who led the research, remarked, "This breakthrough, which allows for the selective editing of five-membered organic ring structures, will open new doors for building libraries of drug candidates, a key challenge in pharmaceuticals. I hope this foundational technology will be used to revolutionize the drug development process."
The significance of this research was highlighted in the Perspective section of Science, a feature where a peer scientist of prominence outside of the project group provides commentary on an impactful research.
This research was supported by the National Research Foundation of Korea’s Creative Research Program, the Cross-Generation Collaborative Lab Project at KAIST, and the POSCO Science Fellowship of the POSCO TJ Park Foundation.
2024.10.11 View 5678
KAIST Changes the Paradigm of Drug Discovery with World's First Atomic Editing
In pioneering drug development, the new technology that enables the easy and rapid editing of key atoms responsible for drug efficacy has been regarded as a fundamental and "dream" technology, revolutionizing the process of discovering potential drug candidates. KAIST researchers have become the first in the world to successfully develop single-atom editing technology that maximizes drug efficacy.
On October 8th, KAIST (represented by President Kwang-Hyung Lee) announced that Professor Yoonsu Park’s research team from the Department of Chemistry successfully developed technology that enables the easy editing and correction of oxygen atoms in furan compounds into nitrogen atoms, directly converting them into pyrrole frameworks, which are widely used in pharmaceuticals.
< Image. Conceptual image illustrating the main idea of the research >
This research was published in the prestigious scientific journal Science on October 3rd under the title "Photocatalytic Furan-to-Pyrrole Conversion."
Many drugs have complex chemical structures, but their efficacy is often determined by a single critical atom. Atoms like oxygen and nitrogen play a central role in enhancing the pharmacological effects of these drugs, particularly against viruses.
This phenomenon, where the introduction of specific atoms into a drug molecule dramatically affects its efficacy, is known as the "Single Atom Effect." In leading-edge drug development, discovering atoms that maximize drug efficacy is key.
However, evaluating the Single Atom Effect has traditionally required multi-step, costly synthesis processes, as it has been difficult to selectively edit single atoms within stable ring structures containing oxygen or nitrogen.
Professor Park’s team overcame this challenge by introducing a photocatalyst that uses light energy. They developed a photocatalyst that acts as a “molecular scissor,” freely cutting and attaching five-membered rings, enabling single-atom editing at room temperature and atmospheric pressure—a world first.
The team discovered a new reaction mechanism in which the excited molecular scissor removes oxygen from furan via single-electron oxidation and then sequentially adds a nitrogen atom.
Donghyeon Kim and Jaehyun You, the study's first authors and candidates in KAIST’s integrated master's and doctoral program in the Department of Chemistry, explained that this technique offers high versatility by utilizing light energy to replace harsh conditions. They further noted that the technology enables selective editing, even when applied to complex natural products or pharmaceuticals. Professor Yoonsu Park, who led the research, remarked, "This breakthrough, which allows for the selective editing of five-membered organic ring structures, will open new doors for building libraries of drug candidates, a key challenge in pharmaceuticals. I hope this foundational technology will be used to revolutionize the drug development process."
The significance of this research was highlighted in the Perspective section of Science, a feature where a peer scientist of prominence outside of the project group provides commentary on an impactful research.
This research was supported by the National Research Foundation of Korea’s Creative Research Program, the Cross-Generation Collaborative Lab Project at KAIST, and the POSCO Science Fellowship of the POSCO TJ Park Foundation.
2024.10.11 View 5678 -
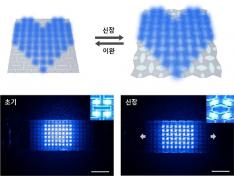 KAIST Develops Stretchable Displays Featuring 25% Expansion Without Image Distortion
Stretchable displays, praised for their spatial efficiency, design flexibility, and human-like flexibility, are seen as the next generation of display technology. A team of Korean researchers has developed a stretchable display that can expand by 25% while maintaining clear image quality without distortion. It can also stretch and contract up to 5,000 times at 15% expansion without any performance degradation, making it the first deformation-free stretchable display with a negative Poisson's ratio* developed in Korea.
*Poisson’s ratio of -1: A ratio where both width and length stretch equally, expressed as a negative value. A positive Poisson's ratio represents the ratio where horizontal stretching leads to vertical contraction, which is the case for most materials.
KAIST (represented by President Kwang-Hyung Lee) announced on the 20th of August that a research team led by Professor Byeong-Soo Bae of the Department of Materials Science and Engineering (Director of the Wearable Platform Materials Technology Center) , in collaboration with the Korea Institute of Machinery & Materials (President Seoghyeon Ryu), successfully developed a stretchable display substrate that suppresses image distortion through omnidirectional stretchability.
Currently, most stretchable displays are made with highly elastic elastomer* materials, but these materials possess a positive Poisson's ratio, causing unavoidable image distortion when the display is stretched.
*Elastomer: A polymer with elasticity similar to rubber.
To address this, the introduction of auxetic* meta-structures has been gaining attention. Unlike conventional materials, auxetic structures have a unique 'negative Poisson's ratio,' expanding in all directions when stretched in just one direction. However, traditional auxetic structures contain many empty spaces, limiting their stability and usability in display substrates.
*Auxetic structure: A special geometric structure that exhibits a negative Poisson's ratio.
To tackle the issue of image distortion, Professor Bae's research team developed a method to create a seamless surface for the auxetic meta-structure, achieving the ideal negative Poisson's ratio of -1 and overcoming the biggest challenge in auxetic meta-structures.
To overcome the second issue of elastic modulus*, the team inserted a textile made of glass fiber bundles with a diameter of just 25 micrometers (a quarter of the thickness of human hair) into the elastomer material. They then filled the empty spaces with the same elastomer, creating a flat and stable integrated film without gaps.
*Elastic Modulus: The ratio that indicates the extent of deformation when force is applied to a material. A higher elastic modulus means that the material is less likely to deform under force.
The research team theoretically identified that the difference in elasticity between the auxetic structure and the elastomer material directly influences the negative Poisson's ratio and successfully achieved an elasticity difference of over 230,000 times, producing a film with a Poisson's ratio of -1, the theoretical limit.
< Figure 2. Deformation of S-AUX film. a) Configurations and visualized principal strain distribution of the optimized S-AUX film at various strain rates. b) Biaxial stretching image. While pristine elastomer shrinks in the directions that were not stretched, S-AUX film developed in this study expands in all directions simultaneously while maintaining its original shape. >
Professor Byeong-Soo Bae, who led the study, explained, "Preventing image distortion using auxetic structures in stretchable displays is a core technology, but it has faced challenges due to the many empty spaces in the surface, making it difficult to use as a substrate. This research outcome is expected to significantly accelerate commercialization through high-resolution, distortion-free stretchable display applications that utilize the entire surface."
This study, co-authored by Dr. Yung Lee from KAIST’s Department of Materials Science and Engineering and Dr. Bongkyun Jang from the Korea Institute of Machinery & Materials, was published on August 20th in the international journal Nature Communications under the title "A seamless auxetic substrate with a negative Poisson's ratio of –1".
The research was supported by the Wearable Platform Materials Technology Center at KAIST, the Korea Institute of Machinery & Materials, and LG Display.
< Figure 3. Structural configuration of the distortion-free display components on the S-AUX film and a contour image of a micro-LED chip transferred onto the S-AUX film. >
< Figure 4. Schematic illustrations and photographic images of the S-AUX film-based image: distortion-free display in its stretched state and released state. >
2024.09.20 View 5946
KAIST Develops Stretchable Displays Featuring 25% Expansion Without Image Distortion
Stretchable displays, praised for their spatial efficiency, design flexibility, and human-like flexibility, are seen as the next generation of display technology. A team of Korean researchers has developed a stretchable display that can expand by 25% while maintaining clear image quality without distortion. It can also stretch and contract up to 5,000 times at 15% expansion without any performance degradation, making it the first deformation-free stretchable display with a negative Poisson's ratio* developed in Korea.
*Poisson’s ratio of -1: A ratio where both width and length stretch equally, expressed as a negative value. A positive Poisson's ratio represents the ratio where horizontal stretching leads to vertical contraction, which is the case for most materials.
KAIST (represented by President Kwang-Hyung Lee) announced on the 20th of August that a research team led by Professor Byeong-Soo Bae of the Department of Materials Science and Engineering (Director of the Wearable Platform Materials Technology Center) , in collaboration with the Korea Institute of Machinery & Materials (President Seoghyeon Ryu), successfully developed a stretchable display substrate that suppresses image distortion through omnidirectional stretchability.
Currently, most stretchable displays are made with highly elastic elastomer* materials, but these materials possess a positive Poisson's ratio, causing unavoidable image distortion when the display is stretched.
*Elastomer: A polymer with elasticity similar to rubber.
To address this, the introduction of auxetic* meta-structures has been gaining attention. Unlike conventional materials, auxetic structures have a unique 'negative Poisson's ratio,' expanding in all directions when stretched in just one direction. However, traditional auxetic structures contain many empty spaces, limiting their stability and usability in display substrates.
*Auxetic structure: A special geometric structure that exhibits a negative Poisson's ratio.
To tackle the issue of image distortion, Professor Bae's research team developed a method to create a seamless surface for the auxetic meta-structure, achieving the ideal negative Poisson's ratio of -1 and overcoming the biggest challenge in auxetic meta-structures.
To overcome the second issue of elastic modulus*, the team inserted a textile made of glass fiber bundles with a diameter of just 25 micrometers (a quarter of the thickness of human hair) into the elastomer material. They then filled the empty spaces with the same elastomer, creating a flat and stable integrated film without gaps.
*Elastic Modulus: The ratio that indicates the extent of deformation when force is applied to a material. A higher elastic modulus means that the material is less likely to deform under force.
The research team theoretically identified that the difference in elasticity between the auxetic structure and the elastomer material directly influences the negative Poisson's ratio and successfully achieved an elasticity difference of over 230,000 times, producing a film with a Poisson's ratio of -1, the theoretical limit.
< Figure 2. Deformation of S-AUX film. a) Configurations and visualized principal strain distribution of the optimized S-AUX film at various strain rates. b) Biaxial stretching image. While pristine elastomer shrinks in the directions that were not stretched, S-AUX film developed in this study expands in all directions simultaneously while maintaining its original shape. >
Professor Byeong-Soo Bae, who led the study, explained, "Preventing image distortion using auxetic structures in stretchable displays is a core technology, but it has faced challenges due to the many empty spaces in the surface, making it difficult to use as a substrate. This research outcome is expected to significantly accelerate commercialization through high-resolution, distortion-free stretchable display applications that utilize the entire surface."
This study, co-authored by Dr. Yung Lee from KAIST’s Department of Materials Science and Engineering and Dr. Bongkyun Jang from the Korea Institute of Machinery & Materials, was published on August 20th in the international journal Nature Communications under the title "A seamless auxetic substrate with a negative Poisson's ratio of –1".
The research was supported by the Wearable Platform Materials Technology Center at KAIST, the Korea Institute of Machinery & Materials, and LG Display.
< Figure 3. Structural configuration of the distortion-free display components on the S-AUX film and a contour image of a micro-LED chip transferred onto the S-AUX film. >
< Figure 4. Schematic illustrations and photographic images of the S-AUX film-based image: distortion-free display in its stretched state and released state. >
2024.09.20 View 5946 -
 KAIST finds ways for Bacteria to produce PET-like materials
Among various eco-friendly polymers, polyhydroxyalkanoates (PHA) stand out for their excellent biodegradability and biocompatibility. They decompose naturally in soil and marine environments and are used in applications such as food packaging and medical products. However, natural PHA produced to date has faced challenges meeting various physical property requirements, such as durability and thermal stability, and has been limited in its commercial application due to low production concentrations. In light of this, KAIST researchers have recently developed a technology that could play a crucial role in solving the environmental pollution problem caused by plastics.
KAIST (represented by President Kwang-Hyung Lee) announced on August 26th that a research team led by Distinguished Professor Sang Yup Lee from the Department of Chemical and Biomolecular Engineering, including Dr. Youngjoon Lee and master's student Minju Kang, has successfully developed a microbial strain that efficiently produces aromatic polyester* using systems metabolic engineering.
※ Aromatic polyester: A polymer containing aromatic compounds (specific carbon ring structures like benzene) and ester bonds.
In this study, the research team used metabolic engineering to enhance the metabolic flux of the biosynthetic pathway for the aromatic monomer phenyllactate (PhLA) in E. coli. They manipulated the metabolic pathway to increase the polymer fraction accumulated within the cells and employed computer simulations to predict the structure of PHA synthase and improve the enzyme based on the structure-function relationship.
Through subsequent fermentation optimization, the team achieved the world’s highest concentration (12.3±0.1 g/L) for the efficient production of poly (PhLA) and successfully produced polyester through a 30L scale fed-batch fermentation, demonstrating the possibility of industrial-level production. The produced aromatic polyesters showed enhanced thermal properties, improved mechanical properties, and potential for use as drug delivery carriers.
< Figure 1. Development schematics of aromatic polyester producing microorganisms >
The research team also demonstrated that an exogenous phasin protein* plays a crucial role in increasing the intracellular polymer accumulation fraction, which is directly related to the economic feasibility and efficiency of non-natural PHA production. They improved PHA synthase using a rational enzyme design approach, predicting the three-dimensional structure of the enzyme through homology modeling (a method of predicting the three-dimensional structure of a new protein based on the structure of similar proteins) followed by molecular docking simulations (simulations that predict how well a monomer can bind to an enzyme) and molecular dynamics simulations (simulations that predict how molecules move and interact over time) to upgrade the enzyme into a mutant enzyme with enhanced monomer polymerization efficiency.
※ Exogenous phasin protein: Phasin is a protein related to PHA production, interacting with the cytoplasmic environment on the surface of granules of PHA, and playing a role in polymer accumulation and controlling the number and size of granules. In this study, genes encoding phasin proteins derived from various natural PHA-producing microorganisms were selected and introduced.
Dr. Youngjoon Lee, co-first author of the paper, explained, "The significance of this study lies in the fact that we have achieved the world's highest concentration of microbial-based aromatic polyester production using eco-friendly materials and methods. This technology is expected to play a crucial role in addressing the environmental pollution caused by plastics." Distinguished Professor Sang Yup Lee added, "This study, which presents various strategies for the high-efficiency production of useful polymers via systems metabolic engineering, is expected to make a significant contribution to solving climate change issues, particularly the recent plastic problem."
< Figure 2. Detailed development strategy for aromatic polyester producing microorganisms >
The research findings were published on August 21st in Trends in Biotechnology, published by Cell, an international academic journal.
※ Paper Title: “Microbial production of an aromatic homopolyester”
※ Author Information: Youngjoon Lee (KAIST, co-first author), Minju Kang (KAIST, co-first author), Woo Dae Jang (KAIST, second author), So Young Choi (KAIST, third author), Jung Eun Yang (KAIST, fourth author), Sang Yup Lee (KAIST, corresponding author), totaling six authors.
This research was supported by the "Development of Next-Generation Biorefinery Platform Technologies for Leading the Bio-based Chemicals Industry" project led by Distinguished Professor Sang Yup Lee at KAIST, under the eco-friendly chemical technology development project aimed at substituting petroleum, funded by the Ministry of Science and ICT. It was also supported by the "Development of Platform Technology for the Production of Novel Aromatic Bioplastic Using Microbial Cell Factories" project (Project Leader: Si Jae Park, Ewha Woman’s University).
2024.08.28 View 6805
KAIST finds ways for Bacteria to produce PET-like materials
Among various eco-friendly polymers, polyhydroxyalkanoates (PHA) stand out for their excellent biodegradability and biocompatibility. They decompose naturally in soil and marine environments and are used in applications such as food packaging and medical products. However, natural PHA produced to date has faced challenges meeting various physical property requirements, such as durability and thermal stability, and has been limited in its commercial application due to low production concentrations. In light of this, KAIST researchers have recently developed a technology that could play a crucial role in solving the environmental pollution problem caused by plastics.
KAIST (represented by President Kwang-Hyung Lee) announced on August 26th that a research team led by Distinguished Professor Sang Yup Lee from the Department of Chemical and Biomolecular Engineering, including Dr. Youngjoon Lee and master's student Minju Kang, has successfully developed a microbial strain that efficiently produces aromatic polyester* using systems metabolic engineering.
※ Aromatic polyester: A polymer containing aromatic compounds (specific carbon ring structures like benzene) and ester bonds.
In this study, the research team used metabolic engineering to enhance the metabolic flux of the biosynthetic pathway for the aromatic monomer phenyllactate (PhLA) in E. coli. They manipulated the metabolic pathway to increase the polymer fraction accumulated within the cells and employed computer simulations to predict the structure of PHA synthase and improve the enzyme based on the structure-function relationship.
Through subsequent fermentation optimization, the team achieved the world’s highest concentration (12.3±0.1 g/L) for the efficient production of poly (PhLA) and successfully produced polyester through a 30L scale fed-batch fermentation, demonstrating the possibility of industrial-level production. The produced aromatic polyesters showed enhanced thermal properties, improved mechanical properties, and potential for use as drug delivery carriers.
< Figure 1. Development schematics of aromatic polyester producing microorganisms >
The research team also demonstrated that an exogenous phasin protein* plays a crucial role in increasing the intracellular polymer accumulation fraction, which is directly related to the economic feasibility and efficiency of non-natural PHA production. They improved PHA synthase using a rational enzyme design approach, predicting the three-dimensional structure of the enzyme through homology modeling (a method of predicting the three-dimensional structure of a new protein based on the structure of similar proteins) followed by molecular docking simulations (simulations that predict how well a monomer can bind to an enzyme) and molecular dynamics simulations (simulations that predict how molecules move and interact over time) to upgrade the enzyme into a mutant enzyme with enhanced monomer polymerization efficiency.
※ Exogenous phasin protein: Phasin is a protein related to PHA production, interacting with the cytoplasmic environment on the surface of granules of PHA, and playing a role in polymer accumulation and controlling the number and size of granules. In this study, genes encoding phasin proteins derived from various natural PHA-producing microorganisms were selected and introduced.
Dr. Youngjoon Lee, co-first author of the paper, explained, "The significance of this study lies in the fact that we have achieved the world's highest concentration of microbial-based aromatic polyester production using eco-friendly materials and methods. This technology is expected to play a crucial role in addressing the environmental pollution caused by plastics." Distinguished Professor Sang Yup Lee added, "This study, which presents various strategies for the high-efficiency production of useful polymers via systems metabolic engineering, is expected to make a significant contribution to solving climate change issues, particularly the recent plastic problem."
< Figure 2. Detailed development strategy for aromatic polyester producing microorganisms >
The research findings were published on August 21st in Trends in Biotechnology, published by Cell, an international academic journal.
※ Paper Title: “Microbial production of an aromatic homopolyester”
※ Author Information: Youngjoon Lee (KAIST, co-first author), Minju Kang (KAIST, co-first author), Woo Dae Jang (KAIST, second author), So Young Choi (KAIST, third author), Jung Eun Yang (KAIST, fourth author), Sang Yup Lee (KAIST, corresponding author), totaling six authors.
This research was supported by the "Development of Next-Generation Biorefinery Platform Technologies for Leading the Bio-based Chemicals Industry" project led by Distinguished Professor Sang Yup Lee at KAIST, under the eco-friendly chemical technology development project aimed at substituting petroleum, funded by the Ministry of Science and ICT. It was also supported by the "Development of Platform Technology for the Production of Novel Aromatic Bioplastic Using Microbial Cell Factories" project (Project Leader: Si Jae Park, Ewha Woman’s University).
2024.08.28 View 6805 -
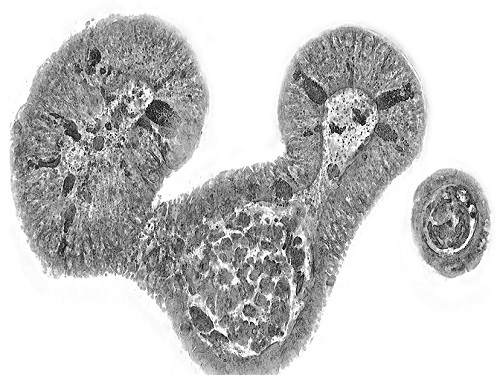 KAIST presents strategies for Holotomography in advanced bio research
Measuring and analyzing three-dimensional (3D) images of live cells and tissues is considered crucial in advanced fields of biology and medicine. Organoids, which are 3D structures that mimic organs, are particular examples that significantly benefits 3D live imaging. Organoids provide effective alternatives to animal testing in the drug development processes, and can rapidly determine personalized medicine. On the other hand, active researches are ongoing to utilize organoids for organ replacement.
< Figure 1. Schematic illustration of holotomography compared to X-ray CT. Similar to CT, they share the commonality of measuring the optical properties of an unlabeled specimen in three dimensions. Instead of X-rays, holotomography irradiates light in the visible range, and provides refractive index measurements of transparent specimens rather than absorptivity. While CT obtains three-dimensional information only through mechanical rotation of the irradiating light, holotomography can replace this by applying wavefront control technology in the visible range. >
Organelle-level observation of 3D biological specimens such as organoids and stem cell colonies without staining or preprocessing holds significant implications for both innovating basic research and bioindustrial applications related to regenerative medicine and bioindustrial applications.
Holotomography (HT) is a 3D optical microscopy that implements 3D reconstruction analogous to that of X-ray computed tomography (CT). Although HT and CT share a similar theoretical background, HT facilitates high-resolution examination inside cells and tissues, instead of the human body. HT obtains 3D images of cells and tissues at the organelle level without chemical or genetic labeling, thus overcomes various challenges of existing methods in bio research and industry. Its potential is highlighted in research fields where sample physiology must not be disrupted, such as regenerative medicine, personalized medicine, and infertility treatment.
< Figure 2. Label-free 3D imaging of diverse live cells. Time-lapse image of Hep3B cells illustrating subcellular morphology changes upon H2O2 treatment, followed by cellular recovery after returning to the regular cell culture medium. >
This paper introduces the advantages and broad applicability of HT to biomedical researchers, while presenting an overview of principles and future technical challenges to optical researchers. It showcases various cases of applying HT in studies such as 3D biology, regenerative medicine, and cancer research, as well as suggesting future optical development. Also, it categorizes HT based on the light source, to describe the principles, limitations, and improvements of each category in detail. Particularly, the paper addresses strategies for deepening cell and organoid studies by introducing artificial intelligence (AI) to HT.
Due to its potential to drive advanced bioindustry, HT is attracting interest and investment from universities and corporates worldwide. The KAIST research team has been leading this international field by developing core technologies and carrying out key application researches throughout the last decade.
< Figure 3. Various types of cells and organelles that make up the imaging barrier of a living intestinal organoid can be observed using holotomography. >
This paper, co-authored by Dr. Geon Kim from KAIST Research Center for Natural Sciences, Professor Ki-Jun Yoon's team from the Department of Biological Sciences, Director Bon-Kyoung Koo's team from the Institute for Basic Science (IBS) Center for Genome Engineering, and Dr. Seongsoo Lee's team from the Korea Basic Science Institute (KBSI), was published in 'Nature Reviews Methods Primers' on the 25th of July. This research was supported by the Leader Grant and Basic Science Research Program of the National Research Foundation, the Hologram Core Technology Development Grant of the Ministry of Science and ICT, the Nano and Material Technology Development Project, and the Health and Medical R&D Project of the Ministry of Health and Welfare.
2024.07.30 View 5699
KAIST presents strategies for Holotomography in advanced bio research
Measuring and analyzing three-dimensional (3D) images of live cells and tissues is considered crucial in advanced fields of biology and medicine. Organoids, which are 3D structures that mimic organs, are particular examples that significantly benefits 3D live imaging. Organoids provide effective alternatives to animal testing in the drug development processes, and can rapidly determine personalized medicine. On the other hand, active researches are ongoing to utilize organoids for organ replacement.
< Figure 1. Schematic illustration of holotomography compared to X-ray CT. Similar to CT, they share the commonality of measuring the optical properties of an unlabeled specimen in three dimensions. Instead of X-rays, holotomography irradiates light in the visible range, and provides refractive index measurements of transparent specimens rather than absorptivity. While CT obtains three-dimensional information only through mechanical rotation of the irradiating light, holotomography can replace this by applying wavefront control technology in the visible range. >
Organelle-level observation of 3D biological specimens such as organoids and stem cell colonies without staining or preprocessing holds significant implications for both innovating basic research and bioindustrial applications related to regenerative medicine and bioindustrial applications.
Holotomography (HT) is a 3D optical microscopy that implements 3D reconstruction analogous to that of X-ray computed tomography (CT). Although HT and CT share a similar theoretical background, HT facilitates high-resolution examination inside cells and tissues, instead of the human body. HT obtains 3D images of cells and tissues at the organelle level without chemical or genetic labeling, thus overcomes various challenges of existing methods in bio research and industry. Its potential is highlighted in research fields where sample physiology must not be disrupted, such as regenerative medicine, personalized medicine, and infertility treatment.
< Figure 2. Label-free 3D imaging of diverse live cells. Time-lapse image of Hep3B cells illustrating subcellular morphology changes upon H2O2 treatment, followed by cellular recovery after returning to the regular cell culture medium. >
This paper introduces the advantages and broad applicability of HT to biomedical researchers, while presenting an overview of principles and future technical challenges to optical researchers. It showcases various cases of applying HT in studies such as 3D biology, regenerative medicine, and cancer research, as well as suggesting future optical development. Also, it categorizes HT based on the light source, to describe the principles, limitations, and improvements of each category in detail. Particularly, the paper addresses strategies for deepening cell and organoid studies by introducing artificial intelligence (AI) to HT.
Due to its potential to drive advanced bioindustry, HT is attracting interest and investment from universities and corporates worldwide. The KAIST research team has been leading this international field by developing core technologies and carrying out key application researches throughout the last decade.
< Figure 3. Various types of cells and organelles that make up the imaging barrier of a living intestinal organoid can be observed using holotomography. >
This paper, co-authored by Dr. Geon Kim from KAIST Research Center for Natural Sciences, Professor Ki-Jun Yoon's team from the Department of Biological Sciences, Director Bon-Kyoung Koo's team from the Institute for Basic Science (IBS) Center for Genome Engineering, and Dr. Seongsoo Lee's team from the Korea Basic Science Institute (KBSI), was published in 'Nature Reviews Methods Primers' on the 25th of July. This research was supported by the Leader Grant and Basic Science Research Program of the National Research Foundation, the Hologram Core Technology Development Grant of the Ministry of Science and ICT, the Nano and Material Technology Development Project, and the Health and Medical R&D Project of the Ministry of Health and Welfare.
2024.07.30 View 5699 -
 Unraveling Mitochondrial DNA Mutations in Human Cells
Throughout our lifetime, cells accumulate DNA mutations, which contribute to genetic diversity, or “mosaicism”, among cells. These genomic mutations are pivotal for the aging process and the onset of various diseases, including cancer. Mitochondria, essential cellular organelles involved in energy metabolism and apoptosis, possess their own DNA, which are susceptible to mutations. However, studies on mtDNA mutations and mosaicism have been limited due to a variety of technical challenges.
Genomic scientists from KAIST have revealed the genetic mosaicism characterized by variations in mitochondrial DNA (mtDNA) across normal human cells. This study provides fundamental insights into understanding human aging and disease onset mechanisms.
The study, “Mitochondrial DNA mosaicism in normal human somatic cells,” was published in Nature Genetics on July 22. It was led by graduate student Jisong An under the supervision of Professor Young Seok Ju from the Graduate School of Medical Science and Engineering.
Researchers from Seoul National University College of Medicine, Yonsei University College of Medicine, Korea University College of Medicine, Washington University School of Medicine National Cancer Center, Seoul National University Hospital, Gangnam Severance Hospital and KAIST faculty startup company Inocras Inc. also participated in this study.
< Figure 1. a. Flow of experiment. b. Schematic diagram illustrating the origin and dynamics of mtDNA alterations across a lifetime. >
The study involved a bioinformatic analysis of whole-genome sequences from 2,096 single cells obtained from normal human colorectal epithelial tissue, fibroblasts, and blood collected from 31 individuals. The study highlights an average of three significant mtDNA differences between cells, with approximately 6% of these variations confirmed to be inherited as heteroplasmy from the mother.
Moreover, mutations significantly increased during tumorigenesis, with some mutations contributing to instability in mitochondrial RNA. Based on these findings, the study illustrates a computational model that comprehensively elucidates the evolution of mitochondria from embryonic development to aging and tumorigenesis.
This study systematically reveals the mechanisms behind mitochondrial DNA mosaicism in normal human cells, establishing a crucial foundation for understanding the impact of mtDNA on aging and disease onset.
Professor Ju remarked, “By systematically utilizing whole-genome big data, we can illuminate previously unknown phenomena in life sciences.” He emphasized the significance of the study, adding, “For the first time, we have established a method to systematically understand mitochondrial DNA changes occurring during human embryonic development, aging, and cancer development.”
This work was supported by the National Research Foundation of Korea and the Suh Kyungbae Foundation.
2024.07.24 View 5470
Unraveling Mitochondrial DNA Mutations in Human Cells
Throughout our lifetime, cells accumulate DNA mutations, which contribute to genetic diversity, or “mosaicism”, among cells. These genomic mutations are pivotal for the aging process and the onset of various diseases, including cancer. Mitochondria, essential cellular organelles involved in energy metabolism and apoptosis, possess their own DNA, which are susceptible to mutations. However, studies on mtDNA mutations and mosaicism have been limited due to a variety of technical challenges.
Genomic scientists from KAIST have revealed the genetic mosaicism characterized by variations in mitochondrial DNA (mtDNA) across normal human cells. This study provides fundamental insights into understanding human aging and disease onset mechanisms.
The study, “Mitochondrial DNA mosaicism in normal human somatic cells,” was published in Nature Genetics on July 22. It was led by graduate student Jisong An under the supervision of Professor Young Seok Ju from the Graduate School of Medical Science and Engineering.
Researchers from Seoul National University College of Medicine, Yonsei University College of Medicine, Korea University College of Medicine, Washington University School of Medicine National Cancer Center, Seoul National University Hospital, Gangnam Severance Hospital and KAIST faculty startup company Inocras Inc. also participated in this study.
< Figure 1. a. Flow of experiment. b. Schematic diagram illustrating the origin and dynamics of mtDNA alterations across a lifetime. >
The study involved a bioinformatic analysis of whole-genome sequences from 2,096 single cells obtained from normal human colorectal epithelial tissue, fibroblasts, and blood collected from 31 individuals. The study highlights an average of three significant mtDNA differences between cells, with approximately 6% of these variations confirmed to be inherited as heteroplasmy from the mother.
Moreover, mutations significantly increased during tumorigenesis, with some mutations contributing to instability in mitochondrial RNA. Based on these findings, the study illustrates a computational model that comprehensively elucidates the evolution of mitochondria from embryonic development to aging and tumorigenesis.
This study systematically reveals the mechanisms behind mitochondrial DNA mosaicism in normal human cells, establishing a crucial foundation for understanding the impact of mtDNA on aging and disease onset.
Professor Ju remarked, “By systematically utilizing whole-genome big data, we can illuminate previously unknown phenomena in life sciences.” He emphasized the significance of the study, adding, “For the first time, we have established a method to systematically understand mitochondrial DNA changes occurring during human embryonic development, aging, and cancer development.”
This work was supported by the National Research Foundation of Korea and the Suh Kyungbae Foundation.
2024.07.24 View 5470 -
 KAIST Develops Microbial Liquid Egg Substitute
A team of researchers published a paper on developing a substitute for eggs using microorganisms, grabbing international attention. It is expected that the development of egg substitutes using non-animal raw materials will solve the problems of factory farming, which causes problems like increased emission of greenhouse gas and waste, and contribute to building a sustainable food system that allows easy protein intake.
KAIST (President Kwang-Hyung Lee) announced that Research Professor Kyeong Rok Choi from the Biological Process Research Center and Distinguished Professor Sang Yup Lee from the Department of Chemical and Biomolecular Engineering have published a paper on the development of an "Eco-Friendly Liquid Egg Substitute Derived from Microorganisms."
Eggs play a crucial role in various culinary applications due to their unique physicochemical properties such as gelling, foaming, and emulsifying, while also providing essential nutrients. However, traditional egg production is not only unethical and resource-intensive but also has significant environmental impacts such as greenhouse gas emissions and waste issues. Additionally, factors such as wars and trade regulations have led to significant increases in egg prices, highlighting food security concerns. In response to these issues, there has been growing interest in egg substitutes made from non-animal sources to establish a sustainable food system.
Although there has been progress in developing non-animal protein-based egg substitutes, no substitute has been able to fully replicate the essential functional properties of liquid eggs, such as gelling and foaming, while also providing complete nutrition. In this context, the research team aimed to develop a liquid egg substitute using microbial biomass, which has a protein content comparable to that of meat per unit dry mass. Various microorganisms, such as yeast, Bacillus, lactic acid bacteria, and other probiotics, have been proven safe through long-term human consumption. Microbial biomass requires fewer resources like water and land during production, and possesses high-quality nutrients, making it a promising sustainable food resource.
< Figure 1. Comparison of heat treatment results of microbial pellets and microbial lysates >
However, the semi-solid microbial biomass recovered through microbial cultivation was observed to turn liquid upon heating, unlike liquid egg. To address this, the research team devised a microbial lysate by breaking down the cell walls and cell membranes of microorganisms, which correspond to the eggshell. They found that the microbial lysate's proteins coagulated when heated and formed a gel similar to that of liquid egg. The gel formed from the heated microbial lysate was found to have microscopic structures and physical properties similar to those of boiled eggs. The addition of microbial-derived edible enzymes or plant-based materials allowed for the adjustment of its properties, enabling the creation of various textures.
Furthermore, the researchers demonstrated that the microbial lysate could form stable foams widely used in baking, such as meringues (made from egg whites). They successfully baked meringue cookies using this lysate, showing its potential as a functional liquid egg substitute.
Distinguished Professor Sang Yup Lee stated, "This substitute has excellent nutritional components, making it suitable for regular food consumption. It is especially promising as emergency food for long-term space travel, wartime situations, and other emergencies. More importantly, it contributes to securing a sustainable food system."
< Figure 2. Example of foaming ability of microbial lysate and meringue cookie production >
< Figure 3. Example of foaming ability of microbial lysate and meringue cookie production >
The paper was published online in the journal npj Science of Food, issued by Nature.
- Paper Title: Microbial lysates repurposed as liquid egg substitutes
- Authors: Kyeong Rok Choi (first author), Da-Hee Ahn, Seok Yeong Jung, YuHyun Lee, and Sang Yup Lee (corresponding author)
This research was supported by the Ministry of Science and ICT's project for developing eco-friendly chemical technologies to replace petroleum (Project Leader: Distinguished Professor Sang Yup Lee, KAIST) and the Rural Development Administration's Agricultural Microorganisms Project Group (Director: Professor Pan-sik Jang, Seoul National University) for developing protein production technology from inorganic substances through microbial metabolic system control (Project Leader: Research Professor Kyeong Rok Choi, KAIST).
2024.07.05 View 7011
KAIST Develops Microbial Liquid Egg Substitute
A team of researchers published a paper on developing a substitute for eggs using microorganisms, grabbing international attention. It is expected that the development of egg substitutes using non-animal raw materials will solve the problems of factory farming, which causes problems like increased emission of greenhouse gas and waste, and contribute to building a sustainable food system that allows easy protein intake.
KAIST (President Kwang-Hyung Lee) announced that Research Professor Kyeong Rok Choi from the Biological Process Research Center and Distinguished Professor Sang Yup Lee from the Department of Chemical and Biomolecular Engineering have published a paper on the development of an "Eco-Friendly Liquid Egg Substitute Derived from Microorganisms."
Eggs play a crucial role in various culinary applications due to their unique physicochemical properties such as gelling, foaming, and emulsifying, while also providing essential nutrients. However, traditional egg production is not only unethical and resource-intensive but also has significant environmental impacts such as greenhouse gas emissions and waste issues. Additionally, factors such as wars and trade regulations have led to significant increases in egg prices, highlighting food security concerns. In response to these issues, there has been growing interest in egg substitutes made from non-animal sources to establish a sustainable food system.
Although there has been progress in developing non-animal protein-based egg substitutes, no substitute has been able to fully replicate the essential functional properties of liquid eggs, such as gelling and foaming, while also providing complete nutrition. In this context, the research team aimed to develop a liquid egg substitute using microbial biomass, which has a protein content comparable to that of meat per unit dry mass. Various microorganisms, such as yeast, Bacillus, lactic acid bacteria, and other probiotics, have been proven safe through long-term human consumption. Microbial biomass requires fewer resources like water and land during production, and possesses high-quality nutrients, making it a promising sustainable food resource.
< Figure 1. Comparison of heat treatment results of microbial pellets and microbial lysates >
However, the semi-solid microbial biomass recovered through microbial cultivation was observed to turn liquid upon heating, unlike liquid egg. To address this, the research team devised a microbial lysate by breaking down the cell walls and cell membranes of microorganisms, which correspond to the eggshell. They found that the microbial lysate's proteins coagulated when heated and formed a gel similar to that of liquid egg. The gel formed from the heated microbial lysate was found to have microscopic structures and physical properties similar to those of boiled eggs. The addition of microbial-derived edible enzymes or plant-based materials allowed for the adjustment of its properties, enabling the creation of various textures.
Furthermore, the researchers demonstrated that the microbial lysate could form stable foams widely used in baking, such as meringues (made from egg whites). They successfully baked meringue cookies using this lysate, showing its potential as a functional liquid egg substitute.
Distinguished Professor Sang Yup Lee stated, "This substitute has excellent nutritional components, making it suitable for regular food consumption. It is especially promising as emergency food for long-term space travel, wartime situations, and other emergencies. More importantly, it contributes to securing a sustainable food system."
< Figure 2. Example of foaming ability of microbial lysate and meringue cookie production >
< Figure 3. Example of foaming ability of microbial lysate and meringue cookie production >
The paper was published online in the journal npj Science of Food, issued by Nature.
- Paper Title: Microbial lysates repurposed as liquid egg substitutes
- Authors: Kyeong Rok Choi (first author), Da-Hee Ahn, Seok Yeong Jung, YuHyun Lee, and Sang Yup Lee (corresponding author)
This research was supported by the Ministry of Science and ICT's project for developing eco-friendly chemical technologies to replace petroleum (Project Leader: Distinguished Professor Sang Yup Lee, KAIST) and the Rural Development Administration's Agricultural Microorganisms Project Group (Director: Professor Pan-sik Jang, Seoul National University) for developing protein production technology from inorganic substances through microbial metabolic system control (Project Leader: Research Professor Kyeong Rok Choi, KAIST).
2024.07.05 View 7011 -
 KAIST Employs Image-recognition AI to Determine Battery Composition and Conditions
An international collaborative research team has developed an image recognition technology that can accurately determine the elemental composition and the number of charge and discharge cycles of a battery by examining only its surface morphology using AI learning.
KAIST (President Kwang-Hyung Lee) announced on July 2nd that Professor Seungbum Hong from the Department of Materials Science and Engineering, in collaboration with the Electronics and Telecommunications Research Institute (ETRI) and Drexel University in the United States, has developed a method to predict the major elemental composition and charge-discharge state of NCM cathode materials with 99.6% accuracy using convolutional neural networks (CNN)*.
*Convolutional Neural Network (CNN): A type of multi-layer, feed-forward, artificial neural network used for analyzing visual images.
The research team noted that while scanning electron microscopy (SEM) is used in semiconductor manufacturing to inspect wafer defects, it is rarely used in battery inspections. SEM is used for batteries to analyze the size of particles only at research sites, and reliability is predicted from the broken particles and the shape of the breakage in the case of deteriorated battery materials.
The research team decided that it would be groundbreaking if an automated SEM can be used in the process of battery production, just like in the semiconductor manufacturing, to inspect the surface of the cathode material to determine whether it was synthesized according to the desired composition and that the lifespan would be reliable, thereby reducing the defect rate.
< Figure 1. Example images of true cases and their grad-CAM overlays from the best trained network. >
The researchers trained a CNN-based AI applicable to autonomous vehicles to learn the surface images of battery materials, enabling it to predict the major elemental composition and charge-discharge cycle states of the cathode materials. They found that while the method could accurately predict the composition of materials with additives, it had lower accuracy for predicting charge-discharge states. The team plans to further train the AI with various battery material morphologies produced through different processes and ultimately use it for inspecting the compositional uniformity and predicting the lifespan of next-generation batteries.
Professor Joshua C. Agar, one of the collaborating researchers of the project from the Department of Mechanical Engineering and Mechanics of Drexel University, said, "In the future, artificial intelligence is expected to be applied not only to battery materials but also to various dynamic processes in functional materials synthesis, clean energy generation in fusion, and understanding foundations of particles and the universe."
Professor Seungbum Hong from KAIST, who led the research, stated, "This research is significant as it is the first in the world to develop an AI-based methodology that can quickly and accurately predict the major elemental composition and the state of the battery from the structural data of micron-scale SEM images. The methodology developed in this study for identifying the composition and state of battery materials based on microscopic images is expected to play a crucial role in improving the performance and quality of battery materials in the future."
< Figure 2. Accuracies of CNN Model predictions on SEM images of NCM cathode materials with additives under various conditions. >
This research was conducted by KAIST’s Materials Science and Engineering Department graduates Dr. Jimin Oh and Dr. Jiwon Yeom, the co-first authors, in collaboration with Professor Josh Agar and Dr. Kwang Man Kim from ETRI. It was supported by the National Research Foundation of Korea, the KAIST Global Singularity project, and international collaboration with the US research team. The results were published in the international journal npj Computational Materials on May 4. (Paper Title: “Composition and state prediction of lithium-ion cathode via convolutional neural network trained on scanning electron microscopy images”)
2024.07.02 View 6884
KAIST Employs Image-recognition AI to Determine Battery Composition and Conditions
An international collaborative research team has developed an image recognition technology that can accurately determine the elemental composition and the number of charge and discharge cycles of a battery by examining only its surface morphology using AI learning.
KAIST (President Kwang-Hyung Lee) announced on July 2nd that Professor Seungbum Hong from the Department of Materials Science and Engineering, in collaboration with the Electronics and Telecommunications Research Institute (ETRI) and Drexel University in the United States, has developed a method to predict the major elemental composition and charge-discharge state of NCM cathode materials with 99.6% accuracy using convolutional neural networks (CNN)*.
*Convolutional Neural Network (CNN): A type of multi-layer, feed-forward, artificial neural network used for analyzing visual images.
The research team noted that while scanning electron microscopy (SEM) is used in semiconductor manufacturing to inspect wafer defects, it is rarely used in battery inspections. SEM is used for batteries to analyze the size of particles only at research sites, and reliability is predicted from the broken particles and the shape of the breakage in the case of deteriorated battery materials.
The research team decided that it would be groundbreaking if an automated SEM can be used in the process of battery production, just like in the semiconductor manufacturing, to inspect the surface of the cathode material to determine whether it was synthesized according to the desired composition and that the lifespan would be reliable, thereby reducing the defect rate.
< Figure 1. Example images of true cases and their grad-CAM overlays from the best trained network. >
The researchers trained a CNN-based AI applicable to autonomous vehicles to learn the surface images of battery materials, enabling it to predict the major elemental composition and charge-discharge cycle states of the cathode materials. They found that while the method could accurately predict the composition of materials with additives, it had lower accuracy for predicting charge-discharge states. The team plans to further train the AI with various battery material morphologies produced through different processes and ultimately use it for inspecting the compositional uniformity and predicting the lifespan of next-generation batteries.
Professor Joshua C. Agar, one of the collaborating researchers of the project from the Department of Mechanical Engineering and Mechanics of Drexel University, said, "In the future, artificial intelligence is expected to be applied not only to battery materials but also to various dynamic processes in functional materials synthesis, clean energy generation in fusion, and understanding foundations of particles and the universe."
Professor Seungbum Hong from KAIST, who led the research, stated, "This research is significant as it is the first in the world to develop an AI-based methodology that can quickly and accurately predict the major elemental composition and the state of the battery from the structural data of micron-scale SEM images. The methodology developed in this study for identifying the composition and state of battery materials based on microscopic images is expected to play a crucial role in improving the performance and quality of battery materials in the future."
< Figure 2. Accuracies of CNN Model predictions on SEM images of NCM cathode materials with additives under various conditions. >
This research was conducted by KAIST’s Materials Science and Engineering Department graduates Dr. Jimin Oh and Dr. Jiwon Yeom, the co-first authors, in collaboration with Professor Josh Agar and Dr. Kwang Man Kim from ETRI. It was supported by the National Research Foundation of Korea, the KAIST Global Singularity project, and international collaboration with the US research team. The results were published in the international journal npj Computational Materials on May 4. (Paper Title: “Composition and state prediction of lithium-ion cathode via convolutional neural network trained on scanning electron microscopy images”)
2024.07.02 View 6884