Sang+Yup+Lee
-
 Natural Rainbow Colorants Microbially Produced
Integrated strategies of systems metabolic engineering and membrane engineering led to the production of natural rainbow colorants comprising seven natural colorants from bacteria for the first time
A research group at KAIST has engineered bacterial strains capable of producing three carotenoids and four violacein derivatives, completing the seven colors in the rainbow spectrum. The research team integrated systems metabolic engineering and membrane engineering strategies for the production of seven natural rainbow colorants in engineered Escherichia coli strains. The strategies will be also useful for the efficient production of other industrially important natural products used in the food, pharmaceutical, and cosmetic industries.
Colorants are widely used in our lives and are directly related to human health when we eat food additives and wear cosmetics. However, most of these colorants are made from petroleum, causing unexpected side effects and health problems. Furthermore, they raise environmental concerns such as water pollution from dyeing fabric in the textiles industry. For these reasons, the demand for the production of natural colorants using microorganisms has increased, but could not be readily realized due to the high cost and low yield of the bioprocesses.
These challenges inspired the metabolic engineers at KAIST including researchers Dr. Dongsoo Yang and Dr. Seon Young Park, and Distinguished Professor Sang Yup Lee from the Department of Chemical and Biomolecular Engineering. The team reported the study entitled “Production of rainbow colorants by metabolically engineered Escherichia coli” in Advanced Science online on May 5. It was selected as the journal cover of the July 7 issue.
This research reports for the first time the production of rainbow colorants comprising three carotenoids and four violacein derivatives from glucose or glycerol via systems metabolic engineering and membrane engineering. The research group focused on the production of hydrophobic natural colorants useful for lipophilic food and dyeing garments. First, using systems metabolic engineering, which is an integrated technology to engineer the metabolism of a microorganism, three carotenoids comprising astaxanthin (red), -carotene (orange), and zeaxanthin (yellow), and four violacein derivatives comprising proviolacein (green), prodeoxyviolacein (blue), violacein (navy), and deoxyviolacein (purple) could be produced. Thus, the production of natural colorants covering the complete rainbow spectrum was achieved.
When hydrophobic colorants are produced from microorganisms, the colorants are accumulated inside the cell. As the accumulation capacity is limited, the hydrophobic colorants could not be produced with concentrations higher than the limit. In this regard, the researchers engineered the cell morphology and generated inner-membrane vesicles (spherical membranous structures) to increase the intracellular capacity for accumulating the natural colorants. To further promote production, the researchers generated outer-membrane vesicles to secrete the natural colorants, thus succeeding in efficiently producing all of seven rainbow colorants. It was even more impressive that the production of natural green and navy colorants was achieved for the first time.
“The production of the seven natural rainbow colorants that can replace the current petroleum-based synthetic colorants was achieved for the first time,” said Dr. Dongsoo Yang. He explained that another important point of the research is that integrated metabolic engineering strategies developed from this study can be generally applicable for the efficient production of other natural products useful as pharmaceuticals or nutraceuticals. “As maintaining good health in an aging society is becoming increasingly important, we expect that the technology and strategies developed here will play pivotal roles in producing other valuable natural products of medical or nutritional importance,” explained Distinguished Professor Lee.
This work was supported by the "Cooperative Research Program for Agriculture Science & Technology Development (Project No. PJ01550602)" Rural Development Administration, Republic of Korea.
-Publication:Dongsoo Yang, Seon Young Park, and Sang Yup Lee. Production of rainbow colorants by metabolically engineered Escherichia coli. Advanced Science, 2100743.
-Profile Distinguished Professor Sang Yup LeeMetabolic &Biomolecular Engineering National Research Laboratoryhttp://mbel.kaist.ac.kr
Department of Chemical and Biomolecular EngineeringKAIST
2021.06.09 View 7072
Natural Rainbow Colorants Microbially Produced
Integrated strategies of systems metabolic engineering and membrane engineering led to the production of natural rainbow colorants comprising seven natural colorants from bacteria for the first time
A research group at KAIST has engineered bacterial strains capable of producing three carotenoids and four violacein derivatives, completing the seven colors in the rainbow spectrum. The research team integrated systems metabolic engineering and membrane engineering strategies for the production of seven natural rainbow colorants in engineered Escherichia coli strains. The strategies will be also useful for the efficient production of other industrially important natural products used in the food, pharmaceutical, and cosmetic industries.
Colorants are widely used in our lives and are directly related to human health when we eat food additives and wear cosmetics. However, most of these colorants are made from petroleum, causing unexpected side effects and health problems. Furthermore, they raise environmental concerns such as water pollution from dyeing fabric in the textiles industry. For these reasons, the demand for the production of natural colorants using microorganisms has increased, but could not be readily realized due to the high cost and low yield of the bioprocesses.
These challenges inspired the metabolic engineers at KAIST including researchers Dr. Dongsoo Yang and Dr. Seon Young Park, and Distinguished Professor Sang Yup Lee from the Department of Chemical and Biomolecular Engineering. The team reported the study entitled “Production of rainbow colorants by metabolically engineered Escherichia coli” in Advanced Science online on May 5. It was selected as the journal cover of the July 7 issue.
This research reports for the first time the production of rainbow colorants comprising three carotenoids and four violacein derivatives from glucose or glycerol via systems metabolic engineering and membrane engineering. The research group focused on the production of hydrophobic natural colorants useful for lipophilic food and dyeing garments. First, using systems metabolic engineering, which is an integrated technology to engineer the metabolism of a microorganism, three carotenoids comprising astaxanthin (red), -carotene (orange), and zeaxanthin (yellow), and four violacein derivatives comprising proviolacein (green), prodeoxyviolacein (blue), violacein (navy), and deoxyviolacein (purple) could be produced. Thus, the production of natural colorants covering the complete rainbow spectrum was achieved.
When hydrophobic colorants are produced from microorganisms, the colorants are accumulated inside the cell. As the accumulation capacity is limited, the hydrophobic colorants could not be produced with concentrations higher than the limit. In this regard, the researchers engineered the cell morphology and generated inner-membrane vesicles (spherical membranous structures) to increase the intracellular capacity for accumulating the natural colorants. To further promote production, the researchers generated outer-membrane vesicles to secrete the natural colorants, thus succeeding in efficiently producing all of seven rainbow colorants. It was even more impressive that the production of natural green and navy colorants was achieved for the first time.
“The production of the seven natural rainbow colorants that can replace the current petroleum-based synthetic colorants was achieved for the first time,” said Dr. Dongsoo Yang. He explained that another important point of the research is that integrated metabolic engineering strategies developed from this study can be generally applicable for the efficient production of other natural products useful as pharmaceuticals or nutraceuticals. “As maintaining good health in an aging society is becoming increasingly important, we expect that the technology and strategies developed here will play pivotal roles in producing other valuable natural products of medical or nutritional importance,” explained Distinguished Professor Lee.
This work was supported by the "Cooperative Research Program for Agriculture Science & Technology Development (Project No. PJ01550602)" Rural Development Administration, Republic of Korea.
-Publication:Dongsoo Yang, Seon Young Park, and Sang Yup Lee. Production of rainbow colorants by metabolically engineered Escherichia coli. Advanced Science, 2100743.
-Profile Distinguished Professor Sang Yup LeeMetabolic &Biomolecular Engineering National Research Laboratoryhttp://mbel.kaist.ac.kr
Department of Chemical and Biomolecular EngineeringKAIST
2021.06.09 View 7072 -
 Prof. Sang Yup Lee Elected as a Foreign Member of the Royal Society
Vice President for Research Distinguished Professor Sang Yup Lee was elected as a foreign member of the Royal Society in the UK. On May 6, the Society announced the list of distinguished new 52 fellows and 10 foreign members who achieved exceptional contributions to science. Professor Lee and Professor V. Narry Kim from Seoul National University are the first foreign members ever elected from Korea.
The Royal Society, established in 1660, is one of the most prestigious national science academies and a fellowship of 1,600 of the world’s most eminent scientists. From Newton to Darwin, Einstein, Hawking, and beyond, pioneers and paragons in their fields are elected by their peers. To date, there are 280 Nobel prize winners among the fellows.
Distinguished Professor Lee from the Department of Chemical and Biomolecular Engineering at KAIST is one of the Highly Cited Researchers (HCRs) who pioneered systems metabolic engineering and developed various micro-organisms for producing a wide range of fuels, chemicals, materials, and natural compounds.
His seminal scholarship and research career have already been recognized worldwide. He is the first Korean ever elected into the National Academy of Inventors (NAI) in the US and one of 13 scholars elected as an International Member of both the National Academy of Sciences (NAS) and the National Academy of Engineering (NAE) in the US. With this fellowship, he added one more accolade of being the first non-US and British Commonwealth scientist elected into the three most prestigious science academies: the NAS, the NAE, and the Royal Society.
2021.05.07 View 7357
Prof. Sang Yup Lee Elected as a Foreign Member of the Royal Society
Vice President for Research Distinguished Professor Sang Yup Lee was elected as a foreign member of the Royal Society in the UK. On May 6, the Society announced the list of distinguished new 52 fellows and 10 foreign members who achieved exceptional contributions to science. Professor Lee and Professor V. Narry Kim from Seoul National University are the first foreign members ever elected from Korea.
The Royal Society, established in 1660, is one of the most prestigious national science academies and a fellowship of 1,600 of the world’s most eminent scientists. From Newton to Darwin, Einstein, Hawking, and beyond, pioneers and paragons in their fields are elected by their peers. To date, there are 280 Nobel prize winners among the fellows.
Distinguished Professor Lee from the Department of Chemical and Biomolecular Engineering at KAIST is one of the Highly Cited Researchers (HCRs) who pioneered systems metabolic engineering and developed various micro-organisms for producing a wide range of fuels, chemicals, materials, and natural compounds.
His seminal scholarship and research career have already been recognized worldwide. He is the first Korean ever elected into the National Academy of Inventors (NAI) in the US and one of 13 scholars elected as an International Member of both the National Academy of Sciences (NAS) and the National Academy of Engineering (NAE) in the US. With this fellowship, he added one more accolade of being the first non-US and British Commonwealth scientist elected into the three most prestigious science academies: the NAS, the NAE, and the Royal Society.
2021.05.07 View 7357 -
 Distinguished Professor Sang Yup Lee Honored with Charles D. Scott Award
Vice President for Research Sang Yup Lee received the 2021 Charles D. Scott Award from the Society for Industrial Microbiology and Biotechnology. Distinguished Professor Lee from the Department of Chemical and Biomolecular Engineering at KAIST is the first Asian awardee.
The Charles D. Scott Award, initiated in 1995, recognizes individuals who have made significant contributions to enable and further the use of biotechnology to produce fuels and chemicals. The award is named in honor of Dr. Charles D. Scott, who founded the Symposium on Biomaterials, Fuels, and Chemicals and chaired the conference for its first ten years.
Professor Lee has pioneered systems metabolic engineering and developed various micro-organisms capable of producing a wide range of fuels, chemicals, materials, and natural compounds, many of them for the first time. Some of the breakthroughs include the microbial production of gasoline, diacids, diamines, PLA and PLGA polymers, and several natural products.
More recently, his team has developed a microbial strain capable of the mass production of succinic acid, a monomer for manufacturing polyester, with the highest production efficiency to date, as well as a Corynebacterium glutamicum strain capable of producing high-level glutaric acid. They also engineered for the first time a bacterium capable of producing carminic acid, a natural red colorant that is widely used for food and cosmetics.
Professor Lee is one of the Highly Cited Researchers (HCR), ranked in the top 1% by citations in their field by Clarivate Analytics for four consecutive years from 2017. He is the first Korean fellow ever elected into the National Academy of Inventors in the US and one of 13 scholars elected as an International Member of both the National Academy of Sciences and the National Academy of Engineering in the USA.
The awards ceremony will take place during the Symposium on Biomaterials, Fuels, and Chemicals held online from April 26.
2021.04.27 View 6401
Distinguished Professor Sang Yup Lee Honored with Charles D. Scott Award
Vice President for Research Sang Yup Lee received the 2021 Charles D. Scott Award from the Society for Industrial Microbiology and Biotechnology. Distinguished Professor Lee from the Department of Chemical and Biomolecular Engineering at KAIST is the first Asian awardee.
The Charles D. Scott Award, initiated in 1995, recognizes individuals who have made significant contributions to enable and further the use of biotechnology to produce fuels and chemicals. The award is named in honor of Dr. Charles D. Scott, who founded the Symposium on Biomaterials, Fuels, and Chemicals and chaired the conference for its first ten years.
Professor Lee has pioneered systems metabolic engineering and developed various micro-organisms capable of producing a wide range of fuels, chemicals, materials, and natural compounds, many of them for the first time. Some of the breakthroughs include the microbial production of gasoline, diacids, diamines, PLA and PLGA polymers, and several natural products.
More recently, his team has developed a microbial strain capable of the mass production of succinic acid, a monomer for manufacturing polyester, with the highest production efficiency to date, as well as a Corynebacterium glutamicum strain capable of producing high-level glutaric acid. They also engineered for the first time a bacterium capable of producing carminic acid, a natural red colorant that is widely used for food and cosmetics.
Professor Lee is one of the Highly Cited Researchers (HCR), ranked in the top 1% by citations in their field by Clarivate Analytics for four consecutive years from 2017. He is the first Korean fellow ever elected into the National Academy of Inventors in the US and one of 13 scholars elected as an International Member of both the National Academy of Sciences and the National Academy of Engineering in the USA.
The awards ceremony will take place during the Symposium on Biomaterials, Fuels, and Chemicals held online from April 26.
2021.04.27 View 6401 -
 Microbial Production of a Natural Red Colorant Carminic Acid
Metabolic engineering and computer-simulated enzyme engineering led to the production of carminic acid, a natural red colorant, from bacteria for the first time
A research group at KAIST has engineered a bacterium capable of producing a natural red colorant, carminic acid, which is widely used for food and cosmetics. The research team reported the complete biosynthesis of carminic acid from glucose in engineered Escherichia coli. The strategies will be useful for the design and construction of biosynthetic pathways involving unknown enzymes and consequently the production of diverse industrially important natural products for the food, pharmaceutical, and cosmetic industries.
Carminic acid is a natural red colorant widely being used for products such as strawberry milk and lipstick. However, carminic acid has been produced by farming cochineals, a scale insect which only grows in the region around Peru and Canary Islands, followed by complicated multi-step purification processes. Moreover, carminic acid often contains protein contaminants that cause allergies so many people are unwilling to consume products made of insect-driven colorants. On that account, manufacturers around the world are using alternative red colorants despite the fact that carminic acid is one of the most stable natural red colorants.
These challenges inspired the metabolic engineering research group at KAIST to address this issue. Its members include postdoctoral researchers Dongsoo Yang and Woo Dae Jang, and Distinguished Professor Sang Yup Lee of the Department of Chemical and Biomolecular Engineering. This study entitled “Production of carminic acid by metabolically engineered Escherichia coli” was published online in the Journal of the American Chemical Society (JACS) on April 2.
This research reports for the first time the development of a bacterial strain capable of producing carminic acid from glucose via metabolic engineering and computer simulation-assisted enzyme engineering. The research group optimized the type II polyketide synthase machinery to efficiently produce the precursor of carminic acid, flavokermesic acid.
Since the enzymes responsible for the remaining two reactions were neither discovered nor functional, biochemical reaction analysis was performed to identify enzymes that can convert flavokermesic acid into carminic acid. Then, homology modeling and docking simulations were performed to enhance the activities of the two identified enzymes. The team could confirm that the final engineered strain could produce carminic acid directly from glucose. The C-glucosyltransferase developed in this study was found to be generally applicable for other natural products as showcased by the successful production of an additional product, aloesin, which is found in aloe leaves.
“The most important part of this research is that unknown enzymes for the production of target natural products were identified and improved by biochemical reaction analyses and computer simulation-assisted enzyme engineering,” says Dr. Dongsoo Yang. He explained the development of a generally applicable C-glucosyltransferase is also useful since C-glucosylation is a relatively unexplored reaction in bacteria including Escherichia coli. Using the C-glucosyltransferase developed in this study, both carminic acid and aloesin were successfully produced from glucose.
“A sustainable and insect-free method of producing carminic acid was achieved for the first time in this study. Unknown or inefficient enzymes have always been a major problem in natural product biosynthesis, and here we suggest one effective solution for solving this problem. As maintaining good health in the aging society is becoming increasingly important, we expect that the technology and strategies developed here will play pivotal roles in producing other valuable natural products of medical or nutritional importance,” said Distinguished Professor Sang Yup Lee.
This work was supported by the Technology Development Program to Solve Climate Changes on Systems Metabolic Engineering for Biorefineries of the Ministry of Science and ICT (MSIT) through the National Research Foundation (NRF) of Korea and the KAIST Cross-Generation Collaborative Lab project; Sang Yup Lee and Dongsoo Yang were also supported by Novo Nordisk Foundation in Denmark.
Publication:
Dongsoo Yang, Woo Dae Jang, and Sang Yup Lee. Production of carminic acid by metabolically engineered Escherichia coli. at the Journal of the American Chemical Society. https://doi.org.10.1021/jacs.0c12406
Profile:
Sang Yup Lee, PhD
Distinguished Professor
leesy@kaist.ac.kr
http://mbel.kaist.ac.kr
Metabolic &Biomolecular Engineering National Research Laboratory
Department of Chemical and Biomolecular Engineering
KAIST
2021.04.06 View 8425
Microbial Production of a Natural Red Colorant Carminic Acid
Metabolic engineering and computer-simulated enzyme engineering led to the production of carminic acid, a natural red colorant, from bacteria for the first time
A research group at KAIST has engineered a bacterium capable of producing a natural red colorant, carminic acid, which is widely used for food and cosmetics. The research team reported the complete biosynthesis of carminic acid from glucose in engineered Escherichia coli. The strategies will be useful for the design and construction of biosynthetic pathways involving unknown enzymes and consequently the production of diverse industrially important natural products for the food, pharmaceutical, and cosmetic industries.
Carminic acid is a natural red colorant widely being used for products such as strawberry milk and lipstick. However, carminic acid has been produced by farming cochineals, a scale insect which only grows in the region around Peru and Canary Islands, followed by complicated multi-step purification processes. Moreover, carminic acid often contains protein contaminants that cause allergies so many people are unwilling to consume products made of insect-driven colorants. On that account, manufacturers around the world are using alternative red colorants despite the fact that carminic acid is one of the most stable natural red colorants.
These challenges inspired the metabolic engineering research group at KAIST to address this issue. Its members include postdoctoral researchers Dongsoo Yang and Woo Dae Jang, and Distinguished Professor Sang Yup Lee of the Department of Chemical and Biomolecular Engineering. This study entitled “Production of carminic acid by metabolically engineered Escherichia coli” was published online in the Journal of the American Chemical Society (JACS) on April 2.
This research reports for the first time the development of a bacterial strain capable of producing carminic acid from glucose via metabolic engineering and computer simulation-assisted enzyme engineering. The research group optimized the type II polyketide synthase machinery to efficiently produce the precursor of carminic acid, flavokermesic acid.
Since the enzymes responsible for the remaining two reactions were neither discovered nor functional, biochemical reaction analysis was performed to identify enzymes that can convert flavokermesic acid into carminic acid. Then, homology modeling and docking simulations were performed to enhance the activities of the two identified enzymes. The team could confirm that the final engineered strain could produce carminic acid directly from glucose. The C-glucosyltransferase developed in this study was found to be generally applicable for other natural products as showcased by the successful production of an additional product, aloesin, which is found in aloe leaves.
“The most important part of this research is that unknown enzymes for the production of target natural products were identified and improved by biochemical reaction analyses and computer simulation-assisted enzyme engineering,” says Dr. Dongsoo Yang. He explained the development of a generally applicable C-glucosyltransferase is also useful since C-glucosylation is a relatively unexplored reaction in bacteria including Escherichia coli. Using the C-glucosyltransferase developed in this study, both carminic acid and aloesin were successfully produced from glucose.
“A sustainable and insect-free method of producing carminic acid was achieved for the first time in this study. Unknown or inefficient enzymes have always been a major problem in natural product biosynthesis, and here we suggest one effective solution for solving this problem. As maintaining good health in the aging society is becoming increasingly important, we expect that the technology and strategies developed here will play pivotal roles in producing other valuable natural products of medical or nutritional importance,” said Distinguished Professor Sang Yup Lee.
This work was supported by the Technology Development Program to Solve Climate Changes on Systems Metabolic Engineering for Biorefineries of the Ministry of Science and ICT (MSIT) through the National Research Foundation (NRF) of Korea and the KAIST Cross-Generation Collaborative Lab project; Sang Yup Lee and Dongsoo Yang were also supported by Novo Nordisk Foundation in Denmark.
Publication:
Dongsoo Yang, Woo Dae Jang, and Sang Yup Lee. Production of carminic acid by metabolically engineered Escherichia coli. at the Journal of the American Chemical Society. https://doi.org.10.1021/jacs.0c12406
Profile:
Sang Yup Lee, PhD
Distinguished Professor
leesy@kaist.ac.kr
http://mbel.kaist.ac.kr
Metabolic &Biomolecular Engineering National Research Laboratory
Department of Chemical and Biomolecular Engineering
KAIST
2021.04.06 View 8425 -
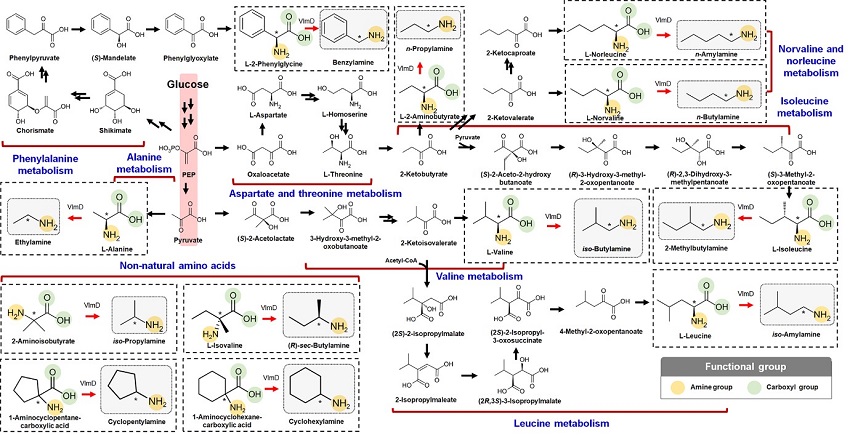 Expanding the Biosynthetic Pathway via Retrobiosynthesis
- Researchers reports a new strategy for the microbial production of multiple short-chain primary amines via retrobiosynthesis. -
KAIST metabolic engineers presented the bio-based production of multiple short-chain primary amines that have a wide range of applications in chemical industries for the first time. The research team led by Distinguished Professor Sang Yup Lee from the Department of Chemical and Biomolecular Engineering designed the novel biosynthetic pathways for short-chain primary amines by combining retrobiosynthesis and a precursor selection step.
The research team verified the newly designed pathways by confirming the in vivo production of 10 short-chain primary amines by supplying the precursors. Furthermore, the platform Escherichia coli strains were metabolically engineered to produce three proof-of-concept short-chain primary amines from glucose, demonstrating the possibility of the bio-based production of diverse short-chain primary amines from renewable resources. The research team said this study expands the strategy of systematically designing biosynthetic pathways for the production of a group of related chemicals as demonstrated by multiple short-chain primary amines as examples.
Currently, most of the industrial chemicals used in our daily lives are produced with petroleum-based products. However, there are several serious issues with the petroleum industry such as the depletion of fossil fuel reserves and environmental problems including global warming. To solve these problems, the sustainable production of industrial chemicals and materials is being explored with microorganisms as cell factories and renewable non-food biomass as raw materials for alternative to petroleum-based products. The engineering of these microorganisms has increasingly become more efficient and effective with the help of systems metabolic engineering – a practice of engineering the metabolism of a living organism toward the production of a desired metabolite. In this regard, the number of chemicals produced using biomass as a raw material has substantially increased.
Although the scope of chemicals that are producible using microorganisms continues to expand through advances in systems metabolic engineering, the biological production of short-chain primary amines has not yet been reported despite their industrial importance. Short-chain primary amines are the chemicals that have an alkyl or aryl group in the place of a hydrogen atom in ammonia with carbon chain lengths ranging from C1 to C7. Short-chain primary amines have a wide range of applications in chemical industries, for example, as a precursor for pharmaceuticals (e.g., antidiabetic and antihypertensive drugs), agrochemicals (e.g., herbicides, fungicides and insecticides), solvents, and vulcanization accelerators for rubber and plasticizers. The market size of short-chain primary amines was estimated to be more than 4 billion US dollars in 2014.
The main reason why the bio-based production of short-chain primary amines was not yet possible was due to their unknown biosynthetic pathways. Therefore, the team designed synthetic biosynthetic pathways for short-chain primary amines by combining retrobiosynthesis and a precursor selection step. The retrobiosynthesis allowed the systematic design of a biosynthetic pathway for short-chain primary amines by using a set of biochemical reaction rules that describe chemical transformation patterns between a substrate and product molecules at an atomic level.
These multiple precursors predicted for the possible biosynthesis of each short-chain primary amine were sequentially narrowed down by using the precursor selection step for efficient metabolic engineering experiments.
“Our research demonstrates the possibility of the renewable production of short-chain primary amines for the first time. We are planning to increase production efficiencies of short-chain primary amines. We believe that our study will play an important role in the development of sustainable and eco-friendly bio-based industries and the reorganization of the chemical industry, which is mandatory for solving the environmental problems threating the survival of mankind,” said Professor Lee.
This paper titled “Microbial production of multiple short-chain primary amines via retrobiosynthesis” was published in Nature Communications. This work was supported by the Technology Development Program to Solve Climate Changes on Systems Metabolic Engineering for Biorefineries from the Ministry of Science and ICT through the National Research Foundation (NRF) of Korea.
-Publication
Dong In Kim, Tong Un Chae, Hyun Uk Kim, Woo Dae Jang, and Sang Yup Lee. Microbial production of multiple short-chain primary amines via retrobiosynthesis. Nature Communications ( https://www.nature.com/articles/s41467-020-20423-6)
-Profile
Distinguished Professor Sang Yup Lee
leesy@kaist.ac.kr
Metabolic &Biomolecular Engineering National Research Laboratory
http://mbel.kaist.ac.kr
Department of Chemical and Biomolecular Engineering
KAIST
2021.01.14 View 8308
Expanding the Biosynthetic Pathway via Retrobiosynthesis
- Researchers reports a new strategy for the microbial production of multiple short-chain primary amines via retrobiosynthesis. -
KAIST metabolic engineers presented the bio-based production of multiple short-chain primary amines that have a wide range of applications in chemical industries for the first time. The research team led by Distinguished Professor Sang Yup Lee from the Department of Chemical and Biomolecular Engineering designed the novel biosynthetic pathways for short-chain primary amines by combining retrobiosynthesis and a precursor selection step.
The research team verified the newly designed pathways by confirming the in vivo production of 10 short-chain primary amines by supplying the precursors. Furthermore, the platform Escherichia coli strains were metabolically engineered to produce three proof-of-concept short-chain primary amines from glucose, demonstrating the possibility of the bio-based production of diverse short-chain primary amines from renewable resources. The research team said this study expands the strategy of systematically designing biosynthetic pathways for the production of a group of related chemicals as demonstrated by multiple short-chain primary amines as examples.
Currently, most of the industrial chemicals used in our daily lives are produced with petroleum-based products. However, there are several serious issues with the petroleum industry such as the depletion of fossil fuel reserves and environmental problems including global warming. To solve these problems, the sustainable production of industrial chemicals and materials is being explored with microorganisms as cell factories and renewable non-food biomass as raw materials for alternative to petroleum-based products. The engineering of these microorganisms has increasingly become more efficient and effective with the help of systems metabolic engineering – a practice of engineering the metabolism of a living organism toward the production of a desired metabolite. In this regard, the number of chemicals produced using biomass as a raw material has substantially increased.
Although the scope of chemicals that are producible using microorganisms continues to expand through advances in systems metabolic engineering, the biological production of short-chain primary amines has not yet been reported despite their industrial importance. Short-chain primary amines are the chemicals that have an alkyl or aryl group in the place of a hydrogen atom in ammonia with carbon chain lengths ranging from C1 to C7. Short-chain primary amines have a wide range of applications in chemical industries, for example, as a precursor for pharmaceuticals (e.g., antidiabetic and antihypertensive drugs), agrochemicals (e.g., herbicides, fungicides and insecticides), solvents, and vulcanization accelerators for rubber and plasticizers. The market size of short-chain primary amines was estimated to be more than 4 billion US dollars in 2014.
The main reason why the bio-based production of short-chain primary amines was not yet possible was due to their unknown biosynthetic pathways. Therefore, the team designed synthetic biosynthetic pathways for short-chain primary amines by combining retrobiosynthesis and a precursor selection step. The retrobiosynthesis allowed the systematic design of a biosynthetic pathway for short-chain primary amines by using a set of biochemical reaction rules that describe chemical transformation patterns between a substrate and product molecules at an atomic level.
These multiple precursors predicted for the possible biosynthesis of each short-chain primary amine were sequentially narrowed down by using the precursor selection step for efficient metabolic engineering experiments.
“Our research demonstrates the possibility of the renewable production of short-chain primary amines for the first time. We are planning to increase production efficiencies of short-chain primary amines. We believe that our study will play an important role in the development of sustainable and eco-friendly bio-based industries and the reorganization of the chemical industry, which is mandatory for solving the environmental problems threating the survival of mankind,” said Professor Lee.
This paper titled “Microbial production of multiple short-chain primary amines via retrobiosynthesis” was published in Nature Communications. This work was supported by the Technology Development Program to Solve Climate Changes on Systems Metabolic Engineering for Biorefineries from the Ministry of Science and ICT through the National Research Foundation (NRF) of Korea.
-Publication
Dong In Kim, Tong Un Chae, Hyun Uk Kim, Woo Dae Jang, and Sang Yup Lee. Microbial production of multiple short-chain primary amines via retrobiosynthesis. Nature Communications ( https://www.nature.com/articles/s41467-020-20423-6)
-Profile
Distinguished Professor Sang Yup Lee
leesy@kaist.ac.kr
Metabolic &Biomolecular Engineering National Research Laboratory
http://mbel.kaist.ac.kr
Department of Chemical and Biomolecular Engineering
KAIST
2021.01.14 View 8308 -
 DeepTFactor Predicts Transcription Factors
A deep learning-based tool predicts transcription factors using protein sequences as inputs
A joint research team from KAIST and UCSD has developed a deep neural network named DeepTFactor that predicts transcription factors from protein sequences. DeepTFactor will serve as a useful tool for understanding the regulatory systems of organisms, accelerating the use of deep learning for solving biological problems.
A transcription factor is a protein that specifically binds to DNA sequences to control the transcription initiation. Analyzing transcriptional regulation enables the understanding of how organisms control gene expression in response to genetic or environmental changes. In this regard, finding the transcription factor of an organism is the first step in the analysis of the transcriptional regulatory system of an organism.
Previously, transcription factors have been predicted by analyzing sequence homology with already characterized transcription factors or by data-driven approaches such as machine learning. Conventional machine learning models require a rigorous feature selection process that relies on domain expertise such as calculating the physicochemical properties of molecules or analyzing the homology of biological sequences. Meanwhile, deep learning can inherently learn latent features for the specific task.
A joint research team comprised of Ph.D. candidate Gi Bae Kim and Distinguished Professor Sang Yup Lee of the Department of Chemical and Biomolecular Engineering at KAIST, and Ye Gao and Professor Bernhard O. Palsson of the Department of Biochemical Engineering at UCSD reported a deep learning-based tool for the prediction of transcription factors. Their research paper “DeepTFactor: A deep learning-based tool for the prediction of transcription factors” was published online in PNAS.
Their article reports the development of DeepTFactor, a deep learning-based tool that predicts whether a given protein sequence is a transcription factor using three parallel convolutional neural networks. The joint research team predicted 332 transcription factors of Escherichia coli K-12 MG1655 using DeepTFactor and the performance of DeepTFactor by experimentally confirming the genome-wide binding sites of three predicted transcription factors (YqhC, YiaU, and YahB).
The joint research team further used a saliency method to understand the reasoning process of DeepTFactor. The researchers confirmed that even though information on the DNA binding domains of the transcription factor was not explicitly given the training process, DeepTFactor implicitly learned and used them for prediction. Unlike previous transcription factor prediction tools that were developed only for protein sequences of specific organisms, DeepTFactor is expected to be used in the analysis of the transcription systems of all organisms at a high level of performance.
Distinguished Professor Sang Yup Lee said, “DeepTFactor can be used to discover unknown transcription factors from numerous protein sequences that have not yet been characterized. It is expected that DeepTFactor will serve as an important tool for analyzing the regulatory systems of organisms of interest.”
This work was supported by the Technology Development Program to Solve Climate Changes on Systems Metabolic Engineering for Biorefineries from the Ministry of Science and ICT through the National Research Foundation of Korea.
-Publication
Gi Bae Kim, Ye Gao, Bernhard O. Palsson, and Sang Yup Lee. DeepTFactor: A deep learning-based tool for the prediction of transcription factors. (https://doi.org/10.1073/pnas202117118)
-Profile
Distinguished Professor Sang Yup Lee
leesy@kaist.ac.kr
Metabolic &Biomolecular Engineering National Research Laboratory
http://mbel.kaist.ac.kr
Department of Chemical and Biomolecular Engineering
KAIST
2021.01.05 View 6440
DeepTFactor Predicts Transcription Factors
A deep learning-based tool predicts transcription factors using protein sequences as inputs
A joint research team from KAIST and UCSD has developed a deep neural network named DeepTFactor that predicts transcription factors from protein sequences. DeepTFactor will serve as a useful tool for understanding the regulatory systems of organisms, accelerating the use of deep learning for solving biological problems.
A transcription factor is a protein that specifically binds to DNA sequences to control the transcription initiation. Analyzing transcriptional regulation enables the understanding of how organisms control gene expression in response to genetic or environmental changes. In this regard, finding the transcription factor of an organism is the first step in the analysis of the transcriptional regulatory system of an organism.
Previously, transcription factors have been predicted by analyzing sequence homology with already characterized transcription factors or by data-driven approaches such as machine learning. Conventional machine learning models require a rigorous feature selection process that relies on domain expertise such as calculating the physicochemical properties of molecules or analyzing the homology of biological sequences. Meanwhile, deep learning can inherently learn latent features for the specific task.
A joint research team comprised of Ph.D. candidate Gi Bae Kim and Distinguished Professor Sang Yup Lee of the Department of Chemical and Biomolecular Engineering at KAIST, and Ye Gao and Professor Bernhard O. Palsson of the Department of Biochemical Engineering at UCSD reported a deep learning-based tool for the prediction of transcription factors. Their research paper “DeepTFactor: A deep learning-based tool for the prediction of transcription factors” was published online in PNAS.
Their article reports the development of DeepTFactor, a deep learning-based tool that predicts whether a given protein sequence is a transcription factor using three parallel convolutional neural networks. The joint research team predicted 332 transcription factors of Escherichia coli K-12 MG1655 using DeepTFactor and the performance of DeepTFactor by experimentally confirming the genome-wide binding sites of three predicted transcription factors (YqhC, YiaU, and YahB).
The joint research team further used a saliency method to understand the reasoning process of DeepTFactor. The researchers confirmed that even though information on the DNA binding domains of the transcription factor was not explicitly given the training process, DeepTFactor implicitly learned and used them for prediction. Unlike previous transcription factor prediction tools that were developed only for protein sequences of specific organisms, DeepTFactor is expected to be used in the analysis of the transcription systems of all organisms at a high level of performance.
Distinguished Professor Sang Yup Lee said, “DeepTFactor can be used to discover unknown transcription factors from numerous protein sequences that have not yet been characterized. It is expected that DeepTFactor will serve as an important tool for analyzing the regulatory systems of organisms of interest.”
This work was supported by the Technology Development Program to Solve Climate Changes on Systems Metabolic Engineering for Biorefineries from the Ministry of Science and ICT through the National Research Foundation of Korea.
-Publication
Gi Bae Kim, Ye Gao, Bernhard O. Palsson, and Sang Yup Lee. DeepTFactor: A deep learning-based tool for the prediction of transcription factors. (https://doi.org/10.1073/pnas202117118)
-Profile
Distinguished Professor Sang Yup Lee
leesy@kaist.ac.kr
Metabolic &Biomolecular Engineering National Research Laboratory
http://mbel.kaist.ac.kr
Department of Chemical and Biomolecular Engineering
KAIST
2021.01.05 View 6440 -
 A Comprehensive Review of Biosynthesis of Inorganic Nanomaterials Using Microorganisms and Bacteriophages
There are diverse methods for producing numerous inorganic nanomaterials involving many experimental variables. Among the numerous possible matches, finding the best pair for synthesizing in an environmentally friendly way has been a longstanding challenge for researchers and industries.
A KAIST bioprocess engineering research team led by Distinguished Professor Sang Yup Lee conducted a summary of 146 biosynthesized single and multi-element inorganic nanomaterials covering 55 elements in the periodic table synthesized using wild-type and genetically engineered microorganisms. Their research highlights the diverse applications of biogenic nanomaterials and gives strategies for improving the biosynthesis of nanomaterials in terms of their producibility, crystallinity, size, and shape.
The research team described a 10-step flow chart for developing the biosynthesis of inorganic nanomaterials using microorganisms and bacteriophages. The research was published at Nature Review Chemistry as a cover and hero paper on December 3.
“We suggest general strategies for microbial nanomaterial biosynthesis via a step-by-step flow chart and give our perspectives on the future of nanomaterial biosynthesis and applications. This flow chart will serve as a general guide for those wishing to prepare biosynthetic inorganic nanomaterials using microbial cells,” explained Dr.Yoojin Choi, a co-author of this research.
Most inorganic nanomaterials are produced using physical and chemical methods and biological synthesis has been gaining more and more attention. However, conventional synthesis processes have drawbacks in terms of high energy consumption and non-environmentally friendly processes. Meanwhile, microorganisms such as microalgae, yeasts, fungi, bacteria, and even viruses can be utilized as biofactories to produce single and multi-element inorganic nanomaterials under mild conditions.
After conducting a massive survey, the research team summed up that the development of genetically engineered microorganisms with increased inorganic-ion-binding affinity, inorganic-ion-reduction ability, and nanomaterial biosynthetic efficiency has enabled the synthesis of many inorganic nanomaterials.
Among the strategies, the team introduced their analysis of a Pourbaix diagram for controlling the size and morphology of a product. The research team said this Pourbaix diagram analysis can be widely employed for biosynthesizing new nanomaterials with industrial applications.Professor Sang Yup Lee added, “This research provides extensive information and perspectives on the biosynthesis of diverse inorganic nanomaterials using microorganisms and bacteriophages and their applications. We expect that biosynthetic inorganic nanomaterials will find more diverse and innovative applications across diverse fields of science and technology.”
Dr. Choi started this research in 2018 and her interview about completing this extensive research was featured in an article at Nature Career article on December 4.
-ProfileDistinguished Professor Sang Yup Lee leesy@kaist.ac.krMetabolic &Biomolecular Engineering National Research Laboratoryhttp://mbel.kaist.ac.krDepartment of Chemical and Biomolecular EngineeringKAIST
2020.12.07 View 7444
A Comprehensive Review of Biosynthesis of Inorganic Nanomaterials Using Microorganisms and Bacteriophages
There are diverse methods for producing numerous inorganic nanomaterials involving many experimental variables. Among the numerous possible matches, finding the best pair for synthesizing in an environmentally friendly way has been a longstanding challenge for researchers and industries.
A KAIST bioprocess engineering research team led by Distinguished Professor Sang Yup Lee conducted a summary of 146 biosynthesized single and multi-element inorganic nanomaterials covering 55 elements in the periodic table synthesized using wild-type and genetically engineered microorganisms. Their research highlights the diverse applications of biogenic nanomaterials and gives strategies for improving the biosynthesis of nanomaterials in terms of their producibility, crystallinity, size, and shape.
The research team described a 10-step flow chart for developing the biosynthesis of inorganic nanomaterials using microorganisms and bacteriophages. The research was published at Nature Review Chemistry as a cover and hero paper on December 3.
“We suggest general strategies for microbial nanomaterial biosynthesis via a step-by-step flow chart and give our perspectives on the future of nanomaterial biosynthesis and applications. This flow chart will serve as a general guide for those wishing to prepare biosynthetic inorganic nanomaterials using microbial cells,” explained Dr.Yoojin Choi, a co-author of this research.
Most inorganic nanomaterials are produced using physical and chemical methods and biological synthesis has been gaining more and more attention. However, conventional synthesis processes have drawbacks in terms of high energy consumption and non-environmentally friendly processes. Meanwhile, microorganisms such as microalgae, yeasts, fungi, bacteria, and even viruses can be utilized as biofactories to produce single and multi-element inorganic nanomaterials under mild conditions.
After conducting a massive survey, the research team summed up that the development of genetically engineered microorganisms with increased inorganic-ion-binding affinity, inorganic-ion-reduction ability, and nanomaterial biosynthetic efficiency has enabled the synthesis of many inorganic nanomaterials.
Among the strategies, the team introduced their analysis of a Pourbaix diagram for controlling the size and morphology of a product. The research team said this Pourbaix diagram analysis can be widely employed for biosynthesizing new nanomaterials with industrial applications.Professor Sang Yup Lee added, “This research provides extensive information and perspectives on the biosynthesis of diverse inorganic nanomaterials using microorganisms and bacteriophages and their applications. We expect that biosynthetic inorganic nanomaterials will find more diverse and innovative applications across diverse fields of science and technology.”
Dr. Choi started this research in 2018 and her interview about completing this extensive research was featured in an article at Nature Career article on December 4.
-ProfileDistinguished Professor Sang Yup Lee leesy@kaist.ac.krMetabolic &Biomolecular Engineering National Research Laboratoryhttp://mbel.kaist.ac.krDepartment of Chemical and Biomolecular EngineeringKAIST
2020.12.07 View 7444 -
 Engineered C. glutamicum Strain Capable of Producing High-Level Glutaric Acid from Glucose
An engineered C. glutamicum strain that can produce the world’s highest titer of glutaric acid was developed by employing systems metabolic engineering strategies
A metabolic engineering research group at KAIST has developed an engineered Corynebacterium glutamicum strain capable of producing high-level glutaric acid without byproducts from glucose. This new strategy will be useful for developing engineered micro-organisms for the bio-based production of value-added chemicals.
Glutaric acid, also known as pentanedioic acid, is a carboxylic acid that is widely used for various applications including the production of polyesters, polyamides, polyurethanes, glutaric anhydride, 1,5-pentanediol, and 5-hydroxyvaleric acid. Glutaric acid has been produced using various petroleum-based chemical methods, relying on non-renewable and toxic starting materials. Thus, various approaches have been taken to biologically produce glutaric acid from renewable resources. Previously, the development of the first glutaric acid producing Escherichia coli by introducing Pseudomonas putida genes was reported by a research group from KAIST, but the titer was low. Glutaric acid production by metabolically engineered Corynebacterium glutamicum has also been reported in several studies, but further improvements in glutaric acid production seemed possible since C. glutamicum has the capability of producing more than 130 g/L of L-lysine.
A research group comprised of Taehee Han, Gi Bae Kim, and Distinguished Professor Sang Yup Lee of the Department of Chemical and Biomolecular Engineering addressed this issue. Their research paper “Glutaric acid production by systems metabolic engineering of an L-lysine-overproducing Corynebacterium glutamicum” was published online in PNAS on November 16, 2020.
This research reports the development of a metabolically engineered C. glutamicum strain capable of efficiently producing glutaric acid, starting from an L-lysine overproducer. The following novel strategies and approaches to achieve high-level glutaric acid production were employed. First, metabolic pathways in C. glutamicum were reconstituted for glutaric acid production by introducing P. putida genes. Then, multi-omics analyses including genome, transcriptome, and fluxome were conducted to understand the phenotype of the L-lysine overproducer strain. In addition to systematic understanding of the host strain, gene manipulation targets were predicted by omics analyses and applied for engineering C. glutamicum, which resulted in the development of an engineered strain capable of efficiently producing glutaric acid. Furthermore, the new glutaric acid exporter was discovered for the first time, which was used to further increase glutaric acid production through enhancing product excretion. Last but not least, culture conditions were optimized for high-level glutaric acid production. As a result, the final engineered strain was able to produce 105.3 g/L glutaric acid, the highest titer ever reported, in 69 hours by fed-batch fermentation.
Professor Sang Yup Lee said, “It is meaningful that we were able to develop a highly efficient glutaric acid producer capable of producing glutaric acid at the world’s highest titer without any byproducts from renewable carbon sources. This will further accelerate the bio-based production of valuable chemicals in pharmaceutical/medical/chemical industries.” This research was supported by the Bio & Medical Technology Development Program of the National Research Foundation and funded by the Ministry of Science and ICT.
-Profile
Distinguished Professor Sang Yup Lee
leesy@kaist.ac.kr
http://mbel.kaist.ac.kr
Department of Chemical and Biomolecular Engineering
KAIST
2020.11.17 View 5034
Engineered C. glutamicum Strain Capable of Producing High-Level Glutaric Acid from Glucose
An engineered C. glutamicum strain that can produce the world’s highest titer of glutaric acid was developed by employing systems metabolic engineering strategies
A metabolic engineering research group at KAIST has developed an engineered Corynebacterium glutamicum strain capable of producing high-level glutaric acid without byproducts from glucose. This new strategy will be useful for developing engineered micro-organisms for the bio-based production of value-added chemicals.
Glutaric acid, also known as pentanedioic acid, is a carboxylic acid that is widely used for various applications including the production of polyesters, polyamides, polyurethanes, glutaric anhydride, 1,5-pentanediol, and 5-hydroxyvaleric acid. Glutaric acid has been produced using various petroleum-based chemical methods, relying on non-renewable and toxic starting materials. Thus, various approaches have been taken to biologically produce glutaric acid from renewable resources. Previously, the development of the first glutaric acid producing Escherichia coli by introducing Pseudomonas putida genes was reported by a research group from KAIST, but the titer was low. Glutaric acid production by metabolically engineered Corynebacterium glutamicum has also been reported in several studies, but further improvements in glutaric acid production seemed possible since C. glutamicum has the capability of producing more than 130 g/L of L-lysine.
A research group comprised of Taehee Han, Gi Bae Kim, and Distinguished Professor Sang Yup Lee of the Department of Chemical and Biomolecular Engineering addressed this issue. Their research paper “Glutaric acid production by systems metabolic engineering of an L-lysine-overproducing Corynebacterium glutamicum” was published online in PNAS on November 16, 2020.
This research reports the development of a metabolically engineered C. glutamicum strain capable of efficiently producing glutaric acid, starting from an L-lysine overproducer. The following novel strategies and approaches to achieve high-level glutaric acid production were employed. First, metabolic pathways in C. glutamicum were reconstituted for glutaric acid production by introducing P. putida genes. Then, multi-omics analyses including genome, transcriptome, and fluxome were conducted to understand the phenotype of the L-lysine overproducer strain. In addition to systematic understanding of the host strain, gene manipulation targets were predicted by omics analyses and applied for engineering C. glutamicum, which resulted in the development of an engineered strain capable of efficiently producing glutaric acid. Furthermore, the new glutaric acid exporter was discovered for the first time, which was used to further increase glutaric acid production through enhancing product excretion. Last but not least, culture conditions were optimized for high-level glutaric acid production. As a result, the final engineered strain was able to produce 105.3 g/L glutaric acid, the highest titer ever reported, in 69 hours by fed-batch fermentation.
Professor Sang Yup Lee said, “It is meaningful that we were able to develop a highly efficient glutaric acid producer capable of producing glutaric acid at the world’s highest titer without any byproducts from renewable carbon sources. This will further accelerate the bio-based production of valuable chemicals in pharmaceutical/medical/chemical industries.” This research was supported by the Bio & Medical Technology Development Program of the National Research Foundation and funded by the Ministry of Science and ICT.
-Profile
Distinguished Professor Sang Yup Lee
leesy@kaist.ac.kr
http://mbel.kaist.ac.kr
Department of Chemical and Biomolecular Engineering
KAIST
2020.11.17 View 5034 -
 Experts to Help Asia Navigate the Post-COVID-19 and 4IR Eras
Risk Quotient 2020, an international conference co-hosted by KAIST and the National University of Singapore (NUS), will bring together world-leading experts from academia and industry to help Asia navigate the post-COVID-19 and Fourth Industrial Revolution (4IR) eras.
The online conference will be held on October 29 from 10 a.m. Korean time under the theme “COVID-19 Pandemic and A Brave New World”. It will be streamed live on YouTube at https://www.youtube.com/c/KAISTofficial and https://www.youtube.com/user/NUScast.
The Korea Policy Center for the Fourth Industrial Revolution (KPC4IR) at KAIST organized this conference in collaboration with the Lloyd's Register Foundation Institute for the Public Understanding of Risk (IPUR) at NUS.
During the conference, global leaders will examine the socioeconomic impacts of the COVID-19 pandemic on areas including digital innovation, education, the workforce, and the economy. They will then highlight digital and 4IR technologies that could be utilized to effectively mitigate the risks and challenges associated with the pandemic, while harnessing the opportunities that these socioeconomic effects may present. Their discussions will mainly focus on the Asian region.
In his opening remarks, KAIST President Sung-Chul Shin will express his appreciation for the Asian populations’ greater trust in and compliance with their governments, which have given the continent a leg up against the coronavirus. He will then emphasize that by working together through the exchange of ideas and global collaboration, we will be able to shape ‘a brave new world’ to better humanity. Welcoming remarks by Prof. Sang Yup Lee (Dean, KAIST Institutes) and Prof. Tze Yun Leong (Director, AI Technology at AI Singapore) will follow.
For the keynote speech, Prof. Lan Xue (Dean, Schwarzman College, Tsinghua University) will share China’s response to COVID-19 and lessons for crisis management. Prof. Danny Quah (Dean, Lee Kuan Yew School of Public Policy, NUS) will present possible ways to overcome these difficult times. Dr. Kak-Soo Shin (Senior Advisor, Shin & Kim LLC, Former Ambassador to the State of Israel and Japan, and Former First and Second Vice Minister of the Ministry of Foreign Affairs of the Republic of Korea) will stress the importance of the international community’s solidarity to ensure peace, prosperity, and safety in this new era.
Panel Session I will address the impact of COVID-19 on digital innovation. Dr. Carol Soon (Senior Research Fellow, Institute of Policy Studies, NUS) will present her interpretation of recent technological developments as both opportunities for our society as a whole and challenges for vulnerable groups such as low-income families. Dr. Christopher SungWook Chang (Managing Director, Kakao Mobility) will show how changes in mobility usage patterns can be captured by Kakao Mobility’s big data analysis. He will illustrate how the data can be used to interpret citizen’s behaviors and how risks can be transformed into opportunities by utilizing technology. Mr. Steve Ledzian’s (Vice President, Chief Technology Officer, FireEye) talk will discuss the dangers caused by threat actors and other cyber risk implications of COVID-19. Dr. June Sung Park (Chairman, Korea Software Technology Association (KOSTA)) will share how COVID-19 has accelerated digital transformations across all industries and why software education should be reformed to improve Korea’s competitiveness.
Panel Session II will examine the impact on education and the workforce. Dr. Sang-Jin Ban (President, Korean Educational Development Institute (KEDI)) will explain Korea’s educational response to the pandemic and the concept of “blended learning” as a new paradigm, and present both positive and negative impacts of online education on students’ learning experiences. Prof. Reuben Ng (Professor, Lee Kuan Yew School of Public Policy, NUS) will present on graduate underemployment, which seems to have worsened during COVID-19. Dr. Michael Fung’s presentation (Deputy Chief Executive (Industry), SkillsFuture SG) will introduce the promotion of lifelong learning in Singapore through a new national initiative known as the ‘SkillsFuture Movement’. This movement serves as an example of a national response to disruptions in the job market and the pace of skills obsolescence triggered by AI and COVID-19.
Panel Session III will touch on technology leadership and Asia’s digital economy and society. Prof. Naubahar Sharif (Professor, Division of Social Science and Division of Public Policy, Hong Kong University of Science and Technology (HKUST)) will share his views on the potential of China in taking over global technological leadership based on its massive domestic market, its government support, and the globalization process. Prof. Yee Kuang Heng (Professor, Graduate School of Public Policy, University of Tokyo) will illustrate how different legal and political needs in China and Japan have shaped the ways technologies have been deployed in responding to COVID-19. Dr. Hayun Kang (Head, International Cooperation Research Division, Korea Information Society Development Institute (KISDI)) will explain Korea’s relative success containing the pandemic compared to other countries, and how policy leaders and institutions that embrace digital technologies in the pursuit of public welfare objectives can produce positive outcomes while minimizing the side effects.
Prof. Kyung Ryul Park (Graduate School of Science and Technology Policy, KAIST) will be hosting the entire conference, whereas Prof. Alice Hae Yun Oh (Director, MARS Artificial Intelligence Research Center, KAIST), Prof. Wonjoon Kim (Dean, Graduate School of Innovation and Technology Management, College of Business, KAIST), Prof. Youngsun Kwon (Dean, KAIST Academy), and Prof. Taejun Lee (Korea Development Institute (KDI) School of Public Policy and Management) are to chair discussions with the keynote speakers and panelists.
Closing remarks will be delivered by Prof. Chan Ghee Koh (Director, NUS IPUR), Prof. So Young Kim (Director, KAIST KPC4IR), and Prof. Joungho Kim (Director, KAIST Global Strategy Institute (GSI)).
“This conference is expected to serve as a springboard to help Asian countries recover from global crises such as the COVID-19 pandemic through active cooperation and joint engagement among scholars, experts, and policymakers,” according to Director So Young Kim.
(END)
2020.10.22 View 9918
Experts to Help Asia Navigate the Post-COVID-19 and 4IR Eras
Risk Quotient 2020, an international conference co-hosted by KAIST and the National University of Singapore (NUS), will bring together world-leading experts from academia and industry to help Asia navigate the post-COVID-19 and Fourth Industrial Revolution (4IR) eras.
The online conference will be held on October 29 from 10 a.m. Korean time under the theme “COVID-19 Pandemic and A Brave New World”. It will be streamed live on YouTube at https://www.youtube.com/c/KAISTofficial and https://www.youtube.com/user/NUScast.
The Korea Policy Center for the Fourth Industrial Revolution (KPC4IR) at KAIST organized this conference in collaboration with the Lloyd's Register Foundation Institute for the Public Understanding of Risk (IPUR) at NUS.
During the conference, global leaders will examine the socioeconomic impacts of the COVID-19 pandemic on areas including digital innovation, education, the workforce, and the economy. They will then highlight digital and 4IR technologies that could be utilized to effectively mitigate the risks and challenges associated with the pandemic, while harnessing the opportunities that these socioeconomic effects may present. Their discussions will mainly focus on the Asian region.
In his opening remarks, KAIST President Sung-Chul Shin will express his appreciation for the Asian populations’ greater trust in and compliance with their governments, which have given the continent a leg up against the coronavirus. He will then emphasize that by working together through the exchange of ideas and global collaboration, we will be able to shape ‘a brave new world’ to better humanity. Welcoming remarks by Prof. Sang Yup Lee (Dean, KAIST Institutes) and Prof. Tze Yun Leong (Director, AI Technology at AI Singapore) will follow.
For the keynote speech, Prof. Lan Xue (Dean, Schwarzman College, Tsinghua University) will share China’s response to COVID-19 and lessons for crisis management. Prof. Danny Quah (Dean, Lee Kuan Yew School of Public Policy, NUS) will present possible ways to overcome these difficult times. Dr. Kak-Soo Shin (Senior Advisor, Shin & Kim LLC, Former Ambassador to the State of Israel and Japan, and Former First and Second Vice Minister of the Ministry of Foreign Affairs of the Republic of Korea) will stress the importance of the international community’s solidarity to ensure peace, prosperity, and safety in this new era.
Panel Session I will address the impact of COVID-19 on digital innovation. Dr. Carol Soon (Senior Research Fellow, Institute of Policy Studies, NUS) will present her interpretation of recent technological developments as both opportunities for our society as a whole and challenges for vulnerable groups such as low-income families. Dr. Christopher SungWook Chang (Managing Director, Kakao Mobility) will show how changes in mobility usage patterns can be captured by Kakao Mobility’s big data analysis. He will illustrate how the data can be used to interpret citizen’s behaviors and how risks can be transformed into opportunities by utilizing technology. Mr. Steve Ledzian’s (Vice President, Chief Technology Officer, FireEye) talk will discuss the dangers caused by threat actors and other cyber risk implications of COVID-19. Dr. June Sung Park (Chairman, Korea Software Technology Association (KOSTA)) will share how COVID-19 has accelerated digital transformations across all industries and why software education should be reformed to improve Korea’s competitiveness.
Panel Session II will examine the impact on education and the workforce. Dr. Sang-Jin Ban (President, Korean Educational Development Institute (KEDI)) will explain Korea’s educational response to the pandemic and the concept of “blended learning” as a new paradigm, and present both positive and negative impacts of online education on students’ learning experiences. Prof. Reuben Ng (Professor, Lee Kuan Yew School of Public Policy, NUS) will present on graduate underemployment, which seems to have worsened during COVID-19. Dr. Michael Fung’s presentation (Deputy Chief Executive (Industry), SkillsFuture SG) will introduce the promotion of lifelong learning in Singapore through a new national initiative known as the ‘SkillsFuture Movement’. This movement serves as an example of a national response to disruptions in the job market and the pace of skills obsolescence triggered by AI and COVID-19.
Panel Session III will touch on technology leadership and Asia’s digital economy and society. Prof. Naubahar Sharif (Professor, Division of Social Science and Division of Public Policy, Hong Kong University of Science and Technology (HKUST)) will share his views on the potential of China in taking over global technological leadership based on its massive domestic market, its government support, and the globalization process. Prof. Yee Kuang Heng (Professor, Graduate School of Public Policy, University of Tokyo) will illustrate how different legal and political needs in China and Japan have shaped the ways technologies have been deployed in responding to COVID-19. Dr. Hayun Kang (Head, International Cooperation Research Division, Korea Information Society Development Institute (KISDI)) will explain Korea’s relative success containing the pandemic compared to other countries, and how policy leaders and institutions that embrace digital technologies in the pursuit of public welfare objectives can produce positive outcomes while minimizing the side effects.
Prof. Kyung Ryul Park (Graduate School of Science and Technology Policy, KAIST) will be hosting the entire conference, whereas Prof. Alice Hae Yun Oh (Director, MARS Artificial Intelligence Research Center, KAIST), Prof. Wonjoon Kim (Dean, Graduate School of Innovation and Technology Management, College of Business, KAIST), Prof. Youngsun Kwon (Dean, KAIST Academy), and Prof. Taejun Lee (Korea Development Institute (KDI) School of Public Policy and Management) are to chair discussions with the keynote speakers and panelists.
Closing remarks will be delivered by Prof. Chan Ghee Koh (Director, NUS IPUR), Prof. So Young Kim (Director, KAIST KPC4IR), and Prof. Joungho Kim (Director, KAIST Global Strategy Institute (GSI)).
“This conference is expected to serve as a springboard to help Asian countries recover from global crises such as the COVID-19 pandemic through active cooperation and joint engagement among scholars, experts, and policymakers,” according to Director So Young Kim.
(END)
2020.10.22 View 9918 -
 E. coli Engineered to Grow on CO₂ and Formic Acid as Sole Carbon Sources
- An E. coli strain that can grow to a relatively high cell density solely on CO₂ and formic acid was developed by employing metabolic engineering. -
Most biorefinery processes have relied on the use of biomass as a raw material for the production of chemicals and materials. Even though the use of CO₂ as a carbon source in biorefineries is desirable, it has not been possible to make common microbial strains such as E. coli grow on CO₂.
Now, a metabolic engineering research group at KAIST has developed a strategy to grow an E. coli strain to higher cell density solely on CO₂ and formic acid. Formic acid is a one carbon carboxylic acid, and can be easily produced from CO₂ using a variety of methods. Since it is easier to store and transport than CO₂, formic acid can be considered a good liquid-form alternative of CO₂.
With support from the C1 Gas Refinery R&D Center and the Ministry of Science and ICT, a research team led by Distinguished Professor Sang Yup Lee stepped up their work to develop an engineered E. coli strain capable of growing up to 11-fold higher cell density than those previously reported, using CO₂ and formic acid as sole carbon sources. This work was published in Nature Microbiology on September 28.
Despite the recent reports by several research groups on the development of E. coli strains capable of growing on CO₂ and formic acid, the maximum cell growth remained too low (optical density of around 1) and thus the production of chemicals from CO₂ and formic acid has been far from realized.
The team previously reported the reconstruction of the tetrahydrofolate cycle and reverse glycine cleavage pathway to construct an engineered E. coli strain that can sustain growth on CO₂ and formic acid. To further enhance the growth, the research team introduced the previously designed synthetic CO₂ and formic acid assimilation pathway, and two formate dehydrogenases.
Metabolic fluxes were also fine-tuned, the gluconeogenic flux enhanced, and the levels of cytochrome bo3 and bd-I ubiquinol oxidase for ATP generation were optimized. This engineered E. coli strain was able to grow to a relatively high OD600 of 7~11, showing promise as a platform strain growing solely on CO₂ and formic acid.
Professor Lee said, “We engineered E. coli that can grow to a higher cell density only using CO₂ and formic acid. We think that this is an important step forward, but this is not the end. The engineered strain we developed still needs further engineering so that it can grow faster to a much higher density.”
Professor Lee’s team is continuing to develop such a strain. “In the future, we would be delighted to see the production of chemicals from an engineered E. coli strain using CO₂ and formic acid as sole carbon sources,” he added.
-Profile:Distinguished Professor Sang Yup Leehttp://mbel.kaist.ac.krDepartment of Chemical and Biomolecular EngineeringKAIST
2020.09.29 View 7958
E. coli Engineered to Grow on CO₂ and Formic Acid as Sole Carbon Sources
- An E. coli strain that can grow to a relatively high cell density solely on CO₂ and formic acid was developed by employing metabolic engineering. -
Most biorefinery processes have relied on the use of biomass as a raw material for the production of chemicals and materials. Even though the use of CO₂ as a carbon source in biorefineries is desirable, it has not been possible to make common microbial strains such as E. coli grow on CO₂.
Now, a metabolic engineering research group at KAIST has developed a strategy to grow an E. coli strain to higher cell density solely on CO₂ and formic acid. Formic acid is a one carbon carboxylic acid, and can be easily produced from CO₂ using a variety of methods. Since it is easier to store and transport than CO₂, formic acid can be considered a good liquid-form alternative of CO₂.
With support from the C1 Gas Refinery R&D Center and the Ministry of Science and ICT, a research team led by Distinguished Professor Sang Yup Lee stepped up their work to develop an engineered E. coli strain capable of growing up to 11-fold higher cell density than those previously reported, using CO₂ and formic acid as sole carbon sources. This work was published in Nature Microbiology on September 28.
Despite the recent reports by several research groups on the development of E. coli strains capable of growing on CO₂ and formic acid, the maximum cell growth remained too low (optical density of around 1) and thus the production of chemicals from CO₂ and formic acid has been far from realized.
The team previously reported the reconstruction of the tetrahydrofolate cycle and reverse glycine cleavage pathway to construct an engineered E. coli strain that can sustain growth on CO₂ and formic acid. To further enhance the growth, the research team introduced the previously designed synthetic CO₂ and formic acid assimilation pathway, and two formate dehydrogenases.
Metabolic fluxes were also fine-tuned, the gluconeogenic flux enhanced, and the levels of cytochrome bo3 and bd-I ubiquinol oxidase for ATP generation were optimized. This engineered E. coli strain was able to grow to a relatively high OD600 of 7~11, showing promise as a platform strain growing solely on CO₂ and formic acid.
Professor Lee said, “We engineered E. coli that can grow to a higher cell density only using CO₂ and formic acid. We think that this is an important step forward, but this is not the end. The engineered strain we developed still needs further engineering so that it can grow faster to a much higher density.”
Professor Lee’s team is continuing to develop such a strain. “In the future, we would be delighted to see the production of chemicals from an engineered E. coli strain using CO₂ and formic acid as sole carbon sources,” he added.
-Profile:Distinguished Professor Sang Yup Leehttp://mbel.kaist.ac.krDepartment of Chemical and Biomolecular EngineeringKAIST
2020.09.29 View 7958 -
 Researchers Present a Microbial Strain Capable of Massive Succinic Acid Production
A research team led by Distinguished Professor Sang Yup Lee reported the production of a microbial strain capable of the massive production of succinic acid with the highest production efficiency to date. This strategy of integrating systems metabolic engineering with enzyme engineering will be useful for the production of industrially competitive bio-based chemicals. Their strategy was described in Nature Communications on April 23.
The bio-based production of industrial chemicals from renewable non-food biomass has become increasingly important as a sustainable substitute for conventional petroleum-based production processes relying on fossil resources. Here, systems metabolic engineering, which is the key component for biorefinery technology, is utilized to effectively engineer the complex metabolic pathways of microorganisms to enable the efficient production of industrial chemicals.
Succinic acid, a four-carbon dicarboxylic acid, is one of the most promising platform chemicals serving as a precursor for industrially important chemicals. Among microorganisms producing succinic acid, Mannheimia succiniciproducens has been proven to be one of the best strains for succinic acid production.
The research team has developed a bio-based succinic acid production technology using the M. succiniciproducens strain isolated from the rumen of Korean cow for over 20 years and succeeded in developing a strain capable of producing succinic acid with the highest production efficiency.
They carried out systems metabolic engineering to optimize the succinic acid production pathway of the M. succiniciproducens strain by determining the crystal structure of key enzymes important for succinic acid production and performing protein engineering to develop enzymes with better catalytic performance.
As a result, 134 g per liter of succinic acid was produced from the fermentation of an engineered strain using glucose, glycerol, and carbon dioxide. They were able to achieve 21 g per liter per hour of succinic acid production, which is one of the key factors determining the economic feasibility of the overall production process. This is the world’s best succinic acid production efficiency reported to date. Previous production methods averaged 1~3 g per liter per hour.
Distinguished professor Sang Yup Lee explained that his team’s work will significantly contribute to transforming the current petrochemical-based industry into an eco-friendly bio-based one.
“Our research on the highly efficient bio-based production of succinic acid from renewable non-food resources and carbon dioxide has provided a basis for reducing our strong dependence on fossil resources, which is the main cause of the environmental crisis,” Professor Lee said.
This work was supported by the Technology Development Program to Solve Climate Changes via Systems Metabolic Engineering for Biorefineries and the C1 Gas Refinery Program from the Ministry of Science and ICT through the National Research Foundation of Korea.
2020.05.06 View 6622
Researchers Present a Microbial Strain Capable of Massive Succinic Acid Production
A research team led by Distinguished Professor Sang Yup Lee reported the production of a microbial strain capable of the massive production of succinic acid with the highest production efficiency to date. This strategy of integrating systems metabolic engineering with enzyme engineering will be useful for the production of industrially competitive bio-based chemicals. Their strategy was described in Nature Communications on April 23.
The bio-based production of industrial chemicals from renewable non-food biomass has become increasingly important as a sustainable substitute for conventional petroleum-based production processes relying on fossil resources. Here, systems metabolic engineering, which is the key component for biorefinery technology, is utilized to effectively engineer the complex metabolic pathways of microorganisms to enable the efficient production of industrial chemicals.
Succinic acid, a four-carbon dicarboxylic acid, is one of the most promising platform chemicals serving as a precursor for industrially important chemicals. Among microorganisms producing succinic acid, Mannheimia succiniciproducens has been proven to be one of the best strains for succinic acid production.
The research team has developed a bio-based succinic acid production technology using the M. succiniciproducens strain isolated from the rumen of Korean cow for over 20 years and succeeded in developing a strain capable of producing succinic acid with the highest production efficiency.
They carried out systems metabolic engineering to optimize the succinic acid production pathway of the M. succiniciproducens strain by determining the crystal structure of key enzymes important for succinic acid production and performing protein engineering to develop enzymes with better catalytic performance.
As a result, 134 g per liter of succinic acid was produced from the fermentation of an engineered strain using glucose, glycerol, and carbon dioxide. They were able to achieve 21 g per liter per hour of succinic acid production, which is one of the key factors determining the economic feasibility of the overall production process. This is the world’s best succinic acid production efficiency reported to date. Previous production methods averaged 1~3 g per liter per hour.
Distinguished professor Sang Yup Lee explained that his team’s work will significantly contribute to transforming the current petrochemical-based industry into an eco-friendly bio-based one.
“Our research on the highly efficient bio-based production of succinic acid from renewable non-food resources and carbon dioxide has provided a basis for reducing our strong dependence on fossil resources, which is the main cause of the environmental crisis,” Professor Lee said.
This work was supported by the Technology Development Program to Solve Climate Changes via Systems Metabolic Engineering for Biorefineries and the C1 Gas Refinery Program from the Ministry of Science and ICT through the National Research Foundation of Korea.
2020.05.06 View 6622 -
 Korea Policy Center for the Fourth Industrial Revolution Opens
The World Economic Forum’s Center for the Fourth Industrial Revolution opened its Korean affiliate center at KAIST on December 10. The Korea Policy Center for the 4th Industrial Revolution (KPC4IR) will develop policy norms and frameworks for accelerating the benefits of emerging technologies.
Many dignitaries including KAIST President Sung-Chul Shin, National Assemblyman Sang-Min Lee, Daejeon City Mayor Her Tae-Jeong, and Managing Director of the WEF Center for the Fourth Industrial Revolution Murat Sonmez attended the opening ceremony.
The center will play a vital role in helping to shape the development of national Fourth Industrial Revolution strategies and public-private initiatives. The Center will actively engage with the government on policy design and piloting activities.
The Center is the result of KAIST’s close partnership with the WEF and its Center for the Fourth Industrial Revolution in San Francisco. KAIST signed an MOU with the WEF in 2017 for this collaboration. Dr. Klaus Schwab expressed his high hopes many times regarding Korea’s potential in responding to the Fourth Industrial Revolution. In addition, he said that KAIST and the City of Daejeon would play a significant role in helping the Fourth Industrial Revolution move forward.
During a meeting with President Moon Jae-In last June, Dr. Schwab expressed his strong desire to collaborate with Korea, and the Korean government designated KAIST as an affiliate center of the WEF.
The KPC4IR had already begun conducting policy research in the areas of block chain and precision medicine even before making a partnership with the WEF. The director of the Center, Distinguished Professor Sang Yup Lee, said, “We have focused on the development of technology but rarely talk about governance. Technology should come with policy. We will conduct policy development on how to ensure inclusive growth capitalizing on emerging technologies. We will also make policy guidelines for technological applications after considering all the ethical perspectives.
President Shin also said in his opening remarks, “Korea has been a fast follower over the past decades in making economic development and innovations. I believe that the Fourth Industrial Revolution gives us the best opportunity to play the role of ‘first mover.’ I look forward to the KPC4IR serving as a ‘Think and Do’ tank, not limiting itself to the role of ‘think tank.’ We will continue to work closely with the WEF in the fields of AI, blockchain, and precision medicine.
2019.12.10 View 6126
Korea Policy Center for the Fourth Industrial Revolution Opens
The World Economic Forum’s Center for the Fourth Industrial Revolution opened its Korean affiliate center at KAIST on December 10. The Korea Policy Center for the 4th Industrial Revolution (KPC4IR) will develop policy norms and frameworks for accelerating the benefits of emerging technologies.
Many dignitaries including KAIST President Sung-Chul Shin, National Assemblyman Sang-Min Lee, Daejeon City Mayor Her Tae-Jeong, and Managing Director of the WEF Center for the Fourth Industrial Revolution Murat Sonmez attended the opening ceremony.
The center will play a vital role in helping to shape the development of national Fourth Industrial Revolution strategies and public-private initiatives. The Center will actively engage with the government on policy design and piloting activities.
The Center is the result of KAIST’s close partnership with the WEF and its Center for the Fourth Industrial Revolution in San Francisco. KAIST signed an MOU with the WEF in 2017 for this collaboration. Dr. Klaus Schwab expressed his high hopes many times regarding Korea’s potential in responding to the Fourth Industrial Revolution. In addition, he said that KAIST and the City of Daejeon would play a significant role in helping the Fourth Industrial Revolution move forward.
During a meeting with President Moon Jae-In last June, Dr. Schwab expressed his strong desire to collaborate with Korea, and the Korean government designated KAIST as an affiliate center of the WEF.
The KPC4IR had already begun conducting policy research in the areas of block chain and precision medicine even before making a partnership with the WEF. The director of the Center, Distinguished Professor Sang Yup Lee, said, “We have focused on the development of technology but rarely talk about governance. Technology should come with policy. We will conduct policy development on how to ensure inclusive growth capitalizing on emerging technologies. We will also make policy guidelines for technological applications after considering all the ethical perspectives.
President Shin also said in his opening remarks, “Korea has been a fast follower over the past decades in making economic development and innovations. I believe that the Fourth Industrial Revolution gives us the best opportunity to play the role of ‘first mover.’ I look forward to the KPC4IR serving as a ‘Think and Do’ tank, not limiting itself to the role of ‘think tank.’ We will continue to work closely with the WEF in the fields of AI, blockchain, and precision medicine.
2019.12.10 View 6126