Sang+Yup+Lee
-
 Distinguished Professor Sang Yup Lee Wins 2025 Global Metabolic Engineering Award
< Distinguished Professor Sang Yup Lee (Senior Vice President for Research) from the Department of Chemical & Biomolecular Engineering >
KAIST announced on the 20th that Professor Sang Yup Lee, who serves as the Vice President for Research and a Distinguished Professor at our university, has been awarded the '2025 Gregory N. Stephanopoulos Award for Metabolic Engineering' by the International Metabolic Engineering Society (IMES). Professor Lee delivered his award lecture at the 16th Metabolic Engineering Conference (ME16), held in Copenhagen, Denmark, from June 15th to 19th.
This award was established through contributions from the American Institute of Chemical Engineers (AIChE) Foundation, as well as fellow colleagues and acquaintances, to honor the achievements of Dr. Gregory Stephanopoulos, widely recognized as one of the pioneers of metabolic engineering. Presented biennially, the award recognizes scientists who have successfully commercialized fundamental research in metabolic engineering or have made outstanding contributions to the quantitative analysis, design, and modeling of metabolic pathways.
Professor Sang Yup Lee boasts an impressive record of over 770 journal papers and more than 860 patents. His groundbreaking research in metabolic engineering and biochemical engineering is highly acclaimed globally.
Throughout his 31 years as a professor at KAIST, Professor Lee has developed various metabolic engineering-based technologies and strategies. These advancements have been transferred to industries, facilitating the production of bulk chemicals, polymers, natural products, pharmaceuticals, and health functional foods. He has also founded companies and actively engages in advisory roles with various enterprises.
The International Metabolic Engineering Society (IMES) defines metabolic engineering as the manipulation of metabolic pathways in microorganisms or cells to produce useful substances (such as pharmaceuticals, biofuels, and chemical products). It utilizes tools like systems biology, synthetic biology, and computational modeling with the aim of enhancing the economic viability and sustainability of bio-based processes.
Furthermore, Professor Lee previously received the Merck Metabolic Engineering Award, a prominent international award in the field, in 2008. In 2018, he was honored with the Eni Award, often referred to as the Nobel Prize in energy, presented by the President of Italy.
Professor Sang Yup Lee remarked, "Metabolic engineering is a discipline that leads the current and future of biotechnology. It is a tremendous honor to receive this meaningful award at a time when the transition to a bio-based economy is accelerating. Together with my students and fellow researchers, we have generated numerous patents and transferred technologies to industry, and also established startups in the fields of biofuels, wound healing, and cosmetics. I will continue to pursue research that encompasses both fundamental research and technological commercialization."
The 'International Metabolic Engineering Society (IMES)' is a specialized society under the American Institute of Chemical Engineers. Its mission is to enable the production of various bio-based products, including pharmaceuticals, food additives, chemicals, and fuels, through metabolic engineering. The society hosts the Metabolic Engineering Conference biennially, offering researchers opportunities for knowledge exchange and collaboration.
2025.06.20 View 623
Distinguished Professor Sang Yup Lee Wins 2025 Global Metabolic Engineering Award
< Distinguished Professor Sang Yup Lee (Senior Vice President for Research) from the Department of Chemical & Biomolecular Engineering >
KAIST announced on the 20th that Professor Sang Yup Lee, who serves as the Vice President for Research and a Distinguished Professor at our university, has been awarded the '2025 Gregory N. Stephanopoulos Award for Metabolic Engineering' by the International Metabolic Engineering Society (IMES). Professor Lee delivered his award lecture at the 16th Metabolic Engineering Conference (ME16), held in Copenhagen, Denmark, from June 15th to 19th.
This award was established through contributions from the American Institute of Chemical Engineers (AIChE) Foundation, as well as fellow colleagues and acquaintances, to honor the achievements of Dr. Gregory Stephanopoulos, widely recognized as one of the pioneers of metabolic engineering. Presented biennially, the award recognizes scientists who have successfully commercialized fundamental research in metabolic engineering or have made outstanding contributions to the quantitative analysis, design, and modeling of metabolic pathways.
Professor Sang Yup Lee boasts an impressive record of over 770 journal papers and more than 860 patents. His groundbreaking research in metabolic engineering and biochemical engineering is highly acclaimed globally.
Throughout his 31 years as a professor at KAIST, Professor Lee has developed various metabolic engineering-based technologies and strategies. These advancements have been transferred to industries, facilitating the production of bulk chemicals, polymers, natural products, pharmaceuticals, and health functional foods. He has also founded companies and actively engages in advisory roles with various enterprises.
The International Metabolic Engineering Society (IMES) defines metabolic engineering as the manipulation of metabolic pathways in microorganisms or cells to produce useful substances (such as pharmaceuticals, biofuels, and chemical products). It utilizes tools like systems biology, synthetic biology, and computational modeling with the aim of enhancing the economic viability and sustainability of bio-based processes.
Furthermore, Professor Lee previously received the Merck Metabolic Engineering Award, a prominent international award in the field, in 2008. In 2018, he was honored with the Eni Award, often referred to as the Nobel Prize in energy, presented by the President of Italy.
Professor Sang Yup Lee remarked, "Metabolic engineering is a discipline that leads the current and future of biotechnology. It is a tremendous honor to receive this meaningful award at a time when the transition to a bio-based economy is accelerating. Together with my students and fellow researchers, we have generated numerous patents and transferred technologies to industry, and also established startups in the fields of biofuels, wound healing, and cosmetics. I will continue to pursue research that encompasses both fundamental research and technological commercialization."
The 'International Metabolic Engineering Society (IMES)' is a specialized society under the American Institute of Chemical Engineers. Its mission is to enable the production of various bio-based products, including pharmaceuticals, food additives, chemicals, and fuels, through metabolic engineering. The society hosts the Metabolic Engineering Conference biennially, offering researchers opportunities for knowledge exchange and collaboration.
2025.06.20 View 623 -
 KAIST Accelerates Synthetic Microbe Design by Discovering Novel Enzymes Using AI
< (From left) Professor Sang Yup Lee of the Department of Chemical and Biomolecular Engineering (top), Hongkeun Ji, PhD candidate of the Department of Chemical and Biomolecular Engineering (top), Ha Rim Kim, PhD candidate of the Department of Chemical and Biomolecular Engineering, and Dr. Gi Bae Kim of the BioProcess Engineering Research Center >
Enzymes are proteins that catalyze biochemical reactions within cells and play a pivotal role in metabolic processes. Accordingly, identifying the functions of novel enzymes is a critical task in the construction of microbial cell factories.
A KAIST research team has leveraged artificial intelligence (AI) to design novel enzymes that do not exist in nature, significantly accelerating microbial cell factory development and boosting the potential for next-generation biotechnological applications such as drug development and biofuel production.
KAIST (represented by President Kwang-Hyung Lee) announced on the 21st of April that Distinguished Professor Sang Yup Lee and his team from the Department of Chemical and Biomolecular Engineering have published a review titled “Enzyme Functional Classification Using Artificial Intelligence,” which outlines the advancement of AI-based enzyme function prediction technologies and analyzes how AI has contributed to the discovery and design of new enzymes.
Professor Lee’s team systematically reviewed the development of enzyme function prediction technologies utilizing machine learning and deep learning, offering a comprehensive analysis.
From sequence similarity-based prediction methods to the integration of convolutional neural networks (CNNs), recurrent neural networks (RNNs), graph neural networks (GNNs), and transformer-based large language models, the paper covers a broad range of AI applications. It analyzes how these technologies extract meaningful information from protein sequences and enhance prediction accuracy.
In particular, enzyme function prediction using deep learning goes beyond simple sequence similarity analysis. By automatically extracting structural and evolutionary features embedded in amino acid sequences, deep learning enables more precise predictions of catalytic functions.
This highlights the unique advantages of AI models compared to traditional bioinformatics approaches.
Moreover, the review suggests that the advancement of generative AI will move future research beyond predicting existing functions to generating entirely new enzymes with functions not found in nature. This shift is expected to profoundly impact the trajectory of biotechnology and synthetic biology.
< Figure 1. Extraction of enzyme characteristics and function prediction using various deep learning structures >
Ha Rim Kim, a Ph.D. candidate and co-first author from the Department of Chemical and Biomolecular Engineering, stated, “AI-based enzyme function prediction and enzyme design are highly important across various fields including metabolic engineering, synthetic biology, and healthcare.”
Distinguished Professor Sang Yup Lee added, “AI-powered enzyme function prediction shows the potential to solve diverse biological problems and will significantly contribute to accelerating research across the entire field.”
The review was published on March 28 in Trends in Biotechnology, a leading biotechnology journal issued by Cell Press.
※ Title: Enzyme Functional Classification Using Artificial Intelligence
※DOI: https://doi.org/10.1016/j.tibtech.2025.03.003
※ Author Information: Ha Rim Kim (KAIST, Co-first author), Hongkeun Ji (KAIST, Co-first author), Gi Bae Kim (KAIST, Third author), Sang Yup Lee (KAIST, Corresponding author)
This research was supported by the Ministry of Science and ICT under the project Development of Core Technologies for Advanced Synthetic Biology to Lead the Bio-Manufacturing Industry (aimed at replacing petroleum-based chemicals), and also by joint support from the Ministry of Science and ICT and the Ministry of Health and Welfare for the project Development of Novel Antibiotic Structures Using Deep Learning-Based Synthetic Biology.
2025.04.07 View 3101
KAIST Accelerates Synthetic Microbe Design by Discovering Novel Enzymes Using AI
< (From left) Professor Sang Yup Lee of the Department of Chemical and Biomolecular Engineering (top), Hongkeun Ji, PhD candidate of the Department of Chemical and Biomolecular Engineering (top), Ha Rim Kim, PhD candidate of the Department of Chemical and Biomolecular Engineering, and Dr. Gi Bae Kim of the BioProcess Engineering Research Center >
Enzymes are proteins that catalyze biochemical reactions within cells and play a pivotal role in metabolic processes. Accordingly, identifying the functions of novel enzymes is a critical task in the construction of microbial cell factories.
A KAIST research team has leveraged artificial intelligence (AI) to design novel enzymes that do not exist in nature, significantly accelerating microbial cell factory development and boosting the potential for next-generation biotechnological applications such as drug development and biofuel production.
KAIST (represented by President Kwang-Hyung Lee) announced on the 21st of April that Distinguished Professor Sang Yup Lee and his team from the Department of Chemical and Biomolecular Engineering have published a review titled “Enzyme Functional Classification Using Artificial Intelligence,” which outlines the advancement of AI-based enzyme function prediction technologies and analyzes how AI has contributed to the discovery and design of new enzymes.
Professor Lee’s team systematically reviewed the development of enzyme function prediction technologies utilizing machine learning and deep learning, offering a comprehensive analysis.
From sequence similarity-based prediction methods to the integration of convolutional neural networks (CNNs), recurrent neural networks (RNNs), graph neural networks (GNNs), and transformer-based large language models, the paper covers a broad range of AI applications. It analyzes how these technologies extract meaningful information from protein sequences and enhance prediction accuracy.
In particular, enzyme function prediction using deep learning goes beyond simple sequence similarity analysis. By automatically extracting structural and evolutionary features embedded in amino acid sequences, deep learning enables more precise predictions of catalytic functions.
This highlights the unique advantages of AI models compared to traditional bioinformatics approaches.
Moreover, the review suggests that the advancement of generative AI will move future research beyond predicting existing functions to generating entirely new enzymes with functions not found in nature. This shift is expected to profoundly impact the trajectory of biotechnology and synthetic biology.
< Figure 1. Extraction of enzyme characteristics and function prediction using various deep learning structures >
Ha Rim Kim, a Ph.D. candidate and co-first author from the Department of Chemical and Biomolecular Engineering, stated, “AI-based enzyme function prediction and enzyme design are highly important across various fields including metabolic engineering, synthetic biology, and healthcare.”
Distinguished Professor Sang Yup Lee added, “AI-powered enzyme function prediction shows the potential to solve diverse biological problems and will significantly contribute to accelerating research across the entire field.”
The review was published on March 28 in Trends in Biotechnology, a leading biotechnology journal issued by Cell Press.
※ Title: Enzyme Functional Classification Using Artificial Intelligence
※DOI: https://doi.org/10.1016/j.tibtech.2025.03.003
※ Author Information: Ha Rim Kim (KAIST, Co-first author), Hongkeun Ji (KAIST, Co-first author), Gi Bae Kim (KAIST, Third author), Sang Yup Lee (KAIST, Corresponding author)
This research was supported by the Ministry of Science and ICT under the project Development of Core Technologies for Advanced Synthetic Biology to Lead the Bio-Manufacturing Industry (aimed at replacing petroleum-based chemicals), and also by joint support from the Ministry of Science and ICT and the Ministry of Health and Welfare for the project Development of Novel Antibiotic Structures Using Deep Learning-Based Synthetic Biology.
2025.04.07 View 3101 -
 KAIST provides a comprehensive resource on microbial cell factories for sustainable chemical production
In silico analysis of five industrial microorganisms identifies optimal strains and metabolic engineering strategies for producing 235 valuable chemicals
Climate change and the depletion of fossil fuels have raised the global need for sustainable chemical production. In response to these environmental challenges, microbial cell factories are gaining attention as eco-friendly platforms for producing chemicals using renewable resources, while metabolic engineering technologies to enhance these cell factories are becoming crucial tools for maximizing production efficiency. However, difficulties in selecting suitable microbial strains and optimizing complex metabolic pathways continue to pose significant obstacles to practical industrial applications.
KAIST (President Kwang-Hyung Lee) announced on 27th of March that Distinguished Professor Sang Yup Lee’s research team in the Department of Chemical and Biomolecular Engineering comprehensively evaluated the production capabilities of various industrial microbial cell factories using in silico simulations and, based on these findings, identified the most suitable microbial strains for producing specific chemicals as well as optimal metabolic engineering strategies.
Previously, researchers attempted to determine the best strains and efficient metabolic engineering strategies among numerous microbial candidates through extensive biological experiments and meticulous verification processes. However, this approach required substantial time and costs. Recently, the introduction of genome-scale metabolic models (GEMs), which reconstruct the metabolic networks within an organism based on its entire genome information, has enabled systematic analysis of metabolic fluxes via computer simulations. This development offers a new way to overcome limitations of conventional experimental approaches, revolutionizing both strain selection and metabolic pathway design.
Accordingly, Professor Lee’s team at the Department of Chemical and Biomolecular Engineering, KAIST, evaluated the production capabilities of five representative industrial microorganisms—Escherichia coli, Saccharomyces cerevisiae, Bacillus subtilis, Corynebacterium glutamicum, and Pseudomonas putida—for 235 bio-based chemicals. Using GEMs, the researchers calculated both the maximum theoretical yields and the maximum achievable yields under industrial conditions for each chemical, thereby establishing criteria to identify the most suitable strains for each target compound.
< Figure 1. Outline of the strategy for improving microbial cell factories using a genome-scale metabolic model (GEM) >
The team specifically proposed strategies such as introducing heterologous enzyme reactions derived from other organisms and exchanging cofactors used by microbes to expand metabolic pathways. These strategies were shown to increase yields beyond the innate metabolic capacities of the microorganisms, resulting in higher production of industrially important chemicals such as mevalonic acid, propanol, fatty acids, and isoprenoids.
Moreover, by applying a computational approach to analyze metabolic fluxes in silico, the researchers suggested strategies for improving microbial strains to maximize the production of various chemicals. They quantitatively identified the relationships between specific enzyme reactions and target chemical production, as well as the relationships between enzymes and metabolites, determining which enzyme reactions should be up- or down-regulated. Through this, the team presented strategies not only to achieve high theoretical yields but also to maximize actual production capacities.
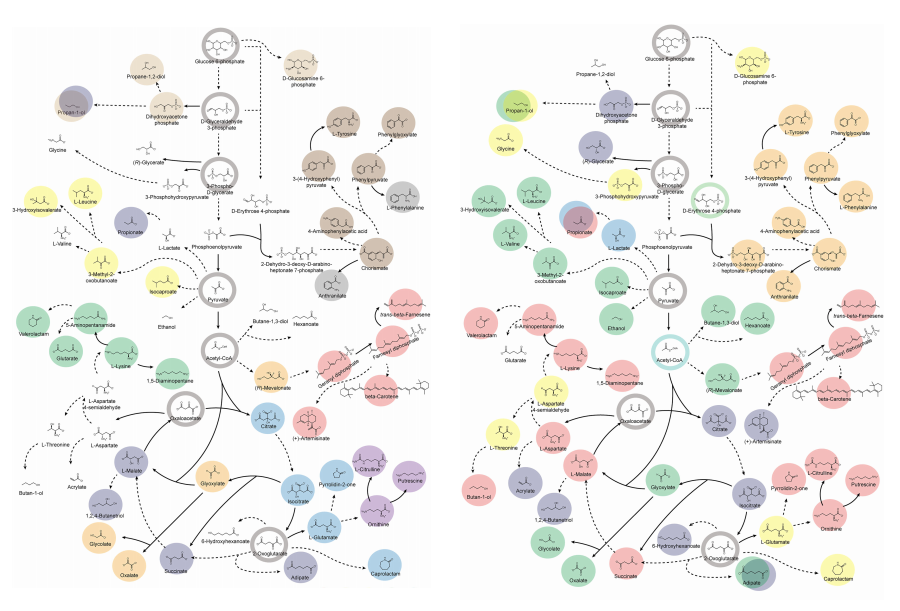
< Figure 2. Comparison of production routes and maximum yields of useful chemicals using representative industrial microorganisms >
Dr. Gi Bae Kim, the first author of this paper from the KAIST BioProcess Engineering Research Center, explained, “By introducing metabolic pathways derived from other organisms and exchanging cofactors, it is possible to design new microbial cell factories that surpass existing limitations. The strategies presented in this study will play a pivotal role in making microbial-based production processes more economical and efficient.” In addition, Distinguished Professor Sang Yup Lee noted, “This research serves as a key resource in the field of systems metabolic engineering, reducing difficulties in strain selection and pathway design, and enabling more efficient development of microbial cell factories. We expect it to greatly contribute to the future development of technologies for producing various eco-friendly chemicals, such as biofuels, bioplastics, and functional food materials.”
This research was conducted with the support from the Development of platform technologies of microbial cell factories for the next-generation biorefineries project and Development of advanced synthetic biology source technologies for leading the biomanufacturing industry project (Project Leader: Distinguished Professor Sang Yup Lee, KAIST) from National Research Foundation supported by the Korean Ministry of Science and ICT.
2025.03.27 View 3586
KAIST provides a comprehensive resource on microbial cell factories for sustainable chemical production
In silico analysis of five industrial microorganisms identifies optimal strains and metabolic engineering strategies for producing 235 valuable chemicals
Climate change and the depletion of fossil fuels have raised the global need for sustainable chemical production. In response to these environmental challenges, microbial cell factories are gaining attention as eco-friendly platforms for producing chemicals using renewable resources, while metabolic engineering technologies to enhance these cell factories are becoming crucial tools for maximizing production efficiency. However, difficulties in selecting suitable microbial strains and optimizing complex metabolic pathways continue to pose significant obstacles to practical industrial applications.
KAIST (President Kwang-Hyung Lee) announced on 27th of March that Distinguished Professor Sang Yup Lee’s research team in the Department of Chemical and Biomolecular Engineering comprehensively evaluated the production capabilities of various industrial microbial cell factories using in silico simulations and, based on these findings, identified the most suitable microbial strains for producing specific chemicals as well as optimal metabolic engineering strategies.
Previously, researchers attempted to determine the best strains and efficient metabolic engineering strategies among numerous microbial candidates through extensive biological experiments and meticulous verification processes. However, this approach required substantial time and costs. Recently, the introduction of genome-scale metabolic models (GEMs), which reconstruct the metabolic networks within an organism based on its entire genome information, has enabled systematic analysis of metabolic fluxes via computer simulations. This development offers a new way to overcome limitations of conventional experimental approaches, revolutionizing both strain selection and metabolic pathway design.
Accordingly, Professor Lee’s team at the Department of Chemical and Biomolecular Engineering, KAIST, evaluated the production capabilities of five representative industrial microorganisms—Escherichia coli, Saccharomyces cerevisiae, Bacillus subtilis, Corynebacterium glutamicum, and Pseudomonas putida—for 235 bio-based chemicals. Using GEMs, the researchers calculated both the maximum theoretical yields and the maximum achievable yields under industrial conditions for each chemical, thereby establishing criteria to identify the most suitable strains for each target compound.
< Figure 1. Outline of the strategy for improving microbial cell factories using a genome-scale metabolic model (GEM) >
The team specifically proposed strategies such as introducing heterologous enzyme reactions derived from other organisms and exchanging cofactors used by microbes to expand metabolic pathways. These strategies were shown to increase yields beyond the innate metabolic capacities of the microorganisms, resulting in higher production of industrially important chemicals such as mevalonic acid, propanol, fatty acids, and isoprenoids.
Moreover, by applying a computational approach to analyze metabolic fluxes in silico, the researchers suggested strategies for improving microbial strains to maximize the production of various chemicals. They quantitatively identified the relationships between specific enzyme reactions and target chemical production, as well as the relationships between enzymes and metabolites, determining which enzyme reactions should be up- or down-regulated. Through this, the team presented strategies not only to achieve high theoretical yields but also to maximize actual production capacities.
< Figure 2. Comparison of production routes and maximum yields of useful chemicals using representative industrial microorganisms >
Dr. Gi Bae Kim, the first author of this paper from the KAIST BioProcess Engineering Research Center, explained, “By introducing metabolic pathways derived from other organisms and exchanging cofactors, it is possible to design new microbial cell factories that surpass existing limitations. The strategies presented in this study will play a pivotal role in making microbial-based production processes more economical and efficient.” In addition, Distinguished Professor Sang Yup Lee noted, “This research serves as a key resource in the field of systems metabolic engineering, reducing difficulties in strain selection and pathway design, and enabling more efficient development of microbial cell factories. We expect it to greatly contribute to the future development of technologies for producing various eco-friendly chemicals, such as biofuels, bioplastics, and functional food materials.”
This research was conducted with the support from the Development of platform technologies of microbial cell factories for the next-generation biorefineries project and Development of advanced synthetic biology source technologies for leading the biomanufacturing industry project (Project Leader: Distinguished Professor Sang Yup Lee, KAIST) from National Research Foundation supported by the Korean Ministry of Science and ICT.
2025.03.27 View 3586 -
 KAIST Develops Eco-Friendly, Nylon-Like Plastic Using Microorganisms
Poly(ester amide) amide is a next-generation material that combines the advantages of PET (polyester) and nylon (polyamide), two widely used plastics. However, it could only be produced from fossil fuels, which posed environmental concerns. Using microorganisms, KAIST researchers have successfully developed a new bio-based plastic to replace conventional plastic.
KAIST (represented by President Kwang Hyung Lee) announced on the 20th of March that a research team led by Distinguished Professor Sang Yup Lee from the Department of Chemical and Biomolecular Engineering has developed microbial strains through systems metabolic engineering to produce various eco-friendly, bio-based poly(ester amide)s. The team collaborated with researchers from the Korea Research Institute of Chemical Technology (KRICT, President Young-Kook Lee) to analyze and confirm the properties of the resulting plastic.
Professor Sang Yup Lee’s research team designed new metabolic pathways that do not naturally exist in microorganisms, and developed a platform microbial strain capable of producing nine different types of poly(ester amide)s, including poly(3-hydroxybutyrate-ran-3-aminopropionate) and poly(3-hydroxybutyrate-ran-4-aminobutyrate).
Using glucose derived from abundant biomass sources such as waste wood and weeds, the team successfully produced poly(ester amide)s in an eco-friendly manner. The researchers also confirmed the potential for industrial-scale production by demonstrating high production efficiency (54.57 g/L) using fed-batch fermentation of the engineered strain.
In collaboration with researchers Haemin Jeong and Jihoon Shin from KRICT, the KAIST team analyzed the properties of the bio-based plastic and found that it exhibited characteristics similar to high-density polyethylene (HDPE). This means the new plastic is not only eco-friendly but also strong and durable enough to replace conventional plastics.
The engineered strains and strategies developed in this study are expected to be useful not only for producing various poly(ester amide)s but also for constructing metabolic pathways for the biosynthesis of other types of polymers.
Professor Sang Yup Lee stated, “This study is the first to demonstrate the possibility of producing poly(ester amide)s (plastics) through a renewable bio-based chemical process rather than relying on the petroleum-based chemical industry. We plan to further enhance the production yield and efficiency through continued research.”
The study was published online on March 17 in the international journal Nature Chemical Biology.
·Title: Biosynthesis of poly(ester amide)s in engineered Escherichia coli
·DOI: 10.1038/s41589-025-01842-2
·Authors: A total of seven authors including Tong Un Chae (KAIST, first author), So Young Choi (KAIST, second author), Da-Hee Ahn (KAIST, third author), Woo Dae Jang (KAIST, fourth author), Haemin Jeong (KRICT, fifth author), Jihoon Shin (KRICT, sixth author), and Sang Yup Lee (KAIST, corresponding author).
This research was supported by the Ministry of Science and ICT (MSIT) under the Eco-Friendly Chemical Technology Development Project as part of the "Next-Generation Biorefinery Technology Development to Lead the Bio-Chemical Industry" initiative (project led by Distinguished Professor Sang Yup Lee at KAIST).
2025.03.24 View 4758
KAIST Develops Eco-Friendly, Nylon-Like Plastic Using Microorganisms
Poly(ester amide) amide is a next-generation material that combines the advantages of PET (polyester) and nylon (polyamide), two widely used plastics. However, it could only be produced from fossil fuels, which posed environmental concerns. Using microorganisms, KAIST researchers have successfully developed a new bio-based plastic to replace conventional plastic.
KAIST (represented by President Kwang Hyung Lee) announced on the 20th of March that a research team led by Distinguished Professor Sang Yup Lee from the Department of Chemical and Biomolecular Engineering has developed microbial strains through systems metabolic engineering to produce various eco-friendly, bio-based poly(ester amide)s. The team collaborated with researchers from the Korea Research Institute of Chemical Technology (KRICT, President Young-Kook Lee) to analyze and confirm the properties of the resulting plastic.
Professor Sang Yup Lee’s research team designed new metabolic pathways that do not naturally exist in microorganisms, and developed a platform microbial strain capable of producing nine different types of poly(ester amide)s, including poly(3-hydroxybutyrate-ran-3-aminopropionate) and poly(3-hydroxybutyrate-ran-4-aminobutyrate).
Using glucose derived from abundant biomass sources such as waste wood and weeds, the team successfully produced poly(ester amide)s in an eco-friendly manner. The researchers also confirmed the potential for industrial-scale production by demonstrating high production efficiency (54.57 g/L) using fed-batch fermentation of the engineered strain.
In collaboration with researchers Haemin Jeong and Jihoon Shin from KRICT, the KAIST team analyzed the properties of the bio-based plastic and found that it exhibited characteristics similar to high-density polyethylene (HDPE). This means the new plastic is not only eco-friendly but also strong and durable enough to replace conventional plastics.
The engineered strains and strategies developed in this study are expected to be useful not only for producing various poly(ester amide)s but also for constructing metabolic pathways for the biosynthesis of other types of polymers.
Professor Sang Yup Lee stated, “This study is the first to demonstrate the possibility of producing poly(ester amide)s (plastics) through a renewable bio-based chemical process rather than relying on the petroleum-based chemical industry. We plan to further enhance the production yield and efficiency through continued research.”
The study was published online on March 17 in the international journal Nature Chemical Biology.
·Title: Biosynthesis of poly(ester amide)s in engineered Escherichia coli
·DOI: 10.1038/s41589-025-01842-2
·Authors: A total of seven authors including Tong Un Chae (KAIST, first author), So Young Choi (KAIST, second author), Da-Hee Ahn (KAIST, third author), Woo Dae Jang (KAIST, fourth author), Haemin Jeong (KRICT, fifth author), Jihoon Shin (KRICT, sixth author), and Sang Yup Lee (KAIST, corresponding author).
This research was supported by the Ministry of Science and ICT (MSIT) under the Eco-Friendly Chemical Technology Development Project as part of the "Next-Generation Biorefinery Technology Development to Lead the Bio-Chemical Industry" initiative (project led by Distinguished Professor Sang Yup Lee at KAIST).
2025.03.24 View 4758 -
 “Cross-Generation Collaborative Labs” for Semiconductor, Chemistry, and Computer Science Opened
< Photo of Professor Hoi-Jun Yoo (center) of the School of Electrical Engineering at the signboard unveiling ceremony >
KAIST held a ceremony to mark the opening of three additional ‘Cross-Generation Collaborative Labs’ on the morning of January 7th, 2025.
The “Next-Generation AI Semiconductor System Lab” by Professor Hoi-Jun Yoo of the School of Electrical Engineering, the “Molecular Spectroscopy and Chemical Dynamics Lab” by Professor Sang Kyu Kim of the Department of Chemistry, and the “Advanced Data Computing Lab” by Professor Sue Bok Moon of the School of Computer Science are the three new labs given the honored titled of the “Cross-Generation Collaborative Lab”.
The Cross-Generation Collaborative Lab is KAIST’s unique system that was set up to facilitate the collaboration between retiring professors and junior professors to continue the achievements and know-how the elders have accumulated over their academic career. Since its introduction in 2018, nine labs have been named to be the Cross-Generation Labs, and this year’s new addition brings the total up to twelve.
The ‘Next-Generation AI Semiconductor System Lab’ led by Professor Hoi-Jun Yoo will be operated by Professor Joo-Young Kim of the same school.
Professor Hoi-Jun Yoo is a world-renowned scholar with outstanding research achievements in the field of on-device AI semiconductor design. Professor Joo-Young Kim is an up-and-coming researcher studying large language models and design of AI semiconductors for server computers, and is currently researching technologies to design PIM (Processing-in-Memory), a core technology in the field of AI semiconductors.
Their research goal is to systematically collaborate and transfer next-generation AI semiconductor design technology, including brain-mimicking AI algorithms such as deep neural networks and generative AI, to integrate core technologies, and to maximize the usability of R&D outputs, thereby further solidifying the position of Korean AI semiconductor companies in the global market.
Professor Hoi-Jun Yoo said, “I believe that, we will be able to present a development direction of for the next-generation AI semiconductors industries at home and abroad through collaborative research and play a key role in transferring and expanding global leadership.”
< Professor Sang Kyu Kim of the Department of Chemistry (middle), at the signboard unveiling ceremony for his laboratory >
The “Molecular Spectroscopy and Chemical Dynamics Laboratory”, where Professor Sang Kyu Kim of the Department of Chemistry is in charge, will be operated by Professor Tae Kyu Kim of the same department, and another professor in the field of spectroscopy and dynamics will join in the future.
Professor Sang Kyu Kim has secured technologies for developing unique experimental equipment based on ultrashort lasers and supersonic molecular beams, and is a world leader who has been creatively pioneering new fields of experimental physical chemistry.
The research goal is to describe chemical reactions and verify from a quantum mechanical perspective and introduce new theories and technologies to pursue a complete understanding of the principles of chemical reactions. In addition, the accompanying basic scientific knowledge will be applied to the design of new materials.
Professor Sang Kyu Kim said, “I am very happy to be able to pass on the research infrastructure to the next generation through this system, and I will continue to nurture it to grow into a world-class research lab through trans-generational collaborative research.”
< Photo of Professor Sue Bok Moon (center) at the signboard unveiling ceremony by the School of Computing >
Lastly, the “Advanced Data Computing Lab” led by Professor Sue Bok Moon is joined by Professor Mee Young Cha of the same school and Professor Wonjae Lee of the Graduate School of Culture Technology.
Professor Sue Bok Moon showed the infinite possibilities of large-scale data-based social network research through Cyworld, YouTube, and Twitter, and had a great influence on related fields beyond the field of computer science.
Professor Mee Young Cha is a data scientist who analyzes difficult social issues such as misinformation, poverty, and disaster detection using big data-based AI. She is the first Korean to be recognized for her achievements as the director of the Max Planck Institute in Germany, a world-class basic science research institute. Therefore, there is high expectation for synergy effects from overseas collaborative research and technology transfer and sharing among the participating professors of the collaborative research lab. Professor Wonjae Lee is researching dynamic interaction analysis between science and technology using structural topic models.
They plan to conduct research aimed at improving the analysis and understanding of negative influences occurring online, and in particular, developing a hateful precursor detection model using emotions and morality to preemptively block hateful expressions.
Professor Sue Bok Moon said, “Through this collaborative research lab, we will play a key role in conducting in-depth collaborative research on unexpected negative influences in the AI era so that we can have a high level of competitiveness worldwide.”
The ceremonies for the unveiling of the new Cross-Generation Collaborative Lab signboard were held in front of each lab from 10:00 AM on the 7th, in the attendance of President Kwang Hyung Lee, Senior Vice President for Research Sang Yup Lee, and other key officials of KAIST and the new staff members to join the laboratories.
2025.01.07 View 4794
“Cross-Generation Collaborative Labs” for Semiconductor, Chemistry, and Computer Science Opened
< Photo of Professor Hoi-Jun Yoo (center) of the School of Electrical Engineering at the signboard unveiling ceremony >
KAIST held a ceremony to mark the opening of three additional ‘Cross-Generation Collaborative Labs’ on the morning of January 7th, 2025.
The “Next-Generation AI Semiconductor System Lab” by Professor Hoi-Jun Yoo of the School of Electrical Engineering, the “Molecular Spectroscopy and Chemical Dynamics Lab” by Professor Sang Kyu Kim of the Department of Chemistry, and the “Advanced Data Computing Lab” by Professor Sue Bok Moon of the School of Computer Science are the three new labs given the honored titled of the “Cross-Generation Collaborative Lab”.
The Cross-Generation Collaborative Lab is KAIST’s unique system that was set up to facilitate the collaboration between retiring professors and junior professors to continue the achievements and know-how the elders have accumulated over their academic career. Since its introduction in 2018, nine labs have been named to be the Cross-Generation Labs, and this year’s new addition brings the total up to twelve.
The ‘Next-Generation AI Semiconductor System Lab’ led by Professor Hoi-Jun Yoo will be operated by Professor Joo-Young Kim of the same school.
Professor Hoi-Jun Yoo is a world-renowned scholar with outstanding research achievements in the field of on-device AI semiconductor design. Professor Joo-Young Kim is an up-and-coming researcher studying large language models and design of AI semiconductors for server computers, and is currently researching technologies to design PIM (Processing-in-Memory), a core technology in the field of AI semiconductors.
Their research goal is to systematically collaborate and transfer next-generation AI semiconductor design technology, including brain-mimicking AI algorithms such as deep neural networks and generative AI, to integrate core technologies, and to maximize the usability of R&D outputs, thereby further solidifying the position of Korean AI semiconductor companies in the global market.
Professor Hoi-Jun Yoo said, “I believe that, we will be able to present a development direction of for the next-generation AI semiconductors industries at home and abroad through collaborative research and play a key role in transferring and expanding global leadership.”
< Professor Sang Kyu Kim of the Department of Chemistry (middle), at the signboard unveiling ceremony for his laboratory >
The “Molecular Spectroscopy and Chemical Dynamics Laboratory”, where Professor Sang Kyu Kim of the Department of Chemistry is in charge, will be operated by Professor Tae Kyu Kim of the same department, and another professor in the field of spectroscopy and dynamics will join in the future.
Professor Sang Kyu Kim has secured technologies for developing unique experimental equipment based on ultrashort lasers and supersonic molecular beams, and is a world leader who has been creatively pioneering new fields of experimental physical chemistry.
The research goal is to describe chemical reactions and verify from a quantum mechanical perspective and introduce new theories and technologies to pursue a complete understanding of the principles of chemical reactions. In addition, the accompanying basic scientific knowledge will be applied to the design of new materials.
Professor Sang Kyu Kim said, “I am very happy to be able to pass on the research infrastructure to the next generation through this system, and I will continue to nurture it to grow into a world-class research lab through trans-generational collaborative research.”
< Photo of Professor Sue Bok Moon (center) at the signboard unveiling ceremony by the School of Computing >
Lastly, the “Advanced Data Computing Lab” led by Professor Sue Bok Moon is joined by Professor Mee Young Cha of the same school and Professor Wonjae Lee of the Graduate School of Culture Technology.
Professor Sue Bok Moon showed the infinite possibilities of large-scale data-based social network research through Cyworld, YouTube, and Twitter, and had a great influence on related fields beyond the field of computer science.
Professor Mee Young Cha is a data scientist who analyzes difficult social issues such as misinformation, poverty, and disaster detection using big data-based AI. She is the first Korean to be recognized for her achievements as the director of the Max Planck Institute in Germany, a world-class basic science research institute. Therefore, there is high expectation for synergy effects from overseas collaborative research and technology transfer and sharing among the participating professors of the collaborative research lab. Professor Wonjae Lee is researching dynamic interaction analysis between science and technology using structural topic models.
They plan to conduct research aimed at improving the analysis and understanding of negative influences occurring online, and in particular, developing a hateful precursor detection model using emotions and morality to preemptively block hateful expressions.
Professor Sue Bok Moon said, “Through this collaborative research lab, we will play a key role in conducting in-depth collaborative research on unexpected negative influences in the AI era so that we can have a high level of competitiveness worldwide.”
The ceremonies for the unveiling of the new Cross-Generation Collaborative Lab signboard were held in front of each lab from 10:00 AM on the 7th, in the attendance of President Kwang Hyung Lee, Senior Vice President for Research Sang Yup Lee, and other key officials of KAIST and the new staff members to join the laboratories.
2025.01.07 View 4794 -
 KAIST Researchers Suggest an Extraordinary Alternative to Petroleum-based PET - Bacteria!
< (From left) Dr. Cindy Pricilia, Ph.D. Candidate Cheon Woo Moon, Distinguished Professor Sang Yup Lee >
Currently, the world is suffering from environmental problems caused by plastic waste. The KAIST research team has succeeded in producing a microbial-based plastic that is biodegradable and can replace existing PET bottles, making it a hot topic.
Our university announced on the 7th of November that the research team of Distinguished Professor Sang Yup Lee of the Department of Chemical and Biomolecular Engineering has succeeded in developing a microbial strain that efficiently produces pseudoaromatic polyester monomer to replace polyethylene terephthalate (PET) using systems metabolic engineering.
Pseudoaromatic dicarboxylic acids have better physical properties and higher biodegradability than aromatic polyester (PET) when synthesized as polymers, and are attracting attention as an eco-friendly monomer* that can be synthesized into polymers. The production of pseudoaromatic dicarboxylic acids through chemical methods has the problems of low yield and selectivity, complex reaction conditions, and the generation of hazardous waste.
*Monomer: A material for making polymers, which is used to synthesize polymers by polymerizing monomers together
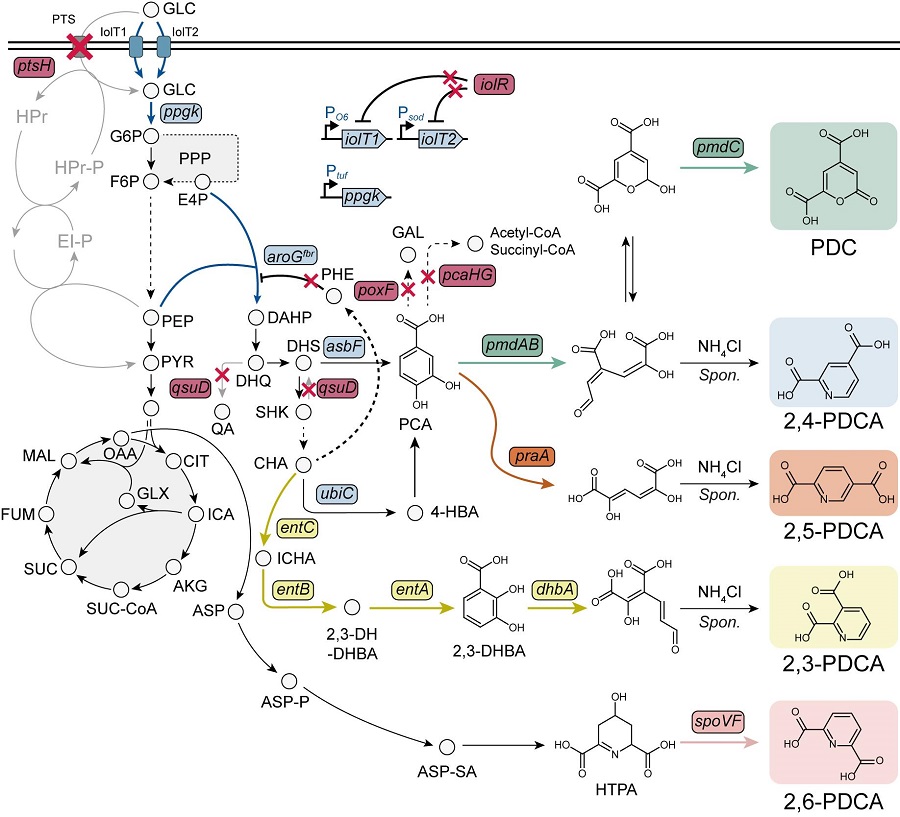
< Figure. Overview of pseudoaromatic dicarboxylic acid production using metabolically engineered C. glutamicum. >
To solve this problem, Professor Sang Yup Lee's research team used metabolic engineering to develop a microbial strain that efficiently produces five types of pseudoaromatic dicarboxylic acids, including 2-pyrone-4,6-dicarboxylic acid and four types of pyridine dicarboxylic acids (2,3-, 2,4-, 2,5-, 2,6-pyridine dicarboxylic acids), in Corynebacterium, a bacterium mainly used for amino acid production.
The research team used metabolic engineering techniques to build a platform microbial strain that enhances the metabolic flow of protocatechuic acid, which is used as a precursor for several pseudoaromatic dicarboxylic acids, and prevents the loss of precursors.
Based on this, the genetic manipulation target was discovered through transcriptome analysis, producing 76.17 g/L of 2-pyrone-4,6-dicarboxylic acid, and by newly discovering and constructing three types of pyridine dicarboxylic acid production metabolic pathways, successfully producing 2.79 g/L of 2,3-pyridine dicarboxylic acid, 0.49 g/L of 2,4-pyridine dicarboxylic acid, and 1.42 g/L of 2,5-pyridine dicarboxylic acid.
In addition, the research team confirmed the production of 15.01 g/L through the construction and reinforcement of the 2,6-pyridine dicarboxylic acid biosynthesis pathway, successfully producing a total of five similar aromatic dicarboxylic acids with high efficiency.
In conclusion, the team succeeded in producing 2,4-, 2,5-, and 2,6-pyridine dicarboxylic acids at the world's highest concentration. In particular, 2,4-, 2,5-pyridine dicarboxylic acid achieved production on the scale of g/L, which was previously produced in extremely small amounts (mg/L).
Based on this study, it is expected that it will be applied to various polyester production industrial processes, and it is also expected that it will be actively utilized in research on the production of similar aromatic polyesters.
Professor Sang Yup Lee, the corresponding author, said, “The significance lies in the fact that we have developed an eco-friendly technology that efficiently produces similar aromatic polyester monomers based on microorganisms,” and “This study will help the microorganism-based bio-monomer industry replace the petrochemical-based chemical industry in the future.”
The results of this study were published in the international academic journal, the Proceedings of the National Academy of Sciences of United States of America (PNAS) on October 30th.
※ Paper title: Metabolic engineering of Corynebacterium glutamicum for the production of pyrone and pyridine dicarboxylic acids
※ Author information: Jae Sung Cho (co-first author), Zi Wei Luo (co-first author), Cheon Woo Moon (co-first author), Cindy Prabowo (co-author), Sang Yup Lee (corresponding author) - a total of 5 people
This study was conducted with the support of the Development of Next-generation Biorefinery Platform Technologies for Leading Bio-based Chemicals Industry Project and the Development of Platform Technologies of Microbial Cell Factories for the Next-generation Biorefineries Project (Project leader: Professor Sang Yup Lee) from the National Research Foundation supported by the Ministry of Science and Technology and ICT of Korea.
2024.11.08 View 8874
KAIST Researchers Suggest an Extraordinary Alternative to Petroleum-based PET - Bacteria!
< (From left) Dr. Cindy Pricilia, Ph.D. Candidate Cheon Woo Moon, Distinguished Professor Sang Yup Lee >
Currently, the world is suffering from environmental problems caused by plastic waste. The KAIST research team has succeeded in producing a microbial-based plastic that is biodegradable and can replace existing PET bottles, making it a hot topic.
Our university announced on the 7th of November that the research team of Distinguished Professor Sang Yup Lee of the Department of Chemical and Biomolecular Engineering has succeeded in developing a microbial strain that efficiently produces pseudoaromatic polyester monomer to replace polyethylene terephthalate (PET) using systems metabolic engineering.
Pseudoaromatic dicarboxylic acids have better physical properties and higher biodegradability than aromatic polyester (PET) when synthesized as polymers, and are attracting attention as an eco-friendly monomer* that can be synthesized into polymers. The production of pseudoaromatic dicarboxylic acids through chemical methods has the problems of low yield and selectivity, complex reaction conditions, and the generation of hazardous waste.
*Monomer: A material for making polymers, which is used to synthesize polymers by polymerizing monomers together
< Figure. Overview of pseudoaromatic dicarboxylic acid production using metabolically engineered C. glutamicum. >
To solve this problem, Professor Sang Yup Lee's research team used metabolic engineering to develop a microbial strain that efficiently produces five types of pseudoaromatic dicarboxylic acids, including 2-pyrone-4,6-dicarboxylic acid and four types of pyridine dicarboxylic acids (2,3-, 2,4-, 2,5-, 2,6-pyridine dicarboxylic acids), in Corynebacterium, a bacterium mainly used for amino acid production.
The research team used metabolic engineering techniques to build a platform microbial strain that enhances the metabolic flow of protocatechuic acid, which is used as a precursor for several pseudoaromatic dicarboxylic acids, and prevents the loss of precursors.
Based on this, the genetic manipulation target was discovered through transcriptome analysis, producing 76.17 g/L of 2-pyrone-4,6-dicarboxylic acid, and by newly discovering and constructing three types of pyridine dicarboxylic acid production metabolic pathways, successfully producing 2.79 g/L of 2,3-pyridine dicarboxylic acid, 0.49 g/L of 2,4-pyridine dicarboxylic acid, and 1.42 g/L of 2,5-pyridine dicarboxylic acid.
In addition, the research team confirmed the production of 15.01 g/L through the construction and reinforcement of the 2,6-pyridine dicarboxylic acid biosynthesis pathway, successfully producing a total of five similar aromatic dicarboxylic acids with high efficiency.
In conclusion, the team succeeded in producing 2,4-, 2,5-, and 2,6-pyridine dicarboxylic acids at the world's highest concentration. In particular, 2,4-, 2,5-pyridine dicarboxylic acid achieved production on the scale of g/L, which was previously produced in extremely small amounts (mg/L).
Based on this study, it is expected that it will be applied to various polyester production industrial processes, and it is also expected that it will be actively utilized in research on the production of similar aromatic polyesters.
Professor Sang Yup Lee, the corresponding author, said, “The significance lies in the fact that we have developed an eco-friendly technology that efficiently produces similar aromatic polyester monomers based on microorganisms,” and “This study will help the microorganism-based bio-monomer industry replace the petrochemical-based chemical industry in the future.”
The results of this study were published in the international academic journal, the Proceedings of the National Academy of Sciences of United States of America (PNAS) on October 30th.
※ Paper title: Metabolic engineering of Corynebacterium glutamicum for the production of pyrone and pyridine dicarboxylic acids
※ Author information: Jae Sung Cho (co-first author), Zi Wei Luo (co-first author), Cheon Woo Moon (co-first author), Cindy Prabowo (co-author), Sang Yup Lee (corresponding author) - a total of 5 people
This study was conducted with the support of the Development of Next-generation Biorefinery Platform Technologies for Leading Bio-based Chemicals Industry Project and the Development of Platform Technologies of Microbial Cell Factories for the Next-generation Biorefineries Project (Project leader: Professor Sang Yup Lee) from the National Research Foundation supported by the Ministry of Science and Technology and ICT of Korea.
2024.11.08 View 8874 -
 KAIST finds ways for Bacteria to produce PET-like materials
Among various eco-friendly polymers, polyhydroxyalkanoates (PHA) stand out for their excellent biodegradability and biocompatibility. They decompose naturally in soil and marine environments and are used in applications such as food packaging and medical products. However, natural PHA produced to date has faced challenges meeting various physical property requirements, such as durability and thermal stability, and has been limited in its commercial application due to low production concentrations. In light of this, KAIST researchers have recently developed a technology that could play a crucial role in solving the environmental pollution problem caused by plastics.
KAIST (represented by President Kwang-Hyung Lee) announced on August 26th that a research team led by Distinguished Professor Sang Yup Lee from the Department of Chemical and Biomolecular Engineering, including Dr. Youngjoon Lee and master's student Minju Kang, has successfully developed a microbial strain that efficiently produces aromatic polyester* using systems metabolic engineering.
※ Aromatic polyester: A polymer containing aromatic compounds (specific carbon ring structures like benzene) and ester bonds.
In this study, the research team used metabolic engineering to enhance the metabolic flux of the biosynthetic pathway for the aromatic monomer phenyllactate (PhLA) in E. coli. They manipulated the metabolic pathway to increase the polymer fraction accumulated within the cells and employed computer simulations to predict the structure of PHA synthase and improve the enzyme based on the structure-function relationship.
Through subsequent fermentation optimization, the team achieved the world’s highest concentration (12.3±0.1 g/L) for the efficient production of poly (PhLA) and successfully produced polyester through a 30L scale fed-batch fermentation, demonstrating the possibility of industrial-level production. The produced aromatic polyesters showed enhanced thermal properties, improved mechanical properties, and potential for use as drug delivery carriers.
< Figure 1. Development schematics of aromatic polyester producing microorganisms >
The research team also demonstrated that an exogenous phasin protein* plays a crucial role in increasing the intracellular polymer accumulation fraction, which is directly related to the economic feasibility and efficiency of non-natural PHA production. They improved PHA synthase using a rational enzyme design approach, predicting the three-dimensional structure of the enzyme through homology modeling (a method of predicting the three-dimensional structure of a new protein based on the structure of similar proteins) followed by molecular docking simulations (simulations that predict how well a monomer can bind to an enzyme) and molecular dynamics simulations (simulations that predict how molecules move and interact over time) to upgrade the enzyme into a mutant enzyme with enhanced monomer polymerization efficiency.
※ Exogenous phasin protein: Phasin is a protein related to PHA production, interacting with the cytoplasmic environment on the surface of granules of PHA, and playing a role in polymer accumulation and controlling the number and size of granules. In this study, genes encoding phasin proteins derived from various natural PHA-producing microorganisms were selected and introduced.
Dr. Youngjoon Lee, co-first author of the paper, explained, "The significance of this study lies in the fact that we have achieved the world's highest concentration of microbial-based aromatic polyester production using eco-friendly materials and methods. This technology is expected to play a crucial role in addressing the environmental pollution caused by plastics." Distinguished Professor Sang Yup Lee added, "This study, which presents various strategies for the high-efficiency production of useful polymers via systems metabolic engineering, is expected to make a significant contribution to solving climate change issues, particularly the recent plastic problem."
< Figure 2. Detailed development strategy for aromatic polyester producing microorganisms >
The research findings were published on August 21st in Trends in Biotechnology, published by Cell, an international academic journal.
※ Paper Title: “Microbial production of an aromatic homopolyester”
※ Author Information: Youngjoon Lee (KAIST, co-first author), Minju Kang (KAIST, co-first author), Woo Dae Jang (KAIST, second author), So Young Choi (KAIST, third author), Jung Eun Yang (KAIST, fourth author), Sang Yup Lee (KAIST, corresponding author), totaling six authors.
This research was supported by the "Development of Next-Generation Biorefinery Platform Technologies for Leading the Bio-based Chemicals Industry" project led by Distinguished Professor Sang Yup Lee at KAIST, under the eco-friendly chemical technology development project aimed at substituting petroleum, funded by the Ministry of Science and ICT. It was also supported by the "Development of Platform Technology for the Production of Novel Aromatic Bioplastic Using Microbial Cell Factories" project (Project Leader: Si Jae Park, Ewha Woman’s University).
2024.08.28 View 6641
KAIST finds ways for Bacteria to produce PET-like materials
Among various eco-friendly polymers, polyhydroxyalkanoates (PHA) stand out for their excellent biodegradability and biocompatibility. They decompose naturally in soil and marine environments and are used in applications such as food packaging and medical products. However, natural PHA produced to date has faced challenges meeting various physical property requirements, such as durability and thermal stability, and has been limited in its commercial application due to low production concentrations. In light of this, KAIST researchers have recently developed a technology that could play a crucial role in solving the environmental pollution problem caused by plastics.
KAIST (represented by President Kwang-Hyung Lee) announced on August 26th that a research team led by Distinguished Professor Sang Yup Lee from the Department of Chemical and Biomolecular Engineering, including Dr. Youngjoon Lee and master's student Minju Kang, has successfully developed a microbial strain that efficiently produces aromatic polyester* using systems metabolic engineering.
※ Aromatic polyester: A polymer containing aromatic compounds (specific carbon ring structures like benzene) and ester bonds.
In this study, the research team used metabolic engineering to enhance the metabolic flux of the biosynthetic pathway for the aromatic monomer phenyllactate (PhLA) in E. coli. They manipulated the metabolic pathway to increase the polymer fraction accumulated within the cells and employed computer simulations to predict the structure of PHA synthase and improve the enzyme based on the structure-function relationship.
Through subsequent fermentation optimization, the team achieved the world’s highest concentration (12.3±0.1 g/L) for the efficient production of poly (PhLA) and successfully produced polyester through a 30L scale fed-batch fermentation, demonstrating the possibility of industrial-level production. The produced aromatic polyesters showed enhanced thermal properties, improved mechanical properties, and potential for use as drug delivery carriers.
< Figure 1. Development schematics of aromatic polyester producing microorganisms >
The research team also demonstrated that an exogenous phasin protein* plays a crucial role in increasing the intracellular polymer accumulation fraction, which is directly related to the economic feasibility and efficiency of non-natural PHA production. They improved PHA synthase using a rational enzyme design approach, predicting the three-dimensional structure of the enzyme through homology modeling (a method of predicting the three-dimensional structure of a new protein based on the structure of similar proteins) followed by molecular docking simulations (simulations that predict how well a monomer can bind to an enzyme) and molecular dynamics simulations (simulations that predict how molecules move and interact over time) to upgrade the enzyme into a mutant enzyme with enhanced monomer polymerization efficiency.
※ Exogenous phasin protein: Phasin is a protein related to PHA production, interacting with the cytoplasmic environment on the surface of granules of PHA, and playing a role in polymer accumulation and controlling the number and size of granules. In this study, genes encoding phasin proteins derived from various natural PHA-producing microorganisms were selected and introduced.
Dr. Youngjoon Lee, co-first author of the paper, explained, "The significance of this study lies in the fact that we have achieved the world's highest concentration of microbial-based aromatic polyester production using eco-friendly materials and methods. This technology is expected to play a crucial role in addressing the environmental pollution caused by plastics." Distinguished Professor Sang Yup Lee added, "This study, which presents various strategies for the high-efficiency production of useful polymers via systems metabolic engineering, is expected to make a significant contribution to solving climate change issues, particularly the recent plastic problem."
< Figure 2. Detailed development strategy for aromatic polyester producing microorganisms >
The research findings were published on August 21st in Trends in Biotechnology, published by Cell, an international academic journal.
※ Paper Title: “Microbial production of an aromatic homopolyester”
※ Author Information: Youngjoon Lee (KAIST, co-first author), Minju Kang (KAIST, co-first author), Woo Dae Jang (KAIST, second author), So Young Choi (KAIST, third author), Jung Eun Yang (KAIST, fourth author), Sang Yup Lee (KAIST, corresponding author), totaling six authors.
This research was supported by the "Development of Next-Generation Biorefinery Platform Technologies for Leading the Bio-based Chemicals Industry" project led by Distinguished Professor Sang Yup Lee at KAIST, under the eco-friendly chemical technology development project aimed at substituting petroleum, funded by the Ministry of Science and ICT. It was also supported by the "Development of Platform Technology for the Production of Novel Aromatic Bioplastic Using Microbial Cell Factories" project (Project Leader: Si Jae Park, Ewha Woman’s University).
2024.08.28 View 6641 -
 KAIST Develops Microbial Liquid Egg Substitute
A team of researchers published a paper on developing a substitute for eggs using microorganisms, grabbing international attention. It is expected that the development of egg substitutes using non-animal raw materials will solve the problems of factory farming, which causes problems like increased emission of greenhouse gas and waste, and contribute to building a sustainable food system that allows easy protein intake.
KAIST (President Kwang-Hyung Lee) announced that Research Professor Kyeong Rok Choi from the Biological Process Research Center and Distinguished Professor Sang Yup Lee from the Department of Chemical and Biomolecular Engineering have published a paper on the development of an "Eco-Friendly Liquid Egg Substitute Derived from Microorganisms."
Eggs play a crucial role in various culinary applications due to their unique physicochemical properties such as gelling, foaming, and emulsifying, while also providing essential nutrients. However, traditional egg production is not only unethical and resource-intensive but also has significant environmental impacts such as greenhouse gas emissions and waste issues. Additionally, factors such as wars and trade regulations have led to significant increases in egg prices, highlighting food security concerns. In response to these issues, there has been growing interest in egg substitutes made from non-animal sources to establish a sustainable food system.
Although there has been progress in developing non-animal protein-based egg substitutes, no substitute has been able to fully replicate the essential functional properties of liquid eggs, such as gelling and foaming, while also providing complete nutrition. In this context, the research team aimed to develop a liquid egg substitute using microbial biomass, which has a protein content comparable to that of meat per unit dry mass. Various microorganisms, such as yeast, Bacillus, lactic acid bacteria, and other probiotics, have been proven safe through long-term human consumption. Microbial biomass requires fewer resources like water and land during production, and possesses high-quality nutrients, making it a promising sustainable food resource.
< Figure 1. Comparison of heat treatment results of microbial pellets and microbial lysates >
However, the semi-solid microbial biomass recovered through microbial cultivation was observed to turn liquid upon heating, unlike liquid egg. To address this, the research team devised a microbial lysate by breaking down the cell walls and cell membranes of microorganisms, which correspond to the eggshell. They found that the microbial lysate's proteins coagulated when heated and formed a gel similar to that of liquid egg. The gel formed from the heated microbial lysate was found to have microscopic structures and physical properties similar to those of boiled eggs. The addition of microbial-derived edible enzymes or plant-based materials allowed for the adjustment of its properties, enabling the creation of various textures.
Furthermore, the researchers demonstrated that the microbial lysate could form stable foams widely used in baking, such as meringues (made from egg whites). They successfully baked meringue cookies using this lysate, showing its potential as a functional liquid egg substitute.
Distinguished Professor Sang Yup Lee stated, "This substitute has excellent nutritional components, making it suitable for regular food consumption. It is especially promising as emergency food for long-term space travel, wartime situations, and other emergencies. More importantly, it contributes to securing a sustainable food system."
< Figure 2. Example of foaming ability of microbial lysate and meringue cookie production >
< Figure 3. Example of foaming ability of microbial lysate and meringue cookie production >
The paper was published online in the journal npj Science of Food, issued by Nature.
- Paper Title: Microbial lysates repurposed as liquid egg substitutes
- Authors: Kyeong Rok Choi (first author), Da-Hee Ahn, Seok Yeong Jung, YuHyun Lee, and Sang Yup Lee (corresponding author)
This research was supported by the Ministry of Science and ICT's project for developing eco-friendly chemical technologies to replace petroleum (Project Leader: Distinguished Professor Sang Yup Lee, KAIST) and the Rural Development Administration's Agricultural Microorganisms Project Group (Director: Professor Pan-sik Jang, Seoul National University) for developing protein production technology from inorganic substances through microbial metabolic system control (Project Leader: Research Professor Kyeong Rok Choi, KAIST).
2024.07.05 View 6883
KAIST Develops Microbial Liquid Egg Substitute
A team of researchers published a paper on developing a substitute for eggs using microorganisms, grabbing international attention. It is expected that the development of egg substitutes using non-animal raw materials will solve the problems of factory farming, which causes problems like increased emission of greenhouse gas and waste, and contribute to building a sustainable food system that allows easy protein intake.
KAIST (President Kwang-Hyung Lee) announced that Research Professor Kyeong Rok Choi from the Biological Process Research Center and Distinguished Professor Sang Yup Lee from the Department of Chemical and Biomolecular Engineering have published a paper on the development of an "Eco-Friendly Liquid Egg Substitute Derived from Microorganisms."
Eggs play a crucial role in various culinary applications due to their unique physicochemical properties such as gelling, foaming, and emulsifying, while also providing essential nutrients. However, traditional egg production is not only unethical and resource-intensive but also has significant environmental impacts such as greenhouse gas emissions and waste issues. Additionally, factors such as wars and trade regulations have led to significant increases in egg prices, highlighting food security concerns. In response to these issues, there has been growing interest in egg substitutes made from non-animal sources to establish a sustainable food system.
Although there has been progress in developing non-animal protein-based egg substitutes, no substitute has been able to fully replicate the essential functional properties of liquid eggs, such as gelling and foaming, while also providing complete nutrition. In this context, the research team aimed to develop a liquid egg substitute using microbial biomass, which has a protein content comparable to that of meat per unit dry mass. Various microorganisms, such as yeast, Bacillus, lactic acid bacteria, and other probiotics, have been proven safe through long-term human consumption. Microbial biomass requires fewer resources like water and land during production, and possesses high-quality nutrients, making it a promising sustainable food resource.
< Figure 1. Comparison of heat treatment results of microbial pellets and microbial lysates >
However, the semi-solid microbial biomass recovered through microbial cultivation was observed to turn liquid upon heating, unlike liquid egg. To address this, the research team devised a microbial lysate by breaking down the cell walls and cell membranes of microorganisms, which correspond to the eggshell. They found that the microbial lysate's proteins coagulated when heated and formed a gel similar to that of liquid egg. The gel formed from the heated microbial lysate was found to have microscopic structures and physical properties similar to those of boiled eggs. The addition of microbial-derived edible enzymes or plant-based materials allowed for the adjustment of its properties, enabling the creation of various textures.
Furthermore, the researchers demonstrated that the microbial lysate could form stable foams widely used in baking, such as meringues (made from egg whites). They successfully baked meringue cookies using this lysate, showing its potential as a functional liquid egg substitute.
Distinguished Professor Sang Yup Lee stated, "This substitute has excellent nutritional components, making it suitable for regular food consumption. It is especially promising as emergency food for long-term space travel, wartime situations, and other emergencies. More importantly, it contributes to securing a sustainable food system."
< Figure 2. Example of foaming ability of microbial lysate and meringue cookie production >
< Figure 3. Example of foaming ability of microbial lysate and meringue cookie production >
The paper was published online in the journal npj Science of Food, issued by Nature.
- Paper Title: Microbial lysates repurposed as liquid egg substitutes
- Authors: Kyeong Rok Choi (first author), Da-Hee Ahn, Seok Yeong Jung, YuHyun Lee, and Sang Yup Lee (corresponding author)
This research was supported by the Ministry of Science and ICT's project for developing eco-friendly chemical technologies to replace petroleum (Project Leader: Distinguished Professor Sang Yup Lee, KAIST) and the Rural Development Administration's Agricultural Microorganisms Project Group (Director: Professor Pan-sik Jang, Seoul National University) for developing protein production technology from inorganic substances through microbial metabolic system control (Project Leader: Research Professor Kyeong Rok Choi, KAIST).
2024.07.05 View 6883 -
 KAIST introduces microbial food as a strategy food production of the future
The global food crisis is increasing due to rapid population growth and declining food productivity to climate change. Moreover, today's food production and supply system emit a huge amount of carbon dioxide, reaching 30% of the total amount emitted by humanity, aggravating climate change. Sustainable and nutritious microbial food is attracting attention as a key to overcoming this impasse.
KAIST (President Kwang Hyung Lee) announced on April 12th that Research Professor Kyeong Rok Choi of the BioProcess Engineering Research Center and Distinguished Professor Sang Yup Lee from the Department of Chemical and Biomolecular Engineering published a paper that proposes a direction of research on ‘microbial food production from sustainable raw materials.’
Microbial food refers to various foods and food ingredients produced using microorganisms. Microbial biomass contains a large amount of protein per unit in dry mass, comparable to that of meat, and emits the smallest amount of carbon dioxide and is required to produce a unit mass compared to various livestock, fish, shellfish, and crops. Since the amount of water and space requirement is small, it can be an eco-friendly, sustainable and highly nutritious food resource.
Fermented foods are the most readily available microbial foods around us. Although the proportion of microbial biomass in fermented foods is small, compounds with relatively low nutritional value, such as carbohydrates, are consumed during the fermentation process, and as microorganisms proliferate, the content of nutrients with higher nutritional value, such as proteins and vitamins, increases.
Various food compounds isolated and purified from biomass or culture media obtained through microbial culture are also a branch of microbial food. Examples that can be found around us include various amino acids, including monosodium glutamate, food proteins, enzymes, flavoring compounds, food colorings, and bioactive substances.
< Figure 1. Schematic diagram portraying various microbial biomass production strategies utlizing sustainable feedstocks >
Lastly, the most ultimate and fundamental form of microbial food can be said to be microbial biomass or extracts produced through microbial culture and foods cooked using them. A representative example is single-cell protein, which collectively refers to microbial biomass or microbial proteins extracted from it.
In this paper, the researchers comprehensively covered various non-edible raw materials and strategies for using them that can be used to produce microbial food in a more sustainable way. Furthermore, it covers various microbial foods that are actually produced in the industry using the relevant raw materials and their characteristics, as well as prospects for the production and generalization of sustainable microbial foods.
Research Professor Kyeong Rok Choi, the first author of this paper, said, “Microbial foods produced from various sustainable raw materials will soon be commonly encountered at our tables.” Second author Seok Yeong Jung, a doctoral student, also said, “Microbial foods of the future will not be limited foods consumed only out of a sense of obligation to the environment, but will be complete foods that are consumed by choice because of their nutritional value and taste.” In addition, Distinguished Professor Sang Yup Lee said, “It is time for the industry and academia, as well as the public and private sectors, to cooperate more closely so that more diverse microbial foods can be developed and supplied in order to create a sustainable society for ourselves and our descendants.”
< Figure 2. Compositions and environmental footprints of animal, plant and microbial biomass. >
This paper was published online on April 9 in ‘Nature Microbiology’ published by Nature.
※ Paper title: From sustainable feedstocks to microbial foods
※ Author information: Kyeong Rok Choi (first author), Seok Yeong Jung (second author) and Sang Yup Lee (corresponding author)
This research was conducted under the development of platform technologies of microbial cell factories for the next-generation biorefineries project (project leader KAIST Distinguished Professor Sang Yup Lee) supported by the Ministry of Science and ICT and the Cooperative Research Program for Agriculture Science and Technology Development (Project leader KAIST Research Professor Kyeong Rok Choi) of the Agricultural Microbiology Project Group (Director, Professor Pahn-Shick Chang) supported by the Rural Development Administration.
2024.04.12 View 7877
KAIST introduces microbial food as a strategy food production of the future
The global food crisis is increasing due to rapid population growth and declining food productivity to climate change. Moreover, today's food production and supply system emit a huge amount of carbon dioxide, reaching 30% of the total amount emitted by humanity, aggravating climate change. Sustainable and nutritious microbial food is attracting attention as a key to overcoming this impasse.
KAIST (President Kwang Hyung Lee) announced on April 12th that Research Professor Kyeong Rok Choi of the BioProcess Engineering Research Center and Distinguished Professor Sang Yup Lee from the Department of Chemical and Biomolecular Engineering published a paper that proposes a direction of research on ‘microbial food production from sustainable raw materials.’
Microbial food refers to various foods and food ingredients produced using microorganisms. Microbial biomass contains a large amount of protein per unit in dry mass, comparable to that of meat, and emits the smallest amount of carbon dioxide and is required to produce a unit mass compared to various livestock, fish, shellfish, and crops. Since the amount of water and space requirement is small, it can be an eco-friendly, sustainable and highly nutritious food resource.
Fermented foods are the most readily available microbial foods around us. Although the proportion of microbial biomass in fermented foods is small, compounds with relatively low nutritional value, such as carbohydrates, are consumed during the fermentation process, and as microorganisms proliferate, the content of nutrients with higher nutritional value, such as proteins and vitamins, increases.
Various food compounds isolated and purified from biomass or culture media obtained through microbial culture are also a branch of microbial food. Examples that can be found around us include various amino acids, including monosodium glutamate, food proteins, enzymes, flavoring compounds, food colorings, and bioactive substances.
< Figure 1. Schematic diagram portraying various microbial biomass production strategies utlizing sustainable feedstocks >
Lastly, the most ultimate and fundamental form of microbial food can be said to be microbial biomass or extracts produced through microbial culture and foods cooked using them. A representative example is single-cell protein, which collectively refers to microbial biomass or microbial proteins extracted from it.
In this paper, the researchers comprehensively covered various non-edible raw materials and strategies for using them that can be used to produce microbial food in a more sustainable way. Furthermore, it covers various microbial foods that are actually produced in the industry using the relevant raw materials and their characteristics, as well as prospects for the production and generalization of sustainable microbial foods.
Research Professor Kyeong Rok Choi, the first author of this paper, said, “Microbial foods produced from various sustainable raw materials will soon be commonly encountered at our tables.” Second author Seok Yeong Jung, a doctoral student, also said, “Microbial foods of the future will not be limited foods consumed only out of a sense of obligation to the environment, but will be complete foods that are consumed by choice because of their nutritional value and taste.” In addition, Distinguished Professor Sang Yup Lee said, “It is time for the industry and academia, as well as the public and private sectors, to cooperate more closely so that more diverse microbial foods can be developed and supplied in order to create a sustainable society for ourselves and our descendants.”
< Figure 2. Compositions and environmental footprints of animal, plant and microbial biomass. >
This paper was published online on April 9 in ‘Nature Microbiology’ published by Nature.
※ Paper title: From sustainable feedstocks to microbial foods
※ Author information: Kyeong Rok Choi (first author), Seok Yeong Jung (second author) and Sang Yup Lee (corresponding author)
This research was conducted under the development of platform technologies of microbial cell factories for the next-generation biorefineries project (project leader KAIST Distinguished Professor Sang Yup Lee) supported by the Ministry of Science and ICT and the Cooperative Research Program for Agriculture Science and Technology Development (Project leader KAIST Research Professor Kyeong Rok Choi) of the Agricultural Microbiology Project Group (Director, Professor Pahn-Shick Chang) supported by the Rural Development Administration.
2024.04.12 View 7877 -
 KAIST Research Team Creates the Scent of Jasmine from Microorganisms
The fragrance of jasmine and ylang-ylang, used widely in the manufacturing of cosmetics, foods, and beverages, can be produced by direct extraction from their respective flowers. In reality, this makes it difficult for production to meet demand, so companies use benzyl acetate, a major aromatic component of the two fragrances that is chemically synthesized from raw materials derived from petroleum.
On February 26, a KAIST research team led by Research Professor Kyeong Rok Choi from the BioProcess Engineering Research Center and Distinguished Professor Sang Yup Lee from the Department of Chemical and Biomolecular Engineering revealed the development of the first microbial process to effectively produce benzyl acetate, an industrially useful compound, from renewable carbon sources such as glucose. The results were published in their paper titled “A microbial process for the production of benzyl acetate”.
< Figure 1. Production of benzyl acetate through co-culture of upstream and downstream strains harboring the benzoic acid-dependent pathway. >
The team, led by Distinguished Professor Lee, aimed to produce benzyl acetate through an environmentally friendly and sustainable method, and developed an Escherichia coli strand to convert glucose into benzyl acetate through system metabolic engineering*.
*System metabolic engineering: a field of research founded by Distinguished Professor Lee to effectively develop microbial cell plants, a core component of the bio-industry that will replace the existing chemical industry, which is highly dependent on petroleum.
The research team developed a metabolic pathway that biosynthesizes benzyl acetate from benzoic acid derived from glucose, and successfully produced benzyl acetate by co-culturing** the strain.
**co-culture: simultaneously synthesizing two or more types of microorganisms in a mixture
However, it has been confirmed that the enzyme used to convert benzoic acid into benzyl acetate in this co-culturing technique acts non-specifically on an intermediate product during benzoic acid biosynthesis, producing a by-product called cinnamyl acetate. This process consumes the intermediate product needed for benzoic acid biosynthesis, thereby reducing the production efficiency of the target compound, benzyl acetate.
To overcome this problem, Distinguished Professor Lee and his team devised a delayed co-culture method, where they first produced benzoic acid in the earlier stages of fermentation by only culturing the top strain that produces benzoic acid from glucose, and later inoculated the bottom strain to convert the accumulated benzoic acid in the culture medium into benzyl acetate.
By applying this co-culture technique, the team suppressed the formation of by-products without further strain improvement or applying additional enzymes, and multiplied the concentration of the target compound by 10 times, producing 2.2 g/L of benzyl acetate. In addition, the team confirmed its potential for the commercial production of benzyl acetate through a technical economic analysis on this microbial process.
< Figure 2. Delayed co-culture of the Bn1 and Bn-BnAc3 strains for improved production of benzyl acetate through the benzoic acid-independent pathway.>
Research Professor Keyong Rok Choi, who was the first author of this paper, said, “This work is significant in that we have developed an effective microbial process to produce the industrially useful compound benzyl acetate, and also in that we have suggested a new approach to overcome the target chemical efficiency diminution and by-product formation issues caused commonly through non-specific enzyme activities during metabolic engineering.”
Distinguished Professor Lee said, “If we can increase the variety and number of microbial processes that produce useful chemicals through sustainable methods and at the same time develop effective strategies to solve chronic and inevitable problems that arise during microbial strain development, we will be able to accelerate the transition from the petrochemical industry into the eco-friendly and sustainable bio-industry.
This work was published online in Nature Chemical Engineering, issued by Nature.
This research was supported by the ‘Implementation of Intelligent Cell Factory Technology (PI: Distinguished Professor Sang Yup Lee) Project by the Ministry of Science and ICT, and the ‘Development of Protein Production Technology from Inorganic Substances through Microbiological Metabolic System Control’ (PI: Research Professor Kyeong Rok Choi) by the Agricultural Microbiological Project Group at the Rural Development Administration.
2024.03.05 View 7190
KAIST Research Team Creates the Scent of Jasmine from Microorganisms
The fragrance of jasmine and ylang-ylang, used widely in the manufacturing of cosmetics, foods, and beverages, can be produced by direct extraction from their respective flowers. In reality, this makes it difficult for production to meet demand, so companies use benzyl acetate, a major aromatic component of the two fragrances that is chemically synthesized from raw materials derived from petroleum.
On February 26, a KAIST research team led by Research Professor Kyeong Rok Choi from the BioProcess Engineering Research Center and Distinguished Professor Sang Yup Lee from the Department of Chemical and Biomolecular Engineering revealed the development of the first microbial process to effectively produce benzyl acetate, an industrially useful compound, from renewable carbon sources such as glucose. The results were published in their paper titled “A microbial process for the production of benzyl acetate”.
< Figure 1. Production of benzyl acetate through co-culture of upstream and downstream strains harboring the benzoic acid-dependent pathway. >
The team, led by Distinguished Professor Lee, aimed to produce benzyl acetate through an environmentally friendly and sustainable method, and developed an Escherichia coli strand to convert glucose into benzyl acetate through system metabolic engineering*.
*System metabolic engineering: a field of research founded by Distinguished Professor Lee to effectively develop microbial cell plants, a core component of the bio-industry that will replace the existing chemical industry, which is highly dependent on petroleum.
The research team developed a metabolic pathway that biosynthesizes benzyl acetate from benzoic acid derived from glucose, and successfully produced benzyl acetate by co-culturing** the strain.
**co-culture: simultaneously synthesizing two or more types of microorganisms in a mixture
However, it has been confirmed that the enzyme used to convert benzoic acid into benzyl acetate in this co-culturing technique acts non-specifically on an intermediate product during benzoic acid biosynthesis, producing a by-product called cinnamyl acetate. This process consumes the intermediate product needed for benzoic acid biosynthesis, thereby reducing the production efficiency of the target compound, benzyl acetate.
To overcome this problem, Distinguished Professor Lee and his team devised a delayed co-culture method, where they first produced benzoic acid in the earlier stages of fermentation by only culturing the top strain that produces benzoic acid from glucose, and later inoculated the bottom strain to convert the accumulated benzoic acid in the culture medium into benzyl acetate.
By applying this co-culture technique, the team suppressed the formation of by-products without further strain improvement or applying additional enzymes, and multiplied the concentration of the target compound by 10 times, producing 2.2 g/L of benzyl acetate. In addition, the team confirmed its potential for the commercial production of benzyl acetate through a technical economic analysis on this microbial process.
< Figure 2. Delayed co-culture of the Bn1 and Bn-BnAc3 strains for improved production of benzyl acetate through the benzoic acid-independent pathway.>
Research Professor Keyong Rok Choi, who was the first author of this paper, said, “This work is significant in that we have developed an effective microbial process to produce the industrially useful compound benzyl acetate, and also in that we have suggested a new approach to overcome the target chemical efficiency diminution and by-product formation issues caused commonly through non-specific enzyme activities during metabolic engineering.”
Distinguished Professor Lee said, “If we can increase the variety and number of microbial processes that produce useful chemicals through sustainable methods and at the same time develop effective strategies to solve chronic and inevitable problems that arise during microbial strain development, we will be able to accelerate the transition from the petrochemical industry into the eco-friendly and sustainable bio-industry.
This work was published online in Nature Chemical Engineering, issued by Nature.
This research was supported by the ‘Implementation of Intelligent Cell Factory Technology (PI: Distinguished Professor Sang Yup Lee) Project by the Ministry of Science and ICT, and the ‘Development of Protein Production Technology from Inorganic Substances through Microbiological Metabolic System Control’ (PI: Research Professor Kyeong Rok Choi) by the Agricultural Microbiological Project Group at the Rural Development Administration.
2024.03.05 View 7190 -
 KAIST to begin Joint Research to Develop Next-Generation LiDAR System with Hyundai Motor Group
< (From left) Jong-Soo Lee, Executive Vice President at Hyundai Motor, Sang-Yup Lee, Senior Vice President for Research at KAIST >
The ‘Hyundai Motor Group-KAIST On-Chip LiDAR Joint Research Lab’ was opened at KAIST’s main campus in Daejeon to develop LiDAR sensors for advanced autonomous vehicles.
The joint research lab aims to develop high-performance and compact on-chip sensors and new signal detection technology, which are essential in the increasingly competitive autonomous driving market. On-chip sensors, which utilize semiconductor manufacturing technology to add various functions, can reduce the size of LiDAR systems compared to conventional methods and secure price competitiveness through mass production using semiconductor fabrication processes.
The joint research lab will consist of about 30 researchers, including the Hyundai-Kia Institute of Advanced Technology Development research team and KAIST professors Sanghyeon Kim, Sangsik Kim, Wanyeong Jung, and Hamza Kurt from KAIST’s School of Electrical Engineering, and will operate for four years until 2028.
KAIST will be leading the specialized work of each research team, such as for the development of silicon optoelectronic on-chip LiDAR components, the fabrication of high-speed, high-power integrated circuits to run the LiDAR systems, and the optimization and verification of LiDAR systems.
Hyundai Motor and Kia, together with Hyundai NGV, a specialized industry-academia cooperation institution, will oversee the operation of the joint research lab and provide support such as monitoring technological trends, suggesting research directions, deriving core ideas, and recommending technologies and experts to enhance research capabilities.
A Hyundai Motor Group official said, "We believe that this cooperation between Hyundai Motor Company and Kia, the leader in autonomous driving technology, and KAIST, the home of world-class technology, will hasten the achievement of fully autonomous driving." He added, "We will do our best to enable the lab to produce tangible results.”
Professor Sanghyeon Kim said, "The LiDAR sensor, which serves as the eyes of a car, is a core technology for future autonomous vehicle development that is essential for automobile companies to internalize."
2024.02.27 View 10995
KAIST to begin Joint Research to Develop Next-Generation LiDAR System with Hyundai Motor Group
< (From left) Jong-Soo Lee, Executive Vice President at Hyundai Motor, Sang-Yup Lee, Senior Vice President for Research at KAIST >
The ‘Hyundai Motor Group-KAIST On-Chip LiDAR Joint Research Lab’ was opened at KAIST’s main campus in Daejeon to develop LiDAR sensors for advanced autonomous vehicles.
The joint research lab aims to develop high-performance and compact on-chip sensors and new signal detection technology, which are essential in the increasingly competitive autonomous driving market. On-chip sensors, which utilize semiconductor manufacturing technology to add various functions, can reduce the size of LiDAR systems compared to conventional methods and secure price competitiveness through mass production using semiconductor fabrication processes.
The joint research lab will consist of about 30 researchers, including the Hyundai-Kia Institute of Advanced Technology Development research team and KAIST professors Sanghyeon Kim, Sangsik Kim, Wanyeong Jung, and Hamza Kurt from KAIST’s School of Electrical Engineering, and will operate for four years until 2028.
KAIST will be leading the specialized work of each research team, such as for the development of silicon optoelectronic on-chip LiDAR components, the fabrication of high-speed, high-power integrated circuits to run the LiDAR systems, and the optimization and verification of LiDAR systems.
Hyundai Motor and Kia, together with Hyundai NGV, a specialized industry-academia cooperation institution, will oversee the operation of the joint research lab and provide support such as monitoring technological trends, suggesting research directions, deriving core ideas, and recommending technologies and experts to enhance research capabilities.
A Hyundai Motor Group official said, "We believe that this cooperation between Hyundai Motor Company and Kia, the leader in autonomous driving technology, and KAIST, the home of world-class technology, will hasten the achievement of fully autonomous driving." He added, "We will do our best to enable the lab to produce tangible results.”
Professor Sanghyeon Kim said, "The LiDAR sensor, which serves as the eyes of a car, is a core technology for future autonomous vehicle development that is essential for automobile companies to internalize."
2024.02.27 View 10995 -
 KAIST presents strategies for environmentally friendly and sustainable polyamides production
- Provides current research trends in bio-based polyamide production
- Research on bio-based polyamides production gains importance for achieving a carbon-neutral society
Global industries focused on carbon neutrality, under the slogan "Net-Zero," are gaining increasing attention. In particular, research on microbial production of polymers, replacing traditional chemical methods with biological approaches, is actively progressing.
Polyamides, represented by nylon, are linear polymers widely used in various industries such as automotive, electronics, textiles, and medical fields. They possess beneficial properties such as high tensile strength, electrical insulation, heat resistance, wear resistance, and biocompatibility. Since the commercialization of nylon in 1938, approximately 7 million tons of polyamides are produced worldwide annually. Considering their broad applications and significance, producing polyamides through bio-based methods holds considerable environmental and industrial importance.
KAIST (President Kwang-Hyung Lee) announced that a research team led by Distinguished Professor Sang Yup Lee, including Dr. Jong An Lee and doctoral candidate Ji Yeon Kim from the Department of Chemical and Biomolecular Engineering, published a paper titled "Current Advancements in Bio-Based Production of Polyamides”. The paper was featured on the cover of the monthly issue of "Trends in Chemistry” by Cell Press.
As part of climate change response technologies, bio-refineries involve using biotechnological and chemical methods to produce industrially important chemicals and biofuels from renewable biomass without relying on fossil resources. Notably, systems metabolic engineering, pioneered by KAIST's Distinguished Professor Sang Yup Lee, is a research field that effectively manipulates microbial metabolic pathways to produce valuable chemicals, forming the core technology for bio-refineries. The research team has successfully developed high-performance strains producing a variety of compounds, including succinic acid, biodegradable plastics, biofuels, and natural products, using systems metabolic engineering tools and strategies.
The research team predicted that if bio-based polyamide production technology, which is widely used in the production of clothing and textiles, becomes widespread, it will attract attention as a future technology that can respond to the climate crisis due to its environment-friendly production technology. In this study, the research team comprehensively reviewed the bio-based polyamide production strategies. They provided insights into the advancements in polyamide monomer production using metabolically engineered microorganisms and highlighted the recent trends in bio-based polyamide advancements utilizing these monomers. Additionally, they reviewed the strategies for synthesizing bio-based polyamides through chemical conversion of natural oils and discussed the biodegradability and recycling of the polyamides. Furthermore, the paper presented the future direction in which metabolic engineering can be applied for the bio-based polyamide production, contributing to environmentally friendly and sustainable society.
Ji Yeon Kim, the co-first author of this paper from KAIST, stated "The importance of utilizing systems metabolic engineering tools and strategies for bio-based polyamides production is becoming increasingly prominent in achieving carbon neutrality."
Professor Sang Yup Lee emphasized, "Amid growing concerns about climate change, the significance of environmentally friendly and sustainable industrial development is greater than ever. Systems metabolic engineering is expected to have a significant impact not only on the chemical industry but also in various fields."
< [Figure 1] A schematic overview of the overall process for polyamides production >
This paper by Dr. Jong An Lee, PhD student Ji Yeon Kim, Dr. Jung Ho Ahn, and Master Yeah-Ji Ahn from the Department of Chemical and Biomolecular Engineering at KAIST was published in the December issue of 'Trends in Chemistry', an authoritative review journal in the field of chemistry published by Cell. It was published on December 7 as the cover paper and featured review.
※ Paper title: Current advancements in the bio-based production of polyamides
※ Author information: Jong An Lee, Ji Yeon Kim, Jung Ho Ahn, Yeah-Ji Ahn, and Sang Yup Lee
This research was conducted with the support from the development of platform technologies of microbial cell factories for the next-generation biorefineries project and C1 gas refinery program by Korean Ministry of Science and ICT.
< [Figure 2] Cover paper of the December issue of Trends in Chemistry >
2023.12.21 View 7326
KAIST presents strategies for environmentally friendly and sustainable polyamides production
- Provides current research trends in bio-based polyamide production
- Research on bio-based polyamides production gains importance for achieving a carbon-neutral society
Global industries focused on carbon neutrality, under the slogan "Net-Zero," are gaining increasing attention. In particular, research on microbial production of polymers, replacing traditional chemical methods with biological approaches, is actively progressing.
Polyamides, represented by nylon, are linear polymers widely used in various industries such as automotive, electronics, textiles, and medical fields. They possess beneficial properties such as high tensile strength, electrical insulation, heat resistance, wear resistance, and biocompatibility. Since the commercialization of nylon in 1938, approximately 7 million tons of polyamides are produced worldwide annually. Considering their broad applications and significance, producing polyamides through bio-based methods holds considerable environmental and industrial importance.
KAIST (President Kwang-Hyung Lee) announced that a research team led by Distinguished Professor Sang Yup Lee, including Dr. Jong An Lee and doctoral candidate Ji Yeon Kim from the Department of Chemical and Biomolecular Engineering, published a paper titled "Current Advancements in Bio-Based Production of Polyamides”. The paper was featured on the cover of the monthly issue of "Trends in Chemistry” by Cell Press.
As part of climate change response technologies, bio-refineries involve using biotechnological and chemical methods to produce industrially important chemicals and biofuels from renewable biomass without relying on fossil resources. Notably, systems metabolic engineering, pioneered by KAIST's Distinguished Professor Sang Yup Lee, is a research field that effectively manipulates microbial metabolic pathways to produce valuable chemicals, forming the core technology for bio-refineries. The research team has successfully developed high-performance strains producing a variety of compounds, including succinic acid, biodegradable plastics, biofuels, and natural products, using systems metabolic engineering tools and strategies.
The research team predicted that if bio-based polyamide production technology, which is widely used in the production of clothing and textiles, becomes widespread, it will attract attention as a future technology that can respond to the climate crisis due to its environment-friendly production technology. In this study, the research team comprehensively reviewed the bio-based polyamide production strategies. They provided insights into the advancements in polyamide monomer production using metabolically engineered microorganisms and highlighted the recent trends in bio-based polyamide advancements utilizing these monomers. Additionally, they reviewed the strategies for synthesizing bio-based polyamides through chemical conversion of natural oils and discussed the biodegradability and recycling of the polyamides. Furthermore, the paper presented the future direction in which metabolic engineering can be applied for the bio-based polyamide production, contributing to environmentally friendly and sustainable society.
Ji Yeon Kim, the co-first author of this paper from KAIST, stated "The importance of utilizing systems metabolic engineering tools and strategies for bio-based polyamides production is becoming increasingly prominent in achieving carbon neutrality."
Professor Sang Yup Lee emphasized, "Amid growing concerns about climate change, the significance of environmentally friendly and sustainable industrial development is greater than ever. Systems metabolic engineering is expected to have a significant impact not only on the chemical industry but also in various fields."
< [Figure 1] A schematic overview of the overall process for polyamides production >
This paper by Dr. Jong An Lee, PhD student Ji Yeon Kim, Dr. Jung Ho Ahn, and Master Yeah-Ji Ahn from the Department of Chemical and Biomolecular Engineering at KAIST was published in the December issue of 'Trends in Chemistry', an authoritative review journal in the field of chemistry published by Cell. It was published on December 7 as the cover paper and featured review.
※ Paper title: Current advancements in the bio-based production of polyamides
※ Author information: Jong An Lee, Ji Yeon Kim, Jung Ho Ahn, Yeah-Ji Ahn, and Sang Yup Lee
This research was conducted with the support from the development of platform technologies of microbial cell factories for the next-generation biorefineries project and C1 gas refinery program by Korean Ministry of Science and ICT.
< [Figure 2] Cover paper of the December issue of Trends in Chemistry >
2023.12.21 View 7326