Bio+and+Brain+Engineering
-
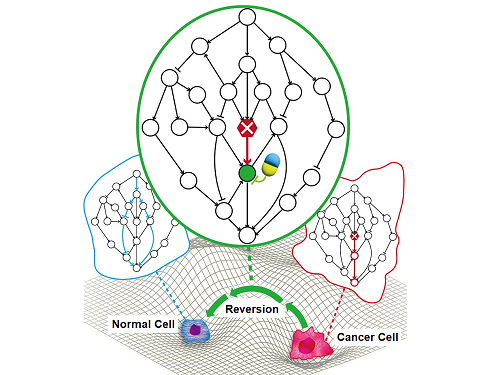 A KAIST Research Team Identifies a Cancer Reversion Mechanism
Despite decades of intensive cancer research by numerous biomedical scientists, cancer still holds its place as the number one cause of death in Korea. The fundamental reason behind the limitations of current cancer treatment methods is the fact that they all aim to completely destroy cancer cells, which eventually allows the cancer cells to acquire immunity. In other words, recurrences and side-effects caused by the destruction of healthy cells are inevitable. To this end, some have suggested anticancer treatment methods based on cancer reversion, which can revert cancer cells back to normal or near-normal cells under certain conditions. However, the practical development of this idea has not yet been attempted.
On June 8, a KAIST research team led by Professor Kwang-Hyun Cho from the Department of Bio and Brain Engineering reported to have successfully identified the fundamental principle of a process that can revert cancer cells back to normal cells without killing the cells.
Professor Cho’s team focused on the fact that unlike normal cells, which react according to external stimuli, cancer cells tend to ignore such stimuli and only undergo uncontrolled cell division. Through computer simulation analysis, the team discovered that the input-output (I/O) relationships that were distorted by genetic mutations could be reverted back to normal I/O relationships under certain conditions. The team then demonstrated through molecular cell experiments that such I/O relationship recovery also occurred in real cancer cells.
The results of this study, written by Dr. Jae Il Joo and Dr. Hwa-Jeong Park, were published in Wiley’s Advanced Science online on June 2 under the title, "Normalizing input-output relationships of cancer networks for reversion therapy."
< Image 1. Input-output (I/O) relationships in gene regulatory networks >
Professor Kwang-Hyun Cho's research team classified genes into four types by simulation-analyzing the effect of gene mutations on the I/O relationship of gene regulatory networks. (Figure A-J) In addition, by analyzing 18 genes of the cancer-related gene regulatory network, it was confirmed that when mutations occur in more than half of the genes constituting each network, reversibility is possible through appropriate control. (Figure K)
Professor Cho’s team uncovered that the reason the distorted I/O relationships of cancer cells could be reverted back to normal ones was the robustness and redundancy of intracellular gene control networks that developed over the course of evolution. In addition, they found that some genes were more promising as targets for cancer reversion than others, and showed through molecular cell experiments that controlling such genes could revert the distorted I/O relationships of cancer cells back to normal ones.
< Image 2. Simulation results of restoration of bladder cancer gene regulation network and I/O relationship of bladder cancer cells. >
The research team classified the effects of gene mutations on the I/O relationship in the bladder cancer gene regulation network by simulation analysis and classified them into 4 types. (Figure A) Through this, it was found that the distorted input-output relationship between bladder cancer cell lines KU-1919 and HCT-1197 could be restored to normal. (Figure B)
< Image 3. Analysis of survival of bladder cancer patients according to reversible gene mutation and I/O recovery experiment of bladder cancer cells. >
As predicted through network simulation analysis, Professor Kwang-Hyun Cho's research team confirmed through molecular cell experiments that the response to TGF-b was normally restored when AKT and MAP3K1 were inhibited in the bladder cancer cell line KU-1919. (Figure A-G) In addition, it was confirmed that there is a difference in the survival rate of bladder cancer patients depending on the presence or absence of a reversible gene mutation. (Figure H)
The results of this research show that the reversion of real cancer cells does not happen by chance, and that it is possible to systematically explore targets that can induce this phenomenon, thereby creating the potential for the development of innovative anticancer drugs that can control such target genes.
< Image 4. Cancer cell reversibility principle >
The research team analyzed the reversibility, redundancy, and robustness of various networks and found that there was a positive correlation between them. From this, it was found that reversibility was additionally inherent in the process of evolution in which the gene regulatory network acquired redundancy and consistency.
Professor Cho said, “By uncovering the fundamental principles of a new cancer reversion treatment strategy that may overcome the unresolved limitations of existing chemotherapy, we have increased the possibility of developing new and innovative drugs that can improve both the prognosis and quality of life of cancer patients.”
< Image 5. Conceptual diagram of research results >
The research team identified the fundamental control principle of cancer cell reversibility through systems biology research. When the I/O relationship of the intracellular gene regulatory network is distorted by mutation, the distorted I/O relationship can be restored to a normal state by identifying and adjusting the reversible gene target based on the redundancy of the molecular circuit inherent in the complex network.
After Professor Cho’s team first suggested the concept of reversion treatment, they published their results for reverting colorectal cancer in January 2020, and in January 2022 they successfully re-programmed malignant breast cancer cells back into hormone-treatable ones. In January 2023, the team successfully removed the metastasis ability from lung cancer cells and reverted them back to a state that allowed improved drug reactivity. However, these results were case studies of specific types of cancer and did not reveal what common principle allowed cancer reversion across all cancer types, making this the first revelation of the general principle of cancer reversion and its evolutionary origins.
This research was funded by the Ministry of Science and ICT of the Republic of Korea and the National Research Foundation of Korea.
2023.06.20 View 4957
A KAIST Research Team Identifies a Cancer Reversion Mechanism
Despite decades of intensive cancer research by numerous biomedical scientists, cancer still holds its place as the number one cause of death in Korea. The fundamental reason behind the limitations of current cancer treatment methods is the fact that they all aim to completely destroy cancer cells, which eventually allows the cancer cells to acquire immunity. In other words, recurrences and side-effects caused by the destruction of healthy cells are inevitable. To this end, some have suggested anticancer treatment methods based on cancer reversion, which can revert cancer cells back to normal or near-normal cells under certain conditions. However, the practical development of this idea has not yet been attempted.
On June 8, a KAIST research team led by Professor Kwang-Hyun Cho from the Department of Bio and Brain Engineering reported to have successfully identified the fundamental principle of a process that can revert cancer cells back to normal cells without killing the cells.
Professor Cho’s team focused on the fact that unlike normal cells, which react according to external stimuli, cancer cells tend to ignore such stimuli and only undergo uncontrolled cell division. Through computer simulation analysis, the team discovered that the input-output (I/O) relationships that were distorted by genetic mutations could be reverted back to normal I/O relationships under certain conditions. The team then demonstrated through molecular cell experiments that such I/O relationship recovery also occurred in real cancer cells.
The results of this study, written by Dr. Jae Il Joo and Dr. Hwa-Jeong Park, were published in Wiley’s Advanced Science online on June 2 under the title, "Normalizing input-output relationships of cancer networks for reversion therapy."
< Image 1. Input-output (I/O) relationships in gene regulatory networks >
Professor Kwang-Hyun Cho's research team classified genes into four types by simulation-analyzing the effect of gene mutations on the I/O relationship of gene regulatory networks. (Figure A-J) In addition, by analyzing 18 genes of the cancer-related gene regulatory network, it was confirmed that when mutations occur in more than half of the genes constituting each network, reversibility is possible through appropriate control. (Figure K)
Professor Cho’s team uncovered that the reason the distorted I/O relationships of cancer cells could be reverted back to normal ones was the robustness and redundancy of intracellular gene control networks that developed over the course of evolution. In addition, they found that some genes were more promising as targets for cancer reversion than others, and showed through molecular cell experiments that controlling such genes could revert the distorted I/O relationships of cancer cells back to normal ones.
< Image 2. Simulation results of restoration of bladder cancer gene regulation network and I/O relationship of bladder cancer cells. >
The research team classified the effects of gene mutations on the I/O relationship in the bladder cancer gene regulation network by simulation analysis and classified them into 4 types. (Figure A) Through this, it was found that the distorted input-output relationship between bladder cancer cell lines KU-1919 and HCT-1197 could be restored to normal. (Figure B)
< Image 3. Analysis of survival of bladder cancer patients according to reversible gene mutation and I/O recovery experiment of bladder cancer cells. >
As predicted through network simulation analysis, Professor Kwang-Hyun Cho's research team confirmed through molecular cell experiments that the response to TGF-b was normally restored when AKT and MAP3K1 were inhibited in the bladder cancer cell line KU-1919. (Figure A-G) In addition, it was confirmed that there is a difference in the survival rate of bladder cancer patients depending on the presence or absence of a reversible gene mutation. (Figure H)
The results of this research show that the reversion of real cancer cells does not happen by chance, and that it is possible to systematically explore targets that can induce this phenomenon, thereby creating the potential for the development of innovative anticancer drugs that can control such target genes.
< Image 4. Cancer cell reversibility principle >
The research team analyzed the reversibility, redundancy, and robustness of various networks and found that there was a positive correlation between them. From this, it was found that reversibility was additionally inherent in the process of evolution in which the gene regulatory network acquired redundancy and consistency.
Professor Cho said, “By uncovering the fundamental principles of a new cancer reversion treatment strategy that may overcome the unresolved limitations of existing chemotherapy, we have increased the possibility of developing new and innovative drugs that can improve both the prognosis and quality of life of cancer patients.”
< Image 5. Conceptual diagram of research results >
The research team identified the fundamental control principle of cancer cell reversibility through systems biology research. When the I/O relationship of the intracellular gene regulatory network is distorted by mutation, the distorted I/O relationship can be restored to a normal state by identifying and adjusting the reversible gene target based on the redundancy of the molecular circuit inherent in the complex network.
After Professor Cho’s team first suggested the concept of reversion treatment, they published their results for reverting colorectal cancer in January 2020, and in January 2022 they successfully re-programmed malignant breast cancer cells back into hormone-treatable ones. In January 2023, the team successfully removed the metastasis ability from lung cancer cells and reverted them back to a state that allowed improved drug reactivity. However, these results were case studies of specific types of cancer and did not reveal what common principle allowed cancer reversion across all cancer types, making this the first revelation of the general principle of cancer reversion and its evolutionary origins.
This research was funded by the Ministry of Science and ICT of the Republic of Korea and the National Research Foundation of Korea.
2023.06.20 View 4957 -
 The cause of disability in aged brain meningeal membranes identified
Due to the increase in average age, studies on changes in the brain following general aging process without serious brain diseases have also become an issue that requires in-depth studies. Regarding aging research, as aging progresses, ‘sugar’ accumulates in the body, and the accumulated sugar becomes a causative agent for various diseases such as aging-related inflammation and vascular disease. In the end, “surplus” sugar molecules attach to various proteins in the body and interfere with their functions.
KAIST (President Kwang Hyung Lee), a joint research team of Professor Pilnam Kim and Professor Yong Jeong of the Department of Bio and Brain Engineering, revealed on the 15th that it was confirmed that the function of being the “front line of defense” for the cerebrocortex of the brain meninges, the layers of membranes that surrounds the brain, is hindered when 'sugar' begins to build up on them as aging progresses.
Professor Kim's research team confirmed excessive accumulation of sugar molecules in the meninges of the elderly and also confirmed that sugar accumulation occurs mouse models in accordance with certain age levels. The meninges are thin membranes that surround the brain and exist at the boundary between the cerebrospinal fluid and the cortex and play an important role in protecting the brain. In this study, it was revealed that the dysfunction of these brain membranes caused by aging is induced by 'excess' sugar in the brain. In particular, as the meningeal membrane becomes thinner and stickier due to aging, a new paradigm has been provided for the discovery of the principle of the decrease in material exchange between the cerebrospinal fluid and the cerebral cortex.
This research was conducted by the Ph.D. candidate Hyo Min Kim and Dr. Shinheun Kim as the co-first authors to be published online on February 28th in the international journal, Aging Cell. (Paper Title: Glycation-mediated tissue-level remodeling of brain meningeal membrane by aging)
The meninges, which are in direct contact with the cerebrospinal fluid, are mainly composed of collagen, an extracellular matrix (ECM) protein, and are composed of fibroblasts, which are cells that produce this protein. The cells that come in contact with collagen proteins that are attached with sugar have a low collagen production function, while the meningeal membrane continuously thins and collapses as the expression of collagen degrading enzymes increases.
Studies on the relationship between excess sugar molecules accumulation in the brain due to continued sugar intake and the degeneration of neurons and brain diseases have been continuously conducted. However, this study was the first to identify meningeal degeneration and dysfunction caused by glucose accumulation with the focus on the meninges itself, and the results are expected to present new ideas for research into approach towards discoveries of new treatments for brain disease.
Researcher Hyomin Kim, the first author, introduced the research results as “an interesting study that identified changes in the barriers of the brain due to aging through a convergent approach, starting from the human brain and utilizing an animal model with a biomimetic meningeal model”.
Professor Pilnam Kim's research team is conducting research and development to remove sugar that accumulated throughout the human body, including the meninges. Advanced glycation end products, which are waste products formed when proteins and sugars meet in the human body, are partially removed by macrophages. However, glycated products bound to extracellular matrix proteins such as collagen are difficult to remove naturally. Through the KAIST-Ceragem Research Center, this research team is developing a healthcare medical device to remove 'sugar residue' in the body.
This study was carried out with the National Research Foundation of Korea's collective research support.
Figure 1. Schematic diagram of proposed mechanism showing aging‐related ECM remodeling through meningeal fibroblasts on the brain leptomeninges. Meningeal fibroblasts in the young brain showed dynamic COL1A1 synthetic and COL1‐interactive function on the collagen membrane. They showed ITGB1‐mediated adhesion on the COL1‐composed leptomeningeal membrane and induction of COL1A1 synthesis for maintaining the collagen membrane. With aging, meningeal fibroblasts showed depletion of COL1A1 synthetic function and altered cell–matrix interaction.
Figure 2. Representative rat meningeal images observed in the study. Compared to young rats, it was confirmed that type 1 collagen (COL1) decreased along with the accumulation of glycated end products (AGE) in the brain membrane of aged rats, and the activity of integrin beta 1 (ITGB1), a representative receptor corresponding to cell-collagen interaction. Instead, it was observed that the activity of discoidin domain receptor 2 (DDR2), one of the tyrosine kinases, increased.
Figure 3. Substance flux through the brain membrane decreases with aging. It was confirmed that the degree of adsorption of fluorescent substances contained in cerebrospinal fluid (CSF) to the brain membrane increased and the degree of entry into the periphery of the cerebral blood vessels decreased in the aged rats. In this study, only the influx into the brain was confirmed during the entry and exit of substances, but the degree of outflow will also be confirmed through future studies.
2023.03.15 View 4591
The cause of disability in aged brain meningeal membranes identified
Due to the increase in average age, studies on changes in the brain following general aging process without serious brain diseases have also become an issue that requires in-depth studies. Regarding aging research, as aging progresses, ‘sugar’ accumulates in the body, and the accumulated sugar becomes a causative agent for various diseases such as aging-related inflammation and vascular disease. In the end, “surplus” sugar molecules attach to various proteins in the body and interfere with their functions.
KAIST (President Kwang Hyung Lee), a joint research team of Professor Pilnam Kim and Professor Yong Jeong of the Department of Bio and Brain Engineering, revealed on the 15th that it was confirmed that the function of being the “front line of defense” for the cerebrocortex of the brain meninges, the layers of membranes that surrounds the brain, is hindered when 'sugar' begins to build up on them as aging progresses.
Professor Kim's research team confirmed excessive accumulation of sugar molecules in the meninges of the elderly and also confirmed that sugar accumulation occurs mouse models in accordance with certain age levels. The meninges are thin membranes that surround the brain and exist at the boundary between the cerebrospinal fluid and the cortex and play an important role in protecting the brain. In this study, it was revealed that the dysfunction of these brain membranes caused by aging is induced by 'excess' sugar in the brain. In particular, as the meningeal membrane becomes thinner and stickier due to aging, a new paradigm has been provided for the discovery of the principle of the decrease in material exchange between the cerebrospinal fluid and the cerebral cortex.
This research was conducted by the Ph.D. candidate Hyo Min Kim and Dr. Shinheun Kim as the co-first authors to be published online on February 28th in the international journal, Aging Cell. (Paper Title: Glycation-mediated tissue-level remodeling of brain meningeal membrane by aging)
The meninges, which are in direct contact with the cerebrospinal fluid, are mainly composed of collagen, an extracellular matrix (ECM) protein, and are composed of fibroblasts, which are cells that produce this protein. The cells that come in contact with collagen proteins that are attached with sugar have a low collagen production function, while the meningeal membrane continuously thins and collapses as the expression of collagen degrading enzymes increases.
Studies on the relationship between excess sugar molecules accumulation in the brain due to continued sugar intake and the degeneration of neurons and brain diseases have been continuously conducted. However, this study was the first to identify meningeal degeneration and dysfunction caused by glucose accumulation with the focus on the meninges itself, and the results are expected to present new ideas for research into approach towards discoveries of new treatments for brain disease.
Researcher Hyomin Kim, the first author, introduced the research results as “an interesting study that identified changes in the barriers of the brain due to aging through a convergent approach, starting from the human brain and utilizing an animal model with a biomimetic meningeal model”.
Professor Pilnam Kim's research team is conducting research and development to remove sugar that accumulated throughout the human body, including the meninges. Advanced glycation end products, which are waste products formed when proteins and sugars meet in the human body, are partially removed by macrophages. However, glycated products bound to extracellular matrix proteins such as collagen are difficult to remove naturally. Through the KAIST-Ceragem Research Center, this research team is developing a healthcare medical device to remove 'sugar residue' in the body.
This study was carried out with the National Research Foundation of Korea's collective research support.
Figure 1. Schematic diagram of proposed mechanism showing aging‐related ECM remodeling through meningeal fibroblasts on the brain leptomeninges. Meningeal fibroblasts in the young brain showed dynamic COL1A1 synthetic and COL1‐interactive function on the collagen membrane. They showed ITGB1‐mediated adhesion on the COL1‐composed leptomeningeal membrane and induction of COL1A1 synthesis for maintaining the collagen membrane. With aging, meningeal fibroblasts showed depletion of COL1A1 synthetic function and altered cell–matrix interaction.
Figure 2. Representative rat meningeal images observed in the study. Compared to young rats, it was confirmed that type 1 collagen (COL1) decreased along with the accumulation of glycated end products (AGE) in the brain membrane of aged rats, and the activity of integrin beta 1 (ITGB1), a representative receptor corresponding to cell-collagen interaction. Instead, it was observed that the activity of discoidin domain receptor 2 (DDR2), one of the tyrosine kinases, increased.
Figure 3. Substance flux through the brain membrane decreases with aging. It was confirmed that the degree of adsorption of fluorescent substances contained in cerebrospinal fluid (CSF) to the brain membrane increased and the degree of entry into the periphery of the cerebral blood vessels decreased in the aged rats. In this study, only the influx into the brain was confirmed during the entry and exit of substances, but the degree of outflow will also be confirmed through future studies.
2023.03.15 View 4591 -
 KAIST team develops smart immune system that can pin down on malignant tumors
A joint research team led by Professor Jung Kyoon Choi of the KAIST Department of Bio and Brain Engineering and Professor Jong-Eun Park of the KAIST Graduate School of Medical Science and Engineering (GSMSE) announced the development of the key technologies to treat cancers using smart immune cells designed based on AI and big data analysis. This technology is expected to be a next-generation immunotherapy that allows precision targeting of tumor cells by having the chimeric antigen receptors (CARs) operate through a logical circuit. Professor Hee Jung An of CHA Bundang Medical Center and Professor Hae-Ock Lee of the Catholic University of Korea also participated in this research to contribute joint effort.
Professor Jung Kyoon Choi’s team built a gene expression database from millions of cells, and used this to successfully develop and verify a deep-learning algorithm that could detect the differences in gene expression patterns between tumor cells and normal cells through a logical circuit. CAR immune cells that were fitted with the logic circuits discovered through this methodology could distinguish between tumorous and normal cells as a computer would, and therefore showed potentials to strike only on tumor cells accurately without causing unwanted side effects.
This research, conducted by co-first authors Dr. Joonha Kwon of the KAIST Department of Bio and Brain Engineering and Ph.D. candidate Junho Kang of KAIST GSMSE, was published by Nature Biotechnology on February 16, under the title Single-cell mapping of combinatorial target antigens for CAR switches using logic gates.
An area in cancer research where the most attempts and advances have been made in recent years is immunotherapy. This field of treatment, which utilizes the patient’s own immune system in order to overcome cancer, has several methods including immune checkpoint inhibitors, cancer vaccines and cellular treatments. Immune cells like CAR-T or CAR-NK equipped with chimera antigen receptors, in particular, can recognize cancer antigens and directly destroy cancer cells.
Starting with its success in blood cancer treatment, scientists have been trying to expand the application of CAR cell therapy to treat solid cancer. But there have been difficulties to develop CAR cells with effective killing abilities against solid cancer cells with minimized side effects. Accordingly, in recent years, the development of smarter CAR engineering technologies, i.e., computational logic gates such as AND, OR, and NOT, to effectively target cancer cells has been underway.
At this point in time, the research team built a large-scale database for cancer and normal cells to discover the exact genes that are expressed only from cancer cells at a single-cell level. The team followed this up by developing an AI algorithm that could search for a combination of genes that best distinguishes cancer cells from normal cells. This algorithm, in particular, has been used to find a logic circuit that can specifically target cancer cells through cell-level simulations of all gene combinations. CAR-T cells equipped with logic circuits discovered through this methodology are expected to distinguish cancerous cells from normal cells like computers, thereby minimizing side effects and maximizing the effects of chemotherapy.
Dr. Joonha Kwon, who is the first author of this paper, said, “this research suggests a new method that hasn’t been tried before. What’s particularly noteworthy is the process in which we found the optimal CAR cell circuit through simulations of millions of individual tumors and normal cells.” He added, “This is an innovative technology that can apply AI and computer logic circuits to immune cell engineering. It would contribute greatly to expanding CAR therapy, which is being successfully used for blood cancer, to solid cancers as well.”
This research was funded by the Original Technology Development Project and Research Program for Next Generation Applied Omic of the Korea Research Foundation.
Figure 1. A schematic diagram of manufacturing and administration process of CAR therapy and of cancer cell-specific dual targeting using CAR.
Figure 2. Deep learning (convolutional neural networks, CNNs) algorithm for selection of dual targets based on gene combination (left) and algorithm for calculating expressing cell fractions by gene combination according to logical circuit (right).
2023.03.09 View 6065
KAIST team develops smart immune system that can pin down on malignant tumors
A joint research team led by Professor Jung Kyoon Choi of the KAIST Department of Bio and Brain Engineering and Professor Jong-Eun Park of the KAIST Graduate School of Medical Science and Engineering (GSMSE) announced the development of the key technologies to treat cancers using smart immune cells designed based on AI and big data analysis. This technology is expected to be a next-generation immunotherapy that allows precision targeting of tumor cells by having the chimeric antigen receptors (CARs) operate through a logical circuit. Professor Hee Jung An of CHA Bundang Medical Center and Professor Hae-Ock Lee of the Catholic University of Korea also participated in this research to contribute joint effort.
Professor Jung Kyoon Choi’s team built a gene expression database from millions of cells, and used this to successfully develop and verify a deep-learning algorithm that could detect the differences in gene expression patterns between tumor cells and normal cells through a logical circuit. CAR immune cells that were fitted with the logic circuits discovered through this methodology could distinguish between tumorous and normal cells as a computer would, and therefore showed potentials to strike only on tumor cells accurately without causing unwanted side effects.
This research, conducted by co-first authors Dr. Joonha Kwon of the KAIST Department of Bio and Brain Engineering and Ph.D. candidate Junho Kang of KAIST GSMSE, was published by Nature Biotechnology on February 16, under the title Single-cell mapping of combinatorial target antigens for CAR switches using logic gates.
An area in cancer research where the most attempts and advances have been made in recent years is immunotherapy. This field of treatment, which utilizes the patient’s own immune system in order to overcome cancer, has several methods including immune checkpoint inhibitors, cancer vaccines and cellular treatments. Immune cells like CAR-T or CAR-NK equipped with chimera antigen receptors, in particular, can recognize cancer antigens and directly destroy cancer cells.
Starting with its success in blood cancer treatment, scientists have been trying to expand the application of CAR cell therapy to treat solid cancer. But there have been difficulties to develop CAR cells with effective killing abilities against solid cancer cells with minimized side effects. Accordingly, in recent years, the development of smarter CAR engineering technologies, i.e., computational logic gates such as AND, OR, and NOT, to effectively target cancer cells has been underway.
At this point in time, the research team built a large-scale database for cancer and normal cells to discover the exact genes that are expressed only from cancer cells at a single-cell level. The team followed this up by developing an AI algorithm that could search for a combination of genes that best distinguishes cancer cells from normal cells. This algorithm, in particular, has been used to find a logic circuit that can specifically target cancer cells through cell-level simulations of all gene combinations. CAR-T cells equipped with logic circuits discovered through this methodology are expected to distinguish cancerous cells from normal cells like computers, thereby minimizing side effects and maximizing the effects of chemotherapy.
Dr. Joonha Kwon, who is the first author of this paper, said, “this research suggests a new method that hasn’t been tried before. What’s particularly noteworthy is the process in which we found the optimal CAR cell circuit through simulations of millions of individual tumors and normal cells.” He added, “This is an innovative technology that can apply AI and computer logic circuits to immune cell engineering. It would contribute greatly to expanding CAR therapy, which is being successfully used for blood cancer, to solid cancers as well.”
This research was funded by the Original Technology Development Project and Research Program for Next Generation Applied Omic of the Korea Research Foundation.
Figure 1. A schematic diagram of manufacturing and administration process of CAR therapy and of cancer cell-specific dual targeting using CAR.
Figure 2. Deep learning (convolutional neural networks, CNNs) algorithm for selection of dual targets based on gene combination (left) and algorithm for calculating expressing cell fractions by gene combination according to logical circuit (right).
2023.03.09 View 6065 -
 KAIST presents a fundamental technology to remove metastatic traits from lung cancer cells
KAIST (President Kwang Hyung Lee) announced on January 30th that a research team led by Professor Kwang-Hyun Cho from the Department of Bio and Brain Engineering succeeded in using systems biology research to change the properties of carcinogenic cells in the lungs and eliminate both drug resistance and their ability to proliferate out to other areas of the body.
As the incidences of cancer increase within aging populations, cancer has become the most lethal disease threatening healthy life. Fatality rates are especially high when early detection does not happen in time and metastasis has occurred in various organs. In order to resolve this problem, a series of attempts were made to remove or lower the ability of cancer cells to spread, but they resulted in cancer cells in the intermediate state becoming more unstable and even more malignant, which created serious treatment challenges.
Professor Kwang-Hyun Cho's research team simulated various cancer cell states in the Epithelial-to-Mesenchymal Transition (EMT) of lung cancer cells, between epithelial cells without metastatic ability and mesenchymal cells with metastatic ability. A mathematical model of molecular network was established, and key regulators that could reverse the state of invasive and drug resistant mesenchymal cells back to the epithelial state were discovered through computer simulation analysis and molecular cell experiments. In particular, this process succeeded in properly reverting the mesenchymal lung cancer cells to a state where they were sensitive to chemotherapy treatment while avoiding the unstable EMT hybrid cell state in the middle process, which had remained a difficult problem.
The results of this research, in which KAIST Ph.D. student Namhee Kim, Dr. Chae Young Hwang, Researcher Taeyoung Kim, and Ph.D. student Hyunjin Kim participated, were published as an online paper in the international journal “Cancer Research” published by the American Association for Cancer Research (AACR) on January 30th. (Paper title: A cell fate reprogramming strategy reverses epithelial-to-mesenchymal transition of lung cancer cells while avoiding hybrid states)
Cells in an EMT hybrid state, which are caused by incomplete transitions during the EMT process in cancer cells, have the characteristics of both epithelial cells and mesenchymal cells, and are known to have high drug resistance and metastatic potential by acquiring high stem cell capacity. In particular, EMT is further enhanced through factors such as transforming growth factor-beta (TGF-β) secreted from the tumor microenvironment (TME) and, as a result, various cell states with high plasticity appear. Due to the complexity of EMT, it has been very difficult to completely reverse the transitional process of the mesenchymal cancer cells to an epithelial cell state in which metastatic ability and drug resistance are eliminated while avoiding the EMT hybrid cell state with high metastatic ability and drug resistance.
Professor Kwang-Hyun Cho's research team established a mathematical model of the gene regulation network that governs the complex process of EMT, and then applied large-scale computer simulation analysis and complex system network control technology to identify and verify 'p53', 'SMAD4', and 'ERK1' and 'ERK 2' (collectively ERKs) through molecular cell experiments as the three key molecular targets that can transform lung cancer cells in the mesenchymal cell state, reversed back to an epithelial cell state that no longer demonstrates the ability to metastasize, while avoiding the EMT hybrid cell state.
In particular, by analyzing the molecular regulatory mechanism of the complex EMT process at the system level, the key pathways were identified that were linked to the positive feedback that plays an important role in completely returning cancer cells to an epithelial cell state in which metastatic ability and drug resistance are removed.
This discovery is significant in that it proved that mesenchymal cells can be reverted to the state of epithelial cells under conditions where TGF-β stimulation are present, like they are in the actual environment where cancer tissue forms in the human body.
Abnormal EMT in cancer cells leads to various malignant traits such as the migration and invasion of cancer cells, changes in responsiveness to chemotherapy treatment, enhanced stem cell function, and the dissemination of cancer. In particular, the acquisition of the metastatic ability of cancer cells is a key determinant factor for the prognosis of cancer patients. The EMT reversal technology in lung cancer cells developed in this research is a new anti-cancer treatment strategy that reprograms cancer cells to eliminate their high plasticity and metastatic potential and increase their responsiveness to chemotherapy.
Professor Kwang-Hyun Cho said, "By succeeding in reversing the state of lung cancer cells that acquired high metastatic traits and resistance to drugs and reverting them to a treatable epithelial cell state with renewed sensitivity to chemotherapy, the research findings propose a new strategy for treatments that can improve the prognosis of cancer patients.”
Professor Kwang-Hyun Cho's research team was the first to present the principle of reversal treatment to revert cancer cells to normal cells, following through with the announcement of the results of their study that reverted colon cancer cells to normal colon cells in January of 2020, and also presenting successful re-programming research where the most malignant basal type breast cancer cells turned into less-malignant luminal type breast cancer cells that were treatable with hormonal therapies in January of 2022. This latest research result is the third in the development of reversal technology where lung cancer cells that had acquired metastatic traits returned to a state in which their metastatic ability was removed and drug sensitivity was enhanced.
This research was carried out with support from the Ministry of Science and ICT and the National Research Foundation of Korea's Basic Research in Science & Engineering Program for Mid-Career Researchers.
< Figure 1. Construction of the mathematical model of the regulatory network to represent the EMT phenotype based on the interaction between various molecules related to EMT.
(A) Professor Kwang-Hyun Cho's research team investigated numerous literatures and databases related to complex EMT, and based on comparative analysis of cell line data showing epithelial and mesenchymal cell conditions, they extracted key signaling pathways related to EMT and built a mathematical model of regulatory network (B) By comparing the results of computer simulation analysis and the molecular cell experiments, it was verified how well the constructed mathematical model simulated the actual cellular phenomena. >
< Figure 2. Understanding of various EMT phenotypes through large-scale computer simulation analysis and complex system network control technology.
(A) Through computer simulation analysis and experiments, Professor Kwang-Hyun Cho's research team found that complete control of EMT is impossible with single-molecule control alone. In particular, through comparison of the relative stability of attractors, it was revealed that the cell state exhibiting EMT hybrid characteristics has unstable properties. (B), (C) Based on these results, Prof. Cho’s team identified two feedbacks (positive feedback consisting of Snail-miR-34 and ZEB1-miR-200) that play an important role in avoiding the EMT hybrid state that appeared in the TGF-β-ON state. It was found through computer simulation analysis that the two feedbacks restore relatively high stability when the excavated p53 and SMAD4 are regulated. In addition, molecular cell experiments demonstrated that the expression levels of E-cad and ZEB1, which are representative phenotypic markers of EMT, changed similarly to the expression profile in the epithelial cell state, despite the TGF-β-ON state. >
< Figure 3. Complex molecular network analysis and discovery of reprogramming molecular targets for intact elimination of EMT hybrid features.
(A) Controlling the expression of p53 and SMAD4 in lung cancer cell lines was expected to overcome drug resistance, but contrary to expectations, chemotherapy responsiveness was not restored. (B) Professor Kwang-Hyun Cho's research team additionally analyzed computer simulations, genome data, and experimental results and found that high expression levels of TWIST1 and EPCAM were related to drug resistance. (C) Prof. Cho’s team identified three key molecular targets: p53, SMAD4 and ERK1 & ERK2. (D), (E) Furthermore, they identified a key pathway that plays an important role in completely reversing into epithelial cells while avoiding EMT hybrid characteristics, and confirmed through network analysis and attractor analysis that high stability of the key pathway was restored when the proposed molecular target was controlled. >
< Figure 4. Verification through experiments with lung cancer cell lines.
When p53 was activated and SMAD4 and ERK1/2 were inhibited in lung cancer cell lines, (A), (B) E-cad protein expression increased and ZEB1 protein expression decreased, and (C) mesenchymal cell status including TWIST1 and EPCAM and gene expression of markers related to stem cell potential characteristics were completely inhibited. In addition, (D) it was confirmed that resistance to chemotherapy treatment was also overcome as the cell state was reversed by the regulated target. >
< Figure 5. A schematic representation of the research results.
Prof. Cho’s research team identified key molecular regulatory pathways to avoid high plasticity formed by abnormal EMT of cancer cells and reverse it to an epithelial cell state through systems biology research. From this analysis, a reprogramming molecular target that can reverse the state of mesenchymal cells with acquired invasiveness and drug resistance to the state of epithelial cells with restored drug responsiveness was discovered.
For lung cancer cells, when a drug that enhances the expression of p53, one of the molecular targets discovered, and inhibits the expression of SMAD4 and ERK1 & ERK2 is administered, the molecular network of genes in the state of mesenchymal cells is modified, eventually eliminating metastatic ability and it is reprogrammed to turn into epithelial cells without the resistance to chemotherapy treatments. >
2023.01.30 View 11771
KAIST presents a fundamental technology to remove metastatic traits from lung cancer cells
KAIST (President Kwang Hyung Lee) announced on January 30th that a research team led by Professor Kwang-Hyun Cho from the Department of Bio and Brain Engineering succeeded in using systems biology research to change the properties of carcinogenic cells in the lungs and eliminate both drug resistance and their ability to proliferate out to other areas of the body.
As the incidences of cancer increase within aging populations, cancer has become the most lethal disease threatening healthy life. Fatality rates are especially high when early detection does not happen in time and metastasis has occurred in various organs. In order to resolve this problem, a series of attempts were made to remove or lower the ability of cancer cells to spread, but they resulted in cancer cells in the intermediate state becoming more unstable and even more malignant, which created serious treatment challenges.
Professor Kwang-Hyun Cho's research team simulated various cancer cell states in the Epithelial-to-Mesenchymal Transition (EMT) of lung cancer cells, between epithelial cells without metastatic ability and mesenchymal cells with metastatic ability. A mathematical model of molecular network was established, and key regulators that could reverse the state of invasive and drug resistant mesenchymal cells back to the epithelial state were discovered through computer simulation analysis and molecular cell experiments. In particular, this process succeeded in properly reverting the mesenchymal lung cancer cells to a state where they were sensitive to chemotherapy treatment while avoiding the unstable EMT hybrid cell state in the middle process, which had remained a difficult problem.
The results of this research, in which KAIST Ph.D. student Namhee Kim, Dr. Chae Young Hwang, Researcher Taeyoung Kim, and Ph.D. student Hyunjin Kim participated, were published as an online paper in the international journal “Cancer Research” published by the American Association for Cancer Research (AACR) on January 30th. (Paper title: A cell fate reprogramming strategy reverses epithelial-to-mesenchymal transition of lung cancer cells while avoiding hybrid states)
Cells in an EMT hybrid state, which are caused by incomplete transitions during the EMT process in cancer cells, have the characteristics of both epithelial cells and mesenchymal cells, and are known to have high drug resistance and metastatic potential by acquiring high stem cell capacity. In particular, EMT is further enhanced through factors such as transforming growth factor-beta (TGF-β) secreted from the tumor microenvironment (TME) and, as a result, various cell states with high plasticity appear. Due to the complexity of EMT, it has been very difficult to completely reverse the transitional process of the mesenchymal cancer cells to an epithelial cell state in which metastatic ability and drug resistance are eliminated while avoiding the EMT hybrid cell state with high metastatic ability and drug resistance.
Professor Kwang-Hyun Cho's research team established a mathematical model of the gene regulation network that governs the complex process of EMT, and then applied large-scale computer simulation analysis and complex system network control technology to identify and verify 'p53', 'SMAD4', and 'ERK1' and 'ERK 2' (collectively ERKs) through molecular cell experiments as the three key molecular targets that can transform lung cancer cells in the mesenchymal cell state, reversed back to an epithelial cell state that no longer demonstrates the ability to metastasize, while avoiding the EMT hybrid cell state.
In particular, by analyzing the molecular regulatory mechanism of the complex EMT process at the system level, the key pathways were identified that were linked to the positive feedback that plays an important role in completely returning cancer cells to an epithelial cell state in which metastatic ability and drug resistance are removed.
This discovery is significant in that it proved that mesenchymal cells can be reverted to the state of epithelial cells under conditions where TGF-β stimulation are present, like they are in the actual environment where cancer tissue forms in the human body.
Abnormal EMT in cancer cells leads to various malignant traits such as the migration and invasion of cancer cells, changes in responsiveness to chemotherapy treatment, enhanced stem cell function, and the dissemination of cancer. In particular, the acquisition of the metastatic ability of cancer cells is a key determinant factor for the prognosis of cancer patients. The EMT reversal technology in lung cancer cells developed in this research is a new anti-cancer treatment strategy that reprograms cancer cells to eliminate their high plasticity and metastatic potential and increase their responsiveness to chemotherapy.
Professor Kwang-Hyun Cho said, "By succeeding in reversing the state of lung cancer cells that acquired high metastatic traits and resistance to drugs and reverting them to a treatable epithelial cell state with renewed sensitivity to chemotherapy, the research findings propose a new strategy for treatments that can improve the prognosis of cancer patients.”
Professor Kwang-Hyun Cho's research team was the first to present the principle of reversal treatment to revert cancer cells to normal cells, following through with the announcement of the results of their study that reverted colon cancer cells to normal colon cells in January of 2020, and also presenting successful re-programming research where the most malignant basal type breast cancer cells turned into less-malignant luminal type breast cancer cells that were treatable with hormonal therapies in January of 2022. This latest research result is the third in the development of reversal technology where lung cancer cells that had acquired metastatic traits returned to a state in which their metastatic ability was removed and drug sensitivity was enhanced.
This research was carried out with support from the Ministry of Science and ICT and the National Research Foundation of Korea's Basic Research in Science & Engineering Program for Mid-Career Researchers.
< Figure 1. Construction of the mathematical model of the regulatory network to represent the EMT phenotype based on the interaction between various molecules related to EMT.
(A) Professor Kwang-Hyun Cho's research team investigated numerous literatures and databases related to complex EMT, and based on comparative analysis of cell line data showing epithelial and mesenchymal cell conditions, they extracted key signaling pathways related to EMT and built a mathematical model of regulatory network (B) By comparing the results of computer simulation analysis and the molecular cell experiments, it was verified how well the constructed mathematical model simulated the actual cellular phenomena. >
< Figure 2. Understanding of various EMT phenotypes through large-scale computer simulation analysis and complex system network control technology.
(A) Through computer simulation analysis and experiments, Professor Kwang-Hyun Cho's research team found that complete control of EMT is impossible with single-molecule control alone. In particular, through comparison of the relative stability of attractors, it was revealed that the cell state exhibiting EMT hybrid characteristics has unstable properties. (B), (C) Based on these results, Prof. Cho’s team identified two feedbacks (positive feedback consisting of Snail-miR-34 and ZEB1-miR-200) that play an important role in avoiding the EMT hybrid state that appeared in the TGF-β-ON state. It was found through computer simulation analysis that the two feedbacks restore relatively high stability when the excavated p53 and SMAD4 are regulated. In addition, molecular cell experiments demonstrated that the expression levels of E-cad and ZEB1, which are representative phenotypic markers of EMT, changed similarly to the expression profile in the epithelial cell state, despite the TGF-β-ON state. >
< Figure 3. Complex molecular network analysis and discovery of reprogramming molecular targets for intact elimination of EMT hybrid features.
(A) Controlling the expression of p53 and SMAD4 in lung cancer cell lines was expected to overcome drug resistance, but contrary to expectations, chemotherapy responsiveness was not restored. (B) Professor Kwang-Hyun Cho's research team additionally analyzed computer simulations, genome data, and experimental results and found that high expression levels of TWIST1 and EPCAM were related to drug resistance. (C) Prof. Cho’s team identified three key molecular targets: p53, SMAD4 and ERK1 & ERK2. (D), (E) Furthermore, they identified a key pathway that plays an important role in completely reversing into epithelial cells while avoiding EMT hybrid characteristics, and confirmed through network analysis and attractor analysis that high stability of the key pathway was restored when the proposed molecular target was controlled. >
< Figure 4. Verification through experiments with lung cancer cell lines.
When p53 was activated and SMAD4 and ERK1/2 were inhibited in lung cancer cell lines, (A), (B) E-cad protein expression increased and ZEB1 protein expression decreased, and (C) mesenchymal cell status including TWIST1 and EPCAM and gene expression of markers related to stem cell potential characteristics were completely inhibited. In addition, (D) it was confirmed that resistance to chemotherapy treatment was also overcome as the cell state was reversed by the regulated target. >
< Figure 5. A schematic representation of the research results.
Prof. Cho’s research team identified key molecular regulatory pathways to avoid high plasticity formed by abnormal EMT of cancer cells and reverse it to an epithelial cell state through systems biology research. From this analysis, a reprogramming molecular target that can reverse the state of mesenchymal cells with acquired invasiveness and drug resistance to the state of epithelial cells with restored drug responsiveness was discovered.
For lung cancer cells, when a drug that enhances the expression of p53, one of the molecular targets discovered, and inhibits the expression of SMAD4 and ERK1 & ERK2 is administered, the molecular network of genes in the state of mesenchymal cells is modified, eventually eliminating metastatic ability and it is reprogrammed to turn into epithelial cells without the resistance to chemotherapy treatments. >
2023.01.30 View 11771 -
 Decoding Brain Signals to Control a Robotic Arm
Advanced brain-machine interface system successfully interprets arm movement directions from neural signals in the brain
Researchers have developed a mind-reading system for decoding neural signals from the brain during arm movement. The method, described in the journal Applied Soft Computing, can be used by a person to control a robotic arm through a brain-machine interface (BMI).
A BMI is a device that translates nerve signals into commands to control a machine, such as a computer or a robotic limb. There are two main techniques for monitoring neural signals in BMIs: electroencephalography (EEG) and electrocorticography (ECoG).
The EEG exhibits signals from electrodes on the surface of the scalp and is widely employed because it is non-invasive, relatively cheap, safe and easy to use. However, the EEG has low spatial resolution and detects irrelevant neural signals, which makes it difficult to interpret the intentions of individuals from the EEG.
On the other hand, the ECoG is an invasive method that involves placing electrodes directly on the surface of the cerebral cortex below the scalp. Compared with the EEG, the ECoG can monitor neural signals with much higher spatial resolution and less background noise. However, this technique has several drawbacks.
“The ECoG is primarily used to find potential sources of epileptic seizures, meaning the electrodes are placed in different locations for different patients and may not be in the optimal regions of the brain for detecting sensory and movement signals,” explained Professor Jaeseung Jeong, a brain scientist at KAIST. “This inconsistency makes it difficult to decode brain signals to predict movements.”
To overcome these problems, Professor Jeong’s team developed a new method for decoding ECoG neural signals during arm movement. The system is based on a machine-learning system for analysing and predicting neural signals called an ‘echo-state network’ and a mathematical probability model called the Gaussian distribution.
In the study, the researchers recorded ECoG signals from four individuals with epilepsy while they were performing a reach-and-grasp task. Because the ECoG electrodes were placed according to the potential sources of each patient’s epileptic seizures, only 22% to 44% of the electrodes were located in the regions of the brain responsible for controlling movement.
During the movement task, the participants were given visual cues, either by placing a real tennis ball in front of them, or via a virtual reality headset showing a clip of a human arm reaching forward in first-person view. They were asked to reach forward, grasp an object, then return their hand and release the object, while wearing motion sensors on their wrists and fingers. In a second task, they were instructed to imagine reaching forward without moving their arms.
The researchers monitored the signals from the ECoG electrodes during real and imaginary arm movements, and tested whether the new system could predict the direction of this movement from the neural signals. They found that the novel decoder successfully classified arm movements in 24 directions in three-dimensional space, both in the real and virtual tasks, and that the results were at least five times more accurate than chance. They also used a computer simulation to show that the novel ECoG decoder could control the movements of a robotic arm.
Overall, the results suggest that the new machine learning-based BCI system successfully used ECoG signals to interpret the direction of the intended movements. The next steps will be to improve the accuracy and efficiency of the decoder. In the future, it could be used in a real-time BMI device to help people with movement or sensory impairments.
This research was supported by the KAIST Global Singularity Research Program of 2021, Brain Research Program of the National Research Foundation of Korea funded by the Ministry of Science, ICT, and Future Planning, and the Basic Science Research Program through the National Research Foundation of Korea funded by the Ministry of Education.
-PublicationHoon-Hee Kim, Jaeseung Jeong, “An electrocorticographic decoder for arm movement for brain-machine interface using an echo state network and Gaussian readout,” Applied SoftComputing online December 31, 2021 (doi.org/10.1016/j.asoc.2021.108393)
-ProfileProfessor Jaeseung JeongDepartment of Bio and Brain EngineeringCollege of EngineeringKAIST
2022.03.18 View 9202
Decoding Brain Signals to Control a Robotic Arm
Advanced brain-machine interface system successfully interprets arm movement directions from neural signals in the brain
Researchers have developed a mind-reading system for decoding neural signals from the brain during arm movement. The method, described in the journal Applied Soft Computing, can be used by a person to control a robotic arm through a brain-machine interface (BMI).
A BMI is a device that translates nerve signals into commands to control a machine, such as a computer or a robotic limb. There are two main techniques for monitoring neural signals in BMIs: electroencephalography (EEG) and electrocorticography (ECoG).
The EEG exhibits signals from electrodes on the surface of the scalp and is widely employed because it is non-invasive, relatively cheap, safe and easy to use. However, the EEG has low spatial resolution and detects irrelevant neural signals, which makes it difficult to interpret the intentions of individuals from the EEG.
On the other hand, the ECoG is an invasive method that involves placing electrodes directly on the surface of the cerebral cortex below the scalp. Compared with the EEG, the ECoG can monitor neural signals with much higher spatial resolution and less background noise. However, this technique has several drawbacks.
“The ECoG is primarily used to find potential sources of epileptic seizures, meaning the electrodes are placed in different locations for different patients and may not be in the optimal regions of the brain for detecting sensory and movement signals,” explained Professor Jaeseung Jeong, a brain scientist at KAIST. “This inconsistency makes it difficult to decode brain signals to predict movements.”
To overcome these problems, Professor Jeong’s team developed a new method for decoding ECoG neural signals during arm movement. The system is based on a machine-learning system for analysing and predicting neural signals called an ‘echo-state network’ and a mathematical probability model called the Gaussian distribution.
In the study, the researchers recorded ECoG signals from four individuals with epilepsy while they were performing a reach-and-grasp task. Because the ECoG electrodes were placed according to the potential sources of each patient’s epileptic seizures, only 22% to 44% of the electrodes were located in the regions of the brain responsible for controlling movement.
During the movement task, the participants were given visual cues, either by placing a real tennis ball in front of them, or via a virtual reality headset showing a clip of a human arm reaching forward in first-person view. They were asked to reach forward, grasp an object, then return their hand and release the object, while wearing motion sensors on their wrists and fingers. In a second task, they were instructed to imagine reaching forward without moving their arms.
The researchers monitored the signals from the ECoG electrodes during real and imaginary arm movements, and tested whether the new system could predict the direction of this movement from the neural signals. They found that the novel decoder successfully classified arm movements in 24 directions in three-dimensional space, both in the real and virtual tasks, and that the results were at least five times more accurate than chance. They also used a computer simulation to show that the novel ECoG decoder could control the movements of a robotic arm.
Overall, the results suggest that the new machine learning-based BCI system successfully used ECoG signals to interpret the direction of the intended movements. The next steps will be to improve the accuracy and efficiency of the decoder. In the future, it could be used in a real-time BMI device to help people with movement or sensory impairments.
This research was supported by the KAIST Global Singularity Research Program of 2021, Brain Research Program of the National Research Foundation of Korea funded by the Ministry of Science, ICT, and Future Planning, and the Basic Science Research Program through the National Research Foundation of Korea funded by the Ministry of Education.
-PublicationHoon-Hee Kim, Jaeseung Jeong, “An electrocorticographic decoder for arm movement for brain-machine interface using an echo state network and Gaussian readout,” Applied SoftComputing online December 31, 2021 (doi.org/10.1016/j.asoc.2021.108393)
-ProfileProfessor Jaeseung JeongDepartment of Bio and Brain EngineeringCollege of EngineeringKAIST
2022.03.18 View 9202 -
 Five Projects Ranked in the Top 100 for National R&D Excellence
Five KAIST research projects were selected as the 2021 Top 100 for National R&D Excellence by the Ministry of Science and ICT and the Korea Institute of Science & Technology Evaluation and Planning.
The five projects are:-The development of E. coli that proliferates with only formic acid and carbon dioxide by Distinguished Professor Sang Yup Lee from the Department of Chemical and Biomolecular Engineering
-An original reverse aging technology that restores an old human skin cell into a younger one by Professor Kwang-Hyun Cho from the Department of Bio and Brain Engineering-The development of next-generation high-efficiency perovskite-silicon tandem solar cells by Professor Byungha Shin from the Department of Materials Science and Engineering-Research on the effects of ultrafine dust in the atmosphere has on energy consumption by Professor Jiyong Eom from the School of Business and Technology Management-Research on a molecular trigger that controls the phase transformation of bio materials by Professor Myungchul Kim from the Department of Bio and Brain Engineering
Started in 2006, an Evaluation Committee composed of experts in industries, universities, and research institutes has made the preliminary selections of the most outstanding research projects based on their significance as a scientific and technological development and their socioeconomic effects. The finalists went through an open public evaluation. The final 100 studies are from six fields: 18 from mechanics & materials, 26 from biology & marine sciences, 19 from ICT & electronics, 10 from interdisciplinary research, and nine from natural science and infrastructure.
The selected 100 studies will receive a certificate and an award plaque from the minister of MSIT as well as additional points for business and institutional evaluations according to appropriate regulations, and the selected researchers will be strongly recommended as candidates for national meritorious awards.
In particular, to help the 100 selected research projects become more accessible for the general public, their main contents will be provided in a free e-book ‘The Top 100 for National R&D Excellence of 2021’ that will be available from online booksellers.
2022.02.17 View 8222
Five Projects Ranked in the Top 100 for National R&D Excellence
Five KAIST research projects were selected as the 2021 Top 100 for National R&D Excellence by the Ministry of Science and ICT and the Korea Institute of Science & Technology Evaluation and Planning.
The five projects are:-The development of E. coli that proliferates with only formic acid and carbon dioxide by Distinguished Professor Sang Yup Lee from the Department of Chemical and Biomolecular Engineering
-An original reverse aging technology that restores an old human skin cell into a younger one by Professor Kwang-Hyun Cho from the Department of Bio and Brain Engineering-The development of next-generation high-efficiency perovskite-silicon tandem solar cells by Professor Byungha Shin from the Department of Materials Science and Engineering-Research on the effects of ultrafine dust in the atmosphere has on energy consumption by Professor Jiyong Eom from the School of Business and Technology Management-Research on a molecular trigger that controls the phase transformation of bio materials by Professor Myungchul Kim from the Department of Bio and Brain Engineering
Started in 2006, an Evaluation Committee composed of experts in industries, universities, and research institutes has made the preliminary selections of the most outstanding research projects based on their significance as a scientific and technological development and their socioeconomic effects. The finalists went through an open public evaluation. The final 100 studies are from six fields: 18 from mechanics & materials, 26 from biology & marine sciences, 19 from ICT & electronics, 10 from interdisciplinary research, and nine from natural science and infrastructure.
The selected 100 studies will receive a certificate and an award plaque from the minister of MSIT as well as additional points for business and institutional evaluations according to appropriate regulations, and the selected researchers will be strongly recommended as candidates for national meritorious awards.
In particular, to help the 100 selected research projects become more accessible for the general public, their main contents will be provided in a free e-book ‘The Top 100 for National R&D Excellence of 2021’ that will be available from online booksellers.
2022.02.17 View 8222 -
 AI Light-Field Camera Reads 3D Facial Expressions
Machine-learned, light-field camera reads facial expressions from high-contrast illumination invariant 3D facial images
A joint research team led by Professors Ki-Hun Jeong and Doheon Lee from the KAIST Department of Bio and Brain Engineering reported the development of a technique for facial expression detection by merging near-infrared light-field camera techniques with artificial intelligence (AI) technology.
Unlike a conventional camera, the light-field camera contains micro-lens arrays in front of the image sensor, which makes the camera small enough to fit into a smart phone, while allowing it to acquire the spatial and directional information of the light with a single shot. The technique has received attention as it can reconstruct images in a variety of ways including multi-views, refocusing, and 3D image acquisition, giving rise to many potential applications.
However, the optical crosstalk between shadows caused by external light sources in the environment and the micro-lens has limited existing light-field cameras from being able to provide accurate image contrast and 3D reconstruction.
The joint research team applied a vertical-cavity surface-emitting laser (VCSEL) in the near-IR range to stabilize the accuracy of 3D image reconstruction that previously depended on environmental light. When an external light source is shone on a face at 0-, 30-, and 60-degree angles, the light field camera reduces 54% of image reconstruction errors. Additionally, by inserting a light-absorbing layer for visible and near-IR wavelengths between the micro-lens arrays, the team could minimize optical crosstalk while increasing the image contrast by 2.1 times.
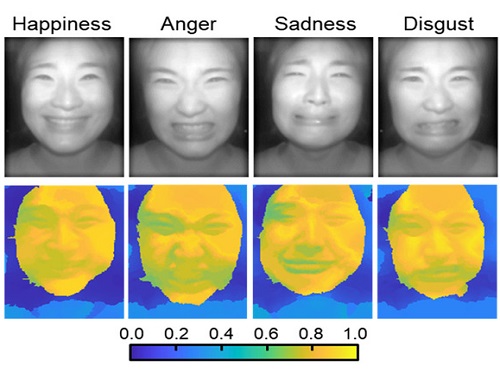
Through this technique, the team could overcome the limitations of existing light-field cameras and was able to develop their NIR-based light-field camera (NIR-LFC), optimized for the 3D image reconstruction of facial expressions. Using the NIR-LFC, the team acquired high-quality 3D reconstruction images of facial expressions expressing various emotions regardless of the lighting conditions of the surrounding environment.
The facial expressions in the acquired 3D images were distinguished through machine learning with an average of 85% accuracy – a statistically significant figure compared to when 2D images were used. Furthermore, by calculating the interdependency of distance information that varies with facial expression in 3D images, the team could identify the information a light-field camera utilizes to distinguish human expressions.
Professor Ki-Hun Jeong said, “The sub-miniature light-field camera developed by the research team has the potential to become the new platform to quantitatively analyze the facial expressions and emotions of humans.” To highlight the significance of this research, he added, “It could be applied in various fields including mobile healthcare, field diagnosis, social cognition, and human-machine interactions.”
This research was published in Advanced Intelligent Systems online on December 16, under the title, “Machine-Learned Light-field Camera that Reads Facial Expression from High-Contrast and Illumination Invariant 3D Facial Images.” This research was funded by the Ministry of Science and ICT and the Ministry of Trade, Industry and Energy.
-Publication“Machine-learned light-field camera that reads fascial expression from high-contrast and illumination invariant 3D facial images,” Sang-In Bae, Sangyeon Lee, Jae-Myeong Kwon, Hyun-Kyung Kim. Kyung-Won Jang, Doheon Lee, Ki-Hun Jeong, Advanced Intelligent Systems, December 16, 2021 (doi.org/10.1002/aisy.202100182)
ProfileProfessor Ki-Hun JeongBiophotonic LaboratoryDepartment of Bio and Brain EngineeringKAIST
Professor Doheon LeeDepartment of Bio and Brain EngineeringKAIST
2022.01.21 View 9604
AI Light-Field Camera Reads 3D Facial Expressions
Machine-learned, light-field camera reads facial expressions from high-contrast illumination invariant 3D facial images
A joint research team led by Professors Ki-Hun Jeong and Doheon Lee from the KAIST Department of Bio and Brain Engineering reported the development of a technique for facial expression detection by merging near-infrared light-field camera techniques with artificial intelligence (AI) technology.
Unlike a conventional camera, the light-field camera contains micro-lens arrays in front of the image sensor, which makes the camera small enough to fit into a smart phone, while allowing it to acquire the spatial and directional information of the light with a single shot. The technique has received attention as it can reconstruct images in a variety of ways including multi-views, refocusing, and 3D image acquisition, giving rise to many potential applications.
However, the optical crosstalk between shadows caused by external light sources in the environment and the micro-lens has limited existing light-field cameras from being able to provide accurate image contrast and 3D reconstruction.
The joint research team applied a vertical-cavity surface-emitting laser (VCSEL) in the near-IR range to stabilize the accuracy of 3D image reconstruction that previously depended on environmental light. When an external light source is shone on a face at 0-, 30-, and 60-degree angles, the light field camera reduces 54% of image reconstruction errors. Additionally, by inserting a light-absorbing layer for visible and near-IR wavelengths between the micro-lens arrays, the team could minimize optical crosstalk while increasing the image contrast by 2.1 times.
Through this technique, the team could overcome the limitations of existing light-field cameras and was able to develop their NIR-based light-field camera (NIR-LFC), optimized for the 3D image reconstruction of facial expressions. Using the NIR-LFC, the team acquired high-quality 3D reconstruction images of facial expressions expressing various emotions regardless of the lighting conditions of the surrounding environment.
The facial expressions in the acquired 3D images were distinguished through machine learning with an average of 85% accuracy – a statistically significant figure compared to when 2D images were used. Furthermore, by calculating the interdependency of distance information that varies with facial expression in 3D images, the team could identify the information a light-field camera utilizes to distinguish human expressions.
Professor Ki-Hun Jeong said, “The sub-miniature light-field camera developed by the research team has the potential to become the new platform to quantitatively analyze the facial expressions and emotions of humans.” To highlight the significance of this research, he added, “It could be applied in various fields including mobile healthcare, field diagnosis, social cognition, and human-machine interactions.”
This research was published in Advanced Intelligent Systems online on December 16, under the title, “Machine-Learned Light-field Camera that Reads Facial Expression from High-Contrast and Illumination Invariant 3D Facial Images.” This research was funded by the Ministry of Science and ICT and the Ministry of Trade, Industry and Energy.
-Publication“Machine-learned light-field camera that reads fascial expression from high-contrast and illumination invariant 3D facial images,” Sang-In Bae, Sangyeon Lee, Jae-Myeong Kwon, Hyun-Kyung Kim. Kyung-Won Jang, Doheon Lee, Ki-Hun Jeong, Advanced Intelligent Systems, December 16, 2021 (doi.org/10.1002/aisy.202100182)
ProfileProfessor Ki-Hun JeongBiophotonic LaboratoryDepartment of Bio and Brain EngineeringKAIST
Professor Doheon LeeDepartment of Bio and Brain EngineeringKAIST
2022.01.21 View 9604 -
 Face Detection in Untrained Deep Neural Networks
A KAIST team shows that primitive visual selectivity of faces can arise spontaneously in completely untrained deep neural networks
Researchers have found that higher visual cognitive functions can arise spontaneously in untrained neural networks. A KAIST research team led by Professor Se-Bum Paik from the Department of Bio and Brain Engineering has shown that visual selectivity of facial images can arise even in completely untrained deep neural networks.
This new finding has provided revelatory insights into mechanisms underlying the development of cognitive functions in both biological and artificial neural networks, also making a significant impact on our understanding of the origin of early brain functions before sensory experiences.
The study published in Nature Communications on December 16 demonstrates that neuronal activities selective to facial images are observed in randomly initialized deep neural networks in the complete absence of learning, and that they show the characteristics of those observed in biological brains.
The ability to identify and recognize faces is a crucial function for social behavior, and this ability is thought to originate from neuronal tuning at the single or multi-neuronal level. Neurons that selectively respond to faces are observed in young animals of various species, and this raises intense debate whether face-selective neurons can arise innately in the brain or if they require visual experience.
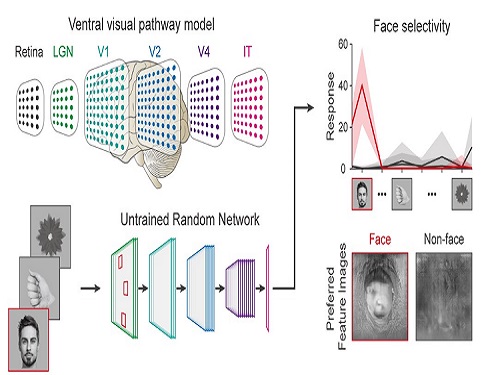
Using a model neural network that captures properties of the ventral stream of the visual cortex, the research team found that face-selectivity can emerge spontaneously from random feedforward wirings in untrained deep neural networks. The team showed that the character of this innate face-selectivity is comparable to that observed with face-selective neurons in the brain, and that this spontaneous neuronal tuning for faces enables the network to perform face detection tasks.
These results imply a possible scenario in which the random feedforward connections that develop in early, untrained networks may be sufficient for initializing primitive visual cognitive functions.
Professor Paik said, “Our findings suggest that innate cognitive functions can emerge spontaneously from the statistical complexity embedded in the hierarchical feedforward projection circuitry, even in the complete absence of learning”.
He continued, “Our results provide a broad conceptual advance as well as advanced insight into the mechanisms underlying the development of innate functions in both biological and artificial neural networks, which may unravel the mystery of the generation and evolution of intelligence.” This work was supported by the National Research Foundation of Korea (NRF) and by the KAIST singularity research project.
-PublicationSeungdae Baek, Min Song, Jaeson Jang, Gwangsu Kim, and Se-Bum Baik, “Face detection in untrained deep neural network,” Nature Communications 12, 7328 on Dec.16, 2021
(https://doi.org/10.1038/s41467-021-27606-9)
-ProfileProfessor Se-Bum PaikVisual System and Neural Network LaboratoryProgram of Brain and Cognitive EngineeringDepartment of Bio and Brain EngineeringCollege of EngineeringKAIST
2021.12.21 View 8337
Face Detection in Untrained Deep Neural Networks
A KAIST team shows that primitive visual selectivity of faces can arise spontaneously in completely untrained deep neural networks
Researchers have found that higher visual cognitive functions can arise spontaneously in untrained neural networks. A KAIST research team led by Professor Se-Bum Paik from the Department of Bio and Brain Engineering has shown that visual selectivity of facial images can arise even in completely untrained deep neural networks.
This new finding has provided revelatory insights into mechanisms underlying the development of cognitive functions in both biological and artificial neural networks, also making a significant impact on our understanding of the origin of early brain functions before sensory experiences.
The study published in Nature Communications on December 16 demonstrates that neuronal activities selective to facial images are observed in randomly initialized deep neural networks in the complete absence of learning, and that they show the characteristics of those observed in biological brains.
The ability to identify and recognize faces is a crucial function for social behavior, and this ability is thought to originate from neuronal tuning at the single or multi-neuronal level. Neurons that selectively respond to faces are observed in young animals of various species, and this raises intense debate whether face-selective neurons can arise innately in the brain or if they require visual experience.
Using a model neural network that captures properties of the ventral stream of the visual cortex, the research team found that face-selectivity can emerge spontaneously from random feedforward wirings in untrained deep neural networks. The team showed that the character of this innate face-selectivity is comparable to that observed with face-selective neurons in the brain, and that this spontaneous neuronal tuning for faces enables the network to perform face detection tasks.
These results imply a possible scenario in which the random feedforward connections that develop in early, untrained networks may be sufficient for initializing primitive visual cognitive functions.
Professor Paik said, “Our findings suggest that innate cognitive functions can emerge spontaneously from the statistical complexity embedded in the hierarchical feedforward projection circuitry, even in the complete absence of learning”.
He continued, “Our results provide a broad conceptual advance as well as advanced insight into the mechanisms underlying the development of innate functions in both biological and artificial neural networks, which may unravel the mystery of the generation and evolution of intelligence.” This work was supported by the National Research Foundation of Korea (NRF) and by the KAIST singularity research project.
-PublicationSeungdae Baek, Min Song, Jaeson Jang, Gwangsu Kim, and Se-Bum Baik, “Face detection in untrained deep neural network,” Nature Communications 12, 7328 on Dec.16, 2021
(https://doi.org/10.1038/s41467-021-27606-9)
-ProfileProfessor Se-Bum PaikVisual System and Neural Network LaboratoryProgram of Brain and Cognitive EngineeringDepartment of Bio and Brain EngineeringCollege of EngineeringKAIST
2021.12.21 View 8337 -
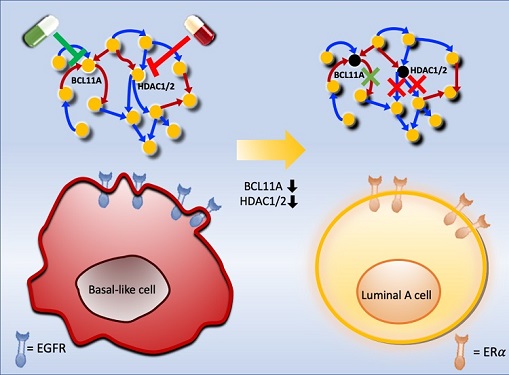 Connecting the Dots to Find New Treatments for Breast Cancer
Systems biologists uncovered new ways of cancer cell reprogramming to treat drug-resistant cancers
Scientists at KAIST believe they may have found a way to reverse an aggressive, treatment-resistant type of breast cancer into a less dangerous kind that responds well to treatment. The study involved the use of mathematical models to untangle the complex genetic and molecular interactions that occur in the two types of breast cancer, but could be extended to find ways for treating many others. The study’s findings were published in the journal Cancer Research.
Basal-like tumours are the most aggressive type of breast cancer, with the worst prognosis. Chemotherapy is the only available treatment option, but patients experience high recurrence rates. On the other hand, luminal-A breast cancer responds well to drugs that specifically target a receptor on their cell surfaces, called estrogen receptor alpha (ERα).
KAIST systems biologist Kwang-Hyun Cho and colleagues analyzed the complex molecular and genetic interactions of basal-like and luminal-A breast cancers to find out if there might be a way to switch the former to the latter and give patients a better chance to respond to treatment.
To do this, they accessed large amounts of cancer and patient data to understand which genes and molecules are involved in the two types. They then input this data into a mathematical model that represents genes, proteins and molecules as dots and the interactions between them as lines. The model can be used to conduct simulations and see how interactions change when certain genes are turned on or off.
“There have been a tremendous number of studies trying to find therapeutic targets for treating basal-like breast cancer patients,” says Cho. “But clinical trials have failed due to the complex and dynamic nature of cancer. To overcome this issue, we looked at breast cancer cells as a complex network system and implemented a systems biological approach to unravel the underlying mechanisms that would allow us to reprogram basal-like into luminal-A breast cancer cells.”
Using this approach, followed by experimental validation on real breast cancer cells, the team found that turning off two key gene regulators, called BCL11A and HDAC1/2, switched a basal-like cancer signalling pathway into a different one used by luminal-A cancer cells. The switch reprograms the cancer cells and makes them more responsive to drugs that target ERα receptors. However, further tests will be needed to confirm that this also works in animal models and eventually humans.
“Our study demonstrates that the systems biological approach can be useful for identifying novel therapeutic targets,” says Cho.
The researchers are now expanding its breast cancer network model to include all breast cancer subtypes. Their ultimate aim is to identify more drug targets and to understand the mechanisms that could drive drug-resistant cells to turn into drug-sensitive ones.
This work was supported by the National Research Foundation of Korea, the Ministry of Science and ICT, Electronics and Telecommunications Research Institute, and the KAIST Grand Challenge 30 Project.
-Publication Sea R. Choi, Chae Young Hwang, Jonghoon Lee, and Kwang-Hyun Cho, “Network Analysis Identifies Regulators of Basal-like Breast Cancer Reprogramming and Endocrine TherapyVulnerability,” Cancer Research, November 30. (doi:10.1158/0008-5472.CAN-21-0621)
-ProfileProfessor Kwang-Hyun ChoLaboratory for Systems Biology and Bio-Inspired EngineeringDepartment of Bio and Brain EngineeringKAIST
2021.12.07 View 7023
Connecting the Dots to Find New Treatments for Breast Cancer
Systems biologists uncovered new ways of cancer cell reprogramming to treat drug-resistant cancers
Scientists at KAIST believe they may have found a way to reverse an aggressive, treatment-resistant type of breast cancer into a less dangerous kind that responds well to treatment. The study involved the use of mathematical models to untangle the complex genetic and molecular interactions that occur in the two types of breast cancer, but could be extended to find ways for treating many others. The study’s findings were published in the journal Cancer Research.
Basal-like tumours are the most aggressive type of breast cancer, with the worst prognosis. Chemotherapy is the only available treatment option, but patients experience high recurrence rates. On the other hand, luminal-A breast cancer responds well to drugs that specifically target a receptor on their cell surfaces, called estrogen receptor alpha (ERα).
KAIST systems biologist Kwang-Hyun Cho and colleagues analyzed the complex molecular and genetic interactions of basal-like and luminal-A breast cancers to find out if there might be a way to switch the former to the latter and give patients a better chance to respond to treatment.
To do this, they accessed large amounts of cancer and patient data to understand which genes and molecules are involved in the two types. They then input this data into a mathematical model that represents genes, proteins and molecules as dots and the interactions between them as lines. The model can be used to conduct simulations and see how interactions change when certain genes are turned on or off.
“There have been a tremendous number of studies trying to find therapeutic targets for treating basal-like breast cancer patients,” says Cho. “But clinical trials have failed due to the complex and dynamic nature of cancer. To overcome this issue, we looked at breast cancer cells as a complex network system and implemented a systems biological approach to unravel the underlying mechanisms that would allow us to reprogram basal-like into luminal-A breast cancer cells.”
Using this approach, followed by experimental validation on real breast cancer cells, the team found that turning off two key gene regulators, called BCL11A and HDAC1/2, switched a basal-like cancer signalling pathway into a different one used by luminal-A cancer cells. The switch reprograms the cancer cells and makes them more responsive to drugs that target ERα receptors. However, further tests will be needed to confirm that this also works in animal models and eventually humans.
“Our study demonstrates that the systems biological approach can be useful for identifying novel therapeutic targets,” says Cho.
The researchers are now expanding its breast cancer network model to include all breast cancer subtypes. Their ultimate aim is to identify more drug targets and to understand the mechanisms that could drive drug-resistant cells to turn into drug-sensitive ones.
This work was supported by the National Research Foundation of Korea, the Ministry of Science and ICT, Electronics and Telecommunications Research Institute, and the KAIST Grand Challenge 30 Project.
-Publication Sea R. Choi, Chae Young Hwang, Jonghoon Lee, and Kwang-Hyun Cho, “Network Analysis Identifies Regulators of Basal-like Breast Cancer Reprogramming and Endocrine TherapyVulnerability,” Cancer Research, November 30. (doi:10.1158/0008-5472.CAN-21-0621)
-ProfileProfessor Kwang-Hyun ChoLaboratory for Systems Biology and Bio-Inspired EngineeringDepartment of Bio and Brain EngineeringKAIST
2021.12.07 View 7023 -
 Hydrogel-Based Flexible Brain-Machine Interface
The interface is easy to insert into the body when dry, but behaves ‘stealthily’ inside the brain when wet
Professor Seongjun Park’s research team and collaborators revealed a newly developed hydrogel-based flexible brain-machine interface. To study the structure of the brain or to identify and treat neurological diseases, it is crucial to develop an interface that can stimulate the brain and detect its signals in real time. However, existing neural interfaces are mechanically and chemically different from real brain tissue. This causes foreign body response and forms an insulating layer (glial scar) around the interface, which shortens its lifespan.
To solve this problem, the research team developed a ‘brain-mimicking interface’ by inserting a custom-made multifunctional fiber bundle into the hydrogel body. The device is composed not only of an optical fiber that controls specific nerve cells with light in order to perform optogenetic procedures, but it also has an electrode bundle to read brain signals and a microfluidic channel to deliver drugs to the brain.
The interface is easy to insert into the body when dry, as hydrogels become solid. But once in the body, the hydrogel will quickly absorb body fluids and resemble the properties of its surrounding tissues, thereby minimizing foreign body response.
The research team applied the device on animal models, and showed that it was possible to detect neural signals for up to six months, which is far beyond what had been previously recorded. It was also possible to conduct long-term optogenetic and behavioral experiments on freely moving mice with a significant reduction in foreign body responses such as glial and immunological activation compared to existing devices.
“This research is significant in that it was the first to utilize a hydrogel as part of a multifunctional neural interface probe, which increased its lifespan dramatically,” said Professor Park. “With our discovery, we look forward to advancements in research on neurological disorders like Alzheimer’s or Parkinson’s disease that require long-term observation.”
The research was published in Nature Communications on June 8, 2021. (Title: Adaptive and multifunctional hydrogel hybrid probes for long-term sensing and modulation of neural activity) The study was conducted jointly with an MIT research team composed of Professor Polina Anikeeva, Professor Xuanhe Zhao, and Dr. Hyunwoo Yook.
This research was supported by the National Research Foundation (NRF) grant for emerging research, Korea Medical Device Development Fund, KK-JRC Smart Project, KAIST Global Initiative Program, and Post-AI Project.
-PublicationPark, S., Yuk, H., Zhao, R. et al. Adaptive and multifunctional hydrogel hybrid probes for long-term sensing and modulation of neural activity. Nat Commun 12, 3435 (2021). https://doi.org/10.1038/s41467-021-23802-9
-ProfileProfessor Seongjun ParkBio and Neural Interfaces LaboratoryDepartment of Bio and Brain EngineeringKAIST
2021.07.13 View 9300
Hydrogel-Based Flexible Brain-Machine Interface
The interface is easy to insert into the body when dry, but behaves ‘stealthily’ inside the brain when wet
Professor Seongjun Park’s research team and collaborators revealed a newly developed hydrogel-based flexible brain-machine interface. To study the structure of the brain or to identify and treat neurological diseases, it is crucial to develop an interface that can stimulate the brain and detect its signals in real time. However, existing neural interfaces are mechanically and chemically different from real brain tissue. This causes foreign body response and forms an insulating layer (glial scar) around the interface, which shortens its lifespan.
To solve this problem, the research team developed a ‘brain-mimicking interface’ by inserting a custom-made multifunctional fiber bundle into the hydrogel body. The device is composed not only of an optical fiber that controls specific nerve cells with light in order to perform optogenetic procedures, but it also has an electrode bundle to read brain signals and a microfluidic channel to deliver drugs to the brain.
The interface is easy to insert into the body when dry, as hydrogels become solid. But once in the body, the hydrogel will quickly absorb body fluids and resemble the properties of its surrounding tissues, thereby minimizing foreign body response.
The research team applied the device on animal models, and showed that it was possible to detect neural signals for up to six months, which is far beyond what had been previously recorded. It was also possible to conduct long-term optogenetic and behavioral experiments on freely moving mice with a significant reduction in foreign body responses such as glial and immunological activation compared to existing devices.
“This research is significant in that it was the first to utilize a hydrogel as part of a multifunctional neural interface probe, which increased its lifespan dramatically,” said Professor Park. “With our discovery, we look forward to advancements in research on neurological disorders like Alzheimer’s or Parkinson’s disease that require long-term observation.”
The research was published in Nature Communications on June 8, 2021. (Title: Adaptive and multifunctional hydrogel hybrid probes for long-term sensing and modulation of neural activity) The study was conducted jointly with an MIT research team composed of Professor Polina Anikeeva, Professor Xuanhe Zhao, and Dr. Hyunwoo Yook.
This research was supported by the National Research Foundation (NRF) grant for emerging research, Korea Medical Device Development Fund, KK-JRC Smart Project, KAIST Global Initiative Program, and Post-AI Project.
-PublicationPark, S., Yuk, H., Zhao, R. et al. Adaptive and multifunctional hydrogel hybrid probes for long-term sensing and modulation of neural activity. Nat Commun 12, 3435 (2021). https://doi.org/10.1038/s41467-021-23802-9
-ProfileProfessor Seongjun ParkBio and Neural Interfaces LaboratoryDepartment of Bio and Brain EngineeringKAIST
2021.07.13 View 9300 -
 Prof. Sang Wan Lee Selected for 2021 IBM Academic Award
Professor Sang Wan Lee from the Department of Bio and Brain Engineering was selected as the recipient of the 2021 IBM Global University Program Academic Award. The award recognizes individual faculty members whose emerging science and technology contains significant interest for universities and IBM.
Professor Lee, whose research focuses on artificial intelligence and computational neuroscience, won the award for his research proposal titled A Neuroscience-Inspired Approach for Metacognitive Reinforcement Learning. IBM provides a gift of $40,000 to the recipient’s institution in recognition of the selection of the project but not as a contract for services.
Professor Lee’s project aims to exploit the unique characteristics of human reinforcement learning. Specifically, he plans to examines the hypothesis that metacognition, a human’s ability to estimate their uncertainty level, serves to guide sample-efficient and near-optimal exploration, making it possible to achieve an optimal balance between model-based and model-free reinforcement learning.
He was also selected as the winner of the Google Research Award in 2016 and has been working with DeepMind and University College London to conduct basic research on decision-making brain science to establish a theory on frontal lobe meta-enhance learning.
"We plan to conduct joint research for utilizing brain-based artificial intelligence technology and frontal lobe meta-enhanced learning technology modeling in collaboration with an international research team including IBM, DeepMind, MIT, and Oxford,” Professor Lee said.
2021.06.25 View 9208
Prof. Sang Wan Lee Selected for 2021 IBM Academic Award
Professor Sang Wan Lee from the Department of Bio and Brain Engineering was selected as the recipient of the 2021 IBM Global University Program Academic Award. The award recognizes individual faculty members whose emerging science and technology contains significant interest for universities and IBM.
Professor Lee, whose research focuses on artificial intelligence and computational neuroscience, won the award for his research proposal titled A Neuroscience-Inspired Approach for Metacognitive Reinforcement Learning. IBM provides a gift of $40,000 to the recipient’s institution in recognition of the selection of the project but not as a contract for services.
Professor Lee’s project aims to exploit the unique characteristics of human reinforcement learning. Specifically, he plans to examines the hypothesis that metacognition, a human’s ability to estimate their uncertainty level, serves to guide sample-efficient and near-optimal exploration, making it possible to achieve an optimal balance between model-based and model-free reinforcement learning.
He was also selected as the winner of the Google Research Award in 2016 and has been working with DeepMind and University College London to conduct basic research on decision-making brain science to establish a theory on frontal lobe meta-enhance learning.
"We plan to conduct joint research for utilizing brain-based artificial intelligence technology and frontal lobe meta-enhanced learning technology modeling in collaboration with an international research team including IBM, DeepMind, MIT, and Oxford,” Professor Lee said.
2021.06.25 View 9208 -
 Ultrafast, on-Chip PCR Could Speed Up Diagnoses during Pandemics
A rapid point-of-care diagnostic plasmofluidic chip can deliver result in only 8 minutes
Reverse transcription-polymerase chain reaction (RT-PCR) has been the gold standard for diagnosis during the COVID-19 pandemic. However, the PCR portion of the test requires bulky, expensive machines and takes about an hour to complete, making it difficult to quickly diagnose someone at a testing site. Now, researchers at KAIST have developed a plasmofluidic chip that can perform PCR in only about 8 minutes, which could speed up diagnoses during current and future pandemics.
The rapid diagnosis of COVID-19 and other highly contagious viral diseases is important for timely medical care, quarantining and contact tracing. Currently, RT-PCR uses enzymes to reverse transcribe tiny amounts of viral RNA to DNA, and then amplifies the DNA so that it can be detected by a fluorescent probe. It is the most sensitive and reliable diagnostic method.
But because the PCR portion of the test requires 30-40 cycles of heating and cooling in special machines, it takes about an hour to perform, and samples must typically be sent away to a lab, meaning that a patient usually has to wait a day or two to receive their diagnosis.
Professor Ki-Hun Jeong at the Department of Bio and Brain Engineering and his colleagues wanted to develop a plasmofluidic PCR chip that could quickly heat and cool miniscule volumes of liquids, allowing accurate point-of-care diagnoses in a fraction of the time. The research was reported in ACS Nano on May 19.
The researchers devised a postage stamp-sized polydimethylsiloxane chip with a microchamber array for the PCR reactions. When a drop of a sample is added to the chip, a vacuum pulls the liquid into the microchambers, which are positioned above glass nanopillars with gold nanoislands. Any microbubbles, which could interfere with the PCR reaction, diffuse out through an air-permeable wall. When a white LED is turned on beneath the chip, the gold nanoislands on the nanopillars quickly convert light to heat, and then rapidly cool when the light is switched off.
The researchers tested the device on a piece of DNA containing a SARS-CoV-2 gene, accomplishing 40 heating and cooling cycles and fluorescence detection in only 5 minutes, with an additional 3 minutes for sample loading. The amplification efficiency was 91%, whereas a comparable conventional PCR process has an efficiency of 98%. With the reverse transcriptase step added prior to sample loading, the entire testing time with the new method could take 10-13 minutes, as opposed to about an hour for typical RT-PCR testing. The new device could provide many opportunities for rapid point-of-care diagnostics during a pandemic, the researchers say.
-Publication
Ultrafast and Real-Time Nanoplasmonic On-Chip Polymerase Chain Reaction for Rapid and Quantitative Molecular Diagnostics
ACS Nano (https://doi.org/10.1021/acsnano.1c02154)
-Professor
Ki-Hun Jeong
Biophotonics Laboratory
https://biophotonics.kaist.ac.kr/
Department of Bio and Brain Engineeinrg
KAIST
2021.06.08 View 8336
Ultrafast, on-Chip PCR Could Speed Up Diagnoses during Pandemics
A rapid point-of-care diagnostic plasmofluidic chip can deliver result in only 8 minutes
Reverse transcription-polymerase chain reaction (RT-PCR) has been the gold standard for diagnosis during the COVID-19 pandemic. However, the PCR portion of the test requires bulky, expensive machines and takes about an hour to complete, making it difficult to quickly diagnose someone at a testing site. Now, researchers at KAIST have developed a plasmofluidic chip that can perform PCR in only about 8 minutes, which could speed up diagnoses during current and future pandemics.
The rapid diagnosis of COVID-19 and other highly contagious viral diseases is important for timely medical care, quarantining and contact tracing. Currently, RT-PCR uses enzymes to reverse transcribe tiny amounts of viral RNA to DNA, and then amplifies the DNA so that it can be detected by a fluorescent probe. It is the most sensitive and reliable diagnostic method.
But because the PCR portion of the test requires 30-40 cycles of heating and cooling in special machines, it takes about an hour to perform, and samples must typically be sent away to a lab, meaning that a patient usually has to wait a day or two to receive their diagnosis.
Professor Ki-Hun Jeong at the Department of Bio and Brain Engineering and his colleagues wanted to develop a plasmofluidic PCR chip that could quickly heat and cool miniscule volumes of liquids, allowing accurate point-of-care diagnoses in a fraction of the time. The research was reported in ACS Nano on May 19.
The researchers devised a postage stamp-sized polydimethylsiloxane chip with a microchamber array for the PCR reactions. When a drop of a sample is added to the chip, a vacuum pulls the liquid into the microchambers, which are positioned above glass nanopillars with gold nanoislands. Any microbubbles, which could interfere with the PCR reaction, diffuse out through an air-permeable wall. When a white LED is turned on beneath the chip, the gold nanoislands on the nanopillars quickly convert light to heat, and then rapidly cool when the light is switched off.
The researchers tested the device on a piece of DNA containing a SARS-CoV-2 gene, accomplishing 40 heating and cooling cycles and fluorescence detection in only 5 minutes, with an additional 3 minutes for sample loading. The amplification efficiency was 91%, whereas a comparable conventional PCR process has an efficiency of 98%. With the reverse transcriptase step added prior to sample loading, the entire testing time with the new method could take 10-13 minutes, as opposed to about an hour for typical RT-PCR testing. The new device could provide many opportunities for rapid point-of-care diagnostics during a pandemic, the researchers say.
-Publication
Ultrafast and Real-Time Nanoplasmonic On-Chip Polymerase Chain Reaction for Rapid and Quantitative Molecular Diagnostics
ACS Nano (https://doi.org/10.1021/acsnano.1c02154)
-Professor
Ki-Hun Jeong
Biophotonics Laboratory
https://biophotonics.kaist.ac.kr/
Department of Bio and Brain Engineeinrg
KAIST
2021.06.08 View 8336